Abstract
Main conclusion
Five poplar CHASE-containing histidine kinase receptors bind cytokinins and display kinase activities. Both endogenous isoprenoid and aromatic cytokinins bind to the receptors in live cell assays.
Abstract
Cytokinins are phytohormones that play key roles in various developmental processes in plants. The poplar species Populus × canadensis, cv. Robusta, is the first organism found to contain aromatic cytokinins. Here, we report the functional characterization of five CHASE-containing histidine kinases from P. × canadensis: PcHK2, PcHK3a, PcHK3b, PcHK4a and PcHK4b. A qPCR analysis revealed high transcript levels of all PcHKs other than PcHK4b across multiple poplar organs. The ligand specificity was determined using a live cell Escherichia coli assay and we provide evidence based on UHPLC-MS/MS data that ribosides can be true ligands. PcHK2 exhibited higher sensitivity to iP-type cytokinins than the other receptors, while PcHK3a and PcHK3b bound these cytokinins much more weakly, because they possess two isoleucine residues that clash with the cytokinin base and destabilize its binding. All receptors display kinase activity but their activation ratios in the presence/absence of cytokinin differ significantly. PcHK4a displays over 400-fold higher kinase activity in the presence of cytokinin, suggesting involvement in strong responses to changes in cytokinin levels. trans-Zeatin was both the most abundant cytokinin in poplar and that with the highest variation in abundance, which is consistent with its strong binding to all five HKs and activation of cytokinin signaling via A-type response regulators. The aromatic cytokinins’ biological significance remains unclear, their levels vary diurnally, seasonally, and annually. PcHK3 and PcHK4 display the strongest binding at pH 7.5 and 5.5, respectively, in line with their putative membrane localization in the endoplasmic reticulum and plasma membrane.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The cytokinins are adenine-derived phytohormones that regulate cell proliferation and differentiation, shoot and root growth, development, and responses to various environmental stimuli including biotic and abiotic stress (Werner and Schmülling 2009; Kieber and Schaller 2014). Two groups of cytokinins, aromatic and isoprenoid, can be distinguished based on the nature of their N-substituents. Isoprenoid cytokinins are ubiquitous in higher plants and include the most biologically active species trans-Zeatin (tZ) and N6-(2-isopentenyl)adenine (iP) as well as less bioactive analogs such as cis-zeatin (cZ) and dihydrozeatin (Spíchal et al. 2004; Lomin et al. 2011; Kuderová et al. 2015). Aromatic cytokinins are rare and may have different biological functions to their isoprenoid counterparts (Strnad 1997). Notable aromatic cytokinins include 6-benzylaminopurine (BAP) and ortho- (oT) and meta-topolin (mT). Aromatic cytokinins were shown to induce changes in chlorophyll a and b levels (Dobránszki and Mendler-Drienyovszki 2014). Both isoprenoid and aromatic cytokinins also exist as ribosides, 9-glucosides, and other derivatives (Mok and Mok 2001). Ribosides, considered in general as a transport form of the hormone (Beveridge et al. 1997; Corbesier et al. 2003; Hirose et al. 2008), can be metabolized by nucleoside-N-ribohydrolase (NRH) to the active form (cytokinin base) in cytosol (Kopečná et al. 2013).
The cytokinin signal is perceived and mediated by a multistep His-Asp phosphorelay cascade, which activates type-B response regulators (RR) that function as transcriptional activators in the nucleus (Hwang and Sheen 2001; Müller and Sheen 2008). Among the genes transcribed upon B-type RRs activation are smaller A-type RRs acting as negative regulators of the initial signal transduction pathway (To et al. 2004). The A-type RRs are cytokinin primary response genes (D’Agostino et al. 2000; Rashotte et al. 2003). Cytokinin receptors are hybrid-type transmembrane sensor histidine kinases (HKs) (Inoue et al. 2001; Suzuki et al. 2001; Ueguchi et al. 2001) that are autophosphorylated at a conserved His residue in the HK domain, and subsequently at an Asp residue in the receiver domain, upon cytokinin binding. The cytokinin binding site is at an N-terminal extracytoplasmic binding domain known as the CHASE (Cyclase/Histidine kinase Associated Sensory Extracellular) domain, which comprises ~ 270 amino acids (Heyl et al. 2007) surrounded by transmembrane (TM) domains of ~ 20–25 amino acids. Receptors are predominantly localized in the membrane of the endoplasmic reticulum (Caesar et al. 2011; Lomin et al. 2011; Wulfetange et al. 2011). The best described cytokinin receptors are three HKs from Arabidopsis (AHK2, AHK3 and AHK4) (Ueguchi et al. 2001). The crystal structure of the CHASE domain of AHK4 (Hothorn et al. 2011) has been solved, revealing that it forms a dimer and explaining previously reported ligand specificity (Spíchal et al. 2004; Romanov et al. 2006). Other organisms with biochemically characterized cytokinin receptors include maize (Yonekura-Sakakibara et al. 2004), rice (Choi et al. 2012), rapeseed (Kuderová et al. 2015), the apple tree (Daudu et al. 2017), and potato (Lomin et al. 2018). The pathogenic bacterium Xanthomonas campestris also senses cytokinin via HKs (Wang et al. 2017), which induces a bacterial response to oxidative stress.
Plant HKs display high redundancy with overlapping specificities, functions, interactions and gene expression, at least in Arabidopsis thaliana (Dortay et al. 2006; Stolz et al. 2011). While ahk2 ahk4 and ahk3 ahk4 double mutants exhibited only minor phenotypic differences from the wild type (Nishimura et al. 2004; Higuchi et al. 2004; Riefler et al. 2006), studies on the dwarfed ahk2 ahk3 double mutant revealed that AHK2 and AHK3 play important roles in leaf development. AHK3 regulates leaf senescence (Kim et al. 2006). Conversely, AHK4 activity is required for the morphogenesis of vascular tissues in roots (Mähönen et al. 2000). Cytokinin signaling is also involved in the formation of legume root nodules: spontaneous nodulation in a Lotus japonicus mutant was linked to a gain-of-function mutation in the HK1 gene (Tirichine et al. 2007). This mutation L266F affected the sequence of the CHASE domain, making the mutant protein constitutively active even without cytokinin binding. Constitutively active variants of the Arabidopsis AHK4 receptor have also been described (Miwa et al. 2007). All these variants feature mutations in a narrow region extending from the TM to the HK domain. Gain-of-function mutants of the AHK2 and AHK3 genes, named repressor of cytokinin deficiency2 (rock2) and rock3, caused increased shoot growth, leaf and flower size, and seed yield (Bartrina et al. 2017). Interestingly, the rock3-1 and rock3-2 mutant genes encode AHK3 variants with point mutations in the CHASE domain (T179I and E182 K, respectively).
Here, we present a study on a family of cytokinin receptors from Populus × canadensis cv. Robusta, a hybrid of P. deltoides × P. nigra. We chose P. × canadensis as a model, because it is the first organism reported to contain aromatic cytokinins: oT (Strnad et al. 1992), ortho-topolin riboside (Hewett and Wareing 1973; Horgan et al. 1973), ortho-topolin-9-glucoside (oT9G) (Strnad et al. 1994), mT, and its sugar conjugates (Strnad et al. 1997) have been detected in extracts of its leaves. We recently analyzed the occurrence of aromatic cytokinins in various poplar species (Jaworek et al. 2019). All five poplar HK genes were cloned and putative cytokinin receptors were analyzed using a live-cell competitive assay to determine ligand specificities and kinase activities. We also investigated the genomic organization of the HK genes and their expression patterns in poplar by qPCR.
Materials and methods
Biological material and growth conditions
The majority of Populus × canadensis (cv. Robusta) samples were collected from a tree growing in the urban environment (Olomouc, CZ, 49°35′44″ N, 17°13′39″ E). Samples for the determination of seasonal changes in cytokinin content were collected at a set time after daybreak between 25 May and 29th September 2017. A total of 48 data points represent values in samples collected every second week in two biological replicates (each replicate was a mixture of five leaves), which were measured in triplicate. Roots were obtained from poplar twigs rooted in a tap water for 2 weeks. Calli were grown in a growth chamber at 23/21 °C (day/night), with a 14 h photoperiod (140 µmol m−2 s−1) on MS medium with vitamins at pH 5.7 solidified with 0.7% (w/v) plant agar (Duchefa) and supplemented with 2% (w/v) sucrose, glutamine (AppliChem) at 10 mg l−1, glycine at 2 mg l−1, tZ at 0.2 mg l−1, and 2,4-dichlorophenoxyacetic acid at 1.0 mg l−1 (Duchefa). Sub-cultivation was performed every 2 weeks. All samples were frozen in liquid nitrogen.
Cloning and construct preparation
Gene-specific primers (Table S1) for amplification of P. × canadensis HK genes, abbreviated as PcHKs, were designed based on previously identified corresponding sequences from Populus trichocarpa (Immanen et al. 2013). The Phytozome 12 (https://phytozome.jgi.doe.gov/) identification numbers of individual sequences are presented in Table 1. Total RNA was extracted from P. × canadensis leaves or calli using the RNAqueous isolation kit and treated with Turbo DNase (Thermo Fisher Scientific). cDNA was synthesized with RevertAid H Minus reverse transcriptase (Thermo Fisher Scientific) for PCR amplification with Q5 DNA polymerase from New England Biolabs (NEB). All PcHK ORFs were ligated into a Zero Blunt vector (Thermo Fisher Scientific), transformed into NEB 10-β competent E. coli cells for plasmid production, and sequenced. A similar procedure was used for genes encoding ubiquitin extension protein 1 (UBQ1; GeneBank XP_002318470) and α-tubulin 2 (TUA2; GeneBank XP_002303998), which were used as a reference genes for qPCR. All HK genes were further subcloned into the pINIIIΔEH vector (kindly provided by David Zalabák, Palacký University, CZ) (Suzuki et al. 2001; Yamada et al. 2001) using its BamHI and SpeI restriction sites and preserving the GATT motif in front of the start codon. The constructs were transformed into Escherichia coli strain KMI001 (kindly provided by David Zalabák, Palacký University, CZ) for use in cytokinin binding and receptor activation assays. Because the natural ORFs of PcHK2, PcHK3a, PcHK3b and PcHK4a contain BamHI or SpeI sites, these were further codon optimized (GenScript).
qPCR analysis of PcHK and PcRR expression
Total RNA from individual tissues and calli of P. × canadensis was extracted using the RNAqueous isolation kit and treated with Turbo DNase (Thermo Fisher Scientific) twice. Expression of A-type response regulator genes (PcRR) was analyzed in freshly cut leaves from mature trees. Leaf stalks were incubated in 1 µM tZ, 0.5 µM oT and 0.5 µM oTR for 4 h. Alternatively, freshly cut young twigs were treated with 2 µM tZ and BAP for 24 h. cDNA was synthesized using the RevertAid H Minus reverse transcriptase (Thermo Fisher Scientific) with random hexamer primers. RNA from a pool of 15–20 poplar organ samples (flowers, buds, leaves, etc.) was transcribed in two independent reactions. All subsequent PCR analyses of cDNA samples were performed in triplicate. The qPCR reaction mixture contained cDNA, 200 nM FAM-TAMRA probe, 400 nM primers, and Luna® universal probe qPCR master mix (NEB). Primer Express 3.0 software (Thermo Fisher Scientific) was used for primer and probe design (Table S1). Data were recorded using the QuantStudio 5 Real-Time PCR System (Thermo Fisher Scientific). PCR efficiencies and specificities of designed probes and primer pairs were verified using plasmid DNA carrying ORFs of the poplar HK genes. Cycle threshold values were normalized with respect to the amplification efficiency of two housekeeping genes UBQ1 and TUA2.
Phylogenetic analysis and cytokinin-binding site comparison
An amino acid alignment of the HK sequences was generated using T-Coffee (Notredame et al. 2000) and then treated with Gblocks (Castresana 2000). A maximum likelihood phylogeny with bootstrap analysis was constructed with PhyML v3.0 using the LG amino acid replacement matrix (Guindon et al. 2010). The phylogeny included HK sequences from Arabidopsis thaliana (AHKs) (Ueguchi et al. 2001), Brassica napus (BnHKs) (Kuderová et al. 2015), Malus domestica (MdHKs) (Daudu et al. 2017), Oryza sativa (OsHKs) (Du et al. 2007), P. × canadensis (PcHKs; Table 1), Solanum tuberosum (StHKs) (Lomin et al. 2018), and Zea mays (ZmHKs) (Yonekura-Sakakibara et al. 2004). A sequence logo was generated using WebLogo from the sequences used in the phylogenetic analysis together with sequences from Prunus persica (Immanen et al. 2013) and sequences from Hordeum vulgare, Kalanchoe laxiflora, Salix purpurea, and Solanum lycopersicum retrieved from the Phytozome 12 (https://phytozome.jgi.doe.gov/) or Uniprot (www.uniprot.org) databases. Models of the poplar receptor sensor domains were constructed with SWISS-MODEL (https://swissmodel.expasy.org) using the dimeric domain of the AHK4 crystal structure (PDB ID 3T4L) (Hothorn et al. 2011) as a template. The resulting models of the PcHK4a and PcHK4b CHASE domains display an average RMSD of 0.15 Å and share 83% sequence identity with that of Arabidopsis AHK4. Models of the PcHK2, PcHK3a and PcHK3b CHASE domains display an average RMSD of 0.25 Å and share 66–68% sequence identity with that of Arabidopsis AHK4. HK sequences from P. × canadensis, P. trichocarpa and P. deltoides were nearly identical as shown in Table S2.
Cytokinin-binding assay
The PcHKs’ binding properties were analyzed using a live cell competitive binding assay (Romanov et al. 2005, 2006) with slight modifications as described previously (Kuderová et al. 2015). Radiolabeled trans-Zeatin ([2-3H]zeatin, 1.3 TBq/mmol) was obtained from the Isotope Laboratory, Institute of Experimental Botany (Prague, CZ). Bacterial cultures were grown in liquid M9 medium supplemented with 0.1% (w/v) casamino acids, 100 μg ml−1 ampicillin and 100 μM IPTG at 25 °C overnight up to an OD600 of 0.7–0.8. Samples of the bacterial suspension (1 ml) were then transferred to Eppendorf tubes, incubated for 30 min, and centrifuged at 6000×g for 6 min. The resulting bacterial pellet was resuspended in 1 ml of a scintillation cocktail (Beckman, USA) and its radioactivity was monitored using a Hidex 300 SL scintillation counter (Hidex). A high excess of unlabeled tZ (at least 3000-fold) was used for competition to discriminate between specific and non-specific binding. Mean KD values were determined based on three independent Scatchard analyses (Scatchard 1949) using GraphPad Prism 5.1 (http://www.graphpad.com/). Competitive assays to determine IC50 values were performed at pH 7.0 using 2.5 nmol l−1 [3H]-tZ for PcHK2, PcHK3a and PcHK3b or 5 nmol l−1 [3H]-tZ for PcHK4a and PcHK4b samples, with or without unlabeled cytokinins at various concentrations, and 0.1% (v/v) dimethylsulphoxide (DMSO). Ki values were calculated using the equation Ki = IC50/(1 + [radioligand]/KD) (Cheng and Prusoff 1973), where KD denotes the radioligand’s affinity for the receptor being studied. The optimal pH was studied using 100 mM MES and MOPS buffers in pH range from 5.5 to 8.0. Prior to measurements, bacterial cultures were resuspended in liquid M9 medium containing tenfold lower concentration of phosphate salts. Riboside stability was studied using the same concentration of iP and iPR (25 nM for PcHK2, 900 nM for PcHK3b and 125 nM for PcHK4a). In parallel, 30 µg of pure maize NRH2b (Kopečná et al. 2013) were added to the reactions with iPR to release free iP either at 4 °C or at room temperature for 30 min.
Receptor activation assay
The E. coli KMI001 strains express the HK receptor → YojN → RcsB → cps::lacZ pathway, enabling reporter gene activation upon cytokinin binding (Takeda et al. 2001). The ability of the PcHK receptors to trigger phosphorelay signaling was verified by monitoring the strains’ β-galactosidase activity in the absence or presence of tZ as described (Spíchal 2011) with slight modifications. Bacterial precultures were grown in liquid M9 medium (pH 7.0) supplemented with 0.1% (w/v) casamino acids and 100 μg ml−1 ampicillin at 25 °C up to an OD600 of 0.6. Precultures were diluted with fresh media containing 100 μM IPTG at ratios of 1:100 for strains expressing PcHK2, 1:5 for PcHK3a, 1:500 for PcHK3b, and 1:5 for PcHK4a and PcHK4b. Each preculture was then split into two batches; solvent was added to one (the control) and tZ at a final concentration of 1 μM to the other. Three aliquots of 200 μl from each culture were transferred to a 96-well plate and incubated for 18 h at 25 °C, with shaking at 250 rpm. At the end of the incubation, the OD600 values of these samples were measured using a Synergy™ H4 hybrid multi-mode microplate reader (Biotek), and a 50 μl subsample from each well was transferred to a new 96-well plate containing 2 μl of 25 mM 4-methyl umbelliferyl galactoside (Sigma–Aldrich) in each well. After a 15 min incubation at 37 °C, the reaction was stopped by adding 100 μl of a pH 10.7 solution (NaOH) containing 130 mM glycine and 80 mM sodium carbonate. The fluorescence of 4-methylumbelliferone was measured using a plate reader at excitation and emission wavelengths of 365 and 460 nm, respectively. The ratio of the fluorescence signal to the product of the OD600 and cultivation time was used to compare the β-galactosidase activity of the controls and tZ-supplemented samples.
Analysis of cytokinin content
Isoprenoid cytokinins were isolated and purified according to (Bar et al. 2016) using 3 pmol of heavy-labeled standards (13C5-tZ, 13C5-cZ, 2H6-iP, 2H6-iPR, 2H5-tZR, 2H5-cZR 2H3-DHZR, 2H6-iPRMP, 2H5-tZ9G, 2H5-cZ9G 2H3-DHZ9G) (OlChemIm, Olomouc, CZ). Cytokinins were isolated by solid-phase extraction with a strong cation exchanger (Agilent Technologies), concentrated under reduced pressure at 37 °C, and analyzed by UHPLC-MS/MS (LCMS-8050, Shimadzu) according to (Novák et al. 2008). The resulting data were processed using the standard isotope dilution method (Rittenberg and Foster 1940). Molar concentrations of cytokinins were calculated by assuming a relative water content of 95% for leaf tissue (Sorrentino et al. 2016) to compare hormone levels in leaves to the receptors’ Ki values.
The identity and levels of iP and iPR incubated with E. coli cell cultures expressing PcHK4a were determined via UHPLC-MS/MS (LCMS-8050, Shimadzu) according to Novák et al. (2008). Samples for analysis were prepared by liquid–liquid extraction of cultivation media with diethyl ether. Three ether aliquots for each sample were pooled and concentrated under reduced pressure at 30 °C prior to analysis.
Accession numbers
Sequence data can be found in the GenBank (www.ncbi.nlm.nih.gov/genbank/) data library under accession numbers MH248795 for PcHK2, MH248793 for PcHK3a, MH248794 for PcHK3b, MH248791 for PcHK4a and MH248792 for PcHK4b. Alternatively spliced PcHK2 sequences were also submitted as MH248796 for PcHK2_v1 and MH248797 for PcHK2_v2.
Results
HK genes in Populus × canadensis cv. Robusta and their phylogeny
A detailed analysis of the Populus trichocarpa genome revealed the existence of five putative cytokinin receptors, all of which are CHASE-containing HKs (Table 1) (Pils and Heyl 2009; Immanen et al. 2013). The high number of these receptors in poplar compared to Arabidopsis is due to a whole-genome duplication event in the Salicaceae family, from which almost 8000 pairs of paralogous genes have survived in poplar to the present day (Tuskan et al. 2006). We cloned all ORFs of the five HKs found in P. × canadensis (for accession numbers, see Table 1) including several splicing variants of PcHK2. The PcHK2 gene lies on chromosome 9 and comprises 13 exons (Fig. S1). The exon number and lengths of HK2 orthologues, including those from Arabidopsis and maize, are conserved except for the first exon of PcHK2, which is about 150 bp longer. Additionally, there are two PcHK3 paralogues on chromosomes 1 and 14 with 10 exons, and two PcHK4 paralogues on chromosomes 8 and 10 with 11 exons. Uniquely, the Arabidopsis AHK4 gene contains an additional first exon and longer second exon that encode an extra membrane-anchored N-terminal helix (Ueguchi et al. 2001; Hothorn et al. 2011). In line with increasing length of N-terminal part of sequences, both PcHK4 comprise two TM segments, both PcHK3 have 3 TM segments and PcHK2 comprises even 4 TM segments (Table S3).
The phylogenetic tree constructed from the HK sequences of the seven studied plant species (which include both monocots and dicots) shows that the receptors cluster into three main clades (Fig. 1). Each clade contains one of the Arabidopsis receptors with established numbering; this pattern is also observed for the poplar, potato (Lomin et al. 2018), and apple tree (Daudu et al. 2017) orthologues. Brassica napus is the only analyzed species (Kuderová et al. 2015) without representation in all clades; it lacks an AHK4 orthologue but has four members of the HK2 clade.
Phylogenetic tree illustrating the relationship of poplar PcHK2, PcHK3a/b and PcHK4a/b to other CHASE-containing HKs. Amino acid alignments of 28 HKs were performed using T-Coffee (Notredame et al. 2000) and treated with Gblocks (Castresana 2000). The maximum likelihood phylogeny with bootstrap analysis was performed with PhyML v3.0 using the LG amino acid replacement matrix (Guindon et al. 2010). The internal labels show bootstrap frequencies (in percentage) for each clade. Scale bar shows an average number of substitutions per site. Only a subset of known HKs was chosen, focusing on those that have been studied previously. The subset includes examples from Arabidopsis thaliana (AtHKs: At5g35750, At1g27320 and At2g01830), Brassica napus (BnHKs: KF621029-33), Malus domestica (MdHKs: KM114879-83), Oryza sativa (OsHKs: Os02g50480, Os03g50860, Os10g21810 and Os01g69920), Solanum tuberosum (StHKs: XM_015303261.1, XM_006352114.2 and XM_006354988.2) and Zea mays (ZmHKs: AB042270, AB102956, AB102957 and AB121445)
Expression profile of poplar HKs
The gene expression of the five PcHKs was analyzed in various poplar organs by qRT-PCR (Fig. 2). Their transcript copy numbers varied between 600 and 2200 per 1 ng of RNA (Table S4). PcHK2 is the most abundant HK gene, and is expressed strongly in all organs. The evolution of paralogous genes often leads to functional diversification, as appears to have happened in the case of the PcHK4 genes based on their distinct expression profiles. The two PcHK3 paralogues exhibit similar expression patterns, with their transcripts being most abundant in young leaves, roots and flowers. However, the only strongly expressed PcHK4 paralogue was PcHK4a, which is mostly expressed in roots and axillary meristems.
Functional analysis of poplar HKs using isoprenoid cytokinins
A functional analysis of all five cloned PcHK protein-encoding genes with conserved CHASE domains was performed to determine whether they can function as genuine cytokinin receptors. A modified direct binding assay using E. coli strains expressing individual full-length ORFs of five PcHKs was used (Romanov et al. 2005, 2006; Kuderová et al. 2015) to obtain saturation curves for [3H]-tZ (Fig. 3a). All five PcHKs display strong tZ-binding affinities with estimated KD values from 1.8 nM for PcHK3a to 5.5 nM for PcHK4b. The effect of pH on binding strength was analyzed in pH range from 5.5 to 8.0 (Fig. 3b). PcHK2 displayed a maximal binding at pH 6.5, while curves of both isoforms PcHK3 as well as of PcHK4 showed opposite trends. Cytokinin binding to PcHK3 receptors increased steadily towards higher pH values. On the other hand, binding to PcHK4 receptors decreased linearly from a maximal value at pH 5.5. The receptors’ kinase activities were also analyzed using tZ in a bacterial activation assay (Suzuki et al. 2001). All five receptors exhibited kinase activity and thus are functional cytokinin receptors. Uniquely, PcHK3b exhibited a relatively high background activity level (60% of that observed in the presence of 1 µM tZ) in the absence of cytokinins (Fig. S2). The fold activation value (i.e. the ratio of kinase activity in the presence and absence of tZ) for PcHK4a was 400, whereas those for PcHK2 and PcHK3b were only 6 and 2, respectively (Fig. 3c).
Cytokinin binding and kinase activity of PcHKs. a Dose-dependent specific binding of [3H]tZ to cytokinin receptor-expressing E. coli clones. Binding was studied at pH 7.0 and cytokinin concentrations of up to 30 nM, and the calculated KD values are given for each receptor. b Effect of pH on binding strength. Binding was analyzed for [3H]-tZ in pH range from 5.5 to 8.0 using 100 mM MES and MOPS buffers. c Ratios of maximal to background kinase activity for individual receptors. Maximal activity was measured in the presence of 1 µM tZ. Absolute detected fluorescence values were normalized against the product of the bacterial cultures’ OD600 values and cultivation times (fluorescence × OD−1600 × h−1). Two values were calculated for each receptor, representing results obtained in the absence and presence, respectively, of 1 µM tZ: 358 ± 16 and 2128 ± 371 for PcHK2, 11 ± 6 and 294 ± 53 for PcHK3a, 3817 ± 477 and 6522 ± 515 for PcHK3b, 6 ± 14 and 2795 ± 83 for PcHK4a, 13 ± 1 and 398 ± 44 for PcHK4b. All measurements were performed twice with three technical replicates
The receptors’ ligand specificities were investigated using isoprenoid and aromatic cytokinins including bases and ribosides (Fig. 4) by determining their IC50 (inhibitory concentration 50%) values, i.e. the concentrations of unlabeled ligands required to reduce the binding of a radioligand by 50%. This enabled the determination of Ki values (Table 2) (Cheng and Prusoff 1973), which are properties of the receptor and unlabeled ligand, whereas IC50 values are only properties of the experiment. The cytokinin binding strengths of all five receptors declined in the order tZ > iP > cZ. PcHK2 had a much higher sensitivity than the other four receptors; it displayed the lowest overall Ki values (the highest being 100 nM) and thus the strongest binding to cytokinin bases and ribosides. The only exception was tZR, which was bound slightly more strongly by the PcHK3a/b isoforms. However, PcHK3a and PcHK3b exhibit weaker binding to iP-type cytokinins, cZ, BAP, and oT than the other HKs. The Ki values for isoprenoid cytokinin bases range from 2 nM (for tZ) to 1.1 µM (for cZ), while those for isoprenoid cytokinin ribosides range from 9 nM (for tZR) to 2.3 µM (for iPR). In general, tZR is a good ligand for all five PcHKs, with Ki values up to 50 nM, while iPR only binds strongly to PcHK2.
Competitive binding curves for various cytokinins versus [3H]-tZ using five poplar cytokinin receptors. The bound radioactivity corresponding to 100% binding was 19500, 11,000, 70,000, 16,500 and 13,500 dpm for experiments with PcHK2, PcHK3a, PcHK3b, PcHK4a and PcHK4b, respectively. Binding was studied at pH 7.0
Binding of cytokinin ribosides to poplar HKs in live cell assay
The fact that cytokinin ribosides can bind to receptors in E. coli assays was further verified by measuring binding stability. E. coli cultures expressing PcHK2, PcHK3b and PcHK4a were incubated with iPR for 2 h at 4 °C compared with the standard 30 min. iP incubated for 30 min was used as a positive control. Ligand concentrations were chosen to reach 50% [3H]-tZ displacement according to Ki values for iPR (25, 900 and 125 nM, Table 2). Obtained data (Fig. 5a) show that receptor saturation was ~ 50% and did not significantly differ over the incubation period. An addition of ZmNRH2b (Kopečná et al. 2013) induced release of iP from iPR resulting in higher receptor saturation.
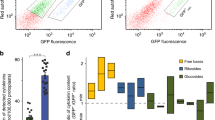
Stability of iPR in cytokinin binding assays with three PcHKs. a The graph shows relative binding of cytokinin base (iP) and riboside (iPR) determined by competitive binding assay with [3H]-tZ at pH 7.0. Concentrations of iP and iPR used in the assay were based on Ki values for each receptor (25 nM for PcHK2, 900 nM for PcHK3b and 125 nM for PcHK4a). Addition of nucleoside N-ribohydrolase (NRH), hydrolyzing ribosides to bases, to E. coli samples resulted in signal increase corresponding to stronger binding of iP. Dashed lines indicate a maximal response if all iPR would be converted to iP. b Concentration of iP and iPR present in the reaction mixture with PcHK4a receptor-expressing E. coli clones determined by UHPLC-MS/MS. Data show cytokinin levels under standard assay conditions and upon addition of NRH; applied 125 nM iPR incubated for 30 min is shown as a dashed line. c Calculated saturation of the PcHK4a receptor by the amount of released iP from iPR as measured by UHPLC-MS/MS in the reaction mixture
Additional UHPLC-MS/MS quantification of iPR and iP incubated at 125 nM (Ki values for iPR) for 30 min in media containing E. coli producing PcHK4a was performed to assess a false positive effect of iP released from iPR (Fig. 5b). A small conversion of iPR to iP was observed (7.1 nM of 125 nM). Such a concentration of iP would saturate the receptor only up to ~ 17% (83% of nondisplaced tritiated tZ) as calculated from the binding curve of iP (Fig. 5c). However, the assay with 125 nM iPR (Fig. 5a, second column) showed saturation of 49.7 ± 3.4%, which must be due to riboside binding. Addition of ZmNRH2b and 30-min incubation at room temperature resulted in 50% conversion of iPR to iP (58.0 nM). This concentration of iP would saturate the receptor up to ~ 60% according to the iP binding curve. We may assume that 65 ± 1.9% PcHK4a receptor saturation (Fig. 5a, last column) can be fully attributed to released iP.
Binding of urea-derived and aromatic cytokinins
The synthetic phenylurea TDZ (Mok et al. 1982) is a similarly good ligand to tZ, with low Ki values (Table 2). Two other TDZ derivatives, namely 1-[1,2,3]thiadiazol-5-yl-3-(3-trifluoromethoxy-phenyl)urea (3FMTDZ) and 1-[2-(2-hydroxy-ethyl)phenyl]-3-(1,2,3-thiadiazol-5-yl)urea (HETDZ) were also tested, because they are better inhibitors of cytokinin oxidase/dehydrogenase (CKX) (Nisler et al. 2016) than TDZ itself. As subtle modulations of cytokinin levels in planta can have positive effects on seed filling and crop yield, it is important to determine whether these new inhibitors can trigger undesired cytokinin signaling. 3FMTDZ and HETDZ are poor ligands for PcHK2, PcHK4a/b and PcHK3a. However, their affinity towards PcHK3b is higher, with Ki values of 0.6 µM and 1.2 µM, respectively. These values are comparable to the inhibition constants for CKX.
Of the tested aromatic cytokinin bases, BAP and mT bind to PcHKs with Ki values between ~ 23 nM and ~ 0.85 µM (Table 2), with mT generally binding more strongly. Conversely, oT is a poor ligand for PcHK3 isoforms; its Ki values were above the range measurable in the assay. Leaves of mature P. × canadensis trees exhibit diurnally fluctuating levels of oT under physiological conditions (Jaworek et al. 2019); its concentration can peak at around 250 nM, which may be sufficient to trigger cytokinin signaling in poplar by activating PcHK2 and (at least partially) PcHK4b. This point was clarified by analyzing the expression of two cytokinin primary fast response genes coding for A-type RRs (Ramírez-Carvajal et al. 2008), namely PcRR1 and PcRR10, by qPCR using FAM-TAMRA probes. PcRR1 gene is strongly (~ 7–9 fold) upregulated while PcRR10 gene response is only moderate (~ 3 fold) after tZ treatment for 4 h (Fig. S3). BAP, which displays much lower Ki values for all five receptors than oT, induces expression of PcRR1 only, although it is still half of that observed with tZ. Activation of cytokinin signaling by oT and oTR (transport form of oT) is inconclusive as only 2 out of 11 type-A RRs were studied and differences in expression are not statistically significant enough.
Cytokinin content in poplar leaves
Finally, we profiled cytokinins in mature leaves of P. × canadensis between the end of May and September in 2017 (Fig. 6). As we had detected oT and oTR in poplar leaves harvested in July 2015 (Jaworek et al. 2019), here, we screened their appearance covering the period of dark green fully expanded leaves till the early stage of senescence. Surprisingly, the levels of aromatic cytokinins were close to the detection limit (oT) or well below those required for appreciable receptor binding (oTR), and are not presented. The main isoprenoid cytokinin was tZ followed by iP, whereas the concentration of cZ was lower by one order of magnitude than that of tZ. Nevertheless, a decrease of total cytokinins was not observed during our experiment. While the levels of tZ and cZ steadily rose from the beginning of August till the end September, those of the remaining cytokinins did not follow any obvious trend (Fig. S4). iP and iPR displayed a positive peak during August.
Seasonal fluctuations of isoprenoid cytokinins in leaves of P. × canadensis. Values for each cytokinin represent a sum of 48 data points and concentrations are reported in units of nM although they were determined in pmol g−1 FW. The conversion assumes a relative leaf water content of 95% (1.0 g–0.95 ml). Values in boxes cover data from upper and lower quartiles (25–75%), lines in the middle of the boxes stand for a median value while the black squares indicate a mean values. Whiskers delineate 9th and 91st percentiles and stars indicate the lowest and highest values
Discussion
The poplar genome contains five putative cytokinin receptor HK genes and to get insight into the cytokinin perception, we cloned all five HKs and analyzed gene expression and binding properties of coded receptor proteins. The data of HK gene expression correlate well with those in the apple tree (Malus domesticus; MdHK) published only recently (Daudu et al. 2017). MdHK2, an orthologue of PcHK2, was the most abundant in leaves and stems; MdHK3a was mainly expressed in roots, and MdHK3b in flowers. In Arabidopsis, AHK2 was reported to be moderately expressed in all tested organs, whereas AHK3 transcripts were even more abundant in leaves and AHK4 transcripts appeared mainly in roots (Ueguchi et al. 2001; Higuchi et al. 2004).
The KD values for tZ and five poplar PcHKs in low nanomolar range observed in this work correlate well with those reported for Arabidopsis (Romanov et al. 2005, 2006; Stolz et al. 2011; Lomin et al. 2015), Brassica (Kuderová et al. 2015), and potato HKs (Lomin et al. 2018). The specificity and sensitivity of PcHK2 are strikingly similar to those for the two orthologous receptors from Brassica (Kuderová et al. 2015). The high sensitivity of PcHK2 parallels that of MdHK2 from apple tree (Daudu et al. 2017). Similarly, the weak binding of iP-type cytokinins by PcHK3a/b is mirrored by that of the Arabidopsis AHK3 receptor (Romanov et al. 2006) and to lesser degree by the AHK3-like Brassica BnHK5 receptor (Kuderová et al. 2015). Both of these receptors also display similar specificity as PcHK3a and PcHK3b. Our experiments also confirmed that cytokinins bind to all five poplar receptors at pH values of their supposed subcellular localization either in endoplasmic reticulum (neutral pH, Martinière et al. 2013) or in apoplast (acidic pH of ~ 5.5) (Fig. 3b). At pH 5.5, both PcHK4 receptors display maximal ligand-binding ability, while PcHK2 and both PcHK3 receptors have their pH maxima at more neutral or basic pH values.
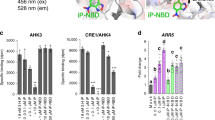
A superposition of the sensor domain of Arabidopsis AHK4 (Hothorn et al. 2011) with those of five PcHK models constructed using SWISS-MODEL (Waterhouse et al. 2018) confirmed the presence of highly conserved and mainly hydrophobic residues that form the cytokinin-binding cavity and are thus essential for receptor activity. The conservation of these residues is also apparent in the analysis of 52 plant HKs (Fig. 7a). All five PcHKs share a conserved aspartate residue (Asp285 in AHK4) that forms two hydrogen bonds to the nitrogen atoms of the cytokinin ligands’ adenine ring, and a conserved threonine residue (Thr317 in AHK4) that binds the hydroxyl group of tZ (Fig. 7b–d). While T317A mutant was still binding tZ, residues interacting with the adenine ring (namely D285, F304, L306 and L307) were found to be essential for cytokinin binding, because the corresponding alanine mutants were inactive (Hothorn et al. 2011).
Amino acid conservation in the cytokinin binding sites of poplar HKs. a Overall conservation of residues forming cytokinin binding sites in 52 plant HKs. Sequence logos were generated using WebLogo (http://weblogo.threeplusone.com). 3D representations of the binding sites for the PcHK4a model (b colored in pink), PcHK3a model (c colored in blue), and PcHK2 model (d colored in green) constructed using SWISS-MODEL with the crystal structure of Arabidopsis AHK4 (PDB ID 3T4L) serving as a template (colored in gray). The numbering follows that of AHK4; substitutions are labeled in red and indicated by red arrows
The primary sphere of interactions of PcHK4a and PcHK4b with the cytokinin ligand comprises only a minor difference compared with that of AHK4 where L274 is replaced by isoleucine in PcHK4a/b (Fig. 7b). The substitution occurs above the cytokinin adenine ring. PcHK2 also exhibits this substitution of L274 for isoleucine, together with the replacement of Y273 with histidine and A345 with serine (Fig. 7d). The latter change introduces a new H-bond donor/acceptor at the base of the binding site, enabling stronger interactions with the cytokinin base’s nitrogen atoms or the hydroxyl groups of a cytokinin riboside, as demonstrated by the Ki values for the PcHK2 and PcHK4a/b isoforms. The binding sites of PcHK3a and PcHK3b differ from that of AHK4 by five substitutions (Fig. 7c). L274, located above the cytokinin adenine ring, is replaced by smaller valine residue. However, two residues below the adenine ring, namely V315 and A345, are replaced by bulkier isoleucine residues. These residues would clash with a cytokinin molecule bound in the position observed in AHK4 (Hothorn et al. 2011) and, therefore, presumably alter the binding position of the cytokinin ligand, shifting its adenine moiety up towards V141 (in the PcHK3a numbering). These changes clearly destabilize the binding of the studied cytokinins (Table 2). The weak binding of iP derivatives can be explained by the missing isoprenoid hydroxyl group, which allows tZ derivatives to establish H-bonds to T317 on the opposite side of the cavity and thus bind more strongly.
Although a crystal structure of cytokinin receptor in complex with cytokinin riboside is not available, an induced fit, rearrangement of the loop close to the nucleobase or eventually a nucleobase flip may occur. In the case of cytokinin oxidase/dehydrogenase, the adenine base of the ligand can be flipped by 180° around its longitudinal axis formed by the isoprenoid side chain as observed for tZ in WT-ZmCKO1 (Malito et al. 2004) and for iP in P427Q-ZmCKO1 (Kopečný et al. 2016). Such a flip in AHK is possible and opens a possibility for riboside binding. Previously, it was demonstrated that cytokinin ribosides are poor ligands in plant assays using microsomal fractions but not intact cells as used in E. coli assay (Lomin et al. 2015). This brings a possibility of contamination with cytosolic enzymes involved in cytokinin metabolism such as NRHs (Kopečná at el. 2013) or adenosine kinases (Schoor et al. 2011) and others. We analyzed the stability of iPR in live cell E. coli assay and found no changes in signal during several hours (Fig. 5). However, all data during competition experiments are always acquired after 30 min incubation at 4 °C. Moreover, our UHPLC-MS/MS data show the presence of only a small amount of iP released from iPR which is not sufficient to displace 50% of [3H]-tZ based on binding curves for iP. These findings confirm that all five poplar HKs bind cytokinin ribosides in live cell assay. In parallel, pure NRH2b (Kopečná et al. 2013) was added to the reactions with iPR to release its free base iP. Our data show that it would take up to 8 h to fully convert iPR and reach the value of iP signal at 4 °C.
Incubation at room temperature resulted in approximately half-conversion of iPR (Fig. 5b). The amount of iP released and the calculated displacement of [3H]-tZ correlate well with the measured response of PcHK4a (displacement of tritiated tZ, Fig. 5a, c). Our finding is further supported by a recent study showing the ability of cytokinin receptors to bind N9-substituted cytokinins fluorescently labeled with 7-nitrobenzofurazan group (Kubiasová et al. 2018). Such compounds cannot be hydrolyzed as NRHs require the presence of the ribose group and, therefore, these derivatives are suitable for in vivo labeling.
The functional analysis of the receptors (Table 2, Fig. 4) suggests that PcHK2 (and to lesser extent also PcHK4a and PcHK4b) are the most likely to be activated by in vivo concentrations of iP. At least partial activation may also be induced by iPR. The cytokinin exhibiting the most dramatic seasonal changes in poplar was tZ (Fig. 6), but its concentration remained above Ki values for all receptors, suggesting that it plays a central role in cytokinin signaling in poplar. The affinity of tZR towards the receptors is lower by approximately one order of magnitude. However, the in vivo concentrations of tZR are more stable than those of tZ and still sufficient to activate all receptors, at least partially. Conversely, the concentration of cZ was much lower than that of the other cytokinins, suggesting that it induces little activation, if any. Levels of isoprenoid and aromatic cytokinins measured from P. × canadensis leaves presented in this work and those gathered 2 years earlier from the same tree (Jaworek et al. 2019) reveal considerable variability between years, as well as seasonal fluctuations. This is consistent with a report on annual and seasonal fluctuations in P. tremula (Edlund et al. 2017) observed in 3 consecutive years. Other factors such as interactions with microbial pathogens or insects must also be accounted for. Activation of cytokinin signaling pathway by both cytokinin types was verified by analyzing the expression of two A-type RRs, which are well known as fast cytokinin response genes. Cytokinin-dependent upregulation of RR1 and weaker activation of RR10 was observed, which slightly differs from findings in P. trichocarpa (Ramírez-Carvajal et al. 2008) where stronger activation was reported for RR10 using semi-quantitative PCR.
PcHK3b displays a significant constitutive background HK activity. The sequence shows no alternations in proximity to the conserved His site (SHE motif) found in the HK domain and conserved Asp site (FMD motif) in the receiver domain. Very recently, the similar activity was reported for both HK3a and HK3b in a study on apple MdHK family (Daudu et al. 2017), which relied on complementation of Δsln1 yeast mutant strain lacking endogenous osmo-sensing HK called SLN1 (Inoue et al. 2001). While MdHK2, MdHK4a and MdHK4b rescued the mutant in cytokinin-dependent manner, both MdHK3a and especially MdHK3b allowed a basal growth of the mutant in absence of cytokinin suggesting a constitutive background HK activity. Similar finding has also been reported for A. thaliana AHK3 and AHK2 (Tran et al. 2007). AHK4 variants equivalent to rock2 and rock3 mutants also suppressed the lethality of Δsln1 even without the addition of cytokinin (Bartrina et al. 2017). Additionally, it is known that among five apple cytokinin receptors only MdHK3a and MdHK3b do not form homodimers (Daudu et al. 2017). Taken together, the constitutive HK activity of HK3 found in woody plants raises the question of either additional sensing function or alternation of kinase activity due to the presence of particular unknown residue and slight structural rearrangements in relation to changes in the receptor’s oligomerization.
Conclusion
We have characterized all five poplar CHASE-containing HKs and verified that they function as genuine cytokinin receptors. The affinity of PcHK2 for cytokinins declines in the order tZ > TDZ > iP > iPR ≥ mT ≥ tZR > BAP > cZ > oT > oTR, whereas the cytokinin affinities of PcHK3a and PcHK3b decline in the order tZ > TDZ > tZR > mT > iP > BAP > cZ ~ iPR > oT > oTR. Those of PcHK4a and PcHK4b decline in the order tZ > iP ~ TDZ > tZR > iPR > mT ~ cZ > BAP > oT > oTR. Although the PcHK2 and PcHK3b genes are expressed quite strongly across various organs in poplar, they have comparatively low kinase activation ratios. Conversely, the kinase activity of PcHK4a increases over 400-fold upon cytokinin binding, suggesting an exceptional role in strong responses to changes in cytokinin levels. The best receptor for aromatic cytokinins is PcHK2, which also displays a strong affinity for the other tested cytokinins. The full set of poplar HK constructs developed in this work could be used as a tool for testing newly developed urea derivatives as CKX inhibitors to identify compounds with strong inhibitory effects that do not activate cytokinin receptors. Such compounds have potential agricultural applications as agents for increasing crop yields (Kopečný et al. 2010; Nisler et al. 2016). It is important to note that cytokinin profiling was performed at the leaf level. Therefore, the quantitative data presented here relate to average cytokinin concentrations in the organs. Future studies should investigate the localization of cytokinin signaling at the sub-organ level.
Author contribution statement
DK and PT designed the research; PJ, TH and DK performed cloning; PJ performed qPCR and binding assays; JN synthesized TDZ derivatives; ŠK and PT measured cytokinin levels in poplar; OV and PT measured cytokinins in E. coli; DK and PJ wrote the paper. All authors reviewed the results and approved the final version of the manuscript.
Abbreviations
- 3FMTDZ:
-
1-[1,2,3]thiadiazol-5-yl-3-(3-trifluoromethoxy-phenyl)urea
- BAP:
-
N6-benzyladenine
- cZ:
-
N6-(cis-4-hydroxy-3-methyl-2-buten-1-yl)adenine, i.e. cis-zeatin
- iP:
-
N6-(2-isopentenyl)adenine
- iPR:
-
N6-(2-isopentenyl)adenosine
- mT:
-
meta-topolin
- HETDZ:
-
1-[2-(2-hydroxy-ethyl)phenyl]-3-(1,2,3-thiadiazol-5-yl)urea
- HK:
-
Histidine kinase
- oT:
-
ortho-Topolin
- RR:
-
Response regulators
- TDZ:
-
N-(1,2,3-thidiazol-5-yl)-N´-phenylurea, i.e. thidiazuron
- tZ:
-
trans-Zeatin
- tZR:
-
Zeatin riboside
References
Bar M, Israeli A, Levy M et al (2016) CLAUSA is a MYB transcription factor that promotes leaf differentiation by attenuating cytokinin signaling. Plant Cell 28:1602–1615. https://doi.org/10.1105/tpc.16.00211
Bartrina I, Jensen H, Novák O et al (2017) Gain-of-function mutants of the cytokinin receptors AHK2 and AHK3 regulate plant organ size, flowering time and plant longevity. Plant Physiol 173:1783–1797. https://doi.org/10.1104/pp.16.01903
Beveridge CA, Murfet IC, Kerhoas L et al (1997) The shoot controls zeatin riboside export from pea roots. Evidence from the branching mutant rms4. Plant J 11:339–345. https://doi.org/10.1046/j.1365-313X.1997.11020339.x
Caesar K, Thamm AMK, Witthöft J et al (2011) Evidence for the localization of the Arabidopsis cytokinin receptors AHK3 and AHK4 in the endoplasmic reticulum. J Exp Bot 62:5571–5580. https://doi.org/10.1093/jxb/err238
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552. https://doi.org/10.1093/oxfordjournals.molbev.a026334
Cheng Y, Prusoff WH (1973) Relationship between the inhibition constant (KI) and the concentration of inhibitor which causes 50 per cent inhibition (I50) of an enzymatic reaction. Biochem Pharmacol 22:3099–3108
Choi J, Lee J, Kim K et al (2012) Functional identification of OsHk6 as a homotypic cytokinin receptor in rice with preferential affinity for iP. Plant Cell Physiol 53:1334–1343. https://doi.org/10.1093/pcp/pcs079
Corbesier L, Prinsen E, Jacqmard A et al (2003) Cytokinin levels in leaves, leaf exudate and shoot apical meristem of Arabidopsis thaliana during floral transition. J Exp Bot 54:2511–2517. https://doi.org/10.1093/jxb/erg276
D’Agostino IB, Deruère J, Kieber JJ (2000) Characterization of the response of the Arabidopsis response regulator gene family to cytokinin. Plant Physiol 124:1706–1717. https://doi.org/10.1104/pp.124.4.1706
Daudu D, Allion E, Liesecke F et al (2017) CHASE-containing histidine kinase receptors in apple tree: from a common receptor structure to divergent cytokinin binding properties and specific functions. Front Plant Sci 8:1614. https://doi.org/10.3389/fpls.2017.01614
Dobránszki J, Mendler-Drienyovszki N (2014) Cytokinin-induced changes in the chlorophyll content and fluorescence of in vitro apple leaves. J Plant Physiol 171:1472–1478. https://doi.org/10.1016/j.jplph.2014.06.015
Dortay H, Mehnert N, Bürkle L et al (2006) Analysis of protein interactions within the cytokinin-signaling pathway of Arabidopsis thaliana. FEBS J 273:4631–4644. https://doi.org/10.1111/j.1742-4658.2006.05467.x
Du L, Jiao F, Chu J et al (2007) The two-component signal system in rice (Oryza sativa L.): a genome-wide study of cytokinin signal perception and transduction. Genomics 89:697–707. https://doi.org/10.1016/j.ygeno.2007.02.001
Edlund E, Novak O, Karady M et al (2017) Contrasting patterns of cytokinins between years in senescing aspen leaves. Plant, Cell Environ 40:622–634. https://doi.org/10.1111/pce.12899
Guindon S, Dufayard J-F, Lefort V et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Hewett EW, Wareing PF (1973) Cytokinins in Populus x robusta (schneid): light effects on endogenous levels. Planta 114:119–129. https://doi.org/10.1007/BF00387470
Heyl A, Wulfetange K, Pils B et al (2007) Evolutionary proteomics identifies amino acids essential for ligand-binding of the cytokinin receptor CHASE domain. BMC Evol Biol 7:62. https://doi.org/10.1186/1471-2148-7-62
Higuchi M, Pischke MS, Mähönen AP et al (2004) In planta functions of the Arabidopsis cytokinin receptor family. Proc Natl Acad Sci 101:8821–8826. https://doi.org/10.1073/pnas.0402887101
Hirose N, Takei K, Kuroha T et al (2008) Regulation of cytokinin biosynthesis, compartmentalization and translocation. J Exp Bot 59:75–83. https://doi.org/10.1093/jxb/erm157
Horgan R, Hewett EW, Purse JG, Wareing PF (1973) A new cytokinin from Populus robusta. Tetrahedron Lett 14:2827–2828. https://doi.org/10.1016/S0040-4039(01)96062-9
Hothorn M, Dabi T, Chory J (2011) Structural basis for cytokinin recognition by Arabidopsis thaliana histidine kinase 4. Nat Chem Biol 7:766–768. https://doi.org/10.1038/nchembio.667
Huang X, Miller W (1991) A time-efficient, linear-space local similarity algorithm. Adv Appl Math 12:337–357. https://doi.org/10.1016/0196-8858(91)90017-D
Hwang I, Sheen J (2001) Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413:383–389. https://doi.org/10.1038/35096500
Immanen J, Nieminen K, Silva HD et al (2013) Characterization of cytokinin signaling and homeostasis gene families in two hardwood tree species: Populus trichocarpa and Prunus persica. BMC Genom 14:885. https://doi.org/10.1186/1471-2164-14-885
Inoue T, Higuchi M, Hashimoto Y et al (2001) Identification of CRE1 as a cytokinin receptor from Arabidopsis. Nature 409:1060–1063. https://doi.org/10.1038/35059117
Jaworek P, Kopečný D, Zalabák D et al (2019) Occurrence and biosynthesis of cytokinins in poplar. Planta 250:229–244. https://doi.org/10.1007/s00425-019-03152-z
Kieber JJ, Schaller GE (2014) Cytokinins. Arab Book. https://doi.org/10.1199/tab.0168
Kim HJ, Ryu H, Hong SH et al (2006) Cytokinin-mediated control of leaf longevity by AHK3 through phosphorylation of ARR2 in Arabidopsis. Proc Natl Acad Sci 103:814–819. https://doi.org/10.1073/pnas.0505150103
Kopečná M, Blaschke H, Kopečný D et al (2013) Structure and function of nucleoside hydrolases from Physcomitrella patens and maize catalyzing the hydrolysis of purine, pyrimidine, and cytokinin ribosides. Plant Physiol 163:1568–1583. https://doi.org/10.1104/pp.113.228775
Kopečný D, Briozzo P, Popelková H et al (2010) Phenyl- and benzylurea cytokinins as competitive inhibitors of cytokinin oxidase/dehydrogenase: a structural study. Biochimie 92:1052–1062. https://doi.org/10.1016/j.biochi.2010.05.006
Kopečný D, Končitíková R, Popelka H et al (2016) Kinetic and structural investigation of the cytokinin oxidase/dehydrogenase active site. FEBS J 283:361–377. https://doi.org/10.1111/febs.13581
Kubiasová K, Mik V, Nisler J et al (2018) Design, synthesis and perception of fluorescently labeled isoprenoid cytokinins. Phytochemistry 150:1–11. https://doi.org/10.1016/j.phytochem.2018.02.015
Kuderová A, Gallová L, Kuricová K et al (2015) Identification of AHK2- and AHK3-like cytokinin receptors in Brassica napus reveals two subfamilies of AHK2 orthologues. J Exp Bot 66:339–353. https://doi.org/10.1093/jxb/eru422
Lomin SN, Yonekura-Sakakibara K, Romanov GA, Sakakibara H (2011) Ligand-binding properties and subcellular localization of maize cytokinin receptors. J Exp Bot 62:5149–5159. https://doi.org/10.1093/jxb/err220
Lomin SN, Krivosheev DM, Steklov MY et al (2015) Plant membrane assays with cytokinin receptors underpin the unique role of free cytokinin bases as biologically active ligands. J Exp Bot 66:1851–1863. https://doi.org/10.1093/jxb/eru522
Lomin SN, Myakushina YA, Kolachevskaya OO et al (2018) Cytokinin perception in potato: new features of canonical players. J Exp Bot 69:3839–3853. https://doi.org/10.1093/jxb/ery199
Mähönen AP, Bonke M, Kauppinen L et al (2000) A novel two-component hybrid molecule regulates vascular morphogenesis of the Arabidopsis root. Genes Dev 14:2938–2943
Malito E, Coda A, Bilyeu KD, Fraaije MW, Mattevi A (2004) Structures of Michaelis and product complexes of plant cytokinin dehydrogenase: implications for flavoenzyme catalysis. J Mol Biol 341:1237–1249. https://doi.org/10.1016/j.jmb.2004.06.083
Martinière A, Bassil E, Jublanc E et al (2013) In vivo intracellular pH measurements in tobacco and Arabidopsis reveal an unexpected pH gradient in the endomembrane system. Plant Cell 25:4028–4043. https://doi.org/10.1105/tpc.113.116897
Miwa K, Ishikawa K, Terada K et al (2007) Identification of amino acid substitutions that render the Arabidopsis cytokinin receptor histidine kinase AHK4 constitutively active. Plant Cell Physiol 48:1809–1814. https://doi.org/10.1093/pcp/pcm145
Mok DW, Mok MC (2001) Cytokinin metabolism and action. Annu Rev Plant Physiol Plant Mol Biol 52:89–118. https://doi.org/10.1146/annurev.arplant.52.1.89
Mok MC, Mok DWS, Armstrong DJ et al (1982) Cytokinin activity of N-phenyl-N′-1,2,3-thiadiazol-5-ylurea (thidiazuron). Phytochemistry 21:1509–1511. https://doi.org/10.1016/S0031-9422(82)85007-3
Müller B, Sheen J (2008) Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 453:1094–1097. https://doi.org/10.1038/nature06943
Nishimura C, Ohashi Y, Sato S et al (2004) Histidine kinase homologs that act as cytokinin receptors possess overlapping functions in the regulation of shoot and root growth in Arabidopsis. Plant Cell 16:1365–1377. https://doi.org/10.1105/tpc.021477
Nisler J, Kopečný D, Končitíková R et al (2016) Novel thidiazuron-derived inhibitors of cytokinin oxidase/dehydrogenase. Plant Mol Biol 92:235–248. https://doi.org/10.1007/s11103-016-0509-0
Notredame C, Higgins DG, Heringa J (2000) T-Coffee: a novel method for fast and accurate multiple sequence alignment. J Mol Biol 302:205–217. https://doi.org/10.1006/jmbi.2000.4042
Novák O, Hauserová E, Amakorová P et al (2008) Cytokinin profiling in plant tissues using ultra-performance liquid chromatography–electrospray tandem mass spectrometry. Phytochemistry 69:2214–2224. https://doi.org/10.1016/j.phytochem.2008.04.022
Pils B, Heyl A (2009) Unraveling the evolution of cytokinin signaling. Plant Physiol 151:782–791. https://doi.org/10.1104/pp.109.139188
Ramírez-Carvajal GA, Morse AM, Davis JM (2008) Transcript profiles of the cytokinin response regulator gene family in Populus imply diverse roles in plant development. New Phytol 177:77–89. https://doi.org/10.1111/j.1469-8137.2007.02240.x
Rashotte AM, Carson SD, To JP, Kieber JJ (2003) Expression profiling of cytokinin action in Arabidopsis. Plant Physiol 132:1998–2011. https://doi.org/10.1104/pp.103.021436
Riefler M, Novak O, Strnad M, Schmülling T (2006) Arabidopsis cytokinin receptor mutants reveal functions in shoot growth, leaf senescence, seed size, germination, root development, and cytokinin metabolism. Plant Cell 18:40–54. https://doi.org/10.1105/tpc.105.037796
Rittenberg D, Foster GL (1940) A new procedure for quantitative analysis by isotope dilution, with application to the determination of amino acids and fatty acids. J Biol Chem 133:737–744
Romanov GA, Spíchal L, Lomin SN et al (2005) A live cell hormone-binding assay on transgenic bacteria expressing a eukaryotic receptor protein. Anal Biochem 347:129–134. https://doi.org/10.1016/j.ab.2005.09.012
Romanov GA, Lomin SN, Schmülling T (2006) Biochemical characteristics and ligand-binding properties of Arabidopsis cytokinin receptor AHK3 compared to CRE1/AHK4 as revealed by a direct binding assay. J Exp Bot 57:4051–4058. https://doi.org/10.1093/jxb/erl179
Scatchard G (1949) The attractions of proteins for small molecules and ions. Ann NY Acad Sci 51:660–672. https://doi.org/10.1111/j.1749-6632.1949.tb27297.x
Schoor S, Farrow S, Blaschke H et al (2011) Adenosine kinase contributes to cytokinin interconversion in Arabidopsis. Plant Physiol 157:659–672. https://doi.org/10.1104/pp.111.181560
Sorrentino G, Haworth M, Wahbi S et al (2016) Abscisic acid induces rapid reductions in mesophyll conductance to carbon dioxide. PLoS One 11:e0148554. https://doi.org/10.1371/journal.pone.0148554
Spíchal L (2011) Bacterial assay to study plant sensor histidine kinases. In: Dissmeyer N, Schnittger A (eds) Plant kinases. Humana Press, Totowa, pp 139–147. https://doi.org/10.1007/978-1-61779-264-9
Spíchal L, Rakova NY, Riefler M et al (2004) Two cytokinin receptors of Arabidopsis thaliana, CRE1/AHK4 and AHK3, differ in their ligand specificity in a bacterial assay. Plant Cell Physiol 45:1299–1305. https://doi.org/10.1093/pcp/pch132
Stolz A, Riefler M, Lomin SN et al (2011) The specificity of cytokinin signalling in Arabidopsis thaliana is mediated by differing ligand affinities and expression profiles of the receptors. Plant J 67:157–168. https://doi.org/10.1111/j.1365-313X.2011.04584.x
Strnad M (1997) The aromatic cytokinins. Physiol Plant 101:674–688. https://doi.org/10.1111/j.1399-3054.1997.tb01052.x
Strnad M, Peters W, Beck E, Kamínek M (1992) Immunodetection and identification of N6-(o-hydroxybenzylamino)purine as a naturally cccurring cytokinin in Populus x canadensis Moench cv Robusta leaves. Plant Physiol 99:74–80
Strnad M, Peters W, Hanuš J, Beck E (1994) Ortho-topolin-9-glucoside, an aromatic cytokinin from Populus x canadensis cv Robusta leaves. Phytochemistry 37:1059–1062. https://doi.org/10.1016/S0031-9422(00)89528-X
Strnad M, Hanus J, Vanek T et al (1997) Meta-topolin, a highly active aromatic cytokinin from poplar leaves (Populus x canadensis moench., cv. Robusta). Phytochemistry 45:213–218. https://doi.org/10.1016/S0031-9422(96)00816-3
Suzuki T, Miwa K, Ishikawa K et al (2001) The Arabidopsis sensor His-kinase, AHK4, can respond to cytokinins. Plant Cell Physiol 42:107–113. https://doi.org/10.1093/pcp/pce037
Takeda S, Fujisawa Y, Matsubara M et al (2001) A novel feature of the multistep phosphorelay in Escherichia coli: a revised model of the RcsC → YojN → RcsB signalling pathway implicated in capsular synthesis and swarming behaviour. Mol Microbiol 40:440–450. https://doi.org/10.1046/j.1365-2958.2001.02393.x
Tirichine L, Sandal N, Madsen LH et al (2007) A gain-of-function mutation in a cytokinin receptor triggers spontaneous root nodule organogenesis. Science 315:104–107. https://doi.org/10.1126/science.1132397
To JP, Haberer G, Ferreira FJ et al (2004) Type-A Arabidopsis response regulators are partially redundant negative regulators of cytokinin signaling. Plant Cell 16:658–671. https://doi.org/10.1105/tpc.018978
Tran LSP, Urao T, Qin F et al (2007) Functional analysis of AHK1/ATHK1 and cytokinin receptor histidine kinases in response to abscisic acid, drought, and salt stress in Arabidopsis. Proc Natl Acad Sci USA 104:20623–20628. https://doi.org/10.1073/pnas.0706547105
Tuskan GA, DiFazio S, Jansson S et al (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604. https://doi.org/10.1126/science.1128691
Ueguchi C, Koizumi H, Suzuki T, Mizuno T (2001) Novel family of sensor histidine kinase genes in Arabidopsis thaliana. Plant Cell Physiol 42:231–235. https://doi.org/10.1093/pcp/pce015
Wang F-F, Cheng S-T, Wu Y et al (2017) A bacterial receptor PcrK senses the plant hormone cytokinin to promote adaptation to oxidative stress. Cell Rep 21:2940–2951. https://doi.org/10.1016/j.celrep.2017.11.017
Waterhouse A, Bertoni M, Bienert S et al (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucl Acids Res 46:W296–W303. https://doi.org/10.1093/nar/gky427
Werner T, Schmülling T (2009) Cytokinin action in plant development. Curr Opin Plant Biol 12:527–538. https://doi.org/10.1016/j.pbi.2009.07.002
Wulfetange K, Lomin SN, Romanov GA et al (2011) The cytokinin receptors of Arabidopsis are located mainly to the endoplasmic reticulum. Plant Physiol 156:1808–1818. https://doi.org/10.1104/pp.111.180539
Yamada H, Suzuki T, Terada K et al (2001) The Arabidopsis AHK4 histidine kinase is a cytokinin-binding receptor that transduces cytokinin signals across the membrane. Plant Cell Physiol 42:1017–1023
Yonekura-Sakakibara K, Kojima M, Yamaya T, Sakakibara H (2004) Molecular characterization of cytokinin-responsive histidine kinases in maize. Differential ligand preferences and response to cis-zeatin. Plant Physiol 134:1654–1661. https://doi.org/10.1104/pp.103.037176
Acknowledgements
This work was supported by the grant 15-16888S from the Czech Science Foundation, institutional support MZE-RO0418 and IGA_PrF_2018_033 (Palacký University, Olomouc, CZ). DK and PT were supported also from ERDF project “Plants as a tool for sustainable global development” (No. CZ.02.1.01/0.0/0.0/16_019/0000827). We acknowledge Zuzana Pěkná and Lukáš Spíchal for their help with the live cell competitive binding assay (Palacký University, Olomouc, CZ) and prof. Peter Hedden for proofreading the manuscript. Poplar calli were derived from dormant leaf buds provided by Dr. Jana Malá from Forestry and Game Management Research Institute (Jíloviště, CZ). The pINIIIΔEH vector and E. coli strain KMI001 were kindly provided by Dr. Zalabák (Palacký University, Olomouc, CZ). The authors are grateful to Sees-editing Ltd. (UK) for editing the manuscript. Populus deltoides sequence data were produced by the US Department of Energy Joint Genome Institute http://www.jgi.doe.gov.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest with the contents of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jaworek, P., Tarkowski, P., Hluska, T. et al. Characterization of five CHASE-containing histidine kinase receptors from Populus × canadensis cv. Robusta sensing isoprenoid and aromatic cytokinins. Planta 251, 1 (2020). https://doi.org/10.1007/s00425-019-03297-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-019-03297-x