Abstract
Main conclusion
This review presents genetic and molecular basis of crop height using a rice crop model. Height is controlled by multiple genes with potential to be manipulated through breeding strategies to improve productivity.
Height is an important factor affecting crop architecture, apical dominance, biomass, resistance to lodging, tolerance to crowding and mechanical harvesting. The impressive increase in wheat and rice yield during the ‘green revolution’ benefited from a combination of breeding for high-yielding dwarf varieties together with advances in agricultural mechanization, irrigation and agrochemical/fertilizer use. To maximize yield under irrigation and high fertilizer use, semi-dwarfing is optimal, whereas extreme dwarfing leads to decreased yield. Rice plant height is controlled by genes that lie in a complex regulatory network, mainly involved in the biosynthesis or signal transduction of phytohormones such as gibberellins, brassinosteroids and strigolactones. Additional dwarfing genes have been discovered that are involved in other pathways, some of which are uncharacterized. This review discusses our current understanding of the regulation of plant height using rice as a well-characterized model and highlights some of the most promising research that could lead to the development of new, high-yielding varieties. This knowledge underpins future work towards the genetic improvement of plant height in rice and other crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Over the past 40 years, the human population has doubled, reaching 7.3 billion people in 2016 and predicted to reach 9.1 billion by 2050. The Food and Agriculture Organisation (FAO) estimates that market demand for food will follow at the same pace. Demand for cereals, for both human consumption and animal feed, is projected to reach some 3 billion tonnes by 2050, up from today’s demand of ~ 2.1 billion tonnes. To meet this demand requires a 70% increase in food production from the levels seen in 2005/07 (FAO 2009, http://www.fao.org/docrep/012/i0680e/i0680e.pdf).
Significant increases in global food production occurred in the 1960s and 1970s during the ‘Green Revolution’, where impressive gains in the yields of wheat and rice doubled production in most parts of the world (Hargrove and Cabanilla 1979; Khush 1999). The biggest contributors to this were the breeding of semi-dwarf, high-yielding varieties, in combination with new fertilizer and pesticide technologies. Dwarfing was attributed to the plants’ genetic inability to synthesize or respond to certain phytohormones, predominantly gibberellins (GAs). GAs are perturbed in recessive mutants of semi dwarfing 1 (sd1) (Monna et al. 2002; Sasaki et al. 2002; Ashikari et al. 2002) as well as one of two reduced height 1 (Rht-B1 and Rht-D1) (Peng et al. 1999; Hedden 2003) (Fig. 1a, b). Alleles of sd1 and Rht1 have been deployed in breeding of commercial rice and wheat varieties, respectively. Dwarfed rice varieties containing sd1 alleles offer several advantages over taller varieties, including more efficient use of fertilizers, resistance to lodging, greater tillering and a higher harvest index (Monna et al. 2002). The GA-insensitive Rht-B1b and Rht-D1b dwarfing mutants in wheat alter plant and canopy height, show reduced crop lodging and greater dry-matter, which results in increase grain number, harvest index and grain yield (Bonnett et al. 2001; Chapman et al. 2007).
The phenotypic effect of key dwarfing genes that were utilized during the ‘green revolution’. a Stature of normal-type (right, Taiwanese indica strain woo-gen) and semi-dwarf type (left, dee-geo-woo-gen, containing a 383-base-pair deletion of sd-1 which introduces a stop codon and produces a truncated, inactive enzyme) rice. The figure is derived from Hedden (2010). b Stature of normal-type (left, cultivar Mercia) and semi-dwarf (centre, Rht-B1b; right, Rht-D1b) near-isogenic wheat lines. The figure is derived from Peng et al. (1999)
However, the benefits of GA-insensitive semi-dwarfs are greatest in favorable environments such as a higher-yielding, irrigated environment (Butler et al. 2005; Chapman et al. 2007). The rising temperatures as a consequence of climate change will have negative impact on GA-insensitive dwarfing mutants (Rebetzke et al. 2012). Moreover, the overuse of limited dwarf genetic materials like sd1 and Rht1 runs the risk of losing genetic diversity. There is a need to expand the range of dwarf source material, to identify new genes by functional genomics, and to exploit these genes in practice. Understanding dwarfing has thus become a major focus of research in crop genetics and genomics, which has led to many great advances (Ikeda et al. 2001; Sasaki et al. 2002, 2003; Hong et al. 2003; Ueguchi-Tanaka et al. 2000, 2005; Tanabe et al. 2005; Arite et al. 2007). Apart from the rice sd1 gene and the wheat Rht1 gene, other genes contributing to both crop height have been cloned in the GA biosynthesis/signaling pathway (Ashikari et al. 1999; Ueguchi-Tanaka et al. 2000; Sasaki et al. 2002, 2003; Itoh et al. 2004) as well as that of other hormones including brassinosteroids (BRs) (Hong et al. 2003; Tanabe et al. 2005) and strigolactones (SLs) (Lin et al. 2009), indole-3-acetic acid (IAA), abscisic acid (ABA), ethylene (ETH) and Jasmonate (JA). Dwarfing non-hormone pathways have also been described (Arite et al. 2007).
Despite high-yielding mutants, some dwarfing mutants defective in hormone biosynthesis or signaling often display pleiotropic phenotypes that negatively affect productivity via altered tillering, stem physiology and fertility. Such affects include small grains, semi-sterility and malformed panicles (Butany et al. 1959; Sazuka et al. 2009; Asano et al. 2010). Thus the productivity phenotype cannot be definitively attributed to the dwarfing effect itself. Nonetheless, despite this potential for unwanted alleles, it is important that the search continues for novel semi-dwarf genes.
In this review, we describe how crop height can be manipulated to improve crop yield. We explain the genetic and physiological basis of crop dwarfing, including the molecular mechanisms that regulate rice plant height. Finally, we show an application of rice plant height genes in crop breeding and offer suggestions for future research towards new strategies to control crop.
Relationship between crop height and yield
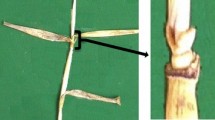
Plants that are dwarfed have been associated with yield increases and yield stability (Peng et al. 1999). There are different categories of dwarfing, namely semi-dwarf, dwarf and extra-dwarf, which refer to varieties that, respectively, attain 50–100%, equal to or slightly less than 50% and substantially less than 50% of the mature height of a wild-type plant (Futsuhara and Kikuchi 1997). Semi-dwarfing increases grain yield per unit area in a variety of ways by allowing (1) planting at higher densities (Mori et al. 2002; Hong et al. 2005), (2) reducing apical dominance and, therefore, promoting tillering (Alder et al. 2012; Waters et al. 2012; Wei et al. 2013), (3) reducing the final biomass and, therefore, increasing the economic coefficient (or harvest index) (Fischer and Quail 1990; Zhang et al. 2014a) and (4) providing greater resistance to lodging (Hedden and Phillips 2000). Enhanced lodging resistance also facilitates mechanical harvesting (Fig. 2).
Excessive dwarfing is often correlated with poor agronomic traits, such as smaller grains, excessive tillering, narrower or rolled leaves and poor disease resistance (Ueguchi-Tanaka et al. 2000; Tanabe et al. 2005; Arite et al. 2007; Li et al. 2009). On the other hand, when a plant is excessively tall, the stems are too weak to support the weight of the head of high-yielding varieties, and they are susceptible to lodging by wind and rain subsequently suffering from diseases and insect pests. Through a combination of these observations, it is assumed that the relationship between crop height and yield follows a bell curve, peaking when the leaf area index (green leaf area per unit ground surface area) reaches the optimal value (Begonia and Begonia 2007). However, the relationship between crop height and yield is affected by a number of factors, such as environment (Botwright et al. 2005; Chapman et al. 2007; Rebetzke et al. 2012), where the bell curve is swayed one way or another. Indeed, there are still situations where tall isolines may yield the same or better than short isolines (Chapman et al. 2007). In general, however, studies indicate wheat yields are maximised within a height range of 70–100 cm (Richards 1992; Flintham et al. 1997; Botwright et al. 2005).
Genetic basis of crop height
Rice plant height is mainly controlled by one to three major genes but modulated by minor genes (Butany et al. 1959; Iwata et al. 1977; Saeda and Kitano 1992; Liu et al. 2008). Most natural or artificially induced dwarf mutants of the ‘indica-type’ varieties are homozygous for the semi-dwarf gene sd1 and are only rarely controlled by two, three or more genes. A similar situation exists with ‘japonica-type’ rice varieties (Butany et al. 1959; Iwata et al. 1977; Saeda and Kitano 1992; Liu et al. 2008). The naming of rice dwarf genes has been standardized, where d denotes dwarf and sd, semi-dwarf (Mitsunaga et al. 1994; Iwata et al. 1995). The genes have been numbered according to the chronology of discovery, with 62 d and 15 sd genes registered (Kinoshita 1995). They can be roughly divided into three categories: (1) the same as the sd1 locus, (2) sharing a complex locus with sd1 and (3) not co-located with sd1 locus (Chang et al. 1984; Murai et al. 1990).
Paradoxically, the study of factors leading to excessive height has contributed to a better understanding of dwarfism. In 1981, the American scholars Rutger and Carnahan reported a mutant rice plant ‘76: 4512’, which had an elongated uppermost internode (elongated uppermost internode stock). The gene causing this phenotype, which is not linked to sd1, was named ELONGATED UPPERMOST INTERNODE (EUI) (Rutger and Carnahan 1981). Crossing experiments demonstrated that EUI was completely or incompletely dominant over sd1 in rice.
Maize plant height is divided into three types: tall (more than 2.5 m), moderate (1.8–2.5 m) and dwarf (less than 1.8 m), controlled by single genes or polygenes. At present, a total of 20 dwarfing genes have been found in maize, but only 17 have been mapped. With the exception of DWARF8 (D8), DWARF9 (D9) and DWARF8 (D11), most of these genes are recessive and are characterized as single-gene traits (Emerson et al. 1935; Neuffer et al. 1997; Lawit et al. 2010; Wang et al. 2013a, b).
Wheat plant height is controlled by major genes and modulated by several other modifier genes. In wheat, the major dwarfing genes belong to the Rht family, wherein 26 dwarf genes have been identified. Nine of these have been mapped, and five are recessive (Mcintosh 1988). Most research has focused on Rht1, Rht2, Rht3, Rht8, Rht9, Rht10, Rht12 and Rht13, with each having a different effect on yield. Multi-varietal comparisons have shown that cultivated semi-dwarf varieties carrying the Rht1 and Rht2 genes are better yielding than the taller stem commercial varieties (Miralles and Slafer 1995; Beharav et al. 1998; Botwright et al. 2005; Rebetzke et al. 2011, 2012; Wang et al. 2014).
In barley, several short-statured genotypes have been identified that possess the semi-dwarf genes uzu and sdw1 and have been widely used in breeding programs to reduce lodging and to improve the harvest index (Sears et al. 1981; Foster and Thompson 1987; Li et al. 2015).
Rapeseed dwarf mutants are also divided into two types. Mutants of one type are controlled by dominant or recessive genes and mutants of the other type are controlled by polygenes. More than ten dwarf or semi-dwarf mutants have been reported in rapeseed, but only one dwarfing gene has been successfully cloned (Foisset et al. 1995; Muangprom and Osborn 2004).
Mechanisms controlling plant height in rice
The mechanisms and genes controlling plant height have been more thoroughly investigated in rice than in any other crop. Some of the mapped genes in other species are homologous to rice genes controlling a similar metabolic pathway; however, the mechanisms controlling plant height of other crops are limited by relative lack of genomic information and research tools. Therefore, this section will focus on examples from rice as a model crop, for broad extrapolation to other species.
The various pathways controlling height are shown in Supplementary Table 1. The phytohormones influencing crop height are GAs, BRs, SLs, IAA, ABA, ETH (Ashikari et al. 1999; Ueguchi-Tanaka et al. 2000; Sasaki et al. 2002, 2003; Hong et al. 2003; Itoh et al. 2004; Tanabe et al. 2005; Lin et al. 2009). Dwarf mutants influenced by phytohormones can be divided into two types according to their responsiveness to exogenous hormones. In loss-of-function dwarf mutants, the hormone synthesis pathway is blocked or inhibited, and normal plant growth can be restored by application of the hormone that is missing. In insensitive dwarf mutants, the endogenous hormone levels are similar to, or even higher than in the wild-type plant, but there are blockages in the hormone signal transduction pathways and normal plant growth cannot be restored by applications of the hormone (Supplementary Table 1). Dwarf mutants influenced by non-hormone factors are mainly involved in several pathways including cell wall development, cytosolic glutamine synthetic pathway, RNA editing, cell division, ubiquitin–proteasome pathway and fatty acid metabolism. In addition, there are genes controlling plant height in rice from as yet uncharacterised pathways.
Genes controlling rice plant height that relate to phytohormone pathways
GA-related pathways
GAs comprise one of the major hormone groups in higher plants and consist of tetracyclic diterpenoid phytohormones, with 136 different compounds currently known (Hedden and Thomas 2012). The GAs play a role in many aspects of plant growth and development, such as stem or internode elongation, flower and fruit development and seed germination (Ross et al. 1997; Ayano et al. 2015). GA-deficient and GA-insensitive mutants are usually much shorter than wild-type plants.
Genes involved in GA biosynthesis
Bioactive GAs, such as GA1 and GA4, are synthesized from trans-geranylgeranyl diphosphate (GGDP) through the action of ent-copaly diphosphate synthases (CPS) and ent-kaurene synthase (KS) in the plastids; by ent-kaurene oxidase (KO), ent-kaurenoic acid oxidase (KAO) and GA 13-oxidase (GA13ox) on the endoplasmic reticulum membrane and through two parallel pathways in the cytoplasm involving GA 20-oxidase (GA20ox) and GA-3β-hydroxylase/GA 3-oxidase (GA3ox) (Hedden and Kamiya 1997; Hedden and Phillips 2000) (Fig. 3).
Principal pathways of gibberellin (GA) biosynthesis and catabolism (see boxes with red dashed lines) in crop species. GA biosynthesis: trans-geranylgeranyl diphosphate (GGDP) is converted to the tetracyclic hydrocarbon ent-kaurene via ent-copalyl diphosphate (CDP) by two kinds of diterpene cyclases in plastids, CDP synthase (CPS) and ent-kaurene synthase (KS). ent-Kaurene is then modified by sequential oxidations to produce GA12 via ent-kaurenoic acid. These steps are catalyzed by two membrane-associated cytochrome P450 monooxygenases, ent-kaurene oxidase (KO) and ent-kaurenoic acid oxidase (KAO). The final stage of bioactive GA synthesis, from GA53/GA12 to GA1/GA4, is catalyzed through two parallel pathways (early-13-hydroxylation and non-13-hydroxylation pathways) by two soluble 2-oxoglutarate-dependent dioxygenases (2ODDs) in the cytosol (GA 20-oxidase (GA20ox) and GA 3-oxidase (GA3ox)) (Sakamoto et al. 2004). GA catabolism: The bioactive GA1/GA4 and their immediate precursors GA20/GA9 are inactivated by a third 2ODD, GA 2-oxidase (GA2ox) (Thomas et al. 1999; Hedden and Phillips 2000; Olszewski et al. 2002; Schomburg et al. 2003; Lee and Zeevaart 2005); GA12, GA9, and GA4 are converted to 16α,17-epoxy derivatives by ELONGATED UPPERMOST INTERNODE (EUI). The epoxides are chemically converted to 16,17-dihydro-16α,17-dihydroxy GAs when the products are purified in the presence of 1% acetic acid (Zhu et al. 2006)
In recent years, dwarf genes involved in the GA biosynthesis and catabolism pathways (CPS, KS, KO, KAO, GA20ox, GA3ox) have been characterized from various plant species (Hedden and Phillips 2000; Luo et al. 2006; Zhu et al. 2006) (Fig. 3; Table 1). The semi-dwarf phenotype of the sd1 mutant (Fig. 1a) is the result of a deficiency of active GAs in the elongating stem resulting from a mutation in the OsGA20ox2 gene (Ashikari et al. 2002; Monna et al. 2002; Sasaki et al. 2002; Spielmeyer et al. 2002). This gene has been extensively used in rice breeding due to its high heritability and minimal impact on other agronomic traits. Most semi-dwarf genes in ‘indica-type’ varieties are sd1 alleles. The d18 and d35 mutants have defective OsGA3ox2 and OsKO2 genes, respectively (Ogawa et al. 1996; Itoh et al. 2001, 2004; Sakamoto et al. 2003), which affect the GA biosynthesis pathway. Sakamoto et al. (2004) performed a large-scale screening of rice mutants, and identified nine novel dwarf mutant lines with recessive alleles, including two alleles of cps, three alleles of ks, one allele of ko2 and three alleles of kao. These mutants are characterized as being GA-deficient as applications of active GA can restore the wild-type phenotypes (Table 1).
Rice plant height can also be altered by modifying the GA catabolism pathways. The aforementioned genes GA20ox and GA3ox function in GA biosynthesis and are involved in feedback regulation, while GA 2-oxidase (GA2ox) functions in GA catabolism and is involved in feedforward regulation by bioactive GAs (Hedden and Phillips 2000; Olszewski et al. 2002) (Fig. 3). Bioactive GA1 and GA4 and their precursors, GA20 and GA9, can be inactivated by GA2ox via 2β hydroxylation (Thomas et al. 1999; Hedden and Phillips 2000; Olszewski et al. 2002; Schomburg et al. 2003; Lee and Zeevaart 2005) and GA2ox mutants do not show a typical dwarf phenotype (Sakamoto et al. 2004). Huang et al. (2010) isolated a CaMV 35S enhancer-labeled mutant in rice, which showed dominant dwarfing with enhanced expression of the Os2ox6 allele gene: endogenous GA activity decreased and applications of GA could recover the phenotype. Moreover, ectopic expression of OsGA2ox1 allele gene driven by the promoter of the GA biosynthesis gene GA3ox2, led to a semi-dwarf phenotype, suggesting its potential for improving yield in rice (Sakamoto et al. 2003; Table 1).
Zhu et al. (2006) demonstrated another GA deactivation mechanism, involving the rice EUI gene (Luo et al. 2006), which encodes a P450 monooxygenase that catalyzes 16α,17-epoxidation of non-13-hydroxylated GAs to generate bio-inactive 16α,17-[OH]2-GAs (Fig. 3). eui is a long stem mutant of rice and the plant accumulates extremely high levels of GA1 and GA4 (Zhu et al. 2006). The constitutive overexpression of this gene in transgenic plants significantly reduces the GA levels, resulting in severe dwarf phenotypes (Zhu et al. 2006). Male sterile cultivars of rice are commonly defective in elongation of the uppermost internode due to inadequate GA production by the empty anthers (Li and Yuan 2000), and the EUI gene is a useful breeding target as mutations to this gene improve the heading performance of rice male sterile cultivars in hybrid rice production (Shen and He 1989; He and Shen 1991; Yang et al. 2002; Table 1).
GA homeostasis is maintained via feedback and feedforward regulation of GA levels by major target genes like GA20ox, GA3ox and GA2ox (Yamaguchi 2008), but how regulation is achieved is not fully understood. The gene products of Oryza sativa YABBY 1 (OsYAB1) directly bind to the promoter of GA3ox2 gene and suppress its expression and, therefore, act negatively in GA feedback regulation. Overexpression of OsYAB1 in transgenic rice results in a semi-dwarf phenotype that can be fully rescued by applications of GA (Dai et al. 2007). Liu et al. (2011) reported that Oryza sativa DWARF RICE WITH OVEREXPRESSION OF GIBBERELLIN-INDUCED GENE (OsDOG) was induced by GA3 and repressed by a GA-synthesis inhibitor. Transgenic lines with constitutively expressed OsDOG have decreased levels of GA1 and retarded cell elongation, leading to dwarfing. The decrease of GA1 in the OsDOG overexpression lines is due to reduced expression of GA3ox2 and increased expression of GA2ox1 and GA2ox3. OsDOG is suggested to have a novel function in feedback regulation of GA homeostasis by binding DNA or protein, through A20 and/or AN1 zinc fingers that interact with GA metabolism genes (Table 1).
Genes involved in GA signaling
Most of the components involved in the GA signaling pathway have been identified in rice through genetic screens during the past decade. The most important components include the GA receptor GIBBERELLIN INSENSITIVE DWARF 1 (GID1), the DELLA growth inhibitors (e.g., SLENDER 1 (SLR1)) and the F-box proteins GIBBERELLIN INSENSITIVE DWARF2 (GID2) (Achard and Genschik 2009). The GA-GID1-DELLA pathway, which is the basic GA signal transduction pathway, has been established (Ikeda et al. 2001; Sasaki et al. 2003; Ueguchi-Tanaka et al. 2005) (Fig. 4a). GID2 is the F-box protein subunit of the SKP-CULLIN-F-BOX (SCF) E3 ubiquitin-ligase complexes (Sasaki et al. 2003). In the presence of SCFGID2, GA triggers the degradation of DELLA proteins through the SCF-mediated 26S proteasome system (Ikeda et al. 2001) (Fig. 4a). As a negative regulator of GA signal transduction, SLR1 interacts with GID1 and GID2, mediating GA signaling to regulate rice growth and development (Sasaki et al. 2003; Ueguchi-Tanaka et al. 2005). The current GA action model proposes that the DELLA proteins inhibit plant growth, while the GA signal promotes plant growth by overcoming this inhibition (Harberd 2003; Achard and Genschik 2009). In other words, GA is an ‘inhibitor of an inhibitor’ (Harberd et al. 2009).
The model of GA signaling pathway and coactivator and corepressor model for GA signaling in rice and Arabidopsis. a The model of GA signaling pathway in rice. GA is perceived by the GA receptor GIBBERELLIN INSENSITIVE DWARF 1 (GID1), and the binding of bioactive GA to GID1 enables DELLA, a repressor of GA signaling, to interact with GID1, eventually forming a GA–GID1–DELLA complex. The GIBBERELLIN INSENSITIVE DWARF 2 (GID2) is the F-box protein subunit of the SKP-CULLIN-F-BOX (SCF) E3 complex (Sasaki et al. 2003), The formation of the GA–GID1–DELLA complex promotes interaction between DELLA and the E3 ubiquitin ligase SCFGID2, leading to polyubiquitylation and then degradation of DELLA through the SCF-mediated 26S proteasome system (Ikeda et al. 2001). In the absence of bioactive GAs, DELLAs repress GA responses. In the presence of bioactive GAs, the receptor GID1 is bound by GA, thus promoting its interaction with a DELLA protein. GA represses DELLA activity in two ways: in the absence of SCFGID2 activity, the GA-GID1 complex that is bound to DELLAs suppresses their transcriptional activity, whereas the presence of SCFGID2 stimulates the degradation of DELLAs (Davière and Achard 2013). The degradation or eventual inactivation of DELLAs relieves DELLA mediated repression of GA responses (Achard and Genschik 2009). b A coactivator and corepressor model for GA signaling in Arabidopsis. Under GA deficient conditions, DELLAs are stable and localized in nuclei. DELLAs titrate PHYTOCHROME INTERCTING FACTORS (PIFs) and BRASSINAZOLE RESISTANT (BZR) transcription factors by inhibiting DNA binding activity, while DELLAs interact with GAI-ASSOCIATED FACTOR 1 (GAF1) and exhibit high transcriptional activity with GAF1. In the presence of GA, DELLAs are degraded via the 26S proteasome pathway. On the one hand, PIF and BZR activate target genes such as PACLOBUTRAZOL RESISTANCE (PRE) and EXPANSIN. On the other hand, the GAF1 TOPLESS RELATED (TPR) complex exhibits transcriptional repression activity. GA-induced functional conversion of GAF1 complex in plants depends on the degradation of coactivator DELLAs (Fukazawa et al. 2014)
GA-response mutants include those with alterations in both GA perception and GA signal transduction. GA perception (i.e. receptor) mutants are GA-insensitive dwarfs, which display a similar dwarf phenotype to GA-deficient mutants, except that they fail to respond to exogenous GA (Ueguchi-Tanaka et al. 2000, 2001, 2005; Sasaki et al. 2003; Harberd et al. 2009; Hirano et al. 2010). While most plants with constitutive GA responses are DELLA loss-of-function mutants, which display a similar phenotype to plants treated with excess GA, like taller stems, paler green leaves and lower fertility, irrespective of bioactive GA levels (Ogawa et al. 2000; Ikeda et al. 2001; Hirano et al. 2012).
The rice mutant dwarf 1 (d1) was the first to be found that was insensitive to applications of GAs; the levels of endogenous GA in the d1 mutant were significantly higher than in the wild-type plant. The d1 mutant is defective in the rice heterotrimeric G PROTEIN ALPHA 1 (RGA1 or Gα protein; Ashikari et al. 1999).
Apart from d1, the most reported GA signaling pathway mutants associated with dwarfing include the high stem mutant slr1 (Ikeda et al. 2001), the dwarf mutant gid1 (Ueguchi-Tanaka et al. 2005) and the dwarf mutant gid2 (Gomi et al. 2004). SLR1 maps to the Oryza Sativa GIBBERELLIN INSENSITIVE (OsGAI) locus in rice (Ogawa et al. 2000), encodes a protein containing a DELLA domain, and has high amino acid similarity to the wheat ‘green revolution’ genes Rht-B1b and Rht-D1b, and Rht-D1a and the maize ‘green revolution’ gene D8. slr1 mutant seedlings are two times taller than the wild-type and are not sensitive to GA synthesis inhibitors (Ikeda et al. 2001). Meanwhile, an slr1 gain-of-function mutant, in which degradation of the SLR1 protein occurs later than in the wild-type, may yield new genetic sources for dwarf breeding (Asano et al. 2009). Analyses of the gid1 mutant have shown that GID1 encodes a soluble GA receptor, which has a high affinity to bioactive GAs. By binding to bioactive GAs, GID1 gains the ability to interact with SLR1, forming a GA–GID1–SLR1 complex and thereby inhibits SLR1 or activates ubiquitin degradation of SLR1 through the SCFGID2 complex pathway (Ueguchi-Tanaka et al. 2005). Analysis of the gid2 mutant has shown that GID2 is an F-box subunit of the SCF E3 complex and that the SCFGID2 complex binds with phosphorylated SLR1, inducing ubiquitin-mediated degradation of SLR1 and transduction of the GA signals. In the gid2 mutant, SLR1 cannot be degraded and GA signaling is inhibited, resulting in a severely dwarfed and sterile phenotype (Sasaki et al. 2003; Gomi et al. 2004; Table 1).
Recently Fukazawa et al. (2015) identified an INDETERMINATE DOMAIN 1 (IDD1) family transcription factor in Arabidopsis called GAI-ASSOCIATED FACTOR 1 (GAF1), which can interact with DELLA and TOPLESS RELATED (TPR) in nuclei. DELLAs and TPR act, respectively, as a coactivator and a corepressor of GAF1. GAs convert the GAF1 complex from transcriptional activator to repressor via degradation of DELLAs (Fukazawa et al. 2014, 2015). While this process is well studied in Arabidopsis rather than in rice (Fig. 4b).
There are three other genes, including CHITIN-INDUCIBLE GIBBERELLIN-RESPONSIVE PROTEIN (CIGR), SPINDLY (SPY) and DWARF AND NARROW-LEAF 1 (DNL1), involved in GA signaling with unclear modes of action: ph1, a major QTL on chromosome 1, increases rice plant height and brings forward flowering. Genome annotation suggests that a gene encoding CIGR is likely to be responsible for the ph1 phenotype, which mapped with a high degree of accuracy to a region close to sd1 (Kovi et al. 2011). Within the GAI, RGA, SCR (GRAS) protein super family, which is a plant-specific transcription factor family, OsCIGR belongs to the PAT1 branch, while OsGAI belongs to the related ‘DELLA’ branch (Bolle 2004). OsCIGR is involved in GA signal transduction but its specific mode of action is unknown. In all of the examined tissues of taller plants, CIGR was more highly expressed than in shorter plants, especially in the young leaf sheath containing the elongating cells, indicating an important role of this gene in regulating rice plant height (Kovi et al. 2011).
The second gene, SPY, encodes an O-fucosyltransferase, which is believed to be a negative regulator of the pathway. In Arabidopsis, O-fucosylation activates DELLA by promoting its interaction with key regulators in BR- and light-signaling pathways, including BRASSINAZOLE RESISTANT 1 (BZR1), PHYTOCHROME INTERCTING FACTORS 3 (PIF3) and PIF4 (Zentella et al. 2017). In the OsSPY antisense or RNAi transgenic rice plants, the increased elongation of lower internodes is correlated with decreased expression levels of OsSPY. The suppression of OsSPY in the GA-insensitive mutant gid2 causes increased phosphorylation of the rice DELLA protein SLR1, but does not change concentrations of SLR1. This indicates that the function of OsSPY in GA signaling is probably suppressing the function of SLR1 instead of changing the amount or stability of SLR1. In addition, OsSPY may play roles both in GA signaling and in the BR pathway (Shimada et al. 2006).
To identify the third gene involved in GA signaling, OsDNL1, Wei et al. (2013) examined a dwarf and narrow-leaf mutant dnl1, which has a thinner culm, a smaller number of cells in the vertical axis of the internode and more tillers than the wild-type but also a smaller yield. Genetic analyses have shown that this phenotype is due to a single recessive gene, OsDNL1, which is located on chromosome 12. dnl1 is a GA-insensitive dwarf mutant, but its role has yet to be elucidated (Wei et al. 2013; Table 1).
BR-related pathways
BRs are a very important family of plant-specific steroids, which are structurally related to insect and animal steroids. Brassinolide (BL), first isolated and purified from the pollen grains of Brassica napus, is the most biologically active of the naturally occurring forms of BR (Grove et al. 1979). More than 50 BL analogues have now been identified (Fujioka and Sakurai 1997). BRs have wide-ranging biological activities and play critical roles in plant development (height, leaf erectness, panicle morphology and seed size) and they are known as a new class of phytohormones (Vriet et al. 2012). Knowledge about BRs has primarily come from studies of BR-deficient and BR-insensitive mutants (Fujioka and Yokota 2003). These mutants typically have dark green, erect leaves and shortened leaf sheaths. Multiple lines of evidence indicate that genetic manipulation of BR biosynthesis or its signaling pathway could significantly improve yield (Choe et al. 2001; Sakamoto et al. 2006; Li et al. 2007; Wu et al. 2008). The BR biosynthesis and signal transduction pathway is much better understood in Arabidopsis than in rice (Wang et al. 2012).
Genes involved in BR biosynthesis
The various BRs are likely derived from a range of common plant sterols that have an appropriate side chain structure, like campesterol, sitosterol, isofucosterol, 24-methylenecholesterol and 24-epicampesterol. However, campesterol is the only compound that has definitively been shown to be a BL progenitor (Yokota 1997). The biosynthesis pathway for BL can be divided into general sterol synthesis, from cycloartenol to campesterol (Clouse 2002), and the BR-specific pathway from campesterol to BL (Clouse 2002; Zhang et al. 2014a) (Fig. 5). To date, almost all of the key enzymes involved in BR biosynthesis have been characterized in both Arabidopsis and rice.
Brassinosteroid (BR)-specific biosynthesis pathway from campesterol to brassinolide and the enzymes involved in each reaction. The pathway shows most of the enzymes from Arabidopsis at the steps indicated, and those in red are enzymes identified only in rice (Hong et al. 2005; Kwon et al. 2005; Tanabe et al. 2005; Vriet et al. 2013). The plant sterol campesterol is converted to brassinolide via castasterone, teasterone, dehydroteasterone, typhasterol and castasteronel. Furthermore, C-6 oxidation could occur before (early C-6 oxidation pathway, the pathway on the left) or after (late C-6 oxidation pathway, the pathway on the right) hydroxylation of the side chain. BR-6-oxidase has been shown to convert 6-deoxo compounds such as 6-deoxocastasterone or 6-deoxotyphasterol to the corresponding 6-oxo compounds in Arabidopsis (Zhang et al. 2014a). CR campesterol, CN campestanol, CT cathasterone, TE teasterone, DT dehydroteasterone, TY typhasterol, CS castasteronel, BL brassinolide
In rice, one mutant with a defective sterol synthesis pathway and five mutants with defective BR-specific pathways have now been identified (Table 2). BR-DEFICIENT DWARF 2 (BRD2) is homologous to Arabidopsis DIMINUTO/DWARF 1 (DIM/DWF1) and encodes a C-24 reductase that is involved in synthesis of campesterol. The mutant of this gene, brd2, produces a semi-dwarf phenotype (Hong et al. 2005). The five genes DWARF 11 (D11), DWARF4, EBISU DWARF (D2), CONSTITUTIVE PHOTOMORPHOGENESIS AND DWARFISM 1 (CPD1), CPD2, and BR-DEFICIENT DWARF 1 (BRD1) encode cytochrome P450 family enzymes that are involved in several steps of the BR-specific pathway (Mori et al. 2002; Hong et al. 2002, 2003; Tanabe et al. 2005). With the exception of OsCPD1 and OsCPD2, mutations of these genes cause typical BR knockdown phenotypes, such as dwarfing, erect leaves, shortened internodes and small seeds; the wild-type phenotype can be rescued by applications of bioactive BR compounds (Mori et al. 2002; Hong et al. 2002, 2003, 2005; Tanabe et al. 2005). CYTOCHROME P450 90B2 (CYP90B2)/OsDWARF4 and CYP724B1/D11 are functionally redundant in C-22 hydroxylation (Sakamoto et al. 2006). The loss-of-function mutant osdwarf4-1 shows a weak BR-deficient phenotype with a slightly dwarfed stature and erect leaves and enhanced grain yields when planted at high densities (Sakamoto et al. 2006). The loss-of-function d11 mutant has a semi-dwarf and short-grained phenotype (Tanabe et al. 2005). D2/CYP90D2 catalyzes the conversion of 6-deoxoteasterone to 3-dehydro-6-deoxoteasterone, as well as teasterone to 3-dehydroteasterone (Hong et al. 2003). The loss of function d2 mutant also has a weak BR-deficient phenotype (Hong et al. 2003; Sakamoto et al. 2006). CYP90A3/OsCPD1 and CYP90A4/OsCPD2 encode C-3 oxidases, previously suggested to act as BR C-23 hydroxylases (Ohnishi et al. 2006, 2012). oscpd1-1, a null mutant of CYP90A3/OsCPD1, failed to deliver a BR-deficient phenotype, implying that OsCPD1 and OsCPD2 are functionally redundant and each is capable of catalyzing BR biosynthesis at the required rate for normal growth (Sakamoto and Matsuoka 2006). The brd1 mutant of rice, which has a defective BR6 OXIDASE (OsBR6OX) gene, has short leaf sheaths, few tillers and is sterile. The BR6 oxidases in tomato and Arabidopsis catalyze multiple C-6 oxidations in BR biosynthesis (Shimada et al. 2001) (Table 2).
Genes involved in BR signaling
The signal pathway for BRs is best understood in Arabidopsis (Wang et al. 2012). In rice, a few components of the BR signal pathway have been characterized by forward or reverse genetics (Fig. 6). Some of them, such as Oryza sativa BRASSINOSTEROID INSENSITIVE 1 (OsBRI1), Oryza sativa BRI1 ASSOCIATED KINASE 1 (OsBAK1), Oryza sativa GLYCOGEN SYNTHASE KINASE3 LIKE GENE 1 (OsGSK1) and OsBZR1, are functional orthologs of known Arabidopsis genes. However, orthologs of PHOSPHATASE 2A (PP2A), BR SIGNALING KINASES (BSKs) and BRI1 SUPPRESSOR 1 (BSU1), which are also BR signaling genes in Arabidopsis, have yet to be identified in rice. Conversely, Oryza sativa TILLER ANGLE INCREASED CONTROLLER (OsLIC), Oryza sativa DWARF AND LOW TILLERING (OsDLT) and Oryza sativa TAIHU DWARF 1 (OsTUD1) are BR signaling genes in rice but there no orthologs in Arabidopsis, indicating that some BR functions are specific to rice (Fig. 6).
BR-signaling pathways in rice and Arabidopsis. The left panel shows the BR signal transduction pathway in Arabidopsis, while the right panel shows that in rice. Orange indicates that the components play positive roles in BR signaling, whereas green indicates that components play negative roles, and a gray dotted line indicates a scenario to be verified (Tong and Chu 2012; Zhang et al. 2014a; Yang et al. 2016; Qiao et al. 2017). In rice, BR binding to BR receptor Oryza sativa BRASSINOSTEROID INSENSITIVE 1 (OsBRI1) promotes association withOryza sativa BRI1 ASSOCIATED KINASE 1 (OsBAK1), and inactivates Oryza sativa GLYCOGEN SYNTHASE KINASE3-LIKE GENE 1 (OsGSK1)/BRASSINOSTEROID INSENSITIVE 2 (OsBIN2) via an unknown mechanism. OsGSK1/OsBIN2 phosphorylates OsBZR1, LIC and DLT, and inhibits their activity. OsBZR1 induces the expression of ILI1, but represses the expression of IBH1, TILLER ANGLE INCREASED CONTROLLER (LIC) and DWARF AND LOW TILLERING (DLT), whereas LIC represses the expression of OsBZR1. Like OsBZR1, OsLIC interacts with and is phosphorylated by OsGSK1/OsBIN2 in order to be retained in the cytoplasm. BR treatment inhibits OsGSK1 activity, thereby inducing OsLIC dephosphorylation and transfer of OsLIC to the nucleus. Nuclear-located OsLIC directly binds to the OsBZR1 promoter, and represses OsBZR1 transcriptional activity. Thus, OsLIC and OsBZR1 represent a pair of antagonistic transcription factors that repress each other during transcription, and their repression strength may depend on the BR level. OsBZR1 promotes signaling in the presence of low levels of BR, whereas OsLIC represses BR signaling predominantly at high levels of BR. Thus, the dynamics of BR responses in rice development are modulated by a positive regulator, OsBZR1, and a negative regulator, OsLIC (Wang et al. 2008; Zhang et al. 2012). Additionally, the heterotrimeric G-protein α subunit, encoded by rice d1 mutant defective gene RICE HETEROTRIMERIC G PROTEIN ALPHA 1 (OsRGA1) and mentioned in the GA pathway before, is involved in another BR-signaling pathway, in which D1/RGA1 and TAIHU DWARF 1 (TUD1) act together to mediate BR signaling (Oki et al. 2009). Oryza sativa OVATE family proteins (OsOFP8) and reduced leaf angle 1 (RLA1)/small organ size 1 (SMOS1) functions as a positive regulator. OsOFP8 is interacted with and phosphorylated by OsGSK2/OsBIN2, thus shuttles from nuclear to the cytoplasm and is targeted for proteasomal degradation (Yang et al. 2016). RLA1/SMOS1 can interact with OsBZR1 to enhance its transcriptional activity. OsGSK2/OsBIN2 can interact with and phosphorylate RLA1/SMOS1 to reduce its stability (Qiao et al. 2017). BR-signaling pathway in Arabidopsis is indicated in text
In Arabidopsis, BR directly interacts with BRI1 and BAK1 to form a BRI1–BR–BAK1 complex on the cell surface. Sequential transphosphorylation between BRI1 and BAK1 activates BRI1, which then phosphorylates BSKs/CONSTITUTIVE DIFFERENTIAL GROWTH 1 (CDG1). The active BSKs/CDG1 phosphorylates BSU1, which dephosphorylates and inactivates BRASSINOSTEROID INSENSITIVE 2 (BIN2). BZR1 and BRI1 EMS SUPPRESSOR 1 (BES1)/BRASSINAZOLE RESISTANT 2 (BZR2) are dephosphorylated by PP2A and move into the nucleus to induce expression of PACLOBUTRAZOL-RESISTANT 1 (PRE1), but repress ILI1 BINDING bHLH 1 (IBH1) expression. PRE1, IBH1 and HOMOLOG OF BEE2 INTERACTING WITH IBH1 (HBI1) form an antagonistic cascade to regulate plant growth. SUPPRESSOR OF BRI 1 (SBI1) is involved in the deactivation of BRI1 through methylation of PP2A (Wang et al. 2012; Zhang et al. 2014a) (Fig. 6). BRI1 is negatively feedback regulated by BZR1 and BES1 at the transcriptional level (Sun et al. 2010; Yu et al. 2011), and BRI1 is also negatively feedback regulated by PP2A at the protein level (Di Rubbo et al. 2011; Wu et al. 2011). In addition, there is a lateral organ boundaries (LOB)-BR feedback loop that limits growth in the organ boundaries. LOB is positively regulated by BRs and is expressed in organ boundaries where it activates expression of PHYB ACTIVATION TAGGED SUPPRESSOR 1 (BAS1), which inactivates BR (Diaz 2016).
In rice, OsBRI1 is the counterpart of the Arabidopsis gene BRI1, which is a cell surface receptor kinase and can perceive the extracellular BR signal. Its loss-of-function mutant, d61, was the first identified BR-insensitive mutant in rice and has erect leaves and dwarf culms (Yamamuro et al. 2000). OsBAK1 (also named Oryza sativa SOMATIC EMBRYOGENESIS RECEPTOR KINASE 1 (OsSERK1) can interact with OsBRI1 and overexpression of a truncated intracellular domain of OsBAK1 caused BR hypersensitivity and partially suppressed mutant phenotypes of the Arabidopsis bri1-5 and rice d61-1 genes (Wang et al. 2007; Li et al. 2009). OsGSK1 and GLYCOGEN SYNTHASE KINASE3-LIKE GENE 3 (GSK3)/SHAGGY-LIKE KINASE (GSK2) (united as OsBIN2 in Fig. 6) are homologues to Arabidopsis BRASSINOSTEROID INSENSITIVE 2 (AtBIN2) and act as negative regulators of rice BR signaling and stress responses (Koh et al. 2007; Tong and Chu 2012). Overexpression of OsGSK1 in Arabidopsis results in a similar dwarf phenotype to the gain-of-function mutant bin2-1 in Arabidopsis. The T-DNA knockout mutant of OsGSK1 is more tolerant to heat, cold, salt and drought stresses (Koh et al. 2007). OsBZR1 is a positive regulator of BR signaling and suppression of OsBZR1 by RNAi causes semi-dwarfism, erect leaves and BR-insensitive phenotypes (Bai et al. 2007; Table 2).
OsLIC, which encodes a novel CCCH-type zinc finger protein, has a negative regulatory role on the BR signal. Antisense-mediated suppression of OsLIC results in larger leaves and tillering angles and decreased numbers of seeds, while a gain-of-function mutant of OsLIC leads to erect leaves and reduced BR sensitivity (Wang et al. 2008; Zhang et al. 2012). OsDLT, encoding a plant-specific GRAS family protein, acts downstream to positively regulate BR responses, and mutants of this gene exhibit a compact semi-dwarf stature, dark-green erect leaves, reduced tillering and late flowering. OsBZR1 directly binds to the promoter of the OsDLT gene and suppresses its transcription (Tong et al. 2009). OsDLT can also physically interact with and is phosphorylated by OsGSK2 (Tong et al. 2009). Oryza sativa INCREASE LEAF INCLINATION 1 (OsILI1) and OsIBH1 are likely to regulate rice leaf angle by antagonistically interacting with each other, and OsBZR1 directly binds to their promoter to induce OsILI1, but represses OsIBH1 expression (Zhang et al. 2009; Table 2).
Rice D1/RGA1, known to be involved in GA signaling (Ueguchi-Tanaka et al. 2000; Suharsono et al. 2002), was also suggested to be a BR-signaling component (Wang et al. 2006; Oki et al. 2009). d1, the loss-of-function mutant of D1/RGA1, is less sensitive to BL and produces plants with longitudinally compressed seeds, dense panicles, erect leaves and a dwarf phenotype (Wang et al. 2006; Oki et al. 2009). Moreover, RGA1 may be involved in a distinct BR signaling pathway from OsBZR1 (Oki et al. 2009). Hu et al. (2013) reported that D1 directly interacts with TUD1, which encodes a U-box E3 ubiquitin ligase, to mediate BR signaling. The dwarf phenotype of the tud1 mutant is mainly due to decreased cell proliferation and disorganized cell patterns in the aerial organs. Various analyses by genetic, phenotypic and physiological methods have shown that tud1 is epistatic to d1 and is less sensitive to BR treatment (Hu et al. 2013) (Table 2, Fig. 6). Tanaka et al. (2009) identified a novel BR-induced gene BRASSINOSTEROID UPREGULATED 1 (BU1), which encodes a helix–loop–helix protein. Rice plants that overexpress BU1 have more pronounced bending of the lamina joint and larger grains and plants in which BU1 has been knocked-out by RNAi have erect leaves. BU1 expression is weaker in both d61 and d1 mutants. The BU1 protein is, therefore, a positive regulator of the BR response and participates in two BR signaling pathways through OsBRI1 and RGA1 (Tanaka et al. 2009) (Table 2, Fig. 6). Zhang et al. (2017) showed that a rice small G protein OsPRA2 (homologous with Pea Pra2 protein) plays a repressive role in the BR signaling pathway by inhibiting dephosphorylation of OsBZR1 and thus inhibiting its function. OsPRA2 co-localized with, and directly bound to OsBRI1 at the plasma membrane, which not only inhibited OsBRI1 autophosphorylation, but also inhibited the phosphorylation of OsBAK1 mediated by OsBRI1 (Zhang et al. 2017).
The transcription factor OsOFP8 is an OVATE FAMILY PROTEIN (OFP) that plays a positive role in BR signaling. OsOFP8 is suggested to be phosphorylated by OsGSK2, the counterpart of Arabidopsis BIN2. Phosphorylated OsOFP8 shuttles from nucleus to the cytoplasm and is targeted for proteasomal degradation (Yang et al. 2016). Qiao et al. (2017) reported another gene, REDUCED LEAF ANGLE 1 (RLA1), which is identical to the previously reported SMALL ORGAN SIZE 1 (SMOS1) induced by IAA (Aya et al. 2014) and encodes a transcription factor with an APETALA2 DNA-binding domain. RLA1/SMOS1 functions as a positive regulator in the BR signaling pathway and can interact with OsBZR1 to enhance its transcriptional activity. GSK2 can interact with and phosphorylate RLA1/SMOS1 to reduce its stability.
There are several other genes involved in the BR signaling pathway with unclear functions. Nakagawa et al. (2012) identified a Short-grain 1 (Sg1) dominant mutant from rice activation-tagged lines. Overexpression of SG1 in rice results in a dwarfed plant with broad, dark-green, erect leaves, short grains and insensitivity to brassinolide in the lamina inclination assay (Nakagawa et al. 2012). Another gene, XIAO, is predicted to encode an LRR kinase in rice, and the T-DNA insertion mutant xiao displays dwarfism and erect leaves, together with lower seed set. The smaller stature of xiao plants is due to a reduced cell division rate and the BR response is enhanced in certain roots and the coleoptile, even though the BR levels are reduced at the whole-plant level. Therefore, XIAO may provide a possible connection between BR signaling and cell-cycle regulation during organ growth (Jiang et al. 2012). SDG725, a member of SET DOMAIN GROUP (SDG) proteins, encodes a H3K36 methyltransferase in rice and its knockdown leads to dwarfism, shortened internodes, erect leaves and smaller seeds. SDG725 depletion results in down-regulation of BR signal-related genes, including D11, BRI1 and BU1 (Sui et al. 2012). Oryza sativa BRASSINOLIDE-ENHANCED GENE 3 (OsBLE3) is a BL up-regulated gene and the BL signal that leads to enhanced expression of OsBLE3 is mediated, at least in part, through OsBRI1. In OsBLE3 antisense transgenic rice plants, growth is retarded and internode cell length is about 70% of that in the control lines. Moreover, it indicates that OsBLE3 is involved in cell elongation through dual regulation of BL and IAA (Yang et al. 2006; Table 2).
SL-related pathways
SLs are a group of terpenoid lactones with high structural diversity. SLs were first known for their function in rhizosphere signaling (Akiyama et al. 2005). More recently, they have been identified as inhibitors of shoot branching (Gomez-Roldan et al. 2008; Umehara et al. 2008; Domagalska and Leyser 2011; Ruyter-Spira et al. 2013; Wen et al. 2015), regulating mesocotyl elongation, stem secondary growth, leaf senescence, seed germination and root development (Ruyter-Spira et al. 2013; Hu et al. 2014; Sun et al. 2015; van Zeijl et al. 2015). Deficiencies in SL production and perception lead to greater shoot branching and reduced rice plant height (Domagalska and Leyser 2011; Ruyter-Spira et al. 2013). Shoot branching plays an important role in determining plant architecture. Tillering is a specialized branch in rice, which is crucial to rice productivity, but its molecular regulation is poorly understood (Wen et al. 2015). Studies with branching mutants have identified key players in SL biosynthesis and signaling.
Genes involved in SL biosynthesis
The SL biosynthesis pathway, beginning with all-trans-β-carotene and finishing with ent-20-epi-5-deoxystrigol, is illustrated in Fig. 7 (Harrison et al. 2015). Rice mutants dwarf 27 (d27), high tillering dwarf 1 (htd1)/semi dwarf-t (sd-t)/dwarf 17 (d17), dwarf 10 (d10), which have defective in D27, Oryza sativa CAROTENOID CLEAVAGE DIOXYGENASE 7 (OsCCD7) and Oryza sativa CAROTENOID CLEAVAGE DIOXYGENASE 8 (OsCCD8) genes, respectively, have reduced SL levels, implying roles in the SL biosynthetic pathway (Umehara et al. 2008; Gomez-Roldan et al. 2008; Lin et al. 2009; Table 3). D27 is a β-carotene isomerase, which works upstream of CCD7 and CCD8 in the biosynthetic pathway (Alder et al. 2012; Waters et al. 2012). OsCCD7 and OsCCD8 are homologous to the Arabidopsis MORE AXILLARY GROWTH 3 (MAX3)/CCD7 and MAX4 genes, respectively, which sequentially cleave carotenoid precursors to produce carlactone. A rice cytochrome P450 CYP711 subfamily member (homologous to Arabidopsis MAX1) then acts as a carlactone oxidase to stereoselectively convert carlactone into ent-2′-epi-5-deoxy-strigol (Zhang et al. 2014b; Table 3). In mutant d27 roots, SLs are undetectable in the root secretions, leading to a high tillering and dwarfed phenotype. This mutant is considered SL-defective as applications of SLs can restore the wild-type phenotype (Lin et al. 2009). In htd1 mutants, the removal of axillary buds can increase height, indicating that the dwarf phenotype results from excessive tillering (Zou et al. 2005). d10 is an enhanced branching rice mutant, with reduced apical dominance (Arite et al. 2007).
The strigolactone (SL) biosynthesis pathway from all-trans-β-carotene to ent-20-epi-5-deoxystrigol. The biosynthetic pathway of SLs begins with the isomerization of all-trans-β-carotene to 9-cis-β-carotene, a reaction that is catalyzed by β-carotene isomerase D27 (Lin et al. 2009; Alder et al. 2012). Oxidative cleavage of 9-cis-β-carotene at the 9′, 10′ position is then catalyzed by carotenoid cleavage dioxygenase 7 (CCD7), to produce 9-cis-β-apo-10′-carotenal (Alder et al. 2012). Carotenoid cleavage dioxygenase 7 (CCD8) catalyzes an unusual double oxygenation of 9-cis-b-apo-10′-carotenal to produce an intermediate known as carlactone (Alder et al. 2012). Carlactone is then oxidized by consecutive P450 enzyme oxidations to give ent-2′-epi-5-deoxy-strigol, the presumed precursor of rice SLs: recombinant Arabidopsis Max1 has been shown to form carlactonoic acid (Abe et al. 2014), whereas a recombinant Max1 homologue from Oryza sativa (called carlactone oxidase) has been shown to form ent-2′-epi-5-deoxystrigol directly (Zhang et al. 2014b)
Genes involved in SL signaling
SL signaling requires the hormone-dependent interaction of DWARF 53 (D53) with DWARF 14 (D14), a probable candidate SL receptor, and DWARF 3 (D3), an F-box component of the SCF E3 ubiquitin ligase complex (Jiang et al. 2013) (Fig. 8a). A model of the OsMADS57 (transcription factors of the MADS-domain family)-mediated network for the control of tillering, as proposed by Guo et al. (2013), is shown in Fig. 8b.
Model of the SL signaling pathway. a A proposed SL signaling pathway that involves SL-dependent degradation of the DWARF 53 (D53) repressor, mediated by the D14–D3 complex (Jiang et al. 2013). In the absence of SLs, D53 is stable and recruits TOPLESS (TPL) or TPL-RELATED PROTEINS (TPRs), which are transcriptional co-repressors, and repress downstream responses. In the presence of SLs, perception of SL leads to SCFD3-mediated ubiquitination of D53 and its subsequent degradation by the proteasome system, in a manner that requires D14 and the SCFD3 ubiquitin ligase, which in turn releases the repression of downstream responses (Jiang et al. 2013). b Model of the OsMADS57 (transcription factors of the MADS-domain family)-mediated network for control of tillering (Guo et al. 2013). OsMADS57 is predicted to be a target of the 21-nt noncoding RNA miR444a, and OsMIR444a-regulated OsMADS57, together with Oryza sativa TEOSINTE BRANCHED 1 (OsTB1), which is synonymous with FINE CULM 1 (FC1), target DWARF 14 (D14) to control tillering. OsMADS57 binds to the promoter and directly suppresses D14 expression, and interaction of OsMADS57 with OsTB1 reduces OsMADS57 inhibition of D14 transcription (Guo et al. 2013)
Some genes responsible for the dwarf and multi-tillering phenotypes of rice are involved in the SL signaling pathway (Ishikawa et al. 2005; Zou et al. 2006; Arite et al. 2007; Lin et al. 2009; Table 3). D3 is homologous to the Arabidopsis MAX2/ORESARA 9 (ORE9) and is involved in tillering and shoot branching (Ishikawa et al. 2005). Recently, a rice mutant high-tillering dwarf 2 (htd2) was used to identify the D14/DWARF 88 (D88)/HTD2 gene, which encodes a novel putative rice hydrolase/esterase and is involved in SL perception or downstream metabolism (Liu et al. 2009; Gao et al. 2009; Leyser 2009; Beveridge and Kyozuka 2010). D14-LIKE (D14L), a homolog of D14, acts similarly to inhibit mesocotyl elongation and functions via an unknown SL independent pathway (Yoshida et al. 2012; Kameoka and Kyozuka 2015). Jiang et al. (2013) described a dominant SL-insensitive rice mutant d53, which exhibits a dwarf and prolific tillering phenotype and using this mutant, cloned the D53 gene, which encodes a substrate of the SCFD3 ubiquitination complex and functions as a repressor of SL signaling. Like the d3 mutant, d53 is resistant to exogenous applications of SLs. Xu et al. (2015) found that the rice gene Oryza sativa TEOSINTE BRANCHED 1 (OsTB1)/Oryza sativa FINE CULM 1 (OsFC1) is required for bud inhibition and is transcriptionally regulated in axillary buds in rice. OsTB1/OsFC1 is a downstream component of the SL signaling pathway. Cytokinin (CK) and SL rapidly reduce and increase the expression of OsTB1/OsFC1 in the buds, respectively, implying that FC1 might be a common target for the CK and SL pathways. The rice mutant tb1/fc1 has greater tillering, which is similar to d14 and the activation-tagged mutant osmads57-1, whereas osmads57-1 has a very weak dwarf phenotype. OsMADS57 is predicted to encode a protein containing a MADS-box domain. OsMADS57 overexpression transgenic lines have increased tillering, while OsMADS57 antisense transgenic lines have reduced tillering (Guo et al. 2013; Table 3).
Genes involved in IAA biosynthesis and signaling
IAA is the predominant auxin in plants and plays an important role in many processes during plant growth and development, including cell division, differentiation and elongation, flower and vascular development, tropism and embryogenesis (Vanneste and Friml 2009). The tryptophan biosynthetic pathway is considered to be one of the critical steps in IAA biosynthesis (Zazimalova and Napier 2003). Tryptophan-deficient dwarf1 (tdd1), a rice IAA-deficient mutant, is embryonic lethal, and the regenerated tdd1 plants exhibit dwarfing, narrow leaves and abnormal flowers. TDD1 encodes an anthranilate synthase β-subunit and works upstream of Trp-dependent IAA biosynthesis (Sazuka et al. 2009; Table 4).
Aux/IAA proteins are short-lived transcriptional regulators, which can interact with ARF transcription factors by preventing them from binding to the promoters of auxin-responsive target genes, thus repressing the auxin response (Song et al. 2009). OsIAA1 is a member of Aux/IAA family in rice and its transcription is induced by various phytohormones including IAA. Overexpression of OsIAA1 in rice results in reduced auxin sensitivity but increased sensitivity to BR. Furthermore, OsIAA1-overexpressing transgenic plants have decreased plant height and loose plant architecture. Yeast two-hybrid analysis suggests that OsIAA1 interacts with OsARF. A T-DNA insertion mutant of OsARF1 is less sensitive to BR application and resembles the phenotype of OsIAA1-overexpression plants (Song et al. 2009). Therefore, OsIAA1–OsARF1 could play a critical role in mediating the auxin and BR signal pathways (Table 4).
Genes involved in interactions between the other relevant dwarfing phytohormones
The interplay of phytohormones controls plant growth and developmental processes. Oryza sativa APETALA 2 (OsAP2)-39 encodes an APETALA-2-like transcription factor and overexpression of this gene leads to a dwarfed, lower yielding phenotype. In vitro and in vivo analyses have demonstrated that transcription of the ABA biosynthesis gene Oryza sativa 9-CIS-EPOXYCAROTENOID DIOXYGENASE-1 (OsNCED-I) and the GA catabolism gene EUI is directly controlled by OsAP2-39. OsAP2-39 is, therefore, likely to regulate the ABA/GA balance in rice, which in turn affects plant growth and seed production (Yaish et al. 2010). Qi et al. (2011) also demonstrated that Oryza sativa ERF PROTEIN ASSOCIATED WITH TILLERING AND BRANCHING (OsEATB), a rice AP2/ETHYLENE-RESPONSIVE ELEMENT BINDING FACTOR (ERF) family gene, restricts ethylene-induced enhancement of GA responsiveness during internode elongation, by downregulating a GA biosynthetic gene CPS. Transgenic plants overexpressing OsEATB have a dwarfed phenotype, indicating that the internodal elongation process is suppressed by cross-talk between ETH and GA (Qi et al. 2011). The F-BOX PROTEIN CORONATINE INSENSITIVE 1 (COI1) encodes a component of the SCF E3 ubiquitin ligase and receptor of JA (Xie et al. 1998; Katsir et al. 2008; Fonseca et al. 2009; Yan et al. 2009). The rice genome contains two closely related COI1 genes, OsCOI1a and OsCOI1b, both involved in crosstalk between the JA and GA signaling pathways in defense against biotic and abiotic stresses (Yang and He 2012). Oryza sativa REMORIN 4.1 (OsREM4.1) gene is transcriptionally regulated by ABA and functions as an OsBRI1 substrate and OsSERK1-interacting protein. This inhibits the formation and subsequent activation of the OsBRI1–OsSERK1 receptor complex, explaining the antagonistic interactions between ABA and BRs that are coordinated in rice (Gui et al. 2016). OsGLU1 encodes a putative membrane-bound endo-1,4-β-d-glucanase and plays an important role in plant cell development. The expression of OsGLU1 is induced by GA and BR, suggesting that it is a component of the GA and BR signaling pathways related to cell wall development. The dwarf mutant glu has reduced cell elongation, and altered cell wall composition, affecting internode elongation (Zhou et al. 2006; Table 4).
Genes related to non-hormone pathways
There are many genes controlling plant height from a range of other pathways besides the GA, BR, SL and IAA-related pathways. This includes those relating to cell wall development, the cytosolic glutamine synthetic pathway, RNA editing, cell division, the ubiquitin–proteasome pathway and fatty acid metabolism (Table 4). In addition to dwarfing, mutations to these genes cause other traits, such as a brittle culm, poor fertility, narrow and curled leaves and short spikes.
The cell wall plays an important role in plant architecture and morphogenesis. Oryza sativa CURLED LEAF AND DWARF 1(OsCD1) is a member of the CELLULOSE SYNTHASE-LIKE D (CSLD) subfamily and is involved in cell wall formation. The rice dwarf mutant cd1 has a defective version of this gene, causing a significant reduction in the cellulose and xylose content of the mature culm. Mutant and wild-type plants all exhibit a constitutive response to GA3, suggesting a mode of action other than phytohormone signal transduction (Luan et al. 2011). Tanaka et al. (2003) found three CELLULOSE SYNTHASE CATALYTIC SUBUNIT (CesA) genes (OsCesA4, OsCesA7 and OsCesA9), and mutation of these genes by Tos17 retrotransposon insertion resulted in a brittle/thin culm, semi-dwarf and low-fertility phenotype. Furthermore, it is indicated that there are no functional overlaps among these genes; therefore, they may form a complex involved in the synthesis of the cell wall (Tanaka et al. 2003; Table 4).
Oryza sativa GLUTAMINE SYNTHETASE 1;1 (OsGS1;1) gene encodes a cytosolic glutamine synthetase, and Tos17 retrotransposon insertional mutants have retarded growth rates, smaller panicles and inhibited grain filling (Tabuchi et al. 2005). A rice mutant opaque and growth retardation 1 (ogr1), defective in seven specific RNA editing sites on five mitochondrial transcripts, displays dwarfism and sterility phenotypes. ogr1, the T-DNA insertion mutant of the OGR1 gene, encodes a pentatricopeptide repeat protein containing a DYW motif (Kim et al. 2009). The phenotype of the dwarf mutant sword shape dwarf 1 (ssd1) exhibits defects in the elongation of various organs, different from typical GA- or BR-related dwarf phenotypes. SSD1 encodes a plant-specific protein of unknown function and probably controls plant growth by regulating cell division (Asano et al. 2010). The single recessive gene, dwarf 162(t) (d162(t)), causes dwarfing of rice and has been mapped to chromosome 3. The candidate gene is considered to encode a U-box domain containing protein, which plays an important role in maintaining cellular homeostasis (Zhang et al. 2011). Oryza sativa FATTY ACID DESATURASE (OsFAD8) encodes rice chloroplast ω-3 fatty acid desaturase and its expression is induced at low temperatures. OsFAD8 T-DNA knockout plants are shorter and have lower levels of trienoic fatty acids after exposure to cold temperatures. OsFAD8 may, therefore, have a role in cold tolerance (Liu et al. 2012). The rice semi-dwarf mutant semi dwarf 37 (sd37), a novel GA/BR independent mutant, is defective in CYP96B4, which is an important regulator of plant growth. sd37 has smaller leaves, panicles and seeds due to a decrease in cell number, especially in the shoots (Zhang et al. 2014c; Table 4).
Genes involved in uncharacterized pathways
The Oryza sativa HOMEOBOX 15 (OSH15) gene is a member of the knotted1-type homeobox gene family and plays a role in the development of internodes by influencing cell division and elongation. Internodes of the OSH15 loss-of-function mutant d6 have abnormally shaped epidermal and hypodermal cells and unusually arranged small vascular bundles (Sato et al. 1999). The high-tillering dwarf 3 (htd3) mutant, which is caused by the single recessive gene HTD3, has a greater number of tillers and a shorter culm. HTD3 has been mapped within 3100 kb of the centromere of chromosome 12, but the identity of this gene has yet to be uncovered (Zhang et al. 2011). Oryza sativa LISSENCEPHALY TYPE-1-LIKE 1 (OsLIS-L1) encodes a protein containing the WD40 motif and plays an important role in elongation of the first internode and male gametophyte formation in rice. The oslis-l1 mutant has an abnormal developmental phenotype, including semi-dwarfing, a shorter panicle length and reduced male fertility (Gao et al. 2012). The DWARF TILLER 1 (DWT1) locus regulates uniform growth of the main shoot and tillers in rice. The main shoot of the dwt1 mutant is of nearly normal height at maturity but the tillers are dwarfed. Thirty-one putative open reading frames (ORFs) have been identified in the 197-kb genomic region covering the DWT1 locus, but none are predicted to be involved in stem elongation or plant architecture. DWT1 may represent a component of an unknown genetic pathway controlling the elongation of tiller culms in rice (Wang et al. 2013a, b; Table 4).
Future perspectives
Most dwarf genes are connected to phytohormone functions and there is a potential for farmers to control crop height using hormone antagonists or analogues. However, the application of chemicals to food crops may not be desirable in the future. Another strategy is to modify hormone content and homeostasis by genetic engineering or pyramiding breeding. Several studies have highlighted the huge potential of modulating expression of GA and BL biosynthesis and signaling genes in plant breeding (Divi and Krishna 2009; Huang et al. 2010). Tillering is crucial to rice productivity. Although there is limited evidence for SL-mediated tillering effects on yield traits, the strigolactone pathway is not fully elucidated and it could potentially be explored to improve productivity. Potential of modulating expression of genes involved in other hormones and non-hormones in plant breeding also remains to be explored.
How can dwarfing genes be better utilized in practice? In rice, for example, plant breeders have mainly utilized weak alleles of dwarf genes which are relatively free of adverse traits. osdwarf4-1 has a slightly dwarfed stature and more erect leaves than the wild-type plant but no other phenotypic differences, which helps to increase grain yield, especially at a high planting density (Sakamoto et al. 2006). In more sophisticated approaches, dwarfing genes could be driven by tissue-specific promoters to ensure no undesirable pleiotropic effects. The advantages of targeted expression have been clearly demonstrated in rice (Sakamoto et al. 2003). When the rice GA-deactivation enzyme gene GA2ox is driven by the strong and constitutive actin promoter, the transgenic plants show severe dwarfing and significantly reduced grain set (Sakai et al. 2003). To avoid this problem, the GA2ox gene was coupled to the promoter of OsGA3ox2 to ensure that GA deactivation only occurred in the vegetative tissues (Sakamoto et al. 2003). The outcome was semi-dwarfed plants with normal grain set, similar to plants containing the sd1 gene. Alternatively, coupling the DWARF4 gene to a vegetative tissue-specific promoter led to plants growing more vigorously than the wild-type with taller stature, better grain filling and larger seeds, leading to improved yields (Wu et al. 2008).
To better utilize crop height-related genes in practice, the following strategies are suggested: (1) As several metabolic pathways involving many genes control crop height, once new genes affecting crop height are identified, it will be necessary to clarify their functions by placing them in specific pathways or networks that regulate phytohormones or other developmental processes, including how they control crop height and whether they control other characteristics. (2) It will be necessary to learn how they interact with their up and downstream genes and determine the phenotype of double mutants. It is crucial to explore the key alleles in the metabolic pathways by assessing their impact on the phenotype in order to manipulate one or more genes to optimize the regulation of crop height. (3) It will be necessary to determine whether there is crosstalk between different hormone pathways, specifically whether common downstream targets exist.
Based on the above information, crop height genes can be divided into two categories depending on whether they have pleiotropic effects or not. If pleiotropic effects are beneficial, such as erect leaves, the genes can be used directly. If other effects are adverse, three strategies are suggested: (1) screen alleles of dwarf genes to ameliorate any pleiotropic adverse effects; (2) employ genome editing strategies for target specificity; (3) limit the expression of the dwarf genes to specific tissues using specific promoters, to avoid undesirable pleiotropic effects; (4) gradually reduce any adverse traits associated with dwarfing by crossing or back-crossing, using molecular markers to monitor inheritance of the dwarfing gene.
Once genes of interest in dwarfism and yield are identified, functional markers can be developed. This could be used to make a genotyping chip for crop height breeding, or as the basis for genome selection breeding. Crop height is one of the most important traits in crop ideotype breeding and each crop height-related gene behaves differently under certain environmental conditions (Botwright et al. 2005; Butler et al. 2005; Chapman et al. 2007; Rebetzke et al. 2012). Therefore, in order to use these genes appropriately, the impact of environmental factors such as light, temperature, water and nutrition needs to be ascertained. This is in agreement with the opinion that rice varieties with super high yield should have different ideal architecture characteristics in different ecological areas (Huang et al. 2009).
Rice is a model plant species, and research to investigate dwarfing will have implications for all crop species. It is also apparent that there are subtle differences between plant species in the metabolic pathways that control crop height, and these difference are expected to become more clear in future work. A multi-disciplinary approach involving plant genomics, physiology, metabolomics and breeding will be needed to untangle the complex genetic and environmental interactions that determine plant architecture. These investigations are so important for our looking ahead a different ‘green revolution’.
Author contribution statement
FL and XZ participated in the writing of this review, and they both performed the modification of manuscript. DF and GW edited and gave suggestions for this review. PW collected important background information and arranged the literature. XL and XY helped to make tables and figures.
References
Abe S, Sado A, Tanaka K, Kisugi T, Asami K, Ota S, Kim HI, Yoneyama K, Xie X, Ohnishi T, Seto Y, Yamaguchi S, Akiyama K, Yoneyama K, Nomura T (2014) Carlactone is converted to carlactonoic acid by MAX1 in Arabidopsis and its methyl ester can directly interact with AtD14 in vitro. Proc Nati Acad Sci USA 11:18084–18089. doi:10.1073/pnas.1410801111
Achard P, Genschik P (2009) Releasing the brakes of plant growth: how GAs shutdown DELLA proteins. J Exp Bot 60:1085–1092. doi:10.1093/jxb/ern301
Akiyama K, Matsuzaki K, Hayashi H (2005) Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435:824–827. doi:10.1038/nature03608
Alder A, Jamil M, Marzorati M, Bruno M, Vermathen M, Bigler P, Ghisla S, Bouwmeester H, Beyer P, Al-Babili S (2012) The path from beta-carotene to carlactone, a strigolactone-like plant hormone. Science (New York, N Y) 335:1348–1351. doi:10.1126/science.1218094
Arite T, Iwata H, Ohshima K, Maekawa M, Nakajima M, Kojima M, Sakakibara H, Kyozuka J (2007) DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice. Plant J 51:1019–1029. doi:10.1111/j.1365-313X.2007.03210.x
Asano K, Hirano K, Ueguchitanaka M, Angelesshim RB, Komura T, Satoh H, Kitano H, Matsuoka M, Ashikari M (2009) Isolation and characterization of dominant dwarf mutants, Slr1-d, in rice. Mol Genet Genom 281:223–231
Asano K, Miyao A, Hirochika H, Kitano H, Matsuoka M, Ashikari M (2010) SSD1, which encodes a plant-specific novel protein, controls plant elongation by regulating cell division in rice. Proc Jpn Acad B Phys 86:265–273
Ashikari M, Wu J, Yano M, Sasaki T, Yoshimura A (1999) Rice gibberellin-insensitive dwarf mutant gene Dwarf 1 encodes the alpha-subunit of GTP-binding protein. Proc Nati Acad Sci USA 96:10284–10289
Ashikari M, Sasaki A, Ueguchi-Tanaka M, Itoh H, Nishimura A, Datta S, Ishiyama K, Saito T, Kobayashi M, Khush GS (2002) Loss-of-function of a rice gibberellin biosynthetic gene, GA20 oxidase (GA20ox-2), led to the rice’ green revolution’. Breed Sci 52:143–150
Ayano M, Kani T, Kojima M, Sakakibara H, Kitaoka T, Kuroha T, Angelesshim RB, Kitano H, Nagai K, Ashikari M (2015) Gibberellin biosynthesis and signal transduction is essential for internode elongation in deepwater rice. Plant Cell Environ 37:2313–2324
Bai MY, Zhang LY, Gampala SS, Zhu SW, Song WY, Chong K, Wang ZY (2007) Functions of OsBZR1 and 14-3-3 proteins in brassinosteroid signaling in rice. Proc Nati Acad Sci USA 104:13839–13844. doi:10.1073/pnas.0706386104
Begonia GB, Begonia MT (2007) Plant photosynthetic production as controlled by leaf growth, phenology, and behavior. Photosynthetica 45:321–333
Beharav A, Cahaner A, Pinthus M (1998) Genetic correlations between culm length, grain yield and seedling elongation within tall (rht1) and semi-dwarf (Rht1) spring wheat (Trificum aestivum L.). Eur J Agron 9:35–40
Beveridge CA, Kyozuka J (2010) New genes in the strigolactone-related shoot branching pathway. Curr Opin Plant Biol 13:34–39. doi:10.1016/j.pbi.2009.10.003
Bolle C (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218:683–692. doi:10.1007/s00425-004-1203-z
Bonnett D, Ellis M, Rebetzke G, Condon A, Spielmeyer W, Richards R (2001) Dwarfing genes in Australian wheat—present and future. In: Proceedings of the 10th Australian wheat breeders assembly, Mildura, Australia, pp 154–157
Botwright TL, Rebetzke GJ, Condon AG, Richards RA (2005) Influence of the gibberellin-sensitive Rht8 dwarfing gene on leaf epidermal cell dimensions and early vigour in wheat (Triticum aestivum L.). Ann Bot 95:631–639. doi:10.1093/aob/mci069
Butany W, Bhattacharyya R, Daiya L (1959) Inheritance of dwarf character in rice and its interrelationship with the occurrence of anthocyanin pigment in various plant parts. Indian J Genet Plant Breed 19:64–72
Butler JD, Byrne PF, Mohammadi V, Chapman PL, Haley SD (2005) Agronomic performance of Rht alleles in a spring wheat population across a range of moisture levels. Crop Sci 45:939–947
Chang T, Zuro C, Marciano R, Loresto G (1984) Semidwarfing genes in rice germplasm collection. Rice Genet Newsl 1:93–94
Chapman SC, Mathews KL, Trethowan RM, Singh RP (2007) Relationships between height and yield in near-isogenic spring wheats that contrast for major reduced height genes. Euphytica 157:391–397
Choe S, Fujioka S, Noguchi T, Takatsuto S, Yoshida S, Feldmann KA (2001) Overexpression of DWARF4 in the brassinosteroid biosynthetic pathway results in increased vegetative growth and seed yield in Arabidopsis. Plant J 26:573–582
Clouse SD (2002) Brassinosteroid signal transduction: clarifying the pathway from ligand perception to gene expression. Mol Cell 10:973–982
Dai M, Zhao Y, Ma Q, Hu Y, Hedden P, Zhang Q, Zhou DX (2007) The rice YABBY1 gene is involved in the feedback regulation of gibberellin metabolism. Plant Physiol 144:121–133. doi:10.1104/pp.107.096586
Davière JM, Achard P (2013) Gibberellin signaling in plants. Development 140:1147–1151
Di Rubbo RS, Irani NG, Russinova E (2011) PP2A phosphatases: the “on–off” regulatory switches of brassinosteroid signaling. Sci Signal 4:pe25
Diaz J (2016) Characterization of the role of LOB-DOMAIN genes and brassinosteroids in rice architecture. UC Riverside Electronic Theses and Dissertations
Divi UK, Krishna P (2009) Brassinosteroid: a biotechnological target for enhancing crop yield and stress tolerance. New Biotechnol 26:131–136. doi:10.1016/j.nbt.2009.07.006
Domagalska MA, Leyser O (2011) Signal integration in the control of shoot branching. Nat Rev Mol Cell Biol 12:211–221. doi:10.1038/nrm3088
Emerson RA, Beadle GW, Fraser AC (1935) A summary of linkage studies in maize. Cornell Univ Agric Exp Stn Mem 180:1–83
Fischer RA, Quail KJ (1990) The effect of major dwarfing genes on yield potential in spring wheats. Euphytica 46:51–56. doi:10.1007/bf00057618
Flintham JE, Borner A, Worland AJ, Gale MD (1997) Optimizing wheat grain yield: effects of Rht (gibberellin-insensitive) dwarfing genes. J Agric Sci 128:11–25
Foisset N, Delourme R, Barret P, Renard M (1995) Molecular tagging of the dwarf BREIZH (Bzh) gene in Brassica napus. Theor Appl Genet 91:756–761. doi:10.1007/bf00220955
Fonseca S, Chini A, Hamberg M, Adie B, Porzel A, Kramell R, Miersch O, Wasternack C, Solano R (2009) (+)-7-iso-Jasmonoyl-l-isoleucine is the endogenous bioactive jasmonate. Nat Chem Biol 5:344–350
Foster AE, Thompson AP (1987) Effects of a semidwarf gene from Jotun on agronomic and quality traits of barley. In: Yasuda S, Kanishi T (eds) Proceedings of the 5th international barley genetics symposium, 1986. Sanyo, Okayama, pp 979–982
Fujioka S, Sakurai A (1997) Brassinosteroids. Nat Prod Rep 14:1–10
Fujioka S, Yokota T (2003) Biosynthesis and metabolism of brassinosteroids. Annu Rev Plant Biol 54:137–164
Fukazawa J, Teramura H, Murakoshi S, Nasuno K, Nishida N, Ito T, Yoshida M, Kamiya Y, Yamaguchi S, Takahashi Y (2014) DELLAs function as coactivators of GAI-ASSOCIATED FACTOR1 in regulation of gibberellin homeostasis and signaling in Arabidopsis. Plant Cell 26:2920–2938
Fukazawa J, Ito T, Kamiya Y, Yamaguchi S, Takahashi Y (2015) Binding of GID1 to DELLAs promotes dissociation of GAF1 from DELLA in GA dependent manner. Plant Sig Beha 10:e1052923-3
Futsuhara Y, Kikuchi F (1997) Inheritance of morphological characters. 2. Inheritance of dwarfism. Sci Rice Plant 3:300–308
Gao Z, Qian Q, Liu X, Yan M, Feng Q, Dong G, Liu J, Han B (2009) Dwarf 88, a novel putative esterase gene affecting architecture of rice plant. Plant Mol Biol 71:265–276. doi:10.1007/s11103-009-9522-x
Gao X, Chen Z, Zhang J, Li X, Chen G, Li X, Wu C (2012) OsLIS-L1 encoding a lissencephaly type-1-like protein with WD40 repeats is required for plant height and male gametophyte formation in rice. Planta 235:713–727. doi:10.1007/s00425-011-1532-7
Gomez-Roldan V, Fermas S, Brewer PB, Puech-Pagès V, Dun EA, Pillot J-P, Letisse F, Matusova R, Danoun S, Portais J-C (2008) Strigolactone inhibition of shoot branching. Nature 455:189–194
Gomi K, Sasaki A, Itoh H, Ueguchi-Tanaka M, Ashikari M, Kitano H, Matsuoka M (2004) GID2, an F-box subunit of the SCF E3 complex, specifically interacts with phosphorylated SLR1 protein and regulates the gibberellin-dependent degradation of SLR1 in rice. Plant J 37:626–634
Grove MD, Spencer GF, Rohwedder WK, Mandava N, Worley JF, Warthen JD, Steffens GL, Flippen-Anderson JL, Cook JC (1979) Brassinolide, a plant growth-promoting steroid isolated from Brassica napus pollen. Nature 281:216–217
Gui J, Zheng S, Liu C, Shen J, Li J, Li L (2016) OsREM4.1 interacts with OsSERK1 to coordinate the interlinking between abscisic acid and brassinosteroid signaling in rice. Dev Cell 38:201–213
Guo S, Xu Y, Liu H, Mao Z, Zhang C, Ma Y, Zhang Q, Meng Z, Chong K (2013) The interaction between OsMADS57 and OsTB1 modulates rice tillering via DWARF14. Nat Commun 4:1566. doi:10.1038/ncomms2542
Harberd NP (2003) Botany. Relieving DELLA restraint. Science (New York, N Y) 299:1853–1854. doi:10.1126/science.1083217
Harberd NP, Belfield E, Yasumura Y (2009) The angiosperm gibberellin-GID1-DELLA growth regulatory mechanism: how an “inhibitor of an inhibitor” enables flexible response to fluctuating environments. Plant Cell 21:1328–1339. doi:10.1105/tpc.109.066969
Hargrove TR, Cabanilla VL (1979) The impact of semidwarf varieties on Asian rice-breeding programs. BioScience 29:731–735
Harrison PJ, Newgas SA, Descombes F, Shepherd SA, Thompson AJ, Bugg TD (2015) Biochemical characterization and selective inhibition of β-carotene cis–trans isomerase D27 and carotenoid cleavage dioxygenase CCD8 on the strigolactone biosynthetic pathway. FEBS J 282:3986–4000
He Z, Shen Z (1991) Inheritance of panicle exsertion and improvement of male sterile line in rice. Chin J Rice Sci 5:1–6
Hedden Peter (2003) The genes of the green revolution. Trends Genet 19:5–9
Hedden P (2010) Green revolution genes. Rothamsted Research, Harpenden, Hertfordshire, UK Essay 20.3
Hedden P, Kamiya Y (1997) GIBBERELLIN BIOSYNTHESIS: enzymes, genes and their regulation. Annu Rev Plant Physiol Plant Mol Biol 48:431–460. doi:10.1146/annurev.arplant.48.1.431
Hedden P, Phillips AL (2000) Gibberellin metabolism: new insights revealed by the genes. Trends Plant Sci 5:523–530
Hedden P, Thomas SG (2012) Gibberellin biosynthesis and its regulation. Biochem J 444:11–25
Hirano K, Asano K, Tsuji H, Kawamura M, Mori H, Kitano H, Ueguchi-Tanaka M, Matsuoka M (2010) Characterization of the molecular mechanism underlying gibberellin perception complex formation in rice. Plant Cell 22:2680–2696. doi:10.1105/tpc.110.075549
Hirano K, Kouketu E, Katoh H, Aya K, Ueguchi-Tanaka M, Matsuoka M (2012) The suppressive function of the rice DELLA protein SLR1 is dependent on its transcriptional activation activity. Plant J 71:443–453. doi:10.1111/j.1365-313X.2012.05000.x
Hong Z, Ueguchi-Tanaka M, Shimizu-Sato S, Inukai Y, Fujioka S, Shimada Y, Takatsuto S, Agetsuma M, Yoshida S, Watanabe Y, Uozu S, Kitano H, Ashikari M, Matsuoka M (2002) Loss-of-function of a rice brassinosteroid biosynthetic enzyme, C-6 oxidase, prevents the organized arrangement and polar elongation of cells in the leaves and stem. Plant J 32:495–508
Hong Z, Ueguchi-Tanaka M, Umemura K, Uozu S, Fujioka S, Takatsuto S, Yoshida S, Ashikari M, Kitano H, Matsuoka M (2003) A rice brassinosteroid-deficient mutant, ebisu dwarf (d2), is caused by a loss of function of a new member of cytochrome P450. Plant Cell 15:2900–2910. doi:10.1105/tpc.014712
Hong Z, Ueguchi-Tanaka M, Fujioka S, Takatsuto S, Yoshida S, Hasegawa Y, Ashikari M, Kitano H, Matsuoka M (2005) The Rice brassinosteroid-deficient dwarf2 mutant, defective in the rice homolog of Arabidopsis DIMINUTO/DWARF1, is rescued by the endogenously accumulated alternative bioactive brassinosteroid, dolichosterone. Plant Cell 17:2243–2254. doi:10.1105/tpc.105.030973
Hu X, Qian Q, Xu T, Zhang Y, Dong G, Gao T, Xie Q, Xue Y (2013) The U-box E3 ubiquitin ligase TUD1 functions with a heterotrimeric G alpha subunit to regulate brassinosteroid-mediated growth in rice. PLoS Genet 9:e1003391. doi:10.1371/journal.pgen.1003391
Hu Z, Yamauchi T, Yang J, Jikumaru Y, TsuchidaMayama T, Ichikawa H, Takamure I, Nagamura Y, Tsutsumi N, Yamaguchi S (2014) Strigolactone and cytokinin act antagonistically in regulating rice mesocotyl elongation in darkness. Plant Cell Physiol 55:30–41
Huang X, Qian Q, Liu Z, Sun H, He S, Luo D, Xia G, Chu C, Li J, Fu X (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41:494–497
Huang J, Tang D, Shen Y, Qin B, Hong L, You A, Li M, Wang X, Yu H, Gu M, Cheng Z (2010) Activation of gibberellin 2-oxidase 6 decreases active gibberellin levels and creates a dominant semi-dwarf phenotype in rice (Oryza sativa L.). J Genet Genom 37:23–36. doi:10.1016/s1673-8527(09)60022-9
Ikeda A, Ueguchi-Tanaka M, Sonoda Y, Kitano H, Koshioka M, Futsuhara Y, Matsuoka M, Yamaguchi J (2001) slender rice, a constitutive gibberellin response mutant, is caused by a null mutation of the SLR1 gene, an ortholog of the height-regulating gene GAI/RGA/RHT/D8. Plant Cell 13:999–1010
Ishikawa S, Maekawa M, Arite T, Onishi K, Takamure I, Kyozuka J (2005) Suppression of tiller bud activity in tillering dwarf mutants of rice. Plant Cell Physiol 46:79–86. doi:10.1093/pcp/pci022
Itoh H, Ueguchi-Tanaka M, Sentoku N, Kitano H, Matsuoka M, Kobayashi M (2001) Cloning and functional analysis of two gibberellin 3 beta -hydroxylase genes that are differently expressed during the growth of rice. Proc Nati Acad Sci USA 98:8909–8914. doi:10.1073/pnas.141239398
Itoh H, Tatsumi T, Sakamoto T, Otomo K, Toyomasu T, Kitano H, Ashikari M, Ichihara S, Matsuoka M (2004) A rice semi-dwarf gene, Tan-Ginbozu (D35), encodes the gibberellin biosynthesis enzyme, ent-kaurene oxidase. Plant Mol Biol 54:533–547. doi:10.1023/b:plan.0000038261.21060.47
Iwata N, Satoh H, Omura T (1977) Linkage studies in rice. Linkage groups for 6 genes newly described. Jpn J Breed 27:250–251
Iwata N, Takamure I, Wu H, Siddinq E, Rutger J (1995) List of genes for various traits (with chromosome and main literature). Rice Genet Newsl 12:61–93
Jiang Y, Bao L, Jeong SY, Kim SK, Xu C, Li X, Zhang Q (2012) XIAO is involved in the control of organ size by contributing to the regulation of signaling and homeostasis of brassinosteroids and cell cycling in rice. Plant J 70:398–408. doi:10.1111/j.1365-313X.2011.04877.x
Jiang L, Liu X, Xiong G, Liu H, Chen F, Wang L, Meng X, Liu G, Yu H, Yuan Y, Yi W, Zhao L, Ma H, He Y, Wu Z, Melcher K, Qian Q, Xu HE, Wang Y, Li J (2013) DWARF 53 acts as a repressor of strigolactone signalling in rice. Nature 504:401–405. doi:10.1038/nature12870
Kameoka H, Kyozuka J (2015) Downregulation of rice DWARF 14 LIKE suppress mesocotyl elongation via a strigolactone independent pathway in the dark. J Genet Genom 42:119–124
Katsir L, Schilmiller AL, Staswick PE, He SY, Howe GA (2008) COI1 is a critical component of a receptor for jasmonate and the bacterial virulence factor coronatine. Proc Nat Acad Sci 105:7100–7105
Khush GS (1999) Green revolution: preparing for the 21st century. Genome 42:646–655
Kim SR, Yang JI, Moon S, Ryu CH, An K, Kim KM, Yim J, An G (2009) Rice OGR1 encodes a pentatricopeptide repeat-DYW protein and is essential for RNA editing in mitochondria. Plant J 59:738–749. doi:10.1111/j.1365-313X.2009.03909.x
Kinoshita T (1995) Report of committee on gene symbolization, nomenclature and linkage groups. Rice Genet Newsl 12:9–153
Koh S, Lee SC, Kim MK, Koh JH, Lee S, An G, Choe S, Kim SR (2007) T-DNA tagged knockout mutation of rice OsGSK1, an orthologue of Arabidopsis BIN2, with enhanced tolerance to various abiotic stresses. Plant Mol Biol 65:453–466. doi:10.1007/s11103-007-9213-4
Kovi MR, Zhang Y, Yu S, Yang G, Yan W, Xing Y (2011) Candidacy of a chitin-inducible gibberellin-responsive gene for a major locus affecting plant height in rice that is closely linked to green revolution gene sd1. Theor Appl Genet 123:705–714. doi:10.1007/s00122-011-1620-x
Kwon M, Fujioka S, Jeon JH, Kim HB, Takatsuto S, Yoshida S, An CS, Choe S (2005) A double mutant for the CYP85A1 and CYP85A2 genes of Arabidopsis exhibits a brassinosteroid dwarf phenotype. J Plant Biol 48:237–244
Lawit SJ, Wych HM, Xu D, Kundu S, Tomes DT (2010) Maize DELLA proteins dwarf plant8 and dwarf plant9 as modulators of plant development. Plant Cell Physiol 51:1854–1868
Lee DJ, Zeevaart JA (2005) Molecular cloning of GA 2-oxidase3 from spinach and its ectopic expression in Nicotiana sylvestris. Plant Physiol 138:243–254. doi:10.1104/pp.104.056499
Leyser O (2009) The control of shoot branching: an example of plant information processing. Plant Cell Environ 32:694–703. doi:10.1111/j.1365-3040.2009.01930.x
Li JM, Yuan LP (2000) Hybrid rice: genetics, breeding, and seed production. Plant Breed Rev 17:15–158
Li F, Asami T, Wu X, Tsang EW, Cutler AJ (2007) A putative hydroxysteroid dehydrogenase involved in regulating plant growth and development. Plant Physiol 145:87–97. doi:10.1104/pp.107.100560
Li M, Xiong G, Li R, Cui J, Tang D, Zhang B, Pauly M, Cheng Z, Zhou Y (2009) Rice cellulose synthase-like D4 is essential for normal cell-wall biosynthesis and plant growth. Plant J 60:1055–1069. doi:10.1111/j.1365-313X.2009.04022.x
Li H, Jiang L, Youn JH, Sun W, Cheng Z, Jin T, Ma X, Guo X, Wang J, Zhang X, Wu F, Wu C, Kim SK, Wan J (2013) A comprehensive genetic study reveals a crucial role of CYP90D2/D2 in regulating plant architecture in rice (Oryza sativa). New Phytol 200:1076–1088. doi:10.1111/nph.12427
Li H, Chen G, Yan W (2015) Molecular Characterization of barley 3H semi-dwarf genes. PLoS One 10:e0120558
Lin H, Wang R, Qian Q, Yan M, Meng X, Fu Z, Yan C, Jiang B, Su Z, Li J, Wang Y (2009) DWARF27, an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth. Plant Cell 21:1512–1525. doi:10.1105/tpc.109.065987
Liu B, Wu Y, Fu X, Qian Q (2008) Characterizations and molecular mapping of a novel dominant semi-dwarf gene Sdd (t) in rice (Oryza sativa). Plant Breed 127:125–130
Liu W, Wu C, Fu Y, Hu G, Si H, Zhu L, Luan W, He Z, Sun Z (2009) Identification and characterization of HTD2: a novel gene negatively regulating tiller bud outgrowth in rice. Planta 230:649–658. doi:10.1007/s00425-009-0975-6
Liu Y, Xu Y, Xiao J, Ma Q, Li D, Xue Z, Chong K (2011) OsDOG, a gibberellin-induced A20/AN1 zinc-finger protein, negatively regulates gibberellin-mediated cell elongation in rice. J Plant Physiol 168:1098–1105. doi:10.1016/j.jplph.2010.12.013
Liu HL, Yin ZJ, Xiao L, Xu YN, le Qu Q (2012) Identification and evaluation of omega-3 fatty acid desaturase genes for hyperfortifying alpha-linolenic acid in transgenic rice seed. J Exp Bot 63:3279–3287. doi:10.1093/jxb/ers051
Lo SF, Yang SY, Chen KT, Hsing YI, Zeevaart JA, Chen LJ, Yu SM (2008) A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 20:2603–2618. doi:10.1105/tpc.108.060913
Luan W, Liu Y, Zhang F, Song Y, Wang Z, Peng Y, Sun Z (2011) OsCD1 encodes a putative member of the cellulose synthase-like D sub-family and is essential for rice plant architecture and growth. Plant Biotechnol J 9:513–524. doi:10.1111/j.1467-7652.2010.00570.x
Luo A, Qian Q, Yin H, Liu X, Yin C, Lan Y, Tang J, Tang Z, Cao S, Wang X, Xia K, Fu X, Luo D, Chu C (2006) EUI1, encoding a putative cytochrome P450 monooxygenase, regulates internode elongation by modulating gibberellin responses in rice. Plant Cell Physiol 47:181–191. doi:10.1093/pcp/pci233
Marcia MP, Zhou XR, Zhu QH, Dennis ES, Upadhyaya NM (2005) Isolation and characterization of a Ds-tagged rice (Oryza sativa L.) GA-responsive dwarf mutant defective in an early step of the gibberellin biosynthesis pathway. Plant Cell Rep 23:819–833. doi:10.1007/s00299-004-0896-6
Mcintosh R, Hart G, Gale M (1988) Catalogue of gene symbols for wheat : 1988 supplement. Cereal Res Commun 121–135
Miralles D, Slafer G (1995) Yield, biomass and yield components in dwarf, semi-dwarf and tall isogenic lines of spring wheat under recommended and late sowing dates. Plant Breed 114:392–396
Mitsunaga S, Tashiro T, Yamaguchi J (1994) Identification and characterization of gibberellin-insensitive mutants selected from among dwarf mutants of rice. TAG Theor Appl Genet 87:705–712. doi:10.1007/bf00222896
Monna L, Kitazawa N, Yoshino R, Suzuki J, Masuda H, Maehara Y, Tanji M, Sato M, Nasu S, Minobe Y (2002) Positional cloning of rice semi-dwarfing gene, sd-1: rice “green revolution gene” encodes a mutant enzyme involved in gibberellin synthesis. DNA Res 9:11–17
Mori M, Nomura T, Ooka H, Ishizaka M, Yokota T, Sugimoto K, Okabe K, Kajiwara H, Satoh K, Yamamoto K, Hirochika H, Kikuchi S (2002) Isolation and characterization of a rice dwarf mutant with a defect in brassinosteroid biosynthesis. Plant Physiol 130:1152–1161. doi:10.1104/pp.007179
Muangprom A, Osborn TC (2004) Characterization of a dwarf gene in Brassica rapa, including the identification of a candidate gene. Theor Appl Genet 108:1378–1384. doi:10.1007/s00122-003-1551-2
Murai M, Shinbashi N, Kusutani A, Hirose S, Takamure I, Kinoshita T (1990) Identification of dwarf genes in rice lines, Yukara dwarf and Fukei 71, and its response to environmental factors. Jpn J Breed 40:33–45
Nakagawa H, Tanaka A, Tanabata T, Ohtake M, Fujioka S, Nakamura H, Ichikawa H, Mori M (2012) Short grain1 decreases organ elongation and brassinosteroid response in rice. Plant Physiol 158:1208–1219. doi:10.1104/pp.111.187567
Neuffer MG, Coe EH, Wessler SR (1997) Mutants of maize. Cold Spring Harbor Laboratory Press, New York, pp 1–255
Ogawa S, Toyomasu T, Yamane H, Murofushi N, Ikeda R, Morimoto Y, Nishimura Y, Omori T (1996) A step in the biosynthesis of gibberellins that is controlled by the mutation in the semi-dwarf rice cultivar Tan-Ginbozu. Plant Cell Physiol 37:363–368
Ogawa M, Kusano T, Katsumi M, Sano H (2000) Rice gibberellin-insensitive gene homolog, OsGAI, encodes a nuclear-localized protein capable of gene activation at transcriptional level. Gene 245:21–29
Ohnishi T, Szatmari AM, Watanabe B, Fujita S, Bancos S, Koncz C, Lafos M, Shibata K, Yokota T, Sakata K, Szekeres M, Mizutani M (2006) C-23 hydroxylation by Arabidopsis CYP90C1 and CYP90D1 reveals a novel shortcut in brassinosteroid biosynthesis. Plant Cell 18:3275–3288. doi:10.1105/tpc.106.045443
Ohnishi T, Godza B, Watanabe B, Fujioka S, Hategan L, Ide K, Shibata K, Yokota T, Szekeres M, Mizutani M (2012) CYP90A1/CPD, a brassinosteroid biosynthetic cytochrome P450 of Arabidopsis, catalyzes C-3 oxidation. J Biol Chem 287:31551–31560. doi:10.1074/jbc.M112.392720
Oki K, Inaba N, Kitagawa K, Fujioka S, Kitano H, Fujisawa Y, Kato H, Iwasaki Y (2009) Function of the alpha subunit of rice heterotrimeric G protein in brassinosteroid signaling. Plant Cell Physiol 50:161–172. doi:10.1093/pcp/pcn182
Olszewski N, T-p Sun, Gubler F (2002) Gibberellin signaling biosynthesis, catabolism, and response pathways. Plant Cell 14:S61–S80
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F (1999) ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 400:256–261
Qiao S, Sun S, Wang L, Wu Z, Li C, Li X, Wang T, Leng L, Tian W, Lu T, Wang X (2017) The RLA1/SMOS1 Transcription Factor Functions with OsBZR1 to Regulate Brassinosteroid Signaling and Rice Architecture. Plant Cell 29:292–309. doi:10.1105/tpc.16.00611
Qi W, Sun F, Wang Q, Chen M, Huang Y, Feng YQ, Luo X, Yang J (2011) Rice ethylene-response AP2/ERF factor OsEATB restricts internode elongation by down-regulating a gibberellin biosynthetic gene. Plant Physiol 157:216–228. doi:10.1104/pp.111.179945
Rebetzke G, Ellis M, Bonnett D, Condon A, Falk D, Richards R (2011) The Rht13 dwarfing gene reduces peduncle length and plant height to increase grain number and yield of wheat. Field Crops Res 124:323–331
Rebetzke G, Bonnett D, Ellis M (2012) Combining gibberellic acid-sensitive and insensitive dwarfing genes in breeding of higher-yielding, sesqui-dwarf wheats. Field Crops Res 127:17–25
Ross JJ, Murfet IC, Reid JB (1997) Gibberellin mutants. Physiol Plant 100:550–560
Rutger J, Carnahan H (1981) A fourth genetic element to facilitate hybrid cereal production—a recessive tall in rice. Crop Sci 21:373–376
Ruyter-Spira C, Al-Babili S, van der Krol S, Bouwmeester H (2013) The biology of strigolactones. Trends Plant Sci 18:72–83
Saeda T, Kitano H (1992) Genetic analysis of a short culm-mutant showing the dm-type internode distribution pattern. Jpn J Breed 42:214–215
Sakai M, Sakamoto T, Saito T, Matsuoka M, Tanaka H, Kobayashi M (2003) Expression of novel rice gibberellin 2-oxidase gene is under homeostatic regulation by biologically active gibberellins. J Plant Res 116:161–164
Sakamoto T, Matsuoka M (2006) Characterization of constitutive photomorphogenesis and DWARFISM homologs in rice (Oryza sativa L.). J Plant Growth Regul 25:245–251
Sakamoto T, Morinaka Y, Ishiyama K, Kobayashi M, Itoh H, Kayano T, Iwahori S, Matsuoka M, Tanaka H (2003) Genetic manipulation of gibberellin metabolism in transgenic rice. Nat Biotechnol 21:909–913. doi:10.1038/nbt847
Sakamoto T, Miura K, Itoh H, Tatsumi T, Ueguchi-Tanaka M, Ishiyama K, Kobayashi M, Agrawal GK, Takeda S, Abe K, Miyao A, Hirochika H, Kitano H, Ashikari M, Matsuoka M (2004) An overview of gibberellin metabolism enzyme genes and their related mutants in rice. Plant Physiol 134:1642–1653. doi:10.1104/pp.103.033696
Sakamoto T, Morinaka Y, Ohnishi T, Sunohara H, Fujioka S, Ueguchi-Tanaka M, Mizutani M, Sakata K, Takatsuto S, Yoshida S, Tanaka H, Kitano H, Matsuoka M (2006) Erect leaves caused by brassinosteroid deficiency increase biomass production and grain yield in rice. Nat Biotechnol 24:105–109. doi:10.1038/nbt1173
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush G (2002) Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 416:701–702
Sasaki A, Itoh H, Gomi K, Ueguchi-Tanaka M, Ishiyama K, Kobayashi M, Jeong DH, An G, Kitano H, Ashikari M, Matsuoka M (2003) Accumulation of phosphorylated repressor for gibberellin signaling in an F-box mutant. Science (New York, N Y) 299:1896–1898. doi:10.1126/science.1081077
Sato Y, Sentoku N, Miura Y, Hirochika H, Kitano H, Matsuoka M (1999) Loss-of-function mutations in the rice homeobox gene OSH15 affect the architecture of internodes resulting in dwarf plants. EMBO J 18:992–1002. doi:10.1093/emboj/18.4.992
Sazuka T, Kamiya N, Nishimura T, Ohmae K, Sato Y, Imamura K, Nagato Y, Koshiba T, Nagamura Y, Ashikari M, Kitano H, Matsuoka M (2009) A rice tryptophan deficient dwarf mutant, tdd1, contains a reduced level of indole acetic acid and develops abnormal flowers and organless embryos. Plant J 60:227–241. doi:10.1111/j.1365-313X.2009.03952.x
Schomburg FM, Bizzell CM, Lee DJ, Zeevaart JA, Amasino RM (2003) Overexpression of a novel class of gibberellin 2-oxidases decreases gibberellin levels and creates dwarf plants. Plant Cell 15:151–163
Sears RG, Kronstad WE, Metzger RJ (1981) Inheritance of dwarf and semi-dwarf plant height in barley. Crop Sci 21:828–833
Shen Z, He Z (1989) Interaction between eui gene and WAMS cytoplasm of rice and improvement of panicle exsertion of MS line. SABRAO J 6:753–756
Shimada Y, Fujioka S, Miyauchi N, Kushiro M, Takatsuto S, Nomura T, Yokota T, Kamiya Y, Bishop GJ, Yoshida S (2001) Brassinosteroid-6-oxidases from Arabidopsis and tomato catalyze multiple C-6 oxidations in brassinosteroid biosynthesis. Plant Physiol 126:770–779
Shimada A, Ueguchi-Tanaka M, Sakamoto T, Fujioka S, Takatsuto S, Yoshida S, Sazuka T, Ashikari M, Matsuoka M (2006) The rice SPINDLY gene functions as a negative regulator of gibberellin signaling by controlling the suppressive function of the DELLA protein, SLR1, and modulating brassinosteroid synthesis. Plant J 48:390–402. doi:10.1111/j.1365-313X.2006.02875.x
Song Y, You J, Xiong L (2009) Characterization of OsIAA1 gene, a member of rice Aux/IAA family involved in auxin and brassinosteroid hormone responses and plant morphogenesis. Plant Mol Biol 70:297–309. doi:10.1007/s11103-009-9474-1
Spielmeyer W, Ellis MH, Chandler PM (2002) Semi-dwarf (sd-1), “green revolution” rice, contains a defective gibberellin 20-oxidase gene. Proc Nati Acad Sci USA 99:9043–9048. doi:10.1073/pnas.132266399
Suharsono U, Fujisawa Y, Kawasaki T, Iwasaki Y, Satoh H, Shimamoto K (2002) The heterotrimeric G protein alpha subunit acts upstream of the small GTPase Rac in disease resistance of rice. Proc Nati Acad Sci USA 99:13307–13312. doi:10.1073/pnas.192244099
Sui P, Jin J, Ye S, Mu C, Gao J, Feng H, Shen WH, Yu Y, Dong A (2012) H3K36 methylation is critical for brassinosteroid-regulated plant growth and development in rice. Plant J 70:340–347. doi:10.1111/j.1365-313X.2011.04873.x
Sun Y, Fan XY, Cao DM, Tang W, He K, Zhu JY, He JX, Bai MY, Zhu S, Oh E (2010) Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev Cell 19:765–777
Sun H, Tao J, Hou M, Huang S, Chen S, Liang Z, Xie T, Wei Y, Xie X, Yoneyama K (2015) A strigolactone signal is required for adventitious root formation in rice. Ann Bot 115:1155–1162
Tabuchi M, Sugiyama K, Ishiyama K, Inoue E, Sato T, Takahashi H, Yamaya T (2005) Severe reduction in growth rate and grain filling of rice mutants lacking OsGS1;1, a cytosolic glutamine synthetase1;1. Plant J 42:641–651. doi:10.1111/j.1365-313X.2005.02406.x
Tanabe S, Ashikari M, Fujioka S, Takatsuto S, Yoshida S, Yano M, Yoshimura A, Kitano H, Matsuoka M, Fujisawa Y, Kato H, Iwasaki Y (2005) A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11, with reduced seed length. Plant Cell 17:776–790. doi:10.1105/tpc.104.024950
Tanaka K, Murata K, Yamazaki M, Onosato K, Miyao A, Hirochika H (2003) Three distinct rice cellulose synthase catalytic subunit genes required for cellulose synthesis in the secondary wall. Plant Physiol 133:73–83
Tanaka A, Nakagawa H, Tomita C, Shimatani Z, Ohtake M, Nomura T, Jiang CJ, Dubouzet JG, Kikuchi S, Sekimoto H, Yokota T, Asami T, Kamakura T, Mori M (2009) Brassinosteroid UPREGULATED1, encoding a helix–loop–helix protein, is a novel gene involved in brassinosteroid signaling and controls bending of the lamina joint in rice. Plant Physiol 151:669–680. doi:10.1104/pp.109.140806
Thomas SG, Phillips AL, Hedden P (1999) Molecular cloning and functional expression of gibberellin 2-oxidases, multifunctional enzymes involved in gibberellin deactivation. Proc Nati Acad Sci USA 96:4698–4703
Tong H, Chu C (2012) Brassinosteroid signaling and application in rice. J Genet Genom 39:3–9. doi:10.1016/j.jgg.2011.12.001
Tong H, Jin Y, Liu W, Li F, Fang J, Yin Y, Qian Q, Zhu L, Chu C (2009) DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J 58:803–816. doi:10.1111/j.1365-313X.2009.03825.x
Toriba T, Harada K, Takamura A, Nakamura H, Ichikawa H, Suzaki T, Hirano HY (2007) Molecular characterization the YABBY gene family in Oryza sativa and expression analysis of OsYABBY1. Mol Genet Genom 277:457–468. doi:10.1007/s00438-006-0202-0
Ueguchi-Tanaka M, Fujisawa Y, Kobayashi M, Ashikari M, Iwasaki Y, Kitano H, Matsuoka M (2000) Rice dwarf mutant d1, which is defective in the alpha subunit of the heterotrimeric G protein, affects gibberellin signal transduction. Proc Nat Acad Sci USA 97:11638–11643. doi:10.1073/pnas.97.21.11638
Ueguchi-Tanaka M, Ashikari M, Nakajima M, Itoh H, Katoh E, Kobayashi M, Chow TY, Hsing YI, Kitano H, Yamaguchi I, Matsuoka M (2005) GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 437:693–698. doi:10.1038/nature04028
Umehara M, Hanada A, Yoshida S, Akiyama K, Arite T, Takeda-Kamiya N, Magome H, Kamiya Y, Shirasu K, Yoneyama K, Kyozuka J, Yamaguchi S (2008) Inhibition of shoot branching by new terpenoid plant hormones. Nature 455:195–200. doi:10.1038/nature07272
van Zeijl A, Liu W, Xiao TT, Kohlen W, Yang WC, Bisseling T, Geurts R (2015) The strigolactone biosynthesis gene DWARF27 is co-opted in rhizobium symbiosis. BMC Plant Biol 15:260
Vanneste S, Friml J (2009) Auxin: a trigger for change in plant development. Cell 136:1005–1016. doi:10.1016/j.cell.2009.03.001
Vriet C, Russinova E, Reuzeau C (2012) Boosting crop yields with plant steroids. Plant Cell 24:842–857. doi:10.1105/tpc.111.094912
Vriet C, Russinova E, Reuzeau C (2013) From squalene to brassinolide: the steroid metabolic and signaling pathways across the plant kingdom. Mol Plant 6:1738–1757
Wang L, Xu YY, Ma QB, Li D, Xu ZH, Chong K (2006) Heterotrimeric G protein alpha subunit is involved in rice brassinosteroid response. Cell Res 16:916–922. doi:10.1038/sj.cr.7310111
Wang H, Ngwenyama N, Liu Y, Walker JC, Zhang S (2007) Stomatal development and patterning are regulated by environmentally responsive mitogen-activated protein kinases in Arabidopsis. Plant Cell 19:63–73. doi:10.1105/tpc.106.048298
Wang L, Xu Y, Zhang C, Ma Q, Joo SH, Kim SK, Xu Z, Chong K (2008) OsLIC, a novel CCCH-type zinc finger protein with transcription activation, mediates rice architecture via brassinosteroids signaling. PLoS One 3:e3521
Wang ZY, Bai MY, Oh E, Zhu JY (2012) Brassinosteroid signaling network and regulation of photomorphogenesis. Annu Rev Genet 46:701–724. doi:10.1146/annurev-genet-102209-163450
Wang Y, Deng D, Ding H, Xu X, Zhang R, Wang S, Bian Y, Yin Z, Chen Y (2013a) Gibberellin biosynthetic deficiency is responsible for maize dominant Dwarf11 (D11) mutant phenotype: physiological and transcriptomic evidence. PLoS One 8:e66466
Wang Y, Sun S, Zhu W, Jia K, Yang H, Wang X (2013b) Strigolactone/MAX2-induced degradation of brassinosteroid transcriptional effector BES1 regulates shoot branching. Dev Cell 27:681–688. doi:10.1016/j.devcel.2013.11.010
Wang Y, Chen L, Du Y, Yang Z, Condon AG, Hu Y-G (2014) Genetic effect of dwarfing gene Rht13 compared with Rht-D1b on plant height and some agronomic traits in common wheat (Triticum aestivum L.). Field Crops Res 162:39–47
Waters MT, Brewer PB, Bussell JD, Smith SM, Beveridge CA (2012) The Arabidopsis ortholog of rice DWARF27 acts upstream of MAX1 in the control of plant development by strigolactones. Plant Physiol 159:1073–1085. doi:10.1104/pp.112.196253
Wei XJ, Tang SQ, Shao GN, Chen ML, Hu YC, Hu PS (2013) Fine mapping and characterization of a novel dwarf and narrow-leaf mutant dnl1 in rice. Genet Mol Res 12:3845–3855. doi:10.4238/2013.September.23.2
Wen C, Xi L, Gao B, Wang K, Lv S, Kou Y, Ma N, Zhao L (2015) Roles of DgD14 in regulation of shoot branching in chrysanthemum (Dendranthema grandiflorum ‘Jinba’). Plant Physiol Biochem 96:241–253
Wu CY, Trieu A, Radhakrishnan P, Kwok SF, Harris S, Zhang K, Wang J, Wan J, Zhai H, Takatsuto S, Matsumoto S, Fujioka S, Feldmann KA, Pennell RI (2008) Brassinosteroids regulate grain filling in rice. Plant Cell 20:2130–2145. doi:10.1105/tpc.107.055087
Wu G, Wang X, Li X, Kamiya Y, Otegui MS, Chory J (2011) Methylation of a phosphatase specifies dephosphorylation and degradation of activated brassinosteroid receptors. Sci Signal 4:ra29
Wu Y, Fu Y, Zhao S, Gu P, Zhu Z, Sun C, Tan L (2016) Clustered primary branch1, a new allele of DWARF11, controls panicle architecture and seed size in rice. Plant Biotechnol J 14:377–386. doi:10.1111/pbi.12391
Xie DX, Feys BF, James S, Nietorostro M, Turner JG (1998) COI1: an Arabidopsis gene required for jasmonate-regulated defense and fertility. Science (New York, N Y) 280:1091–1094
Xu J, Zha M, Li Y, Ding Y, Chen L, Ding C, Wang S (2015) The interaction between nitrogen availability and auxin, cytokinin, and strigolactone in the control of shoot branching in rice (Oryza sativa L.). Plant Cell Rep 34:1647–1662. doi:10.1007/s00299-015-1815-8
Yaish MW, El-Kereamy A, Zhu T, Beatty PH, Good AG, Bi YM, Rothstein SJ (2010) The APETALA-2-like transcription factor OsAP2-39 controls key interactions between abscisic acid and gibberellin in rice. PLoS Genet 6:e1001098. doi:10.1371/journal.pgen.1001098
Yamaguchi S (2008) Gibberellin metabolism and its regulation. Annu Rev Plant Biol 59:225–251
Yamamuro C, Ihara Y, Wu X, Noguchi T, Fujioka S, Takatsuto S, Ashikari M, Kitano H, Matsuoka M (2000) Loss of function of a rice brassinosteroid insensitive1 homolog prevents internode elongation and bending of the lamina joint. Plant Cell 12:1591–1606
Yan J, Zhang C, Gu M, Bai Z, Zhang W, Qi T, Cheng Z, Peng W, Luo H, Nan F, Wang Z, Xie D (2009) The Arabidopsis CORONATINE INSENSITIVE1 protein is a jasmonate receptor. Plant Cell 21:2220–2236. doi:10.1105/tpc.109.065730
Yang DL, He SY (2012) Plant hormone jasmonate prioritizes defense over growth by interfering with gibberellin signaling cascade. Proc Nati Acad Sci USA 109:1192–1200
Yang RC, Zhang SB, Huang RH, Yang SL, Zhang QQ (2002) Breeding technology of eui-hybrids of rice. Agric Sci China 1:359–363
Yang G, Nakamura H, Ichikawa H, Kitano H, Komatsu S (2006) OsBLE3, a brassinolide-enhanced gene, is involved in the growth of rice. Phytochemistry 67:1442–1454. doi:10.1016/j.phytochem.2006.05.026
Yang C, Shen W, He Y, Tian Z, Li J (2016) OVATE family protein 8 positively mediates brassinosteroid signaling through interacting with the GSK3-like kinase in rice. PLoS Genet 12:e1006118
Yasuno N, Yasui Y, Takamure I, Kato K (2007) Genetic interaction between 2 tillering genes, reduced culm number 1 (rcn1) and tillering dwarf gene d3, in rice. J Heredity 98:169–172. doi:10.1093/jhered/esl069
Yokota T (1997) The structure, biosynthesis and function of brassinosteroids. Trends Plant Sci 2:137–143
Yoshida S, Kameoka H, Tempo M, Akiyama K, Umehara M, Yamaguchi S, Hayashi H, Kyozuka J, Shirasu K (2012) The D3 F-box protein is a key component in host strigolactone responses essential for arbuscular mycorrhizal symbiosis. New Phytol 196:1208–1216
Yu X, Li L, Zola J, Aluru M, Ye H, Foudree A, Guo H, Anderson S, Aluru S, Liu P (2011) A brassinosteroid transcriptional network revealed by genome-wide identification of BESI target genes in Arabidopsis thaliana. Plant J 65:634–646
Yuan SJ, Wang T, Yin L, Zhao JG, Wan JM, Li XY (2013) Cloning and expression of gene responsible for high-tillering dwarf phenotype in Indica rice mutant gsor23. Rice Sci 20:320–328
Zazimalova E, Napier RM (2003) Points of regulation for auxin action. Plant Cell Rep 21:625–634. doi:10.1007/s00299-002-0562-9
Zentella R, Sui N, Barnhill B, Hsieh WP, Hu J, Shabanowitz J, Boyce M, Olszewski NE, Zhou P, Hunt DF (2017) The Arabidopsis O-fucosyltransferase SPINDLY activates nuclear growth repressor DELLA. Nat Chem Biol 13:479–485
Zhang LY, Bai MY, Wu J, Zhu JY, Wang H, Zhang Z, Wang W, Sun Y, Zhao J, Sun X, Yang H, Xu Y, Kim SH, Fujioka S, Lin WH, Chong K, Lu T, Wang ZY (2009) Antagonistic HLH/bHLH transcription factors mediate brassinosteroid regulation of cell elongation and plant development in rice and Arabidopsis. Plant Cell 21:3767–3780. doi:10.1105/tpc.109.070441
Zhang S, Li G, Fang J, Chen W, Jiang H, Zou J, Liu X, Zhao X, Li X, Chu C, Xie Q, Jiang X, Zhu L (2010) The interactions among DWARF10, auxin and cytokinin underlie lateral bud outgrowth in rice. J Integr Plant Biol 52:626–638. doi:10.1111/j.1744-7909.2010.00960.x
Zhang B, Tian F, Tan L, Xie D, Sun C (2011) Characterization of a novel high-tillering dwarf 3 mutant in rice. J Genet Genom 38:411–418. doi:10.1016/j.jgg.2011.08.002
Zhang C, Xu Y, Guo S, Zhu J, Huan Q, Liu H, Wang L, Luo G, Wang X, Chong K (2012) Dynamics of brassinosteroid response modulated by negative regulator LIC in rice. PLoS Genet 8:e1002686. doi:10.1371/journal.pgen.1002686
Zhang C, Bai MY, Chong K (2014a) Brassinosteroid-mediated regulation of agronomic traits in rice. Plant Cell Rep 33:683–696
Zhang J, Liu X, Li S, Cheng Z, Li C (2014b) The rice semi-dwarf mutant sd37, caused by a mutation in CYP96B4, plays an important role in the fine-tuning of plant growth. PLoS One 9:e88068. doi:10.1371/journal.pone.0088068
Zhang Y, van Dijk AD, Scaffidi A, Flematti GR, Hofmann M, Charnikhova T, Verstappen F, Hepworth J, van der Krol S, Leyser O, Smith SM, Zwanenburg B, Al-Babili S, Ruyter-Spira C, Bouwmeester HJ (2014c) Rice cytochrome P450 MAX1 homologs catalyze distinct steps in strigolactone biosynthesis. Nat Chem Biol 10:1028–1033. doi:10.1038/nchembio.1660
Zhang X, Song H, Xiaoying C, Rongxiang F (2016) Small G protein as a novel component of the rice bmssinosteroid signal transduction. Mol Plant 9:1260–1271
Zhou HL, He SJ, Cao YR, Chen T, Du BX, Chu CC, Zhang JS, Chen SY (2006) OsGLU1, a putative membrane-bound endo-1,4-beta-d-glucanase from rice, affects plant internode elongation. Plant Mol Biol 60:137–151. doi:10.1007/s11103-005-2972-x
Zhu Y, Nomura T, Xu Y, Zhang Y, Peng Y, Mao B, Hanada A, Zhou H, Wang R, Li P, Zhu X, Mander LN, Kamiya Y, Yamaguchi S, He Z (2006) ELONGATED UPPERMOST INTERNODE encodes a cytochrome P450 monooxygenase that epoxidizes gibberellins in a novel deactivation reaction in rice. Plant Cell 18:442–456. doi:10.1105/tpc.105.038455
Zou J, Chen Z, Zhang S, Zhang W, Jiang G, Zhao X, Zhai W, Pan X, Zhu L (2005) Characterizations and fine mapping of a mutant gene for high tillering and dwarf in rice (Oryza sativa L.). Planta 222:604–612. doi:10.1007/s00425-005-0007-0
Zou J, Zhang S, Zhang W, Li G, Chen Z, Zhai W, Zhao X, Pan X, Xie Q, Zhu L (2006) The rice high-tillering DWARF1 encoding an ortholog of Arabidopsis MAX3 is required for negative regulation of the outgrowth of axillary buds. Plant J 48:687–698. doi:10.1111/j.1365-313X.2006.02916.x
Acknowledgements
Special thanks should go to Dr. Alice Hayward who has put considerable time and effort on improving the clarity and accuracy of this text. This work was supported by a Major Research Project of CAAS Science and the Technology Innovation Program, and by the National Natural Science Foundation of China (Grant numbers: 31400243 and 31201152), and by the Natural Science Foundation of Hubei Province (Grant number: 2013CFB423).
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, F., Wang, P., Zhang, X. et al. The genetic and molecular basis of crop height based on a rice model. Planta 247, 1–26 (2018). https://doi.org/10.1007/s00425-017-2798-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2798-1