Abstract
Main conclusion
We describe a Nicotiana benthamiana system for rapid identification of artificial microRNA (amiRNA) to control cucumber green mottle mosaic virus (CGMMV) disease.
Abstract
Although artificial miRNA technology has been used to control other viral diseases, it has not been applied to reduce severe cucumber green mottle mosaic virus (CGMMV) disease and crop loss in the economically important cucurbits. We used our system to identify three amiRNAs targeting CGMMV RNA (amiR1-CP, amiR4-MP and amiR6-Rep) and show that their expression reduces CGMMV replication and disease in virus-infected plants. This work streamlines the process of generating amiRNA virus-resistant crops and can be broadly applied to identify active antiviral amiRNAs against a broad spectrum of viruses to control disease in diverse crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cucumber green mottle mosaic virus (CGMMV) is a member of the Tobamovirus genus. The sequence of the genomic RNA of CGMMV is 6421 nucleotides long containing at least four open reading frames (ORFs) (Ugaki et al. 1991). The first two ORFs encode the 129 kDa pre-read through and 186 kDa read through proteins. ORF 3 encodes the 29-kDa protein involved in cell to cell movement (Movement protein, MP) and ORF 4 encodes the 17.4 kDa (Coat protein, CP) (Liu et al. 2009). CGMMV is transmitted by mechanical damage, infested soil, contaminated seeds, pollen and plant propagation stocks (Hollings et al. 1975; Choi et al. 2004; Liu et al. 2014; Dombrovsky et al. 2017). Infection by CGMMV results in characteristic mosaic symptoms in leaves of cucurbit plants. Severe infections can cause stunted growth and distorted fruits, imposing severe economic losses on cucurbit crop production (Komuro 1971; Shim et al. 2005). CGMMV has been reported in 43 countries, including UK (Ainsworth 1935), Israel (Antignus et al. 1990), China (Zhang et al. 2009) and United States (Tian et al. 2014; Baker 2016). Seed disinfection has been shown to be insufficient to eliminate CGMMV from stocks (Reingold et al. 2015) and control of CGMMV is dependent on using virus-free propagation materials. Although using virus-tolerant or -resistant cultivars is the desired strategy for CGMMV control, CGMMV-resistant germplasm is limited.
In recent years, microRNA (miRNA)-mediated RNA silencing has been applied to control viral diseases (Khraiwesh et al. 2012). MiRNAs are a class of small non-coding RNAs of 21–24 nucleotides in length that silence gene expression by targeting mRNAs for degradation or translational repression at the post-transcriptional level in eukaryotes (Bartel 2004; Chen 2009; Rogers and Chen 2013). MiRNAs play important roles in regulating genes in plant development (Chen 2009) and response to biotic or abiotic stresses (Sunkar et al. 2006; Xin et al. 2010). Artificial miRNAs (amiRNAs) are produced by replacing the original miRNA:miRNA* duplex region with designed sequences to target and silence individual gene or group of endogenous genes of interest in plants (Schwab et al. 2006; Ossowski et al. 2008). amiRNA technology shows several advantages in comparison with RNA interference (RNAi) and virus-induced gene silencing (VIGS) for engineering virus resistance including the introduction of multiple amiRNAs targeting different virus genes or virus strains (Ai et al. 2011; Sablok et al. 2011; Fahim et al. 2012; Kung et al. 2012; Lafforgue et al. 2013; Tiwari et al. 2014; Kis et al. 2016). Because the amiRNA sequence does not have to be perfectly complementary to the target site, it can be optimized to target only one or, alternatively, several sequence-related genes. Genome-wide expression analyses have shown that amiRNAs have similarly high specificity as endogenous miRNAs that facilitate to assess and hence reduce the potential off-target effects of the expressed small RNA in the donor plant (Schwab et al. 2005, 2006; Kis et al. 2016). Efficient amiRNA-mediated gene silencing has been observed to occur in a quantitative fashion, with stronger promoters often causing higher degrees of gene silencing and it seems that there are few, if any non-autonomous effects (Alvarez et al. 2006; Schwab et al. 2006; Ossowski et al. 2008). In addition, the amiRNA technology offers a way for time-efficient modification of the expression of such genes in any variety. This enables not only rapid knowledge transfer between different varieties, but also the introduction of important traits for improving agronomic performance and/or nutritional value into a broad range of varieties (Warthmann et al. 2008). amiRNAs might also have potential advantages for crop plants, as a single species of sRNA is preferentially generated, the actions of which are much more predictable than those of the collection of small RNAs with diverse sequences produced by hairpin RNAi (hpRNAi) constructs (Ossowski et al. 2008). This property may also help to alleviate regulatory concerns.
Niu et al. (2006) have shown that amiRNA technology can be successfully applied to enhance resistance of Arabidopsis thaliana to turnip yellow mosaic virus and turnip mosaic virus, although other studies showed a small proportion of virus “escape” under amiRNA selection pressure (Lafforgue et al. 2011; Lin et al. 2009; Simon-Mateo and Garcia 2006). Wagaba et al. (2016) have also demonstrated that transgenic plants with amiRNAs enhanced the resistance of Nicotiana benthamiana against cassava brown streak virus and Ugandan cassava brown streak virus. To reduce the risk of viral escape from amiRNA-mediated targeting, several research groups have proposed and used multiple amiRNAs targeting different conserved sequences within viral genomes (Fahim et al. 2012; Kis et al. 2016; Kung et al. 2012; Lafforgue et al. 2013).
AmiRNA-mediated viral resistance has now been successfully deployed in plant to control disease caused by infection of potato virus Y and potato virus X (Ai et al. 2011), cucumber mosaic virus (Zhang et al. 2011), tomato leaf curl New Delhi virus (Vu et al. 2013), wheat dwarf virus (Kis et al. 2016), barley stripe mosaic virus (Jian et al. 2017), cymbidium mosaic virus and odontoglossum ringspot virus (Petchthai et al. 2018), and tomato spotted wilt virus (Carbonell et al. 2019). However, there have been no reports describing amiRNA-mediated CGMMV resistance in model or crop species.
The objective of this study was to develop an effective amiRNA-mediated gene silencing approach using a N. benthamiana–CGMMV pathosystem. Our experiments revealed that amiRNA technology can control the replication of CGMMV in N. benthamiana. This work streamlines the process of generating amiRNA virus-resistant crops and can be broadly applied to identify active antiviral amiRNAs against a broad spectrum of viruses to control disease in diverse crops.
Materials and methods
Plant material
Nicotiana benthamiana reference Nb-1 genotype was used for Agrobacterium infiltration experiments and obtained from the Boyce Thompson Institute. Plants were grown in potting soil in a growth chamber (Versatile Environmental Test Chambers; Sanyo, Tokyo, Japan) under a daily cycle with 16-h light at 24 °C and 8-h dark at 22 °C.
AmiRNA design and cloning, and plant expression vector construction
Active amiRNAs were designed and screened based on the criteria and procedures described by Kis et al. (2016). BLAST (https://www.ncbi.nlm.nih.gov/) was performed to identify conserved coat protein (CP), movement protein (MP) and replicase (Rep) gene sequences of CGMMV GenBank accessions AJ245440.1, AJ243353.1 and AY866427.1. AmiRNAs of 21 nucleotides or more in length were designed according to the WMD3 tool (http://wmd3.weigelworld.org/). AmiRNAs were designed to contain nucleotides A or U at position 1 (5′ terminus), A at position 10 (from the 5′ terminus) and G or C at position 21 (3′ terminus). CGMMV-specific amiRNAs with the lowest free energy (the highest stability) were selected having the least probability of targeting any sequence in the cucumber (Cucumis sativus) or N. benthamiana genome. The positions of amiRNA-target sequences in the conserved regions of CP, MP and Rep in CGMMV genome are shown in Fig. S1.
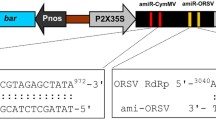
Three A. thaliana precursor miRNA backbones (ath-miR156, ath-miR164 and ath-miR171) were used to generate artificial miRNA precursors (amiRNA precursors) targeting different conserved regions in the CGMMV genome (Fig. 1b). This strategy employed an overlapping PCR approach, according to Li et al. (2012), using overlapping PCR primers (Table S1). The amiRNA precursor sequences obtained from overlapping PCR were cloned into pENTR™/D-TOPO® (Invitrogen, USA) and the clones were confirmed by sequencing. Next, the confirmed sequences were recombined into the binary expression vector pEarlyGate100 (pEG100) by the LR reaction. Expression of all pre-amiRNAs was driven by the cauliflower mosaic virus (CaMV) 35S promoter and terminated with the OCS terminator. Secondary structures of designed pre-amiRNAs were predicted by the Mfold Web Server (http://mfold.rna.albany.edu/?q=mfold/RNA-Folding-Form). The resulting recombinant binary expression plasmids containing the amiRNA precursor were designated pEG100.aMIR1, pEG100.aMIR2, pEG100.aMIR3, pEG100.aMIR4, pEG100.aMIR5 and pEG100.aMIR6 (Fig. S2).
Schematic diagram of amiRNA precursor design and construction for silencing coat protein (CP), movement protein (MP) and replicase (Rep) genes of CGMMV. a Target sites of the amiRNAs relative to positions in CGMMV genome. b Secondary structures of the Arabidopsis miRNA precursors. Structure of ath-miR156 precursor (176 bp), ath-miR164 precursor (176 bp) and ath-miR171 precursor (176 bp), which is presented as a hairpin. c Overlapping polymerase chain reaction (PCR) technique to for amiRNA precursor synthesis. In the first round PCR reaction, intermediates I-12 was synthesized using Primers 1 and 2, and I-345 was synthesized using Primers 3, 4 and 5 in separate reactions (① and ②). In the second round of PCR reaction (③), I-12 and I-345 were used as template and Primers 1 and 5 were used to synthesize amiRNA precursor sequence which contained the amiRNAs from the CGMMV genome. d Original miRNA/miRNA* duplex (highlighted in red) was replaced with the designed amiRNA*/amiRNA sequences (blue) in overlapping PCR
Agrobacterium tumefaciens infiltration
Agrobacterium tumefaciens GV3101 was transformed with the amiRNA precursor (pre-amiRNA) transient expression vectors. GV3101 carrying the above pre-amiRNA expression vectors or control pEarlyGate100 empty vector (EV) were infiltrated into N. benthamiana (two leaves per plant) using a protocol described previously (Li et al. 2012). Briefly, GV3010 was grown to an optical density of 1.0 at 600 nm (OD600) and diluted to an OD600nm of 0.2 and infiltrated into young leaves of 5–6-week-old N. benthamiana. Agro-infiltrations are summarized in Fig. 2a.
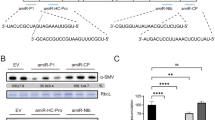
Expression of six amiRNAs at 3 days post agroinfiltration (dpa) in Nicotiana benthamiana plants (from #1 to #7). Sample #8 was infiltrated with an empty vector (EV). a Infiltrated samples from #1 to #6 each were agroinfiltrated with one of the six amiRNAs whilst infiltrated sample #7 was co-agroinflitrated with all six amiRNAs (top panel); Northern blot hybridization of RNA isolated from infiltrated N. benthamiana plants (bottom panel). b Expression of six amiRNAs in infiltrated N. benthamiana plants detected by qRT-PCR. Each RT-qPCR reaction was performed in three biological replicates and three technical replicates for each sample. Error bars represent the mean of three biological replicates ± SD
Virus inoculation
The CGMMV strain No. 2 was collected from Zhejiang Province of China and its identity was confirmed using reverse transcription-polymerase chain reaction (RT-PCR) (Liang et al. 2015). CGMMV strain No. 2 was maintained and propagated by transfer of virus from infected leaves to uninfected cucumber plants. The viral inoculum was prepared by grinding CGMMV-infected cucumber leaves in a mortar with a pestle at a 1:4 w/v ratio in 20-mM sodium phosphate buffer (pH 7.5). For CGMMV inoculation, agro-infiltrated leaves of previously infiltrated N. benthamiana plants at 3 days post agro-infiltration (dpa) were gently rubbed using fingers (with 50-μL CGMMV sap per leaf). Leaf tissues of CGMMV-infected and agro-infiltrated N. benthamiana plants were collected at 3, 10 and 15 days post virus inoculation (dpi) and stored at − 80 °C for later use (Table S2). Control N. benthamiana plants were mock-inoculated with inoculation buffer (20-mM sodium phosphate buffer, pH 7.5) alone. CGMMV-infected plants were scored for virus disease symptoms at 3 dpi, 10 dpi, 15 dpi and 20 dpi.
RNA isolation and cDNA synthesis
Total RNA was extracted from agro-infiltrated N. benthamiana plants at 3 dpa and from CGMMV-infected and agro-infiltrated N. benthamiana plants at 3, 10 and 15 dpi, respectively, using the TRIzol® reagent (Invitrogen, USA) following the manufacturer’s protocol. For amiRNA expression analysis, 0.4 μg of total RNA from agro-infiltrated N. benthamiana leaves was reverse-transcribed into cDNA using the miScript Reverse Transcription Kit (Qiagen, Germany). For analysis of CGMMV RNA, reverse transcription was performed using RNA isolated from leaves of virus-infected or mock-inoculated plants and the Superscript III™ First-Strand Synthesis System (Invitrogen, USA) following the manufacturers protocol after treatment with DNase I (Promega, USA).
Northern blot hybridization analysis
For Northern blotting, 30 μg of total N. benthamiana RNA was heat-treated in a formamide buffer and loaded on a 12% denaturing urea-PAGE gel. Subsequently, RNA samples were transferred to a Hybond-N+ membrane (GE Healthcare Life Sciences, UK) and hybridized with probes for each of the six amiRNAs (Table S3). Hybridization was carried out using a standard protocol (Li et al. 2012). The probes were end labeled using [γ-32P] ATP (PerkinElmer Life Sciences, USA) and purified in an illustra MicroSpin G-25 column (GE Healthcare Life Sciences, UK) according to the supplier’s protocol. The membranes were incubated overnight at 42 °C and then subjected to autoradiography using a TYPHOON phosphor imager (GE Healthcare Life Sciences, UK). The relative intensity of amiRNA band was quantified using Quantity One Basic 4.6.6 (Bio-Rad).
Quantitative real-time PCR (qRT-PCR)
qRT-PCR was performed to measure amiRNA expression using miScript SYBR® Green PCR Kit (Qiagen, Germany) on a Bio-Rad CFX384 instrument (BioRad Laboratories Inc, USA), with designed amiRNA primers (Table S4). A 25-μL reaction mix was prepared using 12.5 μL of 2× QuantiTect SYBR Green PCR Master Mix, 2.5 μL of 10× miScript Universal primer, 2.5 μL of 10× forward primers (amiRNA primers), 6.5 μL of nuclease-free water and 1 μL of RT cDNA product. The thermal cycler was set to the following conditions: initial activation step at 95 °C for 15 min, followed by 40 cycles of 94 °C for 15 s, 55 °C for 30 s and 72 °C for 30 s. The relative expression levels of CP, MP and Rep genes of CGMMV were determined using specific primers (Table S5). The PCR itself was conducted using a 20-μL reaction mix containing 4 μL 5× GoTaq Flexi buffer (Promega, USA), 1.6 μL 25 mM MgCl2+, 2 μL 10× tween-DMSO mix, 1 μL 20× EvaGreen Dye (Biotium, USA), 0.4 μL 10 mM dNTP mix, 0.1 μL GoTaq Flexi (5 U/μL) (Promega, USA), 0.3 μL each of the forward and reverse primers (10 μM), 2 μL cDNA and 8.3 μL RNase-free water, and the following program 95 °C for 5 min, followed by 40 cycles of 95 °C for 15 s, 55 °C for 30 s, and 72 °C for 20 s. Each sample was a composite of leaves from two plants, which was sub-grouped into three technical replicates for qRT-PCR analysis (Table S2). The mean quantification cycle value was used for calculations by the 2−ΔΔCt method (Livak and Schmittgen 2011) for relative normalized expression analysis using 60S ribosomal protein L25 from N. benthamiana as reference gene.
Determination of CGMMV resistance in agro-infiltrated and CGMMV-infected N. benthamiana plants
To determine virus resistance/tolerance in plants with introduced amiRNAs, CGMMV RNA accumulation and virus disease symptoms were determined for CGMMV-infected N. benthamiana plants at 3, 10, 15 and 20 dpi. The resistance ratio based on relative CGMMV RNA accumulation was determined as: [CGMMV gene expression in control empty vector (EV) infiltrated sample#8—CGMMV gene expression in amiRNA infiltrated samples (#1–#7)]/CGMMV gene expression in control empty vector (EV) infiltrated sample #8. In addition, levels of viral resistance were determined based on symptom severity.
Results
Design of amiRNAs targeting CGMMV and construction of vectors
Vectors expressing amiRNAs targeting specific sequences of CP, MP and Rep genes of CGMMV were designed and constructed as described in Fig. 1 and Fig. S1. Six amiRNA precursors were generated by replacing the original miRNA/miRNA* duplex of ath-miR156, ath-miR164 and ath-miR171 backbones (Fig. 1) with expected precursor amiRNAs (pre-amiRNAs) of 168 bp, 159 bp and 163 bp long, respectively (Fig. S3). Six amiRNA plant expression vectors (pEG100.aMIR1 to pEG100.aMIR6) were obtained by oligonucleotide-directed mutagenesis (Fig. S2). Computational prediction of the secondary structure of the amiRNA precursors indicated that the six amiRNAs possessed correct folding parameters and correctly folded (Fig. S4).
Expression of amiRNAs in infiltrated N. benthamiana plants
To determine if amiRNAs were expressed from the six different pre-miRNA expression vectors, each construction was transformed into GV3101 and infiltrated into N. benthamiana plants. The level of transiently expressed amiRNAs was determined by Northern blot hybridization (Fig. 2a) and qRT-PCR analysis of RNA isolated from agro-infiltrated N. benthamiana plants (Fig. 2b). Both Northern blot and qRT-PCR results showed some inconsistent trends in infiltrated one amiRNA or co-infiltrated six amiRNAs N. benthamiana plants. For instance, the level of amiR6-Rep was much lower in infiltrated sample #7 than sample #6 (Fig. 2a), whilst in qRT-PCR assays, amiR6-Rep accumulated to a similar amount in both infiltrated samples (Fig. 2b). The expression level of amiRNA was highest for amiR4-MP, whilst amiR2-CP, amiR3-MP and amiR5-Rep were expressed at lower levels in Northern blot and qRT-PCR results (Fig. 2a, b). The intensity of bands observed in Northern blot hybridization analysis indicated that plants expressing the amiR4-MP construct exhibited the highest level of amiR4-MP expression (Fig. S5), whereas expression of amiR2-CP in infiltrated sample #7 was not detected by Northern blot, but detected by qRT-PCR. Overall, five out of six amiRNAs were processed to the correct size amiRNA (21-nt) and expressed at detectable levels.
Meanwhile, we tested amiRNA expression in CGMMV-infected N. benthamiana plants at 3, 10 and 15 dpi. In general, the level of amiRNA at 10 dpi was slightly higher than at 3 dpi, but quickly dropped at 15 dpi. In addition, amiR4-MP had the highest expression level but amiR2-CP had the lowest (Fig. 3).
Expression of six amiRNAs in infiltrated N. benthamiana plants (from #1 to #7) inoculated with CGMMV at 3, 10, and 15 days post inoculation (dpi). Each RT-qPCR reaction was performed in three biological replicates and three technical replicates for each sample. Error bars represent the mean of three biological replicates ± SD
Correlation between amiRNA expression and CGMMV RNA levels
To determine if the expression of amiRNAs targeting different CGMMV gene sequences affected viral RNA accumulation and virus spread, CGMMV-infection assays were carried out with amiRNA-mediated N. benthamiana-CGMMV pathosystem. The expression levels of CP, MP and Rep genes of CGMMV in infiltrated samples #1, #4 and #6 were lower compared with infiltrated samples #2, #3 and #5 both at 3, 10 and 15 dpi, respectively (Fig. 4). We found that CGMMV RNA levels were negatively correlated with amiRNA expression levels (Figs. 3, 4). Namely, the higher amiRNA expression level, the lower the CGMMV RNA accumulation level.
Accumulation level of CGMMV in infiltrated N. benthamiana plants (from #1 to #8) at 3, 10, and 15 days post inoculation (dpi). The expression level of genes for a coat protein (CP), b movement protein (MP), and c replicase gene (Rep). Each RT-qPCR reaction was performed in three biological replicates and three technical replicates for each sample. Error bars represent the mean of three biological replicates ± SD
Correlation between amiRNA expression and CGMMV resistance
Infiltrated samples #4, #6 and #7 conferred high resistance against CGMMV (resistance ratio > 96%), and sample #1 provided complete resistance (resistance ratio was 100%) (Table 1). The degree of tolerance/resistance was positively correlated with amiRNA expression levels in these single (samples #1 to #6) and all amiRNAs co-infiltrated sample #7 (Fig. 3, Table 1). Overall, agro-infiltrated N. benthamiana plant transformed with amiRNA targeting CGMMV CP gene showed the highest viral resistance with reduced viral accumulation; whereas, plant transformed with amiRNA targeting Rep exhibited only a moderate tolerance to CGMMV infection, plant transformed with amiRNA targeting MP induced the lowest viral resistance.
Resistance phenotyping of agro-infiltrated N. benthamiana plants following CGMMV challenge
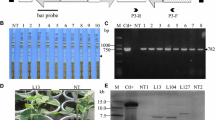
No CGMMV-induced disease symptoms were observed at 20 dpi in individual amiRNA infiltrated samples #1, #4 or #6 expressing amiR1-CP, amiR4-MP or amiR6-Rep and co-infiltrated six amiRNAs sample #7 (Fig. 5b). For samples #2, #3 and #5, viral symptoms started showing up at 15 dpi as virus reached to a high level at 10 and 15 dpi (Fig. 4) and they exhibited a weak tolerance and some mottle were found on the systemic leaves at 20 dpi (Fig. 5b). Sample #8 (infiltrated empty vector) showed typical and severe viral symptoms and a high level of virus RNA accumulation at 10, 15 and 20 dpi (Figs. 4, 5b). Meanwhile, mock-inoculated N. benthamiana was healthy and did not show any symptoms, whereas wild-type N. benthamiana plants inoculated with CGMMV showed characteristic mottle and mosaic symptoms on the systemic leaves at 20 dpi (Fig. 5a).
Determination of viral resistance levels. a Leaves of wild-type (WT) N. benthamiana plants either mock-inoculated (the first panel from left) or cucumber green mottle mosaic virus (CGMMV)-inoculated (the 2nd, 3rd and 4th panels from left) at 20 days post inoculation (dpi). b Systemic symptoms on leaves of infiltrated N. benthamiana plants at 20 days post inoculation (dpi) with CGMMV. Red arrows show typical symptoms of viral infection. Mock-inoculated = wild-type plant inoculated with 20-mM sodium phosphate buffer
Discussion
We designed six amiRNAs that targeted different gene transcripts of CGMMV for gene silencing, and have successfully applied them to N. benthamiana enhance resistance to CGMMV. Theoretically, this system is applicable to different viruses and hosts. As such, the success in N. benthamiana can be transferred to cucumber. Therefore, it can be a potential and practical tool in controlling CGMMV in cucumber. Until now, this study is the first report describing amiRNA-mediated resistance against CGMMV belonging to the Tobamovirus.
The mechanism of miRNAs silencing the target transcripts is related to the degree of their complementarity. Briefly, high levels of complementarity lead to the silencing of target transcripts and their degradation, and low levels of complementarity prevent mRNA from being translated (Vu et al. 2013). In plants, miRNAs usually have near-perfect pairing with their mRNA targets, which induces gene repression through cleavage of the target transcripts (Jones-Rhoades et al. 2006). In this study, there were only one or two mismatched bases between designed amiRNAs and their target sequences (Figs. 1, S1), the targets (virus RNA) were expected to be degraded through the silencing mechanism.
The expression of amiRNAs varies depending on diverse precursor backbones and transformation events (Schwab et al. 2006). In spite of the careful design, the in vivo expression and efficiency of amiRNA candidates can be highly variable (Deveson et al. 2013; Li et al. 2013, 2014). For example, eight (amiR1-8) and two (amiR9-10) amiRNAs were designed to target wheat dwarf virus encoding Rep and/or RepA proteins (C1 and/or C2 genes) and movement protein (MP) (V1 gene), respectively (Kis et al. 2016). The expression levels of amiR3, amiR8, amiR9 and amiR10 were high compared with the moderate accumulation of amiR1, amiR2 and amiR6, while the expression of amiR4, amiR5 and amiR7 could not be detected in the transient assay (Kis et al. 2016). Vu et al. (2013) found that some transgenic plants that contained the amiRNA transgenes could not express mature amiRNAs and that some plants expressed lower levels of the amiRNAs than others are not surprising. In this study, we also found that the expression level of amiRNA was highest for amiR4-MP, whilst amiR2-CP, amiR3-MP and amiR5-Rep were expressed at lower levels. There are several possible explanations for variability of amiRNA expression. Expression levels of mature amiRNAs are influenced by miRNA precursor backbone, complementarity of amiRNA-target sequence, selection of target site and/or the free energy of amiRNA precursor (stem-loop stability) (Ai et al. 2011). Hence, the observations that sample #2 that contained amiR2-CP transgenes expressed lower level of amiR2-CP than others were not surprising.
A high level of virus-specific amiRNA seems to be a crucial determinant for viral resistance level in plants, which has been demonstrated in several studies (Qu et al. 2007; Song et al. 2014; Kis et al. 2016). We found that the accumulation level of amiRNA in resistant infiltrated samples was higher than that in susceptible samples. Wagaba et al. (2016) have also demonstrated that levels of resistance to cassava brown streak disease (CBSD) were correlated with accumulation of detectable amiRNA in N. benthamiana. In addition, there were various degrees of virus RNA degradation in different infiltrated plants. These results again demonstrated that overexpression of amiRNAs led to the silencing and degradation of their targets. In general, CGMMV RNA levels were closely related to the levels of amiRNA expression. However, we don’t deny that other events or pathways might play roles in infiltrated plant resistant to CGMMV infection and accumulation of amiRNAs may be not the only determinant of resistance efficiency.
In infiltrated N. benthamiana plants when challenged with CGMMV, the amount of viral RNA varied. We expected to have a higher resistance by transferring multiple amiRNAs targeting different genes and gene locations. This was demonstrated in most cases, but there were some exceptions. Kung et al. (2012) also observed such a variation. They designed multiple amiRNAs targeting conserved motifs of the L (replicase) gene of watermelon silver mottle virus (WSMoV) using Arabidopsis pre-miRNA159a as the backbone. Single amiRNA transgenic lines showed tolerance to WSMoV by delaying symptom expression. Triple amiRNA lines provided complete resistance, with no symptoms observed at 28 days after inoculation, but the triple amiR-LAB1E line did not provide the same level of resistance (Kung et al. 2012). In our study, resistance level from infiltrated sample #7 expressing six amiRNAs exhibited only some additive effects, probably because the co-infiltrated multiple amiRNA constructs were unfavorable for amiRNA-mediated silencing. This may have to do with steric hindrance, for example, secondary structures of pre-amiRNAs causing physical-spatial constraints to the processing of amiRNA cleaving target mRNA.
We found that virus resistance levels were affected by functional genes of virus to be targeted by amiRNAs or the region of the same functional gene that were targeted. For example, the resistance level was higher when the 5′ end of the Rep gene was targeted by amiR6-Rep; the resistance level was lower when the middle segment of the MP gene was targeted. In fact, there is no equal efficiency for multiple small interference RNAs (siRNAs) against a given mRNA target (Kung et al. 2012). The efficacy of RNA silencing is determined by multiple factors, such as the nature of small RNA sequence, the local structure of target mRNAs, and so on (Luo and Chang 2004; Schubert et al. 2005). It seems that amiRNAs targeting the CP gene of CGMMV induced the highest viral resistance. A higher level of resistance was observed when the 5′ end of CP gene was targeted by amiR1-CP; but the level of resistance was much lower when the 3′ end of the CP gene was targeted by amiR2-CP. However, this result is not consistent with other studies on amiRNA-mediated viral resistance (Jiang et al. 2011). The authors found an amiRNA targeting the 3′ end of CP gene of potato virus Y induced the highest viral resistance. Until we accumulate more evidences, a solid conclusion cannot be drawn.
Plant resistance may be enhanced by incorporating multiple amiRNAs targeting different viral genes. We took this advantage and generated CGMMV-tolerant plants harboring amiRNAs targeting the coat protein, movement protein and replicase genes of CGMMV. The result was promising as expected. This result makes possible of combating devastating effects of various tobamoviruses on diverse host plants.
Song et al. (2014) found that the silencing efficiency of small RNA can be influenced by their secondary structure. Thereinto, two of the determinants are the minimal free energy (∆G) and the number of free nucleotides at one end of the secondary structure (Luo and Chang 2004; Schubert et al. 2005). However, some scholars hold different views, they found no link between RNAi activity and ∆G (Zhang et al. 2013; Chan et al. 2009). In the present study, the predicted secondary structure of six amiRNAs (Table S6) showed a poor correlation (0.2911) between resistance level and ∆G of the amiRNAs. This did not support that the ∆G of the amiRNAs may directly correlate with RNA silencing efficiency. Similarly, we found that the presence of the 5′ or 3′ terminal free nucleotides within the amiRNA structures might not influence RNA silencing efficiency. This was in disagreement with Song et al. (2014).
In conclusion, we have developed a rapid method to identify active antiviral amiRNAs. Infiltrated N. benthamiana plants showed various levels of resistance against CGMMV, and the level of resistance was determined by the expression level of amiRNA. This is the first report describing amiRNA-mediated resistance against CGMMV. Our system can be targeting of different viruses or their strains simultaneously, and could be used not only in N. benthamiana but also in cucumber. We will develop transgenic cucumber using these amiRNA constructs in the future. As a consequence, application of this biotechnological approach may protect plants from disease in pathogen–host interactions and provide resistance sources for crop breeding.
Author contribution statement
CQ, LX and BB conceived and designed the experiments and outline of the article, analyzed data, composed the manuscript and figures. JJ and JQ provided scientific feedback and critical comments and revised the content. All authors read and approved the manuscript.
References
Ai T, Zhang L, Gao Z, Zhu CX, Guo X (2011) Highly efficient virus resistance mediated by artificial microRNAs that target the suppressor of PVX and PVY in plants. Plant Biol 13:304–316. https://doi.org/10.1111/j.1438-8677.2010.00374.x
Ainsworth GC (1935) Mosaic diseases of the cucumber. Ann Appl Biol 22:55–67. https://doi.org/10.1111/j.1744-7348.1935.tb07708.x/full
Alvarez JP, Pekker I, Goldshmidt A, Blum E, Amsellem Z, Eshed Y (2006) Endogenous and synthetic microRNAs stimulate simultaneous, efficient, and localized regulation of multiple targets in diverse species. Plant Cell 18:1134–1151. https://doi.org/10.1105/tpc.105.040725
Antignus Y, Pearlsman M, Ben-Yoseph R, Cohen S (1990) Occurrence of variant of Cucumber green mottle mosaic virus in Israel. Phytoparasitica 18:50–56. https://doi.org/10.1007/BF02980826
Baker C (2016) Cucumber green mottle mosaic virus (CGMMV) found in the United States (California) in melon. Pest alert, Florida Department of Agriculture and Consumer Services, Division of Plant Industry (DACS-P-01863)
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297. https://doi.org/10.1016/S0092-8674(04)00045-5
Carbonell A, López C, Daròs JA (2019) Fast-forward identification of highly effective artificial small RNAs against different Tomato spotted wilt virus isolates. Mol Plant-Microbe Interact 32:142–156. https://doi.org/10.1094/MPMI-05-18-0117-TA
Chan CY, Carmack CS, Long DD, Maliyekkel A, Shao Y, Roninson IB, Ding Y (2009) A structural interpretation of the effect of GC-content on efficiency of RNA interference. BMC Bioinform. https://doi.org/10.1186/1471-2105-10-S1-S33
Chen X (2009) Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25:21–44. https://doi.org/10.1146/annurev.cellbio.042308.113417
Choi GS, Kim JH, Kim JS (2004) Soil transmission of cucumber green mottle virus and its control measures in watermelon. Res Plant Dis 10:44–47. https://doi.org/10.5423/RPD.2004.10.1.044
Deveson I, Li JY, Millar AA (2013) MicroRNAs with analogous target complementarities perform with highly variable efficacies in Arabidopsis. FEBS Lett 587:3703–3708. https://doi.org/10.1016/j.febslet.2013.09.037
Dombrovsky A, Tran-Nguyen LTT, Jones RAC (2017) Cucumber green mottle mosaic virus: rapidly increasing global distribution, etiology, epidemiology, and management. Annu Rev Phytopathol 55:231–256. https://doi.org/10.1146/annurev-phyto-080516-035349
Fahim M, Millar AA, Wood CC, Larkin PJ (2012) Resistance to Wheat streak mosaic virus generated by expression of an artificial polycistronic microRNA in wheat. Plant Biotechnol J 10:150–163. https://doi.org/10.1111/j.1467-7652.2011.00647.x
Hollings M, Komuro Y, Tochihara H (1975) Cucumber green mottle mosaic virus. CMI/AAB Descriptions of Plant Viruses No. 154. Kew, United Kingdom
Jian C, Han R, Chi Q, Wang SJ, Ma M, Liu XL, Zhao HX (2017) Virus-based microRNA silencing and overexpressing in common wheat (Triticum aestivum L.). Front Plant Sci 8:500. https://doi.org/10.3389/fpls.2017.00500
Jiang F, Song YZ, Han QJ, Zhu CX, Wen FJ (2011) The choice of target site is crucial in artificial miRNA-mediated virus resistance in transgenic Nicotiana tabacum. Physiol Mol Plant Pathol 76:2–8. https://doi.org/10.1016/j.pmpp.2011.07.002
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAS and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53. https://doi.org/10.1146/annurev.arplant.57.032905.105218
Khraiwesh B, Zhu JK, Zhu J (2012) Role of miRNAs and siRNAs in biotic and abiotic stress responses of plants. Biochim Biophys Acta 1819:137–148. https://doi.org/10.1016/j.bbagrm.2011.05.001
Kis A, Tholt G, Ivanics M, Varallyay E, Jenes B, Havelda Z (2016) Polycistronic artificial miRNA-mediated resistance to wheat dwarf virus in barley is highly efficient at low temperature. Mol Plant Pathol 17:427–437. https://doi.org/10.1111/mpp.12291
Komuro Y (1971) Cucumber green mottle mosaic virus on cucumber and watermelon and melon necrotic spot virus on muskmelon. J Agric Res Quart 6:41–45
Kung YJ, Lin SS, Huang YL, Chen TC, Harish SS, Chua NH, Yeh SD (2012) Multiple artificial microRNAs targeting conserved motifs of the replicase gene confer robust transgenic resistance to negative-sense single-stranded RNA plant virus. Mol Plant Pathol 13:303–317. https://doi.org/10.1111/j.1364-3703.2011.00747.x
Lafforgue G, Martínez F, Sardanyés J, Iglesia F, Niu QW, Lin SS, Solé RV, Chua NH, Daròs JA, Elena SF (2011) Tempo and mode of plant RNA virus escape from RNA interference-mediated resistance. J Virol 85:9686–9695. https://doi.org/10.1128/JVI.05326-11
Lafforgue G, Martínez F, Niu QW, Chua NH, Daròs JA, Elena SF (2013) Improving the effectiveness of artificial microRNA (amiR)-mediated resistance against Turnip Mosaic Virus by combining two amiRs or by targeting highly conserved viral genomic regions. J Virol 87:8254–8256. https://doi.org/10.1128/JVI.00914-13
Li F, Pignatta D, Bendix C, Brunkard JO, Cohn MM, Tung J, Sun HY, Kumar P, Baker B (2012) MicroRNA regulation of plant innate immune receptors. Proc Natl Acad Sci USA 109:1790–1795. https://doi.org/10.1073/pnas.1118282109
Li JF, Chung HS, Niu YJ, Bush J, McCormack M, Sheen J (2013) Comprehensive protein-based artificial microRNA screens for effective gene silencing in plants. Plant Cell 25:1507–1522. https://doi.org/10.1105/tpc.113.112235
Li JF, Zhang DD, Sheen J (2014) Epitope-tagged protein-based artificial miRNA screens for optimized gene silencing in plants. Nat Protoc 9:939–949. https://doi.org/10.1038/nprot.2014.061
Liang CQ, Yan M, Luo LX, Liu PF, Li JQ (2015) Phylogenetic and bioinformatics analysis of replicase gene sequence of Cucumber green mottle mosaic virus. Chin J Virol 31:620–628. https://doi.org/10.13242/j.cnki.bingduxuebao.002821 (in Chinese)
Lin SS, Wu HW, Elena SF, Chen KC, Niu QW, Yeh SD, Chen CC, Chua NH (2009) Molecular evolution of a viral non-coding sequence under the selective pressure of amiRNA-mediated silencing. PLoS Pathog 5:e1000312. https://doi.org/10.1371/journal.ppat.1000312
Liu Y, Wang YN, Wang XF, Zhou GH (2009) Molecular characterization and distribution of Cucumber green mottle mosaic virus in China. J Phytopathol 157:393. https://doi.org/10.1111/j.1439-0434.2008.01509.x
Liu HW, Luo LX, Li JQ, Liu PF, Chen XY, Hao JJ (2014) Pollen and seed transmission of Cucumber green mottle mosaic virus in cucumber. Plant Pathol 63:72–77. https://doi.org/10.1111/ppa.12065
Livak KJ, Schmittgen TD (2011) Analysis of relative gene expression data using real-time quantitative PCR and the 2−∆∆CT method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Luo KQ, Chang DC (2004) The gene-silencing efficiency of siRNA is strongly dependent on the local structure of mRNA at the targeted region. Biochem Biophys Res Commun 318:303–310. https://doi.org/10.1016/j.bbrc.2004.04.027
Niu QW, Lin SS, Reyes JL, Chen KC, Wu HW, Yeh SD, Chua NH (2006) Expression of artificial microRNAs in transgenic Arabidopsis thaliana confers virus resistance. Nat Biotechnol 24:1420–1428. https://doi.org/10.1038/nbt1255
Ossowski S, Schwab R, Weigel D (2008) Gene silencing in plants using artificial microRNAs and other small RNAs. Plant J 53:674–690. https://doi.org/10.1111/j.1365-313X.2007.03328.x
Petchthai U, Yee CSL, Wong S (2018) Resistance to CymMV and ORSV in artificial microRNA transgenic Nicotiana benthamiana plants. Sci Rep 8:9958. https://doi.org/10.1038/s41598-018-28388-9
Qu J, Ye J, Fang RX (2007) Artificial microRNA-mediated virus resistance in plants. J Virol 81:6690–6699. https://doi.org/10.1128/JVI.02457-06
Reingold V, Lachman O, Blaosov E, Dombrovsky A (2015) Seed disinfection treatments do not sufficiently eliminate the infectivity of Cucumber green mottle mosaic virus (CGMMV) on cucurbit seeds. Plant Pathol 64:245–255. https://doi.org/10.1111/ppa.12260
Rogers K, Chen X (2013) Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell 25:2383–2399. https://doi.org/10.1105/tpc.113.113159
Sablok G, Pérez-Quintero ÁL, Hassan M, Tatarinova TV, López C (2011) Artificial microRNAs (amiRNAs) engineering-on how microRNA-based silencing methods have affected current plant silencing research. Biochem Biophys Res Commun 406:315–319. https://doi.org/10.1016/j.bbrc.2011.02.045
Schubert S, Grünweller A, Erdmann VA, Kurreck J (2005) Local RNA target structure influences siRNA efficacy: systematic analysis of intentionally designed binding regions. J Mol Biol 348:883–893. https://doi.org/10.1016/j.jmb.2005.03.011
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on the plant transcriptome. Dev Cell 8:517–527. https://doi.org/10.1016/j.devcel.2005.01.018
Schwab R, Ossowski S, Riester M, Warthmann N, Weigel D (2006) Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18:1121–1133. https://doi.org/10.1105/tpc.105.039834
Shim CK, Han KS, Lee JH, Bae DW, Kim DK, Kim HK (2005) Isolation and characterization of watermelon isolate of Cucumber green mottle mosaic virus (CGMMV-HY1) from watermelon plants with severe mottle mosaic symptoms. J Plant Pathol 21:167–171. https://doi.org/10.5423/PPJ.2005.21.2.167
Simon-Mateo C, Garcia JA (2006) MicroRNA-guided processing impairs Plum pox virus replication, but the virus readily evolves to escape this silencing mechanism. J Virol 80:2429–2436. https://doi.org/10.1128/JVI.80.5.2429-2436.2006
Song YZ, Han QJ, Jiang F, Sun RZ, Fan ZH, Zhu CX, Wen FJ (2014) Effects of the sequence characteristics of miRNAs on multi-viral resistance mediated by single amiRNAs in transgenic N. benthamiana. Plant Physiol Biochem 77:90–98. https://doi.org/10.1016/j.plaphy.2014.01.008
Sunkar R, Kapoor A, Zhu J (2006) Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis in mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 18:2051–2065. https://doi.org/10.1105/tpc.106.041673
Tian T, Posis K, Maroon-Lango CJ, Mavrodieva V, Haymes S, Pitman TL, Falk BW (2014) First report of Cucumber green mottle mosaic virus on melon in the United States. Plant Dis 98:1163. https://doi.org/10.1094/PDIS-02-14-0176-PDN
Tiwari M, Sharma D, Trivedi PK (2014) Artificial microRNA mediated gene silencing in plants: progress and perspectives. Plant Mol Biol 86:1–18. https://doi.org/10.1007/s11103-014-0224-7
Ugaki M, Tomiyama M, Kakutani T, Hidaka S, Kiguchi T, Nagata R, Sato T, Motoyoshi F, Nishiguchi M (1991) The complete nucleotide sequence of cucumber green mottle mosaic virus (SH strain) genomic RNA. J Gen Virol 72:1487–1495. https://doi.org/10.1099/0022-1317-72-7-1487
Vu TV, Choudhury NR, Mukherjee SK (2013) Transgenic tomato plants expressing artificial microRNAs for silencing the pre-coat and coat proteins of a begomovirus, Tomato leaf curl New Delhi virus, show tolerance to virus infection. Virus Res 172:35–45. https://doi.org/10.1016/j.virusres.2012.12.008
Wagaba H, Patil BL, Mukasa S, Alicai T, Fauquet CM, Taylor NJ (2016) Artificial microRNA-derived resistance to Cassava brown streak disease. J Virol Methods 231:38–43. https://doi.org/10.1016/j.jviromet.2016.02.004
Warthmann N, Chen H, Ossowski S, Weigel D, Herve P (2008) Highly specific gene silencing by artificial miRNAs in rice. PLoS One 3:e1829. https://doi.org/10.1371/journal.pone.0001829
Xin MM, Wang Y, Yao YY, Xie CJ, Peng HR, Ni ZF, Sun QX (2010) Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol 10:123. https://doi.org/10.1186/1471-2229-10-123
Zhang YJ, Li GF, Li MF (2009) Occurrence of Cucumber green mottle mosaic virus on cucurbitaceous plants in China. Plant Dis 93:200. https://doi.org/10.1094/PDIS-93-2-0200C
Zhang XH, Li HX, Zhang JH, Zhang CJ, Gong PJ, Ziaf K, Xiao FM, Ye ZB (2011) Expression of artificial microRNAs in tomato confers efficient and stable virus resistance in a cell-autonomous manner. Transgenic Res 20:569–581. https://doi.org/10.1007/s11248-010-9440-3
Zhang L, Xie X, Song Y, Jiang F, Zhu C, Wen F (2013) siRNA-mediated viral resistance depends on the sequence similarity and mismatched sites between the target sequence and siRNA. Biol Plant 57:547–554. https://doi.org/10.1007/s10535-013-0314-4
Acknowledgements
This work was partially supported by the National Key Research and Development Program of China (2017YFD0201601), the National Science Foundation of China (NSFC) project (31371910) and China Scholarship Council (CSC) (201606350070).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Fig. S1
The positions of amiRNAs target conserved regions of genes for coat protein (CP), movement protein (MP) and replicase (Rep) in CGMMV genome. The purple boxes denote the positions of the target sites of amiRNAs (PNG 881 kb)
Fig. S2
The construction strategy of the amiRNA plant expression vectors (PDF 479 kb)
Fig. S3
Products of overlapping PCR for six amiRNA precursors. I-12 was synthesized using Primers 1 and 2 in PCR reaction ①. I-345 was synthesized using Primers 3, 4 and 5 in PCR reaction ②. I-12 and I-345 were used as template and Primers 1 and 5 were used to synthesize amiRNA precursor sequence in PCR reaction ③ (TIFF 185 kb)
Fig. S4
Secondary structure of amiRNA precursors by bioinformatic analysis using MFOLD program (PDF 41 kb)
Fig. S5
The relative amiRNA band intensities with respect to the corresponding miR172 bands, estimated using Quantity One Basic, are shown as a column graph (TIFF 64 kb)
Table S1.
Primers used to synthesize amiRNA precursors by overlapping PCR. Table S2. Detail of repeat setting of the experiment. Table S3. Probes used to detect amiRNA expression by northern blot hybridization analysis. Table S4. Primers used for qRT-PCR in the detection of amiRNA overexpression. Table S5. Primers used for qRT-PCR in the detection of CGMMV. Table S6. Prediction of secondary structure of amiRNA (XLSX 20 kb)
Rights and permissions
About this article
Cite this article
Liang, C., Hao, J., Li, J. et al. Artificial microRNA-mediated resistance to cucumber green mottle mosaic virus in Nicotiana benthamiana. Planta 250, 1591–1601 (2019). https://doi.org/10.1007/s00425-019-03252-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-019-03252-w