Abstract
Main conclusion
We demonstrate that rbcL underwent strong positive selection during the C 3 –C 4 photosynthetic transitions in PACMAD grasses, in particular the 3′ end of the gene. In contrast, selective pressures on other plastid genes vary widely and environmental drivers remain to be identified.
Plastid genomes have been widely used to infer phylogenetic relationships among plants, but the selective pressures driving their evolution have not been systematically investigated. In our study, we analyse all protein-coding plastid genes from 113 species of PACMAD grasses (Poaceae) to evaluate the selective pressures driving their evolution. Our analyses confirm that the gene encoding the large subunit of RubisCO (rbcL) evolved under strong positive selection after C3–C4 photosynthetic transitions. We highlight new codons in rbcL that underwent parallel changes, in particular those encoding the C-terminal part of the protein. C3–C4 photosynthetic shifts did not significantly affect the evolutionary dynamics of other plastid genes. Instead, while two-third of the plastid genes evolved under purifying selection or neutrality, 25 evolved under positive selection across the PACMAD clade. This set of genes encode for proteins involved in diverse functions, including self-replication of plastids and photosynthesis. Our results suggest that plastid genes widely adapt to changing ecological conditions, but factors driving this evolution largely remain to be identified.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plastids (including chloroplasts) first appeared as cyanobacterial endosymbionts of eukaryotic organisms 1 billion years ago (Mereschkowski 1905; McFadden and van Dooren 2004). This symbiosis allowed eukaryotic organisms to capture solar energy through photosynthesis. The cyanobacterial genome was maintained as an extranuclear genome through the endosymbiosis, but over evolutionary times it underwent significant changes, with numerous genes being transferred to the nuclear genome (Stegemann et al. 2003; Olejniczak et al. 2016). However, over 80 protein-coding genes usually still persist in the plastids of plants, including those encoding elements for the genetic apparatus and the proteins involved in photosynthesis (Bock 2007; Olejniczak et al. 2016).
Markers extracted from plastid genomes (hereafter plastomes) have been widely used in studies of plant phylogenetic relationships, biogeography and species identification (e.g. Schaal et al. 1998; APG III 2009; Hollingsworth et al. 2009; Moore et al. 2010; Nguyen et al. 2015). Their high copy number makes them easy to isolate and sequence. Indeed, one plant cell usually possesses tens of plastids, and each contains multiple copies of the plastome, resulting in a relatively high number of copies per cell compared to the nuclear genome (Burgess 1989). Plastid genes encode proteins and RNA molecules that are crucial to the functioning of the plant metabolism, and can consequently undergo selective pressures. Purifying selection acts to maintain protein functions, while positive selection may come into play in response to environmental changes.
The plastid gene most studied in terms of selective pressures is probably the one encoding the large subunit of RubisCO, rbcL. Previous studies on various plant groups have highlighted positive selection on rbcL in relation to temperature, drought, and carbon dioxide concentration, particularly following C4 photosynthesis evolution (e.g. Miller 2003; Kapralov and Filatov 2007; Christin et al. 2008; Iida et al. 2009; Kapralov et al. 2011, 2012; Wang et al. 2011; Young et al. 2012; Galmés et al. 2014a, b, 2015; Orr et al. 2016). C4 photosynthesis is a novel assemblage of biochemical and anatomical components that result in an increased CO2 concentration around RubisCO (Hatch 1987; von Caemmerer and Furbank 2003). This reduces the selective pressure for higher substrate specificity, allowing the evolution of more efficient versions of the enzyme at the expense of CO2 specificity (Tcherkez et al. 2006; Nisbet et al. 2007; Christin et al. 2008; Whitney et al. 2011b; Studer et al. 2014). Despite this clear evidence for non-neutral substitutions in rbcL, investigations of the selective pressures driving the evolutionary diversification of other plastid genes are still lacking. In this study, we address this gap by assessing the selective pressures acting on all protein-coding genes of the grass plastome.
The grass family (Poaceae), which encompasses more than 12,000 species, is one of the most important families of flowering plants, both economically and ecologically (Gibson 2009; Kellogg 2015). Grasses dominate most open biomes across the world, and constitute a major food source for numerous animals, including humans (Gibson 2009). Over the last 65 million years, grasses have come to dominate a variety of habitats, and these transitions were accompanied by major functional innovations (Gibson 2009). Among the two main clades of Poaceae, the so-called PACMAD clade includes all 22–24 known origins of C4 photosynthesis in grasses (GPWG II 2012). C4 grasses now dominate large open biomes such as savannas across the globe thanks to C4 photosynthesis, which increases photosynthetic efficiency in warm conditions (Griffith et al. 2015; Atkinson et al. 2016). Other PACMAD lineages adapted to a variety of contrasted conditions, from the shade of tropical forests, to arid deserts and cold climates (e.g. Humphreys and Linder 2013; Taylor et al. 2014). These functional and ecological conditions alter the quantity of light available to the plant as well as the amount of CO2 received by RubisCO. Indeed, CO2 availability decreases with temperature, aridity and salinity, which lead to stomata closure, limiting gas exchange and entailing CO2 depletion inside the leaves (Galmés et al. 2005; Sage et al. 2012). It is therefore possible that the optimum of the photosynthetic apparatus of grasses shifted with changing environments. In this case, adaptation of proteins encoded by plastid genes would have left traces in the form of adaptive amino-acid substitutions.
In this study, we used codon-substitution models to assess the past selective pressures acting on protein-coding genes from complete plastomes of grasses. These models can detect the fingerprint of episodes of adaptive evolution along specific branches of a phylogenetic tree (Yang et al. 2005). The hypothesis that C3–C4 transitions were accompanied by adaptive changes in plastid genes was tested. We also asked whether there were adaptive changes in the plastome of PACMAD grasses that are unrelated to the evolution of C4 photosynthesis. We thus analyzed all protein-encoding plastid genes of 113 grass species of the PACMAD clade. For each of the 76 protein-coding genes present in grass plastomes, we tested the hypothesis of non-neutral evolution, and specifically of positive selection leading to an excess of functionally adaptive amino-acid substitutions. We first evaluated the occurrence of positive selection across the whole phylogeny, without prior expectations regarding the selective drivers. We then specifically tested whether the evolution of C4 photosynthesis altered the selective pressures on plastid genes in addition to rbcL. Overall, our efforts provide the first systematic evaluation of selective pressures across plastomes in a functionally and ecologically diverse group of plants. We were able to detect widespread positive selection acting on plastid protein-coding genes, with important consequences for our understanding of the adaptive significance of plastid genes and their usefulness as phylogenetic markers.
Materials and methods
Taxon sampling and dataset assembly
Complete plastomes or complete sets of protein-coding plastid genes (76 genes) were retrieved from GenBank on June 2016, and completed with the sequencing and assembly of plastomes of 21 additional species (Table S1). The final sampling includes 113 grass species (or subspecies in the case of Alloteropsis semialata, the only known species with C3 and C4 lineages) belonging to the PACMAD clade (Table S1). Of these, 77 are taxa performing C4 photosynthesis. Among these C4 accessions, three belong to Aristidoideae, two to Micrairoideae, 14 to Chloridoideae and 58 to Panicoideae. Our sampling covers 16 separate C4 lineages reported in the PACMAD clade (GPWG II 2012) and was thus particularly appropriate for testing the impact of C3–C4 photosynthetic transitions on the selective pressures acting on protein-coding genes.
The newly sequenced plastomes were assembled from shotgun data (following the “genome skimming” approach; Straub et al. 2012), as described by Besnard et al. (2013). Briefly, we generated millions of paired-end sequence reads with HiSeq 2000 or HiSeq 3000 (Illumina Inc., San Diego). Plastomes were then assembled using the Organelle Assembler package, version 00.01.000 (https://pythonhosted.org/ORG.asm/) in Python. Protein-coding genes of Arabidopsis thaliana as available in the package were used as seeds to initiate plastome assembly. Then, assembled plastomes were annotated using Geneious v.6.1.8 (Biomatters Ltd., Auckland; Kearse et al. 2012) by transferring annotations from Zea mays (NC_001666.2). All protein-coding genes were extracted from the different plastomes and each dataset was aligned separately as codons using MUSCLE (Edgar 2004).
The 76 gene alignments were concatenated, and the best substitution model was determined for each gene (partition) using PartitionFinder v.1.1.1 (Lanfear et al. 2014). In all cases, the general time reversible model of nucleotide substitution with a gamma shape parameter (GTR + G) was selected, with or without a correction for invariant sites (+ I) (Table S2). A phylogenetic tree was estimated from this alignment using Bayesian inference as implemented in MrBayes v.3.2.4 (Huelsenbeck and Ronquist 2001). The dataset was partitioned per group of genes identified by PartitionFinder, and each partition was allowed to have different parameter values. Three independent Markov Chain Monte Carlo analyses, each with four parallel chains, were run for 100 million generations, sampling a tree every 20,000 generations. The first 25% were discarded as burn-in, after visual inspection of the log files in Tracer v1.6 (Rambaut et al. 2014), and a consensus tree was computed from the posterior distribution. We rooted the tree with Aristidoideae as outgroup, following GPWG II (2012).
Tests for positive selection
Past selective pressures were inferred using different codon-substitution models, as implemented in the PAML package v.4 (Yang 2007). In these models, the rate of fixation of non-synonymous substitutions (d N) is compared to the rate of fixation of synonymous substitutions (d S). Under neutrality, the ratio d N/d S (ω) will be equal to one, while purifying selection will remove non-synonymous substitutions and therefore lead to ω < 1. Conversely, the preferential fixation of adaptive, non-synonymous substitutions under positive selection will inflate d N, leading to ω > 1. The phylogeny inferred above was used for the different models. Note that the maximum likelihood is calculated on unrooted trees, so that the choice of the outgroup does not influence the outputs.
To statistically test for positive selection, we compared the performance of four codon models, individually for each gene. The null site model M1a allows codons to evolve under either purifying selection or neutrality, while the alternative site model M2a adds a third category corresponding to positive selection. In addition, we used branch-site models allowing variation among branches of the phylogenetic tree. In the null model MA’, some sites evolve under purifying selection or neutrality across the whole tree, while others switch to neutrality on a priori defined branches (i.e. foreground branches). The alternative model MA is identical except that sites switch to positive selection on foreground branches. It therefore specifically tests for positive selection on these branches. In our case, we specifically tested for positive selection in C4 grasses, so all branches within monophyletic C4 clades were set as foreground branches. Because the different models are not nested, they cannot be compared using likelihood ratio tests. Instead, the fit of the different models was compared using the Akaike information criterion (AIC). A ΔAIC threshold of 2 was used to accept alternative models. Posterior probabilities of belonging to the different classes of codons were estimated by the Bayes Empirical Bayes procedure (Yang et al. 2005).
Results
Twenty-one new plastomes were released in our study. All plastomes show a similar organization, except the one of Loudetia simplex whom harbors a long inversion in the Large Single Copy (Fig. S1). In addition, we also detected the insertion of a mitochondrial region in the plastome of Paspalum paniculatum (Fig. S1) as already reported in this lineage by Burke et al. (2016). Seventy-six protein-coding plastid genes were detected in all plastomes.
Positive selection tests
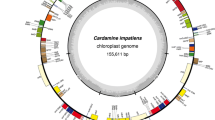
The phylogenetic tree inferred from the 76 protein-coding plastid genes of 113 PACMAD species (Figs. 2, S2) is congruent with previous topologies inferred from a reduced number of markers or taxa (e.g. GPWG II 2012; Burke et al. 2016). Every node is well supported, with posterior probabilities of 1 except for a few nodes (Fig. S2). Based on this topology, codon-substitution models were then tested separately on the 76 protein-coding plastid genes of the 113 PACMAD species. These analyses revealed significant signatures of positive selection on 27 genes (Fig. 1; Table 1). These positively selected genes do not appear to be clustered in particular regions of the plastome (Fig. 1).
First, the site model M2a, allowing positive selection on every branch of the phylogenetic tree, was found to be the best fitting model for 25 of those genes (Tables 1, S3). These genes encode proteins involved in diverse plastid functions: post-transcriptional modification, subunits of the NAD(P)H dehydrogenase complex (hereafter ‘NDH complex’) involved in the light-dependent photosynthesis phase (nine out of the 11 ndh genes), photosystem II, transcription, proteins involved in the light-independent phase of photosynthesis, and translation, more precisely large and small ribosomal proteins. All plastid genes encoding proteins involved in transcription and post-transcriptional modification (matK, rpoA, rpoB, rpoC1 and rpoC2) were thus found to evolve under positive selection across the phylogeny.
Second, the branch-site model MA, allowing positive selection only on C4 branches of the phylogenetic tree, was found to best represent sequence evolution for the two additional genes, rbcL and psaJ (Table 1). For rbcL, the branch-site model MA better fits the data than the site model M2a (ΔAIC = 2.2), the site model M1a (ΔAIC = 143.7) and the branch–site model MA’ (ΔAIC = 136.1; Table 1). Ten codons were identified as evolving under positive selection in C4 species, and these were recurrently mutated in independent C4 lineages, yet mostly conserved in C3 species (Fig. 2; Table 2). Three sites were detected in the 3′ end (sites 468–477) that encodes the C-terminal part. In addition, all species belonging to the C4 tribe Andropogoneae and the C4 species Tristachya humbertii show a codon insertion at position 469, which encodes different amino acids (i.e. Thr, Ser, Asn, or Gly) among the two lineages (Fig. S2). The last codon (site 477) is also missing in a few C4 species and was independently lost at least four times (Fig. S2). For psaJ, the branch–site model MA fitted better than site model M2a (ΔAIC = 6.0), site model M1a (ΔAIC = 3.1), and branch–site model MA’ (ΔAIC = 2.6; Table 1). However, the site model M2a did not perform better than site model M1a. Only one codon (at position 2 numbered following the psaJ coding sequence of Z. mays) was found to be under positive selection on C4 branches with posterior probability > 0.95. Overall, evidence of positive selection on psaJ is weak as differences between all models are low.
Codon sites of rbcL (relative to Zea mays NC_001666.2) identified to have evolved under positive selection in C4 species plotted against the phylogeny inferred from 76 protein-coding plastid genes for 113 PACMAD species. Codon changes are highlighted in black. Branches leading to C4 species are in red. The scale represents substitutions per site. The phylogeny is well supported; for precise support values and classification according to GPWG II (2012), refer to Fig. S2
Finally, the branch-site model MA’, assuming a shift to neutrality on C4 branches, performed better than the null site model M1a for nine genes (atpA, atpB, clpP, ndhE, psbD, rps3, rps8, rps16 and ycf4) and the model MA did not perform significantly worse (Table 1). These genes therefore switched to different selective pressures in C4 taxa, but the nature of the new selective pressure cannot be established with confidence. They encode for proteins involved in the light-dependent phase of photosynthesis phase, translation and enzyme modification.
Discussion
Positive selection on grass plastomes
Our codon models indicate that positive selection acted on some sites of roughly one-third of all protein-coding plastid genes (25 out of 76) across the whole PACMAD tree (Tables 1, S3). These genes are involved in different plastid functions, including self-replication (subunits of the ribosome, post-transcriptional processes and RNA polymerase) and photosynthesis (photosystems II, subunit of the NDH complex, cytochrome assembly and chloroplast membrane protein). Interestingly, all plastid genes encoding DNA-dependent RNA polymerases (rpo) evolved under positive selection in the PACMAD clade. These proteins are involved in transcription processes and therefore control gene expression. Six out of the 21 genes encoding for subunits of the ribosome (rpl and rps), carrying out translation, were found to evolve under positive selection across the whole tree. Along with matK, involved in post-transcriptional modifications, these genes have tremendous importance in self-replication and gene expression. Nine of the 11 genes encoding NDH subunits were also found to evolve under positive selection. In chloroplasts, NDH subunits form the NDH complex and are involved in chlororespiration and photosystem I cyclic electron transport. Additionally, four psb genes encoding for different subunits of the photosystem II evolved under positive selection. While the importance of the NDH complex remains obscure in plastids (Martín and Sabater 2010), proteins of the photosystem II are necessary for the light-dependent phase of photosynthesis. The last two genes found to evolve under positive selection across the whole tree were ccsA involved in cytochrome assembly (Xie and Merchant 1996) and cemA, a chloroplast envelope membrane protein whose exact role in plastids is not clearly determined.
The factors driving the detected selective pressures on these genes are so far unknown. Positive selection acting on these genes is probably evidence for adaptation to novel ecological conditions. Another likely hypothesis is that positive selection results from coevolutionary processes, as identified in arms races (e.g. Burri et al. 2010). Indeed, coevolution of nuclear and plastid genes can occur when different subunits of the same enzyme are encoded by each of the two genomes (Zhang et al. 2015; Weng et al. 2016). However, there is no reason to expect that positive selection would be sustained throughout the whole tree. Instead, we hypothesize that selective shifts occurred in disparate branches across the tree, after changes in the catalytic context of photosynthesis following alteration of the external environment (i.e. ecological shifts) or internal conditions (i.e. metabolomic shifts). Since our branch model only tested for C4-specific shifts, positive selection on other sets of branches would favour the model assuming phylogeny-wide adaptive evolution. Specification of the foreground branches in this analysis was focused on C3 versus C4 comparisons, but it is likely that other environmental conditions (e.g. low temperatures or shaded habitats) have also driven the adaptive evolution of plastid genes, not only in C4 species. Testing such hypotheses, with adequate species sampling and foreground branch specification, could be the subject of future studies.
C4 photosynthesis only affects rbcL
Plastid genes code for proteins involved in vital functions (De Las Rivas et al. 2002). In C3 species, photosynthesis is greatly optimized and plastid genes were, until recently, believed to evolve under purifying selection in order to eliminate any deleterious mutations (Clegg et al. 1994; Bock et al. 2014). However, C3–C4 photosynthetic transitions led to important anatomical and biochemical modifications in leaves of C4 species and to changes in ecological conditions (Christin and Osborne 2014). Our hypothesis was that these changes could have relaxed selective pressures acting on some amino-acid sites under purifying selection and led to their adaptive evolution to face novel environmental conditions. Despite significant changes, the evolution of C4 photosynthesis was not accompanied by obvious adaptive evolution on protein-coding plastid genes in grasses, except for rbcL which encodes the large subunit of RubisCO. Even though the model of positive selection on C4 branches (MA) was the best model to represent the data for psaJ, small AIC differences between each model do not show clear evidence of positive selection acting on this plastid gene in PACMAD grasses. For rbcL, AIC differences between alternative models (M2a and MA) and null models (M1a and MA’) are clear, demonstrating strong positive selection acting on this gene. However, small differences between the two alternative models (M2a and MA) suggest that C3–C4 photosynthetic transition is not the only factor driving positive selection on rbcL and other ecological conditions should also drive the adaptive evolution of this gene.
For the ten codons evolving under positive selection in rbcL of PACMAD grasses, the same codon substitutions appear in different C4 lineages, suggesting convergent evolution. However, not every C4 species shows substitutions at the same site or in the same codon. This pattern indicates that not all C4 species evolved alike and that several evolutionary pathways exist to adapt to C4 photosynthesis. Our identification of the putative amino acids positively selected in C4 species is not fully consistent with the results of a related study of rbcL on grasses and sedges (Christin et al. 2008). Six sites out of eight were still detected evolving under positive selection (Table 2). Four additional amino acids are found here to evolve under positive selection, and three of them are located at the end of the gene (positions 468–477), a region not included in the study of Christin et al. (2008) due to the difficulty to amplify the 3′ end of rbcL by PCR on a large sample of species. Other difference in the identification of positively selected amino-acid sites can also be explained by sampling differences, as we performed our analysis on 113 grass species of the PACMAD clade whereas Christin et al. (2008) used more than 200 species, covering the entire grass (Poaceae) and sedge (Cyperaceae) families.
The rbcL gene encodes the large subunit of the RubisCO enzyme which fixes CO2 to RuBP in the Calvin cycle. This enzyme is subject to a trade-off between its specificity for CO2 and its catalytic activity (Tcherkez et al. 2006). Thanks to the carbon concentrating mechanism in C4 species, CO2 specificity of RubisCO may have decreased, allowing its catalytic activity to increase (Sage 2002). Selective pressure changes would have led to the relaxation of the purifying selection acting on amino-acid sites of RubisCO that were important for the maintenance of CO2 specificity. This change would have allowed the directional evolution of some amino-acid sites and especially those having a role in the catalytic activity of RubisCO which evolved under positive selection (Whitney et al. 2011b; Studer et al. 2014). It is known that the C-terminal part of RbcL (positions after 460), and in particular site 471 (referred to as 470 in Burisch et al. 2007), is involved in the opening/closing mechanism of the active site of the RubisCO enzyme (Burisch et al. 2007). Amino-acid changes in this protein region affect RubisCO’s catalytic activity by acting on the opening speed of the active site (Gutteridge et al. 1993; Burisch et al. 2007). Shorter opening time of the active site allow for higher CO2 specificity, because CO2 fixes to the binding niche more rapidly than O2 (Schlitter and Wildner 2000). Positively selected substitutions at three codons (468, 471 and 476; Fig. 2) plus recurrent insertions (site 469) and deletions (site 477) in the 3′ end of rbcL should thus be major determinants to optimize RubisCO in a C4 catalytic context. These changes likely modify the energy needed to open the active site of the RubisCO enzyme (Burisch et al. 2007), leading to an increased catalytic activity to the expense of CO2 specificity. The precise role and adaptive evolution of these terminal sites in rbcL need to be confirmed with biochemical analyses but hold potential for biotechnological applications such as crop improvement for global change adaptation (Withney et al. 2011a; Carmo-Silva et al. 2015; Olejniczak et al. 2016).
Consequences of adaptive selection for the study of plant evolution
Our study revealed widespread positive selection on plastid protein-coding genes of grasses. These genes are widely used in plant phylogenetics and phylogeography. In particular, three genes (matK, ndhF, and rbcL) that were shown here to evolve under positive selection, either across the whole PACMAD phylogeny, or specifically on C4 branches, are among the most widely used in phylogenetic studies, both within grasses (GPWG II 2012) and in other groups (e.g. APG III 2009). It has been shown previously that adaptive evolution can bias phylogenetic reconstructions if it leads to the convergent fixation of some amino-acid residues against a background of limited neutral substitutions (Kellogg and Giullano 1997; Christin et al. 2012). Since the plastid substitution rate is usually much lower than the nuclear substitution rate (Wolfe et al. 1987), we hypothesize that selection could alter inferred phylogenetic relationships, especially when studying closely related species with a few markers. The risk of spurious groupings is decreased by considering multiple genes, a practice that is becoming widespread with the accumulation of complete plastomes (e.g. Moore et al. 2010; Washburn et al. 2015; Burke et al. 2016). For inferences of phylogenetic relationships among close relatives based on a limited number of sites, however, we suggest to minimize the effects of non-neutral evolution by favoring non-coding plastid genes.
Conclusion
In this study, we provide the first analysis of the selective pressures acting across plastomes in grasses. Our results suggest that positive selection is driving the evolution of about one-third of all protein-coding plastid genes. These encode for proteins involved in vital functions such as self-replication of plastids and photosynthesis and are therefore of tremendous importance for plant survival. However, factors driving this adaptive evolution are still unknown and new hypotheses regarding ecological adaptation are needed to define appropriate foreground branches and taxon sampling. Secondly, we found that the establishment of C4 photosynthesis in grasses did not lead to major adaptive evolution of protein-coding plastid genes except for the one encoding the large subunit of RubisCO, rbcL. Finally, despite the common thinking that plastid markers evolve slowly and are well conserved, we suggest that the evolution of plastid genes was shaped, to a great extent, by changing ecological conditions. Given their adaptive potential, plastid genes provide powerful genetic markers to study plant evolution, but possible bias in phylogenetic analyses needs to be taken into account.
Author contribution statement
PAC and GB conceived and designed research. GB prepared DNA for shotgun sequencing. AP, JH and GB analyzed data. AP wrote the manuscript with the help of all co-authors.
References
Apg III (2009) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG III. Bot J Linn Soc 161:105–121
Atkinson RRL, Mockford EJ, Bennett C, Christin PA, Spriggs E, Freckleton RP, Thompson K, Rees M, Osborne CP (2016) C4 photosynthesis boost growth via altered physiology, allocation and size. Nat Plant 2:16038
Besnard G, Christin PA, Malé PJG, Coissac E, Ralimanana H, Vorontsova MS (2013) Phylogenomics and taxonomy of Lecomtelleae (Poaceae), an isolated panicoid lineage from Madagascar. Ann Bot 112:1057–1066
Bock R (2007) Structure, function, and inheritance of plastid genomes. In: Bock R (ed) Cell and molecular biology of plastids, topics in current genetics, vol 19. Springer, Berlin, pp 29–63
Bock DG, Andrew RL, Rieseberg LH (2014) On the adaptive value of cytoplasmic genomes in plants. Mol Ecol 23:4899–4911
Burgess J (1989) An introduction to plant cell development. Cambridge University Press, Cambridge
Burisch C, Wildner GF, Schlitter J (2007) Bioinformatic tools uncover the C-terminal strand of Rubisco’s large subunit as hot-spot for specificity-enhancing mutations. FEBS Lett 581:741–748
Burke SV, Wysocki WP, Zuloaga FO, Craine JM, Pires JC, Edger PP, Mayfield-Jones D, Clark LG, Kelchner SA, Duvall MR (2016) Evolutionary relationships in Panicoid grasses based on plastome phylogenomics (Panicoideae; Poaceae). BMC Plant Biol 16:140
Burri R, Salamin N, Studer RA, Roulin A, Fumagalli L (2010) Adaptive divergence of ancient gene duplicates in the avian MHC class II beta. Mol Biol Evol 27:2360–2374
Carmo-Silva E, Scales JC, Madgwick PJ, Parry MAJ (2015) Optimizing Rubisco and its regulation for greater resource use efficiency. Plant Cell Environ 38:1817–1832
Christin PA, Osborne CP (2014) The evolutionary ecology of C4 plants. New Phytol 204:765–781
Christin PA, Salamin N, Muasya AM, Roalson EH, Russier F, Besnard G (2008) Evolutionary switch and genetic convergence on rbcL following the evolution of C4 photosynthesis. Mol Biol Evol 25:2361–2368
Christin PA, Besnard G, Edwards EJ, Salamin N (2012) Effect of genetic convergence on phylogenetic inference. Mol Phylogenet Evol 62:921–927
Clegg MT, Gaut BS, Learn GH, Morton BR (1994) Rates and patterns of chloroplast DNA evolution. Proc Natl Acad Sci USA 91:6795–6801
De Las Rivas J, Lozano JJ, Ortiz AR (2002) Comparative analysis of chloroplast genomes: functional annotation, genome-based phylogeny, and deduced evolutionary patterns. Genome Res 12:567–583
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Galmés J, Flexas J, Keys AJ, Cifre J, Mitchell RAC, Madgwick PJ, Haslam RP, Medrano H, Parry MAJ (2005) Rubisco specificity factor tends to be larger in plant species from drier habitats and in species with persistent leaves. Plant Cell Environ 28:571–579
Galmés J, Andralojc PJ, Kapralov MV, Flexas J, Keys AJ, Molins A, Parry MAJ, Conesa MÀ (2014a) Environmentally driven evolution of Rubisco and improved photosynthesis and growth within the C3 genus Limonium (Plumbaginaceae). New Phytol 203:989–999
Galmés J, Kapralov MV, Andralojc PJ, Conesa MÀ, Keys AJ, Parry MAJ, Flexas J (2014b) Expanding knowledge of the Rubisco kinetics variability in plant species: environmental and evolutionary trends. Plant Cell Environ 37:1989–2001
Galmés J, Kapralov MV, Copolovic LO, Hermida-Carrera C, Niinemets Ü (2015) Temperature responses of the Rubisco maximum carboxylase activity across domains of life: phylogenetic signals, trade-offs, and importance for carbon gain. Photosynth Res 123:183
Gibson DJ (2009) Grasses and grassland ecology. Oxford University Press Inc, New York
Gpwg II (2012) New grass phylogeny resolves deep evolutionary relationships and discovers C4 origins. New Phytol 193:304–312
Griffith DM, Anderson TM, Osborne CP, Strömberg CAE, Forrestel EJ, Still CJ (2015) Biogeographically distinct controls on C3 and C4 grass distributions: merging community and physiological ecology. Glob Ecol Biogeogr 24:304–313
Gutteridge S, Rhades D, Herrmann C (1993) Site-specific mutations in a loop region of the C-terminal domain of the large subunit of ribulose bisphosphate carboxylase/oxygenase that influence substrate partitioning. J Biol Chem 268:7818–7824
Hatch MD (1987) C4 photosynthesis: a unique blend of modified biochemistry, anatomy and ultrastructure. Biochim Biophys Acta 895:81–106
Hollingsworth ML, Clark AA, Forrest LL, Richardson J, Pennington RT, Long DG, Cowan R, Chase MW, Gaudeul M, Hollingsworth PM (2009) Selecting barcoding loci for plants: evaluation of seven candidate loci with species-level sampling in three divergent groups of land plants. Mol Ecol Res 9:439–457
Huelsenbeck JP, Ronquist F (2001) MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 17:754–755
Humphreys AM, Linder HP (2013) Evidence for recent evolution of cold tolerance in grasses suggests current distribution is not limited by (low) temperature. New Phytol 198:1261–1273
Iida S, Miyagi A, Aoki S, Ito M, Kadono Y, Kosuge K (2009) Molecular adaptation of rbcL in the heterophyllous aquatic plant Potamogeton. PLoS One 4:e4633
Kapralov MV, Filatov DA (2007) Widespread positive selection in the photosynthetic Rubisco enzyme. BMC Evol Biol 7:73
Kapralov MV, Kubien DS, Andersson I, Filatov DA (2011) Changes in Rubisco kinetics during the evolution of C4 photosynthesis in Flaveria (Asteraceae) are associated with positive selection on genes encoding the enzyme. Mol Biol Evol 28:1491–1503
Kapralov MV, Smith JAC, Filatov DA (2012) Rubisco evolution in C4 eudicots: an analysis of Amaranthaceae sensu lato. PLoS One 7:e52974
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Kellogg EA (2015) Flowering plants. Monocots: Poaceae. In: Kellogg EA (ed) The families and genera of vascular plants, vol 13. Springer, New York
Kellogg EA, Giullano ND (1997) The structure and function of RuBisCO and their implications for systematic studies. Am J Bot 84:413–428
Lanfear R, Calcott B, Kainer D, Mayer C, Stamatakis A (2014) Selecting optimal partitioning schemes for phylogenomic datasets. BMC Evol Biol 14:82
Martín M, Sabater B (2010) Plastid ndh genes in plant evolution. Plant Physiol Biochem 48:636–645
McFadden G, van Dooren GG (2004) Evolution: red algal genome affirms a common origin of all plastids. Curr Biol 14:D514–D516
Mereschkowski C (1905) Übernatur und ursprung der chromatophoren im pflanzenreiche. Biol Centralb 25:593–604
Miller SR (2003) Evidence for the adaptive evolution of the carbon fixation gene rbcL during diversification in temperature tolerance of a clade of hot spring cyanobacteria. Mol Ecol 12:1237–1246
Moore MJ, Soltis PS, Bell CD, Burleigh JG, Soltis DE (2010) Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proc Natl Acad Sci USA 107:4623–4628
Nguyen PAT, Kim JS, Kim JH (2015) The complete chloroplast genome of colchicine plants (Colchicum autumnale L. and Gloriosa superba L.) and its application for identifying the genus. Planta 242:223–237
Nisbet EG, Grassineau NV, Howe CJ, Abell PI, Regelous M, Nisbet RER (2007) The age of Rubisco: the evolution of oxygenic photosynthesis. Geobiology 5:311–335
Olejniczak SA, Łojewska E, Kowalczyk T, Sakowicz T (2016) Chloroplasts: state of research and practical applications of plastome sequencing. Planta 244:517–527
Orr DJ, Alcântara A, Kapralov MV, Andralojc PJ, Carmo-Silva E, Parry MAJ (2016) Surveying Rubisco diversity and temperature response to improve crop photosynthetic efficiency. Plant Physiol 172:707–717
Rambaut A, Suchard MA, Xie D, Drummond AJ (2014) Tracer v1.6. http://beast.bio.ed.ac.uk/Tracer. Accessed Jan 2017
Sage RF (2002) Variation in the k cat of Rubisco in C3 and C4 plants and some implications for photosynthetic performance at high and low temperature. J Exp Bot 53:609–620
Sage RF, Sage TL, Kocacinar F (2012) Photorespiration and the evolution of C4 photosynthesis. Annu Rev Plant Biol 63:19–47
Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474
Schlitter J, Wildner GF (2000) The kinetics of conformation change as determinant of Rubisco’s specificity. Photosynth Res 65:7–13
Stegemann S, Hartmann S, Ruf S, Bock R (2003) High-frequency gene transfer from the chloroplast genome to the nucleus. Proc Natl Acad Sci USA 100:8828–8833
Straub SCK, Parks M, Weitemier K, Fishbein M, Cronn RC, Liston A (2012) Navigating the tip of the genomic iceberg: next-generation sequencing for plant systematics. Am J Bot 99:349–364
Studer R, Christin PA, Williams MA, Orengo C (2014) Stability-activity tradeoffs constrain the adaptive evolution of RubisCO. Proc Natl Acad Sci USA 111:2223–2228
Taylor SH, Ripley BS, Martin T, De-Wet LA, Woodward FI, Osborne CP (2014) Physiological advantages of C4 grasses in the field: a comparative experiment demonstrating the importance of drought. Glob Change Biol 20:1992–2003
Tcherkez GBB, Farquhar GD, Andrews TJ (2006) Despite slow catalysis and confused substrate specificity, all ribulose bisphosphate carboxylases may be nearly perfectly optimized. Proc Natl Acad Sci USA 103:7246–7251
von Caemmerer S, Furbank RT (2003) The C4 pathway: an efficient CO2 pump. Photosynth Res 77:191–207
Wang M, Kapralov MV, Anisimova M (2011) Coevolution of amino acid residues in the key photosynthetic enzyme Rubisco. BMC Evol Biol 11:266
Washburn JD, Schnable JC, Davidse G, Pires JC (2015) Phylogeny and photosynthesis of the grass tribe Paniceae. Am J Bot 102:1493–1505
Weng ML, Ruhlman TA, Jansen RK (2016) Plastid-nuclear interaction and accelerated coevolution in plastid ribosomal genes in Geraniaceae. Genome Biol Evol 8:1824–1838
Whitney SM, Houtz RL, Alonso H (2011a) Advancing our understanding and capacity to engineer nature’s CO2-sequestering enzyme, Rubisco. Plant Physiol 155:27–35
Whitney SM, Sharwood RE, Orr D, White SJ, Alonso H, Galmés J (2011b) Isoleucine 309 acts as a C4 catalytic switch that increases ribulose-1,5-bisphosphate carboxylase/oxygenase (Rubisco) carboxylation rate in Flaveria. Proc Natl Acad Sci USA 108:14688–14693
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA 84:9054–9058
Xie Z, Merchant S (1996) The plastid-encoded ccsA gene is required for heme attachment to chloroplast c-type cytochromes. J Biol Chem 271:4632–4639
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang ZH, Wong WSW, Nielsen R (2005) Bayes empirical Bayes inference of amino acids sites under positive selection. Mol Biol Evol 22:1107–1118
Young JN, Rickaby REM, Kapralov MV, Filatov DA (2012) Adaptive signals in algal Rubisco reveal a history of ancient atmospheric carbon dioxide. Philos Trans R Soc B 367:483–492
Zhang J, Ruhlman TA, Sabir J, Blazier JC, Jansen RK (2015) Coordinated rates of evolution between interacting plastid and nuclear genes in Geraniaceae. Plant Cell 27:563–573
Acknowledgements
JH and GB are members of the Laboratoire Evolution and Diversité Biologique (EDB) part of the LABEX “TULIP” managed by Agence Nationale de la Recherche (ANR-10-LABX-0041). We also acknowledge an Investissement d’Avenir grant of the Agence Nationale de la Recherche (CEBA: ANR-10-LABX-25-01). PAC is funded by a Royal Society University Research Fellowship (Grant number URF120119). This study received support from the PhyloAlps project, and we thank M. Boleda, H. Holota and A. Iribar. We also thank Maria S. Vorontsova for providing plant material. We thank JD Washburn for kindly sharing data before their accessibility on GenBank, and RC Hall for sharing shotgun data for two grass species.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Piot, A., Hackel, J., Christin, PA. et al. One-third of the plastid genes evolved under positive selection in PACMAD grasses. Planta 247, 255–266 (2018). https://doi.org/10.1007/s00425-017-2781-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2781-x