Abstract
Main conclusion
The complete chloroplast genome of two colchicine medicinal plants is reported for the first time. Deletion of ycf 15 gene occurred only in Colchicum but not in Gloriosa and suggests this as a potential marker for delineating the two species.
Colchicum autumnale L. and Gloriosa superba L. are well-known sources of colchicine, a type of alkaloid and an ancient anti-inflammatory drug used to prevent gout. Accordingly, this alkaloid has been used as a chemical marker for identifying the expanded Colchicaceae family. In the present study, we report the complete chloroplast genome (cpDNA) sequence of two colchicine medicinal plants (G. superba and C. autumnale) that belong to the tribe Colchiceae of the Colchicaceae family. In C. autumnale, the circular double-stranded cpDNA sequence of 156,462 bp consists of two inverted repeat (IR) regions of 27,741 bp each, a large single-copy region (LSC) of 84,246 bp, and a small single-copy region (SSC) of 16,734 bp. The cpDNA sequence of G. superba is longer than that of C. autumnale (157,924 bp), which consists of two IRs (28,063 bp), an SSC (16,786 bp), and an LSC (85,012 bp). Significant structural differences between them were observed in the ycf15 gene. ycf15 gene was absent from C. autumnale cpDNA and affected the length of the chloroplast genome between the species. Furthermore, this gene loss event was specific to the expanded genus of Colchicum sensu Vinnersten and Manning. Therefore, this gene may be an effective and powerful molecular marker for identifying the Colchicum genus within the family.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Chloroplasts, responsible for photosynthesis in green plants, are cytoplasmic organelles beside the nucleus and the mitochondrion in plant cells. A typical chloroplast genome in angiosperm is double-stranded circular DNA including 120–130 genes and ranging from 120 to 170 kb (Odintsova and Yurina 2006; Ruhlman and Jansen 2014). This genome is generally composed of two inverted repeat copies (IRa and IRb) ranging 25 kb in size separated by one large single-copy region (LSC) and a small single-copy region (SSC) with dissimilar size ~80 and ~20 kb, respectively (Palmer 1991; Raubeson and Jansen 2005; Ruhlman and Jansen 2014). The evolutionary of chloroplast genomes in angiosperm are provided by structure, gene content and organization such as plastid genome rearrangement of Fabaceae, Asteraceae, Poaceae, Oleaceae, and Gesneriaceae (Saski et al. 2005; Kim et al. 2005; Jansen and Palmer 1987; Doyle et al. 1992; Hachtel et al. 1991; Lee et al. 2007; Hoot and Palmer 1994). During the evolution of land plants, many genes were lost and transferred to the nucleus or became pseudogenes. For example, the chloroplast genomes of Orchidaceae species revealed that the ndh genes were deleted or truncated (Wu et al. 2009). Interestingly, the ndh genes were also nonfunctional in diverse plant lineages including Pinus thunbergii, Keteleeria davidiana, Ephedra equisetina, and Welwitschia mirabilis (Wakasugi et al. 1994; McCoy et al. 2008). In addition, ycf15 and ycf68 are nonfunctional in several chloroplast genomes (Guo et al. 2007; Steane 2005; Schmitz-Linneweber et al. 2001). Chloroplast DNA has been a potential candidate for plant evolutionary studies because of its simple structure, highly conserved sequence, and maternal inheritance (Tian and Li 2002).
Colchicum autumnale L. and Gloriosa superba L. are commonly important sources of colchicine which was originally extracted from bulbs and seeds. It is a type of alkaloid used for the treatment of gout and rheumatism, painful muscles, inflammation, and patients with familial Mediterranean fever (FMF) (Kim et al. 2003; Lange et al. 2001; Touitou et al. 2008; Ade and Rai 2010). Since 2009, colchicine has been used in drugs under approval of Food and Drug Administration (FDA, USA). On the other hand, colchicine has been considered a chemical marker for identifying the Colchicaceae family within Liliales (Vinnersten and Larsson 2010). Based on phylogenetic studies, both species belong to the same tribe Colchiceae of subfamily Wurmbeoideae (Vinnersten and Manning 2007; Nguyen et al. 2013). C. autumnale L., a native plant to South and Central Europe and Africa, is herbaceous perennial and has a corm consisting of a brown membrane or scales (Baker 2001). G. superba L. is a perennial flowering plant with tubers (underground stem) and distributed throughout Southern Africa and tropical Asia including foothills of Himalayas, Burma, Indonesia, and so on (Gec et al. 2002; Ade and Rai 2010).
Our understanding of the chloroplast genome organization and evolution has improved due to advances in next-generation sequencing (NGS) techniques. Currently, over 300 chloroplast genomes are available in The Chloroplast Genome Database (http://chloroplast.ocean.washington.edu/tools/cpbase/run). However, few genomics studies have been conducted on medicinal plants while natural plant products are commonly used for drug development. Up to date, there were a few studies and projects focusing on chloroplast genomes of medicinal plants such as Korean Ginseng Panax ginseng C. A. Mey (Kim and Lee 2004), Chinese sage Salvia miltiorrhiza Bunge (Qian et al. 2013), and Mongolia medicine Artemisia frigida (Liu et al. 2013). The traditional Chinese medicine project has established a foundation for the development of natural medicines and the selection of cultivars with good traits based on genomic research (Chen et al. 2011). Additionally, 12 medicinal plant species were examined within the Medicinal Plant Genomics Resource project (http://medicinalplantgenomics.msu.edu) to obtain genomic data and provide novel information to the community regarding genes and markers for medicinal compound synthesis.
Although C. autumnale and G. superba play an importantly commercial role especially in medicinal industries and genetic engineering (Brown 1995), their genomic studies have not been performed. Therefore, here we analyzed the complete chloroplast genomes sequences of both species and it is the first record in colchicine produced plants and Colchicaceae family. The genome organization, gene contents, and order were compared between both species and among previously reported cpDNA within Liliales to understand the genomic feature of Colchicaceae in Liliales. Especially, we discussed about the finding of gene loss only occurred in Colchicum sensu Vinnersten and Manning, and suggested it as a useful molecular marker which can be easily detected in this genus.
Materials and methods
DNA extraction, sequencing and assembly of chloroplast genome sequences
Plant materials used in this study were collected through KNRRC (Medicinal Plant Resources Bank NRF-2010-0005790), supported by Korea Research Foundation, to which resources were provided by the Ministry of Education, Science and Technology in 2013. Fresh young leaves were collected from C. autumnale and G. superba cultivated in a greenhouse, and a voucher specimen was deposited in the herbarium of Gachon University (GCU). Total genomic DNA was isolated using the DNeasy Plant Mini Kit (Qiagen, Seoul, South Korea) from 1 g fresh leaves. The DNA concentration was determined using a UV–visible spectrophotometer (BioSpec-nano; Shimadzu Corp. Japan). High-quality DNA was used as a template for Hiseq 2000. Geneious (version 6.1, Biomatters Ltd. Auckland, New Zealand) was used to assemble pair-end reads from the NGS data based on default setting. We used Alstroemeria aurea (KC968976) (Kim and Kim 2013) as a reference sequence to align the contigs and identify gaps. To fill the gap, Sanger methods were applied and used to identify the borders of the IR, LSC, and SSC regions. PCR products were purified using the PCRquick-spin Kit according to the manufacturer’s protocol (Intron Biotechnology, Korea). We sequenced PCR products using the BigDye Terminator Cycle Sequencing Kit (Perkin Elmer Applied Biosystems). Summary of the sequencing data was described in Supplementary Table 1.
Annotation, codon usage, and comparative analysis
The complete sequences of two species were annotated using Geneious ver. 6.1 program, and manual correction was performed to identify the gene and exon boundaries. All tRNAs were confirmed using the tRNA scan-SE search server (Schattner et al. 2005). Other protein coding genes were verified by BLAST search on the NCBI website (http://blast.ncbi.nlm.nih.gov/), and manual correction for start and stop codons were conducted. GenomeVx software (Conant and Wolfe 2008) was used to construct the visual cpDNA map. The codon usage for all exons of protein coding genes excluding pseudogenes was examined using MEGA5 (Tamura et al. 2011). To calculate the ratio of synonymous (dS) and nonsynonymous (dN) substitutions among reported cpDNA in Liliales, BioEdit (Hall 1999) and DnaSPs software (Librado and Rozas 2009) were used. For the comparative analysis, we used six and four complete chloroplast genome sequences from Liliales and other monocot orders, respectively (Table 1).
To examine the gene loss of ycf15, a pair of primers was designed to amplify regions from ycf2 to trnL-CAA using Primer3 software (Untergrasser et al. 2012) based on the sequence assembly results of C. autumnale and G. superba. PCR amplifications were performed in 25-µl reactions containing 10× PCR reaction buffer, 0.5 U e-Taq DNA polymerase (SolGent Co. Ltd, Korea), 25 mM magnesium chloride (MgCl2), 2.5 mM deoxynucleotide triphosphates (dNTPs), 10 pM primer, and 50–100 ng template DNA. The PCR reactions consisted of one initial denaturation step of 3 min at 94 °C followed by 25 cycles of denaturation at 94 °C for 1 min, annealing at 56 °C for 30 s, extension at 72 °C for 20 s, and a final extension of 5 min at 72 °C. The PCR products were sequenced and aligned using MUSCLE (Edgar 2004), and further manual adjustments were made if necessary.
Examination of repeat units
Repeat sequences were analyzed using REPuter (Kurtz and Schleiermacher 1999) in both genomes using a minimum repeat size of 18–20 bp. The GRAMENE Ssrtool (Temnykh et al. 2001) program was used to identify SSRs.
Results
Chloroplast genome assembly and features
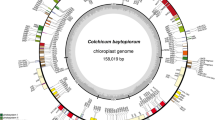
For C. autumnale, a total of 9,314,386 reads with an average length of 91 bp were generated from the Illumina HiSeq 2000, and 715,966 reads (7.68 %) were assembled to the A. aurea reference genome (KC 968976). The cpDNA of C. autumnale was 156,462 bp (KP 125337, Fig. 1) in length and composed of a pair of inverted repeats (IRs) of 27,741 bp, which were divided by SSC and LSC of 84,246 and 16,734 bp, respectively (Fig. 1; Table 1). The cpDNA sequence consisted of 132 coding genes, including 87 protein coding genes, 37 transfer RNA (tRNA) genes, and eight ribosomal RNA (rRNA) genes. Among the 132 genes, four rRNA, eight tRNA, and seven coding genes were duplicated in the IR regions (Supplementary Table 2).
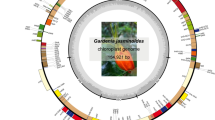
Gene maps of the C. autumnale L. and G. superba L. chloroplast genomes. Genes shown outside of the outer circle are transcribed counter-clockwise, whereas those shown inside are transcribed clockwise. The thick lines in small circles indicate the inverted repeats (IR) regions. The asterisks indicate genes containing intron(s). The arrow indicate ycf15 gene which is not present in the C. autumnale chloroplast genome
The same NGS platform was applied in the sequencing of the G. superba chloroplast genome, which generated 6,464,451 reads with an average length of 91 bp, of which 19,973 reads (0.3 %) were assembled for genome sequencing. The chloroplast genome size of G. superba was 157,924 bp (KP 125338) and displayed the typical angiosperm cpDNA structure, consisting of the LSC (85,012 bp), SSC (16,786 bp), and two IR copies (28,063 bp) (Fig. 1; Table 1). It also included 134 coding genes, of which 89 were protein coding genes, and 37 and 8 were distinct tRNA and rRNA genes, respectively. The IR regions contained 20 duplicates including four rRNA, eight tRNA, and eight coding genes (Supplementary Table 2).
Comparison of the chloroplast genome sequences between C. autumnale and G. superba revealed that the overall A+T and G+C contents in both whole chloroplast genomes were similar 62.4 and 37.6 %, respectively (Table 1). These percentages are within the range of typical monocots 61–63 % for AT content and 31–38 % for GC content. In both genomes, 19 genes contained introns, 16 of which contained one intron such as atpF, ndhA, ndhB, petB, petD, rpl16, rpl2, rpoC1, rps12, rps16, trnA-UGC, trnG-GCC, trnI-GAU, trnK-UUU, trnL-UAA or trnV-UAC, while two genes (clpP and ycf3) were possessing two introns. For the rps12 gene, the first exon was located in the LSC region, and the second exon was duplicated in the IR regions. Among the genes containing an intron, the trnK-UUU gene had the largest intron (2675–2711 bp) containing matK.
The gene contents were similar between both genomes, excluding ycf15 gene loss found in C. autumnale cpDNA. The infA gene was present as a pseudogene in both genomes because of several stop codons. In addition, an incomplete duplicated copy of ycf1 located at the junction between IRb and SSC was present in both genomes. In the chloroplast genome of G. superba, ycf68 and ycf15 in the IR were thought to be pseudogenes based on the presence of internal stop codons. However, in the C. autumnale chloroplast genome, ycf68 did not contain internal stop codons, and ycf15 was completely deleted (Fig. 2).
The genome size of G. superba was slightly larger than that of C. autumnale (1,154 bp) affected by the presence of the ycf15 gene, and variable lengths of ycf2 and ycf1. Among the available cpDNA sequences of the order Liliales, G. superba had the largest genome size. The A+T and G+C contents in the complete genomes were quite stable and similar among Liliales species (Table 1).
To confirm this hypothetical protein coding gene loss in the tribe Colchiceae of family Colchicaceae, we designed a pair of primers to amplify the region from ycf2 to trnL-CAA (for-TGGATCAAATGACAAAGACA; rev-CTAAAGAGCGTGGAGGTTCG; Fig. 3a). We observed that two different length PCR products, shorter (400–500 bp) and longer (1000–1100 bp) were generated from the same primer set in the Colchicaceae. Sequences of both PCR products indicated that only Colchicum and its alliances had a shorter PCR product and lack ycf15 in their chloroplast genomes against the remaining taxa of tribe Colchiceae (Hexacyrtis, Ornithoglossum, Sandersonia, Gloriosa) and the related tribe Angullarieae (Wurmbea) which had a ycf15 gene on their genomes (Fig. 3b, c). Alignment of this region was shown in more detail, and the location of ycf15 loss was highlighted in Fig. 4.
Analyses of the missing ycf15 gene. a Primer positions were used for amplifying the ycf15 region (psi pseudogenes). b Results of PCR amplification of ycf15. c Modified phylogenetic tree of the Colchicaceae family from Nguyen et al. (2013) result. Bootstrap and posterior probability (PP) values are shown above and below each branch, respectively. The numbers beside taxa name indicate the number of stop codons present in ycf15 sequences. The asterisk beside taxa name indicate the ycf15 gene missing in that species. L, Ladder (100-bp ladder, BIOFACT); W, Wurmbea; G, Gloriosa; S, Sandersonia; O, Ornithoglossum; H, Hexacyrtis; A, Androcymbium; M, Merendera; B, Bulbocodium; C1, C2 and C3, Colchicum
Codon usage
All 78 protein coding genes of C. autumnale and G. superba in the chloroplast genome were encoded by 22,905 and 22,806 codons, respectively (Table 2). The highest codon usage was commonly recorded for isoleucine (especially the ATT codon), and the two genomes shared the same number of stop codons (36—TAA; 24—TAG; 18—UGA). For the start codon, ATG was the most common among coding genes, with the exception of ATC for ndhD, ACG for rpl2, ACT for rps2, GTG for rps19, and ATT for ycf15.
Simple sequence repeats (SSRs) and long repetitive sequences
There was no significant difference in SSRs between C. autumnale and G. superba chloroplast genomes in which 58 and 56 SSRs with lengths of at least 10 bp were found, respectively (Table 3). The majority of mononucleotide repeats were A–T rich, which were common in the two species, while polyG or polyT repeats were rare. Generally, the number of dinucleotide repeat units was slightly higher than those of other repeat units, such as tri-, tetra-, penta- and hexa-nucleotides, observed in the chloroplast genomes of colchicine plants. SSRs were more abundant in noncoding regions than in protein coding genes.
Based on the repeat sequences in C. autumnale and G. superba plastid genomes (Table 4), they were divided into three categories including tandem repeat, forward repeat, and palindromic repeat. Three tandem repeats, four forward repeats, and three palindromic sequences were detected in the C. autumnale chloroplast genome. Similar to C. autumnale, one tandem sequence, four forward repeats, six palindromic sequences, and one reverse repeat were found in G. superba cpDNA. By comparing these plastid genomes, three palindromic sequence repeats were identified at the same location namely, ccsA-ndhD, trnH-GUG-psbA, and psbT-psbN regions with similar lengths from 42 to 62 bp. In addition, three palindromic sequences were detected in G. superba at the rrn16-trnI-GAU, accD-psaI, and rpl32-trnL-UAG spacer. Three tandem repeats were found in C. autumnale with a maximum length of 24 bp, and there was only one long tandem repeat (55 bp) in G. superba. We also found the same type of forward repeats (25 bp) at the same location in the IGS trnS-GCU and trnS-UGA in both plastid genomes.
Base substitution ratios among Liliales species
Nonsynonymous (dN) and synonymous (dS) substitution rates (dN/dS) are indicators of evolutionary rate and natural selection (Yang and Nielsen 2000). In this report, the substitution ratios among Liliales members were calculated (Fig. 5). We used Dioscorea elephantipes (Dioscoreaceae, Dioscoreales) as a reference instead of Asparagales, because our genomic understanding of Asparagales is restricted to Orchidaceae, which possesses many pseudogenes or lacks the ndh gene (Wu et al. 2009). The majority of genes showed dN/dS ratios of less than 1 (except ycf2). In the SSC region, the highest and lowest ratios were found in ndhD and psaC, respectively. The other ndh genes were stable from 0.1 to 0.3. The highest dN/dS ratios were observed at psbK and rps8 in the LSC and were approximately 0.8–0.9. Additionally, the substitution rates were zero in psbI, psbM, psbJ, psbL, psbE, petG, atpH, all of which are photosynthetic genes, in both C. autumnale and G. superba.
Comparison of IR junction
We compared the IR junction among the Liliales (Fig. 6) and found variable gene composition especially in the IR-LSC junction. However, C. autumnale and G. superba chloroplast genomes showed a similar pattern among them. The IRa-LSC junction was located at a part of rps19 in both genomes, and ndhF and ycf1 gene was overlapped at IRb-SSC junction (Fig. 6).
Discussion
First records of the complete chloroplast genome in colchicine medicinal plants
We firstly analyzed the complete chloroplast genomes of two colchicine plants, C. autumnale and G. superba, and discussed about evolutionary implication and the probability for usefulness of this data for identifying the related taxa from the result. Both chloroplast genomes were very similar in the structure except the gene loss of ycf15 in C. autumnale.
The gene infA encodes translation initiation factor 1, which has been lost completely or is present as a pseudogene in the majority of angiosperm (Millen et al. 2001). In this study, we identified two and three internal stop codons in the infA sequence of C. autumnale and G. superba chloroplast genomes, respectively, suggesting that infA is a pseudogene (Fig. 2a). This has also been observed in the chloroplast genomes of Lilium longiflorum (Kim and Kim 2013) and Veratrum patulum (Do et al. 2013). However, compared with other Liliales members, the infA gene was absent in Smilax china (Liu et al. 2012) and A. aurea (Kim and Kim 2013). In contrast, the complete sequence of the infA gene without any internal stop codons was present in Chionographis japonica (Bodin et al. 2013) and Paris verticillata (Do et al. 2014). Therefore, further studies on the evolution of the infA gene in Liliales are required to improve our understanding of this observation. We also observed that the two hypothetical coding genes of ycf68 and ycf15 were truncated by five and three internal stop codons, respectively, in G. superba (Fig. 2b, c). This situation was not only detected in G. superba, but also in L. longiflorum (Kim and Kim 2013), C. japonica (Bodin et al. 2013), and V. patulum (Do et al. 2013) of Liliales and in Cymbidium of Asparagales (Yang et al. 2013). However, in C. autumnale cpDNA, ycf68 was present as a protein coding gene without internal stop codon (Fig. 2b). This in-frame sequence of this gene was present and could represent a functional protein coding gene in the Nymphaeales (Raubeson et al. 2007) and in the gymnosperms P. thunbergii (Wakasugi et al. 1994) and P. koraiensis (AY228468). ycf15 was absent in the C. autumnale chloroplast genome (Fig. 4). The role of ycf15 as a protein coding gene remains unclear and requires further study.
ycf15 gene loss reflecting the phylogenetic relationship of colchicine plants
Remarkable difference of gene content and genome size between Gloriosa and Colchicum was the loss of ycf15. Using the expanded sampling, we confirmed that ycf15 gene loss was occurred only in the expanded Colchicum genus including Colchicum, Androcymbium, Bulbocodium and Merendera, whereas it is still remained in Ornithoglossum, Sandersonia, Hexacyrtis and Gloriosa, as well as Wurmbea (tribe Anguilarieae) (Fig. 3c). As a result, missing ycf15 may be an effective molecular marker for distinguishing this expanded genus recognized Vinnersten and Manning (2007) from the Colchicaceae family. Four closely related genera of Colchicum, Androcymbium, Merendera and Bulbocodium have been considered to be combined into one genus Colchicum sensu lato based on molecular evidence (Vinnersten and Manning 2007; Nguyen et al. 2013). However, until now this suggestion has not been completely agreed (del-Hoyo and Pedrola-Monfort 2008). Therefore, a finding of specific deletion of ycf15 on chloroplast genome provides important evidence at molecular level to support their phylogenetic relationship and circumscription.
Implications of microsatellites in both chloroplast genomes
SSRs are short repeat motifs (of at least 10 bp) found in DNA sequences (Katti et al. 2001; Shanker et al. 2007). Identification of SSRs is important to develop molecular markers and to map traits of economic, medical, or ecological interest. Early studies on SSRs of chloroplast genome in rice showed that they are abundant in non-coding regions through the genome (Rajendrakumar et al. 2007). In the G. superba chloroplast genome, multiple repeat sequences are found in the trnC-GCA-petN intergenic spacer and ycf1 gene (three repeat sequences for each). Similarly, in the C. autumnale plastid genome, the clpP intron 2 and ycf1 possess three and five repeat units, respectively. In addition, these colchicine plants chloroplast genomes share several SSRs distributed in the same region. For example, mononucleotides were found in trnK-UUU-rps16, rpoB-trnC-GCA, overlapped ycf1-ndhF, ndhF-rpl32, and ycf1; dinucleotides were detected in ndhH and ycf2, and trinucleotides (TCC) were found in the rpoA gene (Table 3). It is interesting that both chloroplast genomes shared several chloroplast SSRs co-located in noncoding regions and it may be variable in other Colchicaceae species. This information of SSRs can be used as a reference when the molecular identification marker development and application in the population genetics are tried in future studies.
IR expansion and its evolutionary implications in Liliales
The boundaries of IR regions with LSC and SSC play a critical role in expansion and contraction of the genome; thus, the gene contents of IRs vary significantly (Goulding et al. 1996; Plunkett and Downie 2000). Eight complete plastid genome sequences including the data generated in the present study are available currently and are represented from five families in the order Liliales, namely, C. autumnale and G. superba (Colchicaceae), A. aurea (Alstroemeriacae), L. longiflorum (Liliaceae), S. china (Smilacaceae), C. japonica, V. patulum and P. verticillata (Melanthiaceae). The IR boundaries between the C. autumnale and G. superba chloroplast genomes showed a similar pattern (Fig. 6). The IRB-LSC junction is located within the sequence of rps19 in both genomes, and part of the rps19 gene was copied in IRA and its 3′ end adjacent psbA in LSC, which is common in the typical monocot chloroplast genome structure. This junction was similar to A. aurea, which shares a close relationship with the Colchicaceae family and L. longiflorum. However, it was expanded to include the whole rps19 gene and part of rpl22 in S. china. In case of family Melanthiaceae, which is a sister clade of Liliaceae and Smilacaceae, it was varied within the family. In V. patulum, it was located in the intergenic spacer between rps19 and trnH-GUG (Fig. 6). In contrast, it was expanded to part of rps3, and the entire rps19 and rpl22 genes in the chloroplast genomes of C. japonica and P. verticillata. Comparative studies among cpDNAs from eight species in Liliales indicated that they share the same structures and gene orders in the SSC region and IR-SSC junction (JLA, JLB) at the position of ycf1 region. However, the overlapped length between the ycf1 pseudogene and ndhF (JSB) varied among the species; e.g., C. autumnale (39 bp), G. superba (53 bp), A. aurea (78 bp), L. longiflorum (17 bp), and C. japonica (5 bp). The overlap of these genes was also observed in the other chloroplast genomes of angiosperm species (Yang et al. 2010).
In addition, the trnH-GUG-psbA spacer region located in the junction between IRA and LSC is common in monocots. The trnH-GUG-psbA chloroplast intergenetic spacer region has been used in many DNA barcoding studies (Pang et al. 2012) and has become a frequently used marker for molecular phylogenetic studies at the lower taxonomic level (Degtjareva et al. 2012). However, this study showed various gene compositions in these regions among the species of Liliales. Therefore, the trnH-GUG-psbA intergenetic spacer may not be a good candidate for resolving phylogenetic relationships (at least in this order), because this spacer showed significantly different gene contents and lengths, which cause the complicate alignment and analysis.
Author contribution statement
Nguyen PAT carried out the analyses and drafted the manuscript. Kim JS participated in the design and coordination of the study, advised to analyze the data, and drafted the manuscript. Kim JH participated in the coordination of the study and helped to draft the manuscript. All authors read and approved the final manuscript.
References
Ade R, Rai MK (2010) Review: colchicine, current advances and future prospects. Bioscience 2(2):90–96
Baker J (2001) The medicinal flora of Britain and Northwestern Europe. Winter Press, UK, p 507
Bodin SS, Kim JS, Kim JH (2013) Complete chloroplast genome of Chionographis japonica (Willd.) Maxim (Melanthiaceae): comparative genomics and evaluation of universal primers of Liliales. Plant Mol Biol Rep 31(6):1407–1421
Brown D (1995) Encyclopedia of herbs and their uses. Dorling kindersley limited, London, pp 423–425
Chen S, Li X, Guo X, Li Q (2011) An introduction to the medicinal plant genome project. Front Med 5(2):178–184
Conant GC, Wolfe KH (2008) GenomeVx: simple web-based creation of editable circular chromosome maps. Bioinformatics 24:861–862
Degtjareva G, Logacheva M, Samigullin T, Terentieva E, Valiejo-Roman C (2012) Organization of chloroplast psbA-trnH intergenic spacer in dicotyledonous angiosperms of the family Umbelliferae. Biochemistry (Mosc) 77(9):1056–1064
del-Hoyo A, Pedrola-Monfort J (2008) Phylogeny of Androcymbium (Colchicaceae) based on morphology and DNA sequences. Plant Syst Evol 273:151–167
Do HDK, Kim JS, Kim JH (2013) Comparative genomics of four Liliales families inferred from the complete chloroplast genome sequence of Veratrum patulum O. Loes. (Melanthiaceae). Gene 530(3):229–235
Do HDK, Kim JS, Kim JH (2014) A trnI-CAU triplication event in the complete chloroplast genome of Paris verticillata M. Bieb. (Melanthiaceae, Liliales). Genome Biol Evol 6(7):1699–1706
Doyle JJ, Davis JI, Soreng RJ, Garvin D, Anderson MJ (1992) Chloroplast DNA inversions and the origin of the grass family (Poaceae). Proc Natl Acad Sci USA 89:7722–7726
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughout. Nucleic Acids Res 32(5):1792–1797
GEC, GUIDE, MSU (2002) Conservation of rare and endangered biodiversity of Gujarat. A final report by Gujarat ecological commission. Gujarat, India, pp 428
Goulding S, Olmstead R, Morden C, Wolfe K (1996) Ebb and flow of the chloroplast inverted repeat. Mol Gen Genet 252:195–206
Guo X, Castillo-Ramirez S, Gonzalez V et al (2007) Rapid evolutionary change of common bean (Phaseolus vulgaris L.) plastome, and the genomic diversification of legume chloroplasts. BMC Genom 8:228
Hachtel W, Neuss A, Vomstein J (1991) A Chloroplast DNA inversion marks an evolutionary split in the genus Oenothera. Evolution 45:1050–1052
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hoot SB, Palmer JD (1994) Structural rearrangements, including parallel inversions, within the chloroplast genome of anemone and related genera. J Mol Evol 38:274–281
Jansen RK, Palmer JD (1987) A chloroplast DNA inversion marks an ancient evolutionary split in the sunflower family (Asteraceae). Proc Natl Acad Sci USA 84:5818–5822
Katti MV, Ranjekar PK, Gupta VS (2001) Differential distribution of simple sequence repeats in eukaryotic genome sequences. Mol Biol Ecol 18:1161–1167
Kim JS, Kim JH (2013) Comparative genome analysis and phylogenetic relationship of order Liliales insight from the complete plastid genome sequences of two lilies (Lilium longiflorum and Alstroemeria aurea). PLoS One 8(6):e68180
Kim KJ, Lee HL (2004) Complete chloroplast genome sequences from Korean ginseng (Panax schinseng Nees) and comparative analysis of sequence evolution among 17 vascular plants. DNA Res 11:247–261
Kim KY, Schumacher HR, Hunsche E, Wertheimer AI, Kong SX (2003) A literature review of the epidemiology and treatment of acute gout. Clin Ther 25:1593–1617
Kim KJ, Choi KS, Jansen RK (2005) Two chloroplast DNA inversions originated simultaneously during the early evolution of the sunflower family (Asteraceae). Mol Biol Evol 22:1783–1792
Kurtz S, Schleiermacher C (1999) REPuter: fast computation of maximal repeats in complete genomes. Bioinformatics 15:426–427
Lange U, Schumann C, Schmidt KL (2001) Current aspects of colchicine therapy, classical indications and new therapeutic uses. Eur J Med Res 6:150–160
Lee HL, Jansen RK, Chumley TW, Kim KJ (2007) Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol Biol Evol 24:1161–1180
Librado P, Rozas J (2009) DnaSP v5. A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Liu J, Qi ZC, Zhao YP, Fu CX, Xiang QY (2012) Complete cpDNA genome sequence of Smilax china and phylogenetic placement of Liliales—influences of gene partitions and taxon sampling. Mol Phylogenet Evol 64(3):545–562
Liu Y, Huo N, Dong L, Wang Y, Zhang S et al (2013) Complete chloroplast genome sequences of Mongolia medicine Artemisia frigida and phylogenetic relationships with other plants. PLoS One 8(2):e57533
McCoy S, Kuehl J, Boore J, Raubeson L (2008) The complete plastid genome sequence of Welwitschia mirabilis: an unusually compact plastome with accelerated divergence rates. BMC Evol Biol 8:130–146
Millen RS, Olmstead RG, Adams KL, Palmer JD, Lao NT et al (2001) Many parallel losses of infA from chloroplast DNA during angiosperm evolution with multiple independent transfers to the nucleus. Plant Cell 13(3):645–658
Nguyen PAT, Kim JS, Kim JH (2013) Molecular phylogenetic relationship and implications for the circumscription of Colchicaceae (Liliales). Bot J Linn Soc 172:255–269
Odintsova MS, Yurina NP (2006) Chloroplast genomics of land plants and algae, biotechnological applications of photosynthetic proteins: biochips, biosensors and biodevices. Springer, US, pp 57–72
Palmer JD (1991) Plastid chromosomes: structure and evolution. In: Bogorad L (ed) Molecular biology of plastids. Academic Press, San Diego, pp 5–53
Pang X, Liu C, Shi L, Liu R, Liang D et al (2012) Utility of the trnH–psbA intergenic spacer region and its combinations as plant DNA barcodes: a meta-analysis. PLoS One 7(11):e48833
Plunkett G, Downie S (2000) Expansion and contraction of the chloroplast inverted repeat in Apiaceae subfamily Apioideae. Syst Bot 25:648–667
Qian J, Song J, Gao H, Zhu Y, Xu J et al (2013) The complete chloroplast genome sequence of the medicinal plant Salvia miltiorrhiza. PLoS One 8(2):e57607
Rajendrakumar P, Biswal AK, Balachandran SM, Srinivasarao K, Sundaram RM (2007) Simple sequence repeats in organellar genomes of rice: frequency and distribution in genic and intergenic regions. Bioinformatics 23:1–4
Raubeson LA, Jansen RK (2005) Chloroplast genomes of plants. In: Henry RJ (ed) Plant diversity and evolution: genotypic and phenotypic variation in higher plants. CABI, Cambridge, pp 45–68
Raubeson LA, Peery R, Chumley TW, Dziubek C, Fourcade HM, Boore JL, Jansen RK (2007) Comparative chloroplast genomics: analyses including new sequences from the angiosperms Nuphar advena and Ranunculus macranthus. BMC Genomics 8(1):174
Ruhlman TA, Jansen (2014) The plastid genomes of flowering plant. Methods Mol Biol 1132:3–38
Saski C, Lee SB, Daniell H, Wood TC, Tomkins J, Kim HG, Jansen RK (2005) Complete chloroplast genome sequence of Glycine max and comparative analyses with other legume genomes. Plant Mol Biol 59:309–322
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:686–689
Schmitz-Linneweber C, Maier RM, Alcaraz JP et al (2001) The plastid chromosome of spinach (Spinacia oleracea): complete nucleotide sequence and gene organization. Plant Mol Biol 45(3):307–315
Shanker A, Singh A, Sharma V (2007) In silico mining in expressed sequences of Neurospora crassa for identification and abundance of microsatellites. Microbiol Res 162:250–256
Steane DA (2005) Complete nucleotide sequence of the chloroplast genome from the Tasmanian blue gum, Eucalyptus globulus (Myrtaceae). DNA Res 12(3):215–220
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Temnykh et al (2001) Computational and experimental analysis of microsatellites in rice (Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11:1441–1452
Tian X, Li DZ (2002) Application of DNA sequences in plant phylogenetic study. Acta Bot Yunnan 24:170–184
Touitou S, Savic RJ, Mathews G, Grateau MF, McDermott (2008) Fifth international congress on familial Mediterranean fever and systemic auto inflammatory diseases. Expert Rev Clin Immunol 4:425–428
Untergrasser A et al (2012) Primer3-new capabilities and interfaces. Nucleic Acids Res 40(15):e115
Vinnersten A, Larsson S (2010) Colchicine is still a chemical marker for the expanded Colchicaceae. Biochem Syst Ecol 38:1193–1198
Vinnersten A, Manning J (2007) A new classification of Colchicaceae. Taxon 56:171–178
Wakasugi T, Tsudzuki J, Ito S, Nakashima K, Tsudzuki T et al (1994) Loss of all ndh genes as determined by sequencing the entire chloroplast genome of the black pine Pinus thunbergii. Proc Natl Acad Sci USA 91:9794–9798
Wu CS, Lai YT, Lin CP, Wang YN, Chaw SM (2009) Evolution of reduced and compact chloroplast genomes (cpDNAs) in gnetophytes: selection toward a lower-cost strategy. Mol Phylogenet Evol 52:115–124
Yang ZH, Nielsen R (2000) Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol Biol Evol 17:32–43
Yang M, Zhang X, Liu G, Yin Y, Chen K, Yun Q, Zhao D, Al-Mssallem IS, Yu J (2010) The complete chloroplast genome sequence of date palm (Phoenix dactylifera L.). PLoS One 5(9):e12762
Yang JB, Tang M, Li HT, Zhang JR, Li DJ (2013) Complete chloroplast genome of the genus Cymbidium: lights into the species identification, phylogenetic implications and population genetic analyses. BMC Evol Biol 13:84
Acknowledgments
This study was supported by the National Research Foundation of Korea (NRF) Grant Foundation (MEST 2010-002913).
Author information
Authors and Affiliations
Corresponding author
Additional information
P. A. T. Nguyen and J. S. Kim contributed equally to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Nguyen, P.A.T., Kim, J.S. & Kim, JH. The complete chloroplast genome of colchicine plants (Colchicum autumnale L. and Gloriosa superba L.) and its application for identifying the genus. Planta 242, 223–237 (2015). https://doi.org/10.1007/s00425-015-2303-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-015-2303-7