Abstract
RNA silencing is an innate immune mechanism of plants against invasion by viral pathogens. Artificial microRNA (amiRNA) can be engineered to specifically induce RNA silencing against viruses in transgenic plants and has great potential for disease control. Here, we describe the development and application of amiRNA-based technology to induce resistance to soybean mosaic virus (SMV), a plant virus with a positive-sense single-stranded RNA genome. We have shown that the amiRNA targeting the SMV P1 coding region has the highest antiviral activity than those targeting other SMV genes in a transient amiRNA expression assay. We transformed the gene encoding the P1-targeting amiRNA and obtained stable transgenic Nicotiana benthamiana lines (amiR-P1-3-1-2-1 and amiR-P1-4-1-2-1). Our results have demonstrated the efficient suppression of SMV infection in the P1-targeting amiRNA transgenic plants in an expression level-dependent manner. In particular, the amiR-P1-3-1-2-1 transgenic plant showed high expression of amiR-P1 and low SMV accumulation after being challenged with SMV. Thus, a transgenic approach utilizing the amiRNA technology appears to be effective in generating resistance to SMV.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Artificial microRNAs (amiRNAs) function by mimicking the natural microRNA molecules in plants (Schwab et al. 2006). These engineered molecules are designed to target specific messenger RNAs for degradation or translational inhibition, thereby effectively regulating gene expression (Park et al. 2009). The amiRNA has been successfully engineered in plants to introduce virus resistance (Carbonell et al. 2019; Jian et al. 2017; Miao et al. 2021; Qu et al. 2007; Vu et al. 2013). This approach offers potential advantages over antiviral strategies based on hairpin RNA structures (Mitter and Dietzgen 2012). Notably, the amiRNA sequences are only 21–24 nucleotides (nt) long, which minimizes the risk of unintended interference with plant genes. Furthermore, unlike siRNAs, the generation of miRNAs remains unaffected by low temperatures (Szittya et al. 2003). Transgenic expression of amiRNAs has demonstrated high efficacy in preventing infection by the target viruses, particularly at low temperatures (Kis et al. 2016).
Not every randomly selected 21–24 nt sequence complementary to the viral genome can confer resistance to plant viruses. The differential inhibitory activities of various amiRNAs against viruses arise at different levels. Viral RNAs are highly structured (Simon 2015; Xu et al. 2022) and exhibit different levels of accessibility to amiRNAs complementary to the viral genome, leading to differential efficiency in virus inhibition (Duan et al. 2008). The sequence itself and the hybridization energy of virus-derived amiRNA can also influence its effectiveness in translational repression (Miao et al. 2021). Therefore, experimental assessment of engineered amiRNAs is essential before establishing stable transgenic lines to combat different viruses.
Soybean mosaic virus (SMV) is a prevalent and major viral pathogen that infects soybean plants and causes substantial losses in soybean production worldwide (Hajimorad et al. 2018). SMV transmission occurs through seeds during soybean growth seasons, with aphids facilitating short-distance transmission in the fields (Hajimorad et al. 2018). The current management of SMV disease involves controlling the aphid vector and using resistant soybean genotypes (Hong et al. 2024). However, due to limited resources for resistance and the emergence of resistance-breaking SMV isolates (Ishibashi et al. 2019; Yin et al. 2021), new strategies to control soybean mosaic disease are crucial.
Our study investigated four amiRNAs targeting different SMV genes. We found that amiRNA targeting the SMV P1 gene most effectively inhibits SMV infection. Stable transgenic N. benthamiana lines expressing the P1-targeting amiRNA were generated and exhibited SMV resistance.
Materials and methods
Plant material and virus inoculation
Nicotiana benthamiana plants were grown in 14 h light/10 h dark cycles at 22 °C with ~ 50% relative humidity. Virus inoculation in N. benthamiana was performed by infiltrating 5-week-old plants with A. tumefaciens strain EHA 105 (OD600 = 0.2) carrying the SMV infectious clone, which was described previously (Yin et al. 2021).
Construction of artificial miRNAs
The amiRNAs designed to target the genome of SMV SC7 strain (Yin et al. 2021) were designed as following, amiR-P1 (5’- UGGUUAAAGAUGAUCGCUCAU -3’), amiR-HcPro (5’- UCGCUUUGAAUGCCUGCCACG -3’), amiR-NIb (5’- UGUCUCUGCAAUAUAUGGUGC -3’), and amiR-CP (5’- UAUCUCUCAAAUUCCUCAGUA -3’). The primers of artificial miRNA were designed by WMD3-Web MicroRNA designer (Schwab et al. 2006). The precursor structure of amiRNAs was obtained by replacing the 21nt of the Arabidopsis thaliana miR319a as well as the partially complementary region of the miR319a* with four site-directed mutant primers and two primers complement to the template plasmid sequence. The pRS300 plasmid, containing the precursor sequence of at-miR319a, was used as the template for PCR. Primer A and amiRNAs* reverse primer amplify the 5’-terminus of the amiRNAs precursor. AmiRNAs* forward primer and amiRNAs reverse primer amplify the middle part of the precursor. The 3’-terminus was amplified with amiRNAs forward primer and primer B. The three resulting fragments were gel purified and subjected to overlapping PCR with Primer A and Primer B. The fused PCR product was gel purified, cleaved by restriction enzymes XhoI and XbaI, and inserted into vector pGD. Further, the amiRNA precursor sequences were amplified using primers #927 and #928 from the pGD-35S-based templates and ligated into the vector pCambia1301 digested by NcoI and BstEII, resulting in the plant expression vector for transient expression assay and plant transformation. The primer sequences are provided in Supplementary Table 1.
Agrobacterium-mediated gene expression
Agrobacterium tumefaciens (reclassified as Rhizobium radiobacter) strain EHA105 carrying the appropriate plasmid was cultured in LB broth supplemented with Kanamycin and Rifampicin overnight at 28 °C with shaking. The cells were then centrifuged at 4500 g for 10 min and resuspended in MMA buffer (10 mM MES, 10 mM MgCl2, and 200 μM acetosyringone). The cell suspensions, adjusted to an optical density of 1.0 (OD600 = 1), were incubated for 3 h at room temperature before infiltration. For the experiments, the cultures were mixed according to the requirements and infiltrated into the underside of leaves of 5-week-old N. benthamiana plants.
Plant transformation
To generate stable N. benthamiana transgenic lines, the plants were transformed with pCambin1301-amiRNA-P1 using the A. tumefaciens-mediated leaf disc transformation method (Gallois and Marinho 1995). Briefly, the leaves of N. benthamiana plants of 4 weeks old were incubated in 70% ethanol solution for 30 s followed by incubation in 2.5% sodium hypochlorite solution for 10 min. The leaves were spliced into leaf discs of about 1 cm in diameter, followed by incubation with A. tumefaciens strain GV3101 carrying pCambia1301-amiR-P1 on regeneration medium (Murashige and Skoog medium supplemented with 3 mg/L 6-benzylaminopurine, 0.2 mg/L 1-naphthaleneacetic acid, and 100 μM acetosyringone) for 2 days, followed by transfer to selection medium (Murashige and Skoog medium supplemented with 3 mg/L 6-benzylaminopurine, 0.2 mg/L 1-naphthaleneacetic acid, 50 mg/mL hygromycin B, and 250 mg/L carbenicillin) for the generation of transgenic shoots. Subsequently, shoots were excised with some explant to induce roots in the rooting medium (1/2 Murashige and Skoog medium). Fully regenerated T0 plants were then transplanted into the soil for seed collection.
Protein extraction and western blotting analysis
N. benthamiana leaf disks were rapidly frozen in liquid nitrogen and subsequently ground into a fine powder using a pestle. The samples were then mixed with SDS loading buffer and incubated at 85 °C for 15 min. Following this, the protein samples were subjected to electrophoresis through SDS-PAGE gels. After separation, the gels containing total leaf proteins were transferred onto PVDF membranes for immunodetection. A rabbit polyclonal antibody specific to soybean mosaic virus coat protein (SMV CP) was employed to detect SMV accumulation. Band quantification was performed using ImageQuant TL software (Cytiva, USA).
RNA extraction and stem-loop RT-qPCR
N. benthamiana leaf disks were ground to fine powders using a pestle after freezing in liquid nitrogen. Subsequently, 250 μl of RNA extraction buffer (0.1 M glycine, 0.1 M NaCl, 0.01 M EDTA, and 1% SDS, pH 9.0) and 250 μl RNA phenol were immediately added. The tube was vortexed for 30 s and centrifuged at 15,000 rpm for 5 min at 4 °C. Approximately 200 μl of the upper aqueous phase was pipetted into a fresh 1.5 ml tube containing one volume of saturated phenol–chloroform. The mixture was vortexed for 30 s and centrifuged at 15,000 rpm for 5 min at 4 °C. About 160 μl of the upper aqueous phase was pipetted into a fresh 1.5 ml tube containing 3 volumes of cold 100% ethanol with 2% (v/v) 3 M sodium acetate (pH 5.6). The mixture was then incubated overnight at -20 °C. The precipitated total RNA was subjected to centrifugation at 15,000 rpm for 20 min at 4 °C, washed with 70% cold ethanol, and dissolved in 30 µl H2O. The concentration of the total RNA was measured using an ultraviolet spectrophotometer. For reverse transcription, approximately 1 μg of total RNA was used with the HiScript III 1st Strand cDNA Synthesis Kit (Vazyme Biotech Co. Ltd., Nanjing, China), employing stem-loop RT primer for amiR-P1 and oligo (dT) 20 for NbTubulin, which served as an internal control. ChamQ SYBR qRCR Master Mix (Vazyme Biotech Co. Ltd., Nanjing, China) was utilized for the relative quantification of the cDNA. Data analysis was performed using the 2 − ΔΔCT method, and corresponding graphs were plotted using Prism 9 (GraphPad).
Viral infection and plant phenotype observation
Plants infected by SMV-GFP (Yin et al. 2021) were observed under a blue-laser hand-hold lamp LUYOR-3415RG (Luyor, Shanghai, China), and the images showing the SMV infection indicated by GFP fluorescence were taken with a camera installed with an LP510 filter.
Statistical analysis
Data were analyzed using one-way analysis of variance (ANOVA) with Tukey’s multiple comparison test. To establish the prerequisites of analysis, Shapiro–Wilk tests were used to check the data normality, and Brown-Forsythe tests were used to assess the equality of variances. Results did not justify rejecting the null hypotheses that the data are normally distributed and that the variances are equal, supporting the statistical methods used.
Results and discussion
Functional assessment of amiRNAs targeting SMV genes
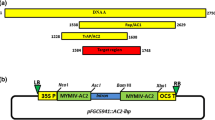
We analyzed the sequences of 11 Chinese SMV isolates and selected four 21 nt-long sequence sites in the P1, Hc-Pro, NIb, and CP coding regions, which are conserved in all SMV isolates analyzed. Based on these four sites, four amiRNAs were designed: amiR-P1, amiR-HcPro, amiR-NIb, and amiR-CP (See amiRNA sequences in the Materials and Methods). These four amiRNAs fully complement the SMV reference sequences (Fig. 1A) and have uridine at position 1 and adenine at position 10, a common feature of highly efficient siRNAs and plant miRNAs (Mallory et al. 2004; Schwab et al. 2006). We used the Arabidopsis thaliana miR319a backbone vector (Schwab et al. 2006) to construct the artificial microRNA expression genes, which were later individually inserted into the plant expression vector pCambia1301, which drives the miRNAs expression under a 35S promoter. We agroinfiltrated these amiRNAs together with an infectious clone of SMV into the leaves of N. benthamiana. At 5.5 days post-infiltration (dpi), total proteins were extracted from the infiltrated leaf areas and subjected to Western blotting analysis using polyclonal antibodies against SMV CP. We found that the accumulation of SMV CP reduced to ~ 36% of that of the empty vector control when amiR-P1 was coexpressed (Fig. 1B, C). Meanwhile, amiR-Hc-Pro and amiR-NIb coexpression reduced the accumulation of SMV CP to ~ 46% and ~ 76% of the level in the empty vector control, respectively. SMV accumulation was not reduced during amiR-CP coexpression. These data suggest that amiR-P1 is the most effective in inhibiting SMV accumulation. Interestingly, amiRNAs targeting the 5’-end genes of the viral genome are generally more effective than those targeting the 3’-end genes. One possible explanation for this difference is that the 5’-end regions of the viral genome are more accessible than the 3’-end regions, where “kissing loop” structures are more frequently found (Bassett et al. 2022; Simon 2015). Experiments are needed to test whether preexisting viral RNA structures hide the recognition sites for amiR-CP recognition.
The screen of artificial microRNAs inhibiting soybean mosaic virus replication in N. benthamiana leaves. A A scheme showing the amiRNA sequences and their target genes on the SMV genome. The designed amiRNAs are fully complementary to the targeted SMV genomic RNA. B Western blotting analysis shows various levels of inhibition on SMV replication by transient expression of amiRNAs. A. tumefaciens cells carrying the vector for the expression of amiRNAs and the SMV infectious clone were mixed at an optical density 600 (OD600) of 0.8 and 0.2, followed by agroinfiltration into N. benthamiana leaves. The empty vector (EV) for amiRNA expression was co-agroinfiltrated with SMV infectious clones as a control. The SMV-specific antibodies detected SMV CP accumulation at 5.5 dpi. The Rubisco large subunit (RbcL), visualized by coomassie blue-staining, was shown as a loading control. C Quantification of SMV CP accumulation under different treatments. Error bars represent the standard deviation of two or three biological repeats performed using leaves from different plants. The significant level was calculated by the one-way ANOVA test (Chatzi and Doody 2023). ****denotes p-value less than 0.0001, **denotes p-value less than 0.005, ns means statistically non-significant
Transgenic N. benthamiana lines stably expressing amiR-P1
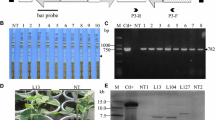
Since amiR-P1 is the most effective amiRNA, we transformed N. benthamiana using the above-mentioned pCambia1301-based plant expression vector carrying amiR-P1. A total of 13 transgenic lines were obtained, of which two lines were selected based on the level of SMV resistance in the initial screen. They were advanced to the T4 generation. We could not detect any further segregation of the transgenes based on PCR analysis, indicating that the T4 lines are homozygous. We measured the amiRNA expression level by stem-loop RT-qPCR (Varkonyi-Gasic et al. 2007). The transgenic line amiR-P1-3-1-2–2-1 showed a significantly higher miR-P1 expression level than the line amiR-P1-4-1-1-2-1 (Fig. 2). We used these two lines in further studies to test whether the transgenic expression of amiR-P1 can inhibit SMV infection.
Expression level of amiR-P1 in N. benthamiana transgenic lines 4-1-1-2-1 and 3-1-2-2-1. Total RNAs were extracted from lines 4-1-1-2-1, 3-1-2–2-1, and non-transgenic N. benthamiana plants and subjected to stem-loop RT-qPCR analysis. The expression of the N. benthamiana β-tubulin coding gene was used as an internal control to calculate relative miR-P1 expression. Error bars represent the standard deviation of three biological repeats performed using different plants. The significant level was calculated by the one-way ANOVA test (Chatzi and Doody 2023). **** denotes p-value less than 0.0001, *denotes p-value less than 0.05
Transgenic expression of amiR-P1 suppresses SMV infection
To test whether the transgenic plants are resistant to SMV infection, we inoculated 5-week-old plants with an infectious SMV clone carrying a GFP cistron (SMV-GFP) (Yin et al. 2021). At 10 dpi, green fluorescence was observed on the upper leaves of the non-transgenic plants under a blue laser lamp, suggesting successful SMV-GFP infection (Fig. 3A). In contrast, the amiR-P1 transgenic lines, especially line 3-1-2-2-1 that had higher amiR-P1 expression, displayed significantly reduced green fluorescence (Fig. 3A). Western blotting analysis, detecting the accumulation of SMV CP in the upper leaves of the inoculated plants, confirmed the reduced level of SMV infection in the transgenic lines (Fig. 3B, C). Specifically, the transgenic line 3–1-2–2-1 suppressed the SMV level to ~ 5%, while the SMV level was reduced to 20% in line 4-1-1-2-1, as compared to that of the non-transgenic plants (Fig. 3B, C). The level of SMV accumulation was negatively correlated with the miR-P1 expression level in the two transgenic lines (Fig. 2). These data demonstrate that transgenic expression of amiR-P1 can efficiently suppress SMV infection in an expression level-dependent manner.
Transgenic expression of amiR-P1 inhibits SMV-GFP infection. A Representative images of N. benthamiana plants infected by SMV-GFP. Magnified regions show different levels of SMV-GFP infection in systemically infected leaves, as suggested by the green fluorescence. In line 3-1-2–2-1, no obvious green fluorescence signals can be observed. Scale bar, 4 cm. B Western blotting analysis shows decreased levels of SMV replication in systemically infected leaves. The SMV-specific antibodies detected SMV CP accumulation at 10 dpi. The rubisco large subunit (RbcL), visualized by Ponceau S Staining, was shown as a loading control. C Quantification of SMV CP accumulation. Error bars represent the standard deviation of three biological repeats performed using different plants. The significant level was calculated by the one-way ANOVA test. ****denotes p-value less than 0.0001, *denotes p-value less than 0.05
Conclusion
Designing artificial microRNAs targeting SMV genes, especially the P1 gene, appears to be an effective strategy to restrict SMV infection. The differential effects of amiRNAs targeting different regions of the plant viral genome were also reported for cucumber mosaic virus, cucumber green mottle mosaic virus, and tomato leaf curl virus (Miao et al. 2021; Qu et al. 2007; Vu et al. 2013). The amiRNAs hybridizing to the cucumber mosaic virus 3’ terminal tRNA structure region showed little effect inhibiting the viral replication, likely due to its attenuated accessibility to the highly structured region (Duan et al. 2008). We also observed that amiRNAs targeting the 3’ region are less effective. However, the RNA structure of SMV has not yet been determined. Whether it is the amiRNA accessibility to the viral RNA needs further demonstration.
Previous attempts to obtain SMV-resistant plants through an RNA silencing-based transgenic approach were only reported using Hc-Pro gene-derived hairpin RNAs (Gao et al. 2015; Kim et al. 2016). This work added a new avenue for developing SMV-resistant plants utilizing the RNA silencing mechanism and could be employed in future agricultural practices.
Data availability
All data are available within the article.
References
Bassett M, Salemi M, Rife Magalis B (2022) Lessons learned and yet-to-be learned on the importance of RNA structure in SARS-CoV-2 replication. Microbiol Mol Biol Rev 86:e0005721. https://doi.org/10.1128/mmbr.00057-21
Carbonell A, Lopez C, Daros JA (2019) Fast-forward identification of highly effective artificial small RNAs against different tomato spotted wilt virus isolates. Mol Plant Microbe Interact 32:142–156. https://doi.org/10.1094/MPMI-05-18-0117-TA
Chatzi A, Doody O (2023) The one-way ANOVA test explained. Nurse Res 31:8–14. https://doi.org/10.7748/nr.2023.e1885
Duan CG, Wang CH, Fang RX, Guo HS (2008) Artificial microRNAs highly accessible to targets confer efficient virus resistance in plants. J Virol 82:11084–11095. https://doi.org/10.1128/JVI.01377-08
Gallois P, Marinho P (1995) Leaf disk transformation using Agrobacterium tumefaciens-expression of heterologous genes in tobacco. Methods Mol Biol 49:39–48. https://doi.org/10.1385/0-89603-321-X:39
Gao L, Ding X, Li K, Liao W, Zhong Y, Ren R, Liu Z, Adhimoolam K, Zhi H (2015) Characterization of Soybean mosaic virus resistance derived from inverted repeat-SMV-HC-Pro genes in multiple soybean cultivars. Theor Appl Genet 128:1489–1505. https://doi.org/10.1007/s00122-015-2522-0
Hajimorad MR, Domier LL, Tolin SA, Whitham SA, Saghai Maroof MA (2018) Soybean mosaic virus: a successful potyvirus with a wide distribution but restricted natural host range. Mol Plant Pathol 19:1563–1579. https://doi.org/10.1111/mpp.12644
Hong X, Li S, Cheng X, Zhi H, Yin J, Xu K (2024) Searching for plant NLR immune receptors conferring resistance to potyviruses. Crop J 12:28–44. https://doi.org/10.1016/j.cj.2023.11.010
Ishibashi K, Saruta M, Shimizu T, Shu M, Anai T, Komatsu K, Yamada N, Katayose Y, Ishikawa M, Ishimoto M, Kaga A (2019) Soybean antiviral immunity conferred by dsRNase targets the viral replication complex. Nat Commun 10:4033. https://doi.org/10.1038/s41467-019-12052-5
Jian C, Han R, Chi Q, Wang S, Ma M, Liu X, Zhao H (2017) Virus-based microRNA silencing and overexpressing in common wheat (Triticum aestivum L.). Front Plant Sci 8:500. https://doi.org/10.3389/fpls.2017.00500
Kim HJ, Kim M-J, Pak JH, Im HH, Lee DH, Kim K-H, Lee J-H, Kim D-H, Choi HK, Jung HW, Chung Y-S (2016) RNAi-mediated Soybean mosaic virus (SMV) resistance of a Korean soybean cultivar. Plant Biotechnol Rep 10:257–267. https://doi.org/10.1007/s11816-016-0402-y
Kis A, Tholt G, Ivanics M, Varallyay E, Jenes B, Havelda Z (2016) Polycistronic artificial miRNA-mediated resistance to wheat dwarf virus in barley is highly efficient at low temperature. Mol Plant Pathol 17:427–437. https://doi.org/10.1111/mpp.12291
Mallory AC, Reinhart BJ, Jones-Rhoades MW, Tang G, Zamore PD, Barton MK, Bartel DP (2004) MicroRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5’ region. EMBO J 23:3356–3364. https://doi.org/10.1038/sj.emboj.7600340
Miao S, Liang C, Li J, Baker B, Luo L (2021) Polycistronic artificial microrna-mediated resistance to cucumber green mottle mosaic virus in cucumber. Int J Mol Sci. https://doi.org/10.3390/ijms222212237
Mitter N, Dietzgen RG (2012) Use of hairpin RNA constructs for engineering plant virus resistance. Methods Mol Biol 894:191–208. https://doi.org/10.1007/978-1-61779-882-5_13
Park W, Zhai J, Lee JY (2009) Highly efficient gene silencing using perfect complementary artificial miRNA targeting AP1 or heteromeric artificial miRNA targeting AP1 and CAL genes. Plant Cell Rep 28:469–480. https://doi.org/10.1007/s00299-008-0651-5
Qu J, Ye J, Fang R (2007) Artificial microRNA-mediated virus resistance in plants. J Virol 81:6690–6699. https://doi.org/10.1128/JVI.02457-06
Schwab R, Ossowski S, Riester M, Warthmann N, Weigel D (2006) Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18:1121–1133. https://doi.org/10.1105/tpc.105.039834
Simon AE (2015) 3’UTRs of carmoviruses. Virus Res 206:27–36. https://doi.org/10.1016/j.virusres.2015.01.023
Szittya G, Silhavy D, Molnar A, Havelda Z, Lovas A, Lakatos L, Banfalvi Z, Burgyan J (2003) Low temperature inhibits RNA silencing-mediated defence by the control of siRNA generation. EMBO J 22:633–640. https://doi.org/10.1093/emboj/cdg74
Varkonyi-Gasic E, Wu R, Wood M, Walton EF, Hellens RP (2007) Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 3:12. https://doi.org/10.1186/1746-4811-3-12
Vu TV, Choudhury NR, Mukherjee SK (2013) Transgenic tomato plants expressing artificial microRNAs for silencing the pre-coat and coat proteins of a begomovirus, tomato leaf curl New Delhi virus, show tolerance to virus infection. Virus Res 172:35–45. https://doi.org/10.1016/j.virusres.2012.12.008
Xu B, Zhu Y, Cao C, Chen H, Jin Q, Li G, Ma J, Yang SL, Zhao J, Zhu J, Ding Y, Fang X, Jin Y, Kwok CK, Ren A, Wan Y, Wang Z, Xue Y, Zhang H, Zhang QC, Zhou Y (2022) Recent advances in RNA structurome. Sci China Life Sci 65:1285–1324. https://doi.org/10.1007/s11427-021-2116-2
Yin J, Wang L, Jin T, Nie Y, Liu H, Qiu Y, Yang Y, Li B, Zhang J, Wang D, Li K, Xu K, Zhi H (2021) A cell wall-localized NLR confers resistance to Soybean mosaic virus by recognizing viral-encoded cylindrical inclusion protein. Mol Plant 14:1881–1900. https://doi.org/10.1016/j.molp.2021.07.013
Acknowledgements
The authors acknowledge Dr. Caiji Gao from South China Normal University for providing pRS300 vector.
Funding
This work was supported by the National Natural Science Foundation of China (32370158), Jiangsu Province's Innovation Program (JSSCTD202142).
Author information
Authors and Affiliations
Contributions
KX conceived the project. MFL, JT, WZ, WY, TZ, WL, YQ, XD, and XZ performed the experiments. KX, JY, JKK, and TZ analyzed the data and wrote the manuscript. The authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
The authors understand the ethics disclosure statement. Any approval of ethics is not required for the work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Latif, M.F., Tan, J., Zhang, W. et al. Transgenic expression of artificial microRNA targeting soybean mosaic virus P1 gene confers virus resistance in plant. Transgenic Res 33, 149–157 (2024). https://doi.org/10.1007/s11248-024-00388-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11248-024-00388-8