Abstract
Main conclusion
The present review compiles the up-to-date knowledge on vanillin biosynthesis in plant systems to focus principally on the enzymatic reactions of in planta vanillin biosynthetic pathway and to find out its impact and prospect in future research in this field.
Vanillin, a very popular flavouring compound, is widely used throughout the world. The principal natural resource of vanillin is the cured vanilla pods. Due to the high demand of vanillin as a flavouring agent, it is necessary to explore its biosynthetic enzymes and genes, so that improvement in its commercial production can be achieved through metabolic engineering. In spite of significant advancement in elucidating vanillin biosynthetic pathway in the last two decades, no conclusive demonstration had been reported yet for plant system. Several biosynthetic enzymes have been worked upon but divergences in published reports, particularly in characterizing the crucial biochemical steps of vanillin biosynthesis, such as side-chain shortening, methylation, and glucoside formation and have created a space for discussion. Recently, published reviews on vanillin biosynthesis have focused mainly on the biotechnological approaches and bioconversion in microbial systems. This review, however, aims to compile in brief the overall vanillin biosynthetic route and present a comparative as well as comprehensive description of enzymes involved in the pathway in Vanilla planifolia and other plants. Special emphasis has been given on the key enzymatic biochemical reactions that have been investigated extensively. Finally, the present standpoint and future prospects have been highlighted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Use of flavours and fragrances is an important area of food, pharmaceutical, and cosmetic industries. Among all the other natural flavouring agents, vanillin (4-hydroxy-3-methoxybenzaldehyde) is one of the most admired and widely used flavouring natural products throughout the world. Vanillin biosynthesis has caught plant biologist’s interest for several decades due to its direct usage in food, beverages, medicines, and cosmetics as a flavouring agent, as well as its use as a preservative. However, its biosynthetic route remained unexplored until some excellent work was conducted in the recent years. Outstanding reviews were published previously on extraction, isolation, and quantification of biovanillin and its production through biotechnological approaches in microbial systems (Sinha et al. 2008; Kaur and Chakraborty 2013). In a recent advanced review, latest biotechnological advances of microbial bioconversion of ferulic acid, eugenol, isoeugenol, and sugars have been discussed thoroughly (Gallage and Møller 2015), but a review of studies on the enzymatic biosynthesis of vanillin in plants is yet to be done. Therefore, the aim of this review is to focus principally on the overall pathway of vanillin biosynthesis in plants, characteristics of biochemical conversions through enzymes involved in this pathway, and the knowledge gaps due to differences in research evidences regarding the key enzymes involved in vanillin production in plant system.
The main source of natural vanillin is the beans of orchid Vanilla planifolia and to a lesser extent of Vanilla tahitienis and Vanilla pompona (Zamzuri and Abd-Aziz 2012). In other plants, including commercial plants (e.g., tobacco), vanillin is found in trace amounts (Makkar and Beeker 1994). In the green beans, vanillin is found exclusively in conjugated form as β-d-glucoside. This conjugated form does not have any characteristic vanilla flavour. The vanilla flavour is only developed during the ‘curing’ process. Usually, vanilla green beans are harvested after 6–8 months of pollination. After harvesting the beans, the ‘curing’ process takes more than 6 months (Walton et al. 2003). During the ‘curing’ process, vanillin-β-d-glucoside comes in contact with β-d-glucosidase and releases free vanillin. Other β-d-glucosides, which are present together with vanillin-β-d-glucoside, are also converted to their free forms (e.g., 4-hydroxybenzaldehyde) (Kanisawa et al. 1994; Dignum et al. 2001).
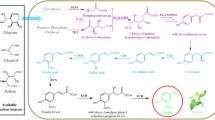
A number of biosynthetic routes for vanillin biosynthesis have been suggested so far and all of them use different phenylpropanoids as precursors. The proposals on metabolic routes that lead to vanillin biosynthesis have been described so far as followed: (1) using coniferin as precursor through C3 side-chain cleavage and hydrolysis of glucoside (Anwar 1963); (2) through CoA-dependent β-oxidative pathway (Zenk 1965); (3) formation of glucovanillin from 4-hydroxybenzyl alcohol glucoside (Tokoro et al. 1990; Kanisawa et al. 1994); (4) through non-CoA-dependent non-β-oxidative pathway (Yazaki et al. 1991); (5) CoA-dependent non-β-oxidative pathway (Mitra et al. 1999); and (6) direct enzymatic conversion of ferulic acid to vanillin (Gallage et al. 2014). Recently, an artificial pathway was constructed in Escherichia coli cells, where cheap and accessible carbon sources (e.g., glucose, glycerol, and xylose) were used as primary precursors for vanillin production. In this pathway, the natural plant-based vanillin biosynthetic pathway was partially followed, but instead of phenylpropanoid pathway, the E. coli strain followed the common aromatic pathway of l-tyrosine production, which was then converted to 4-coumaric acid and, thereafter, entered into the ferulic acid mediated vanillin biosynthetic route (Ni et al. 2015).
Different groups of investigators have accepted that vanillin biosynthesis in plants is preceded by the synthesis of phenylalanine via shikimate pathway in green vanilla beans or in cell culture of V. planifolia and that it follows central phenylpropanoid pathway from this point (Dignum et al. 2001). Though several compounds (e.g., caffeic acid, hydroxycinnamic acid, ferulic acid, and 4-coumaric acid) were used as precursors in vanillin biosynthesis experiments, all of them were intermediates of phenylpropanoid pathway. Therefore, it was comprehended that vanillin biosynthesis was parallel to the biosynthesis of other phenylpropanoids mediated molecules, such as benzoic acid and lignin. The critical ambiguities were remaining in the chain shortening step of the vanillin biosynthesis and in the reactions that produce vanillin-β-d-glucoside via vanillin (Walton et al. 2003). The differences in proposals of vanillin biosynthesis suggest that the biosynthetic pathways might be different in diverse conditions or in different systems, but it has to be investigated which route is followed predominantly, and which one will be advantageous in metabolic engineering for vanillin biosynthesis. Up till now, vanillin production from Vanilla orchids has not been cost effective and this kind of production is laborious; only 1 kg of vanillin can be extracted from about 500 kg of Vanilla pods (Gallage et al. 2014). Surprisingly, only 1% of global vanillin production comes from Vanilla orchids and majority is produced by chemical conversion from fossil fuels or by acid hydrolysis of lignin (Walton et al. 2003). Application of metabolic engineering by heterologous expression of native vanillin biosynthetic genes in the microbial system is a prospective biotechnological way to overcome the challenges to natural vanillin production, but it has not been achieved, yet as most parts of the biosynthetic pathways leading to vanillin production are unknown (Gallage et al. 2014). This communication aims to review all the aspects of vanillin biosynthesis in plants, the enzymes involved and future scopes of research on the basis of the published data hitherto, so that it may contribute to the future research on biosynthetic routes and the genes that could be utilized in biotechnological approaches for commercial vanillin production through plants.
Relation between phenylpropanoid pathway and vanillin biosynthesis
Central phenylpropanoid pathway is a vast and diverse pathway that leads to productions of several benzoates and hydroxybenzoates. One of the main reasons of investigations on biosynthetic pathway of vanillin was its relationship with the central phenylpropanoid pathway and benzoates biosynthesis including 4-hydroxybenzoic acid and salicylic acid (2-hydroxybenzoic acid) (Walton et al. 2003). There are some reports, which suggested that vanillin is synthesised from l-phenylalnine via monomeric lignin precursors, such as cinnamic acid, 4-coumaric acid, caffeic acid, and ferulic acid (Gallage et al. 2014). Many of the key phenylpropanoid pathway enzymes are possibly being involved in these catalytic reactions, viz. phenylalanine ammonia lyase (PAL) (Fritz et al. 1976), cinnamic acid-4-hydroxylase (C4H) (Ro et al. 2001), and 4-coumaric acid-4-hydroxylase (C3H) (Schoch et al. 2001). The probable intermediates in these reactions are CoA derivatives as well as shikimate- and quinate esters of coumaric acid and caffeic acid (Gallage et al. 2014). Caffeic acid can be converted to ferulic acid by an O-methyltransferase (4-COMT) (Lam et al. 2007). All these probable intermediates lead to the idea of most crucial step of C2 side-chain shortening (Fig. 1). According to Gallage et al. (2014), side-chain shortening of ferulic acid results in the formation of vanillin by a direct conversion. Recently, a gene Vp VAN has been identified in V. planifolia that encodes an enzyme having C2 side-chain shortening activity, which can convert ferulic acid and its glucoside to vanillin and its glucoside (Gallage et al. 2014). Previously, feeding of radio-labelled ferulic acid and vanillic acid in V. planifolia exhibited the conversion of ferulic acid to vanillin via CoA-dependent β-oxidative cleavage-mediated pathway, where ferulic acid formes feruloyl-CoA, which is further oxidized to vanilloyl-CoA and finally reduction of vanilloyl-CoA leads to vanillin formation (Zenk 1965) (Fig. 2). In this report, involved enzymes were not identified.
Vanillin biosynthetic web in plants (grey arrows are showing vanillin biosynthetic scheme in Pseudomonassp HR199 strain). Key to enzymes: PAL phenylalanine ammonia lyase; C4H cinnamic acid-4-hydroxylase; 4CL 4-hydroxycinnamoyl-CoA ligase; HCT hydroxycinnamoyl transferase; C3′H coumaroyl ester 3′-hydroxylase; COMT caffeic acid/5-hydroxyferulic acid O-methyltransferase; HBS hydroxybenzaldehyde synthase; OMT O-methyltransferase
Apart from the proposed ferulic acid mediated pathway (Gallage et al. 2014; Zenk 1965), a more complex pathway was delineated on the basis of the results of feeding radio-labelled compounds to V. planifolia tissue cultures (Funk and Brodelius 1990a, b). This report suggested that caffeic acid (3,4-dihydroxycinnamic acid) produced via phenylpropanoid pathway is methylated at 4′ position to form isoferulic acid (3-hydroxy-4-methoxycinnamic acid), which is further methylated at 3′ position to produce 3,4-dimethoxycinnamic acid. Thereafter, this compound is consecutively demethylated, glucosilated, and subjected to a late-stage side-chain shortening (Fig. 2) to produce vanillic acid or its β-d-glucoside. This vanillic acid or its glucoside is further reduced to vanillin or its glucoside. Though the enzymes of this biosynthetic route were not explored, this pathway suggested another aspect of phenylpropanoid-mediated vanillin biosynthesis.
In another report, a non-β-oxidative route for vanillin biosynthesis was proposed, where 4-coumaric acid is converted to 4-hydroxybenzaldehyde by 4-hydroxybenzaldehyde synthase (4-HBS), a possible immediate precursor of vanillin or vanillin glucoside biosynthesis by a C2 side-chain shortening enzymatic reaction (Podstolski et al. 2002) (discussed in the third section) (Fig. 3e). After biosynthesis of 4-hydroxybenzaldehyde, hydroxylation occurs at 3′ position that results in 3,4-dihydroxybenzaldehyde or protocatechuic aldehyde formation. This 3′ hydroxyl group is then methylated by an O-methyltransferase to produce vanillin (Pak et al. 2004) (Fig. 2).
Roles of C2 side-chain shortening enzymes in vanillin biosynthesis
Non-β-oxidative pathway is an important step not only for vanillin biosynthesis but also for benzoates biosynthesis. In this pathway, side-chain shortening reactions occur to produce C6–C1 compounds from C6–C3 compounds (such as cinnamates and coumarates) (Fig. 1). The products of these kinds of reaction are usually benzoates or benzaldehydes. This step is also involved in vanillin biosynthetic pathway; several investigations were carried out to identify and characterize the enzymes involved in conversion of C6–C3 to C6–C1 compounds through ‘chain shortening’ mechanism, as it is one of the crucial steps of benzoic acid, hydroxybenzoic acid, and vanillin biosynthesis. However, discrepancy was observed in this ‘chain shortening’ step and in production of benzoates via benzaldehydes formation (Walton et al. 2003). In some cases, hydroxybenzoates were produced via hydroxybenzaldehyde formation. For example, a conversion of 4-coumaric acid to 4-hydroxybenzoic acid was observed in potato tubers, where an intermediate step of non-β-oxidative conversion of 4-coumaric acid to 4-hydroxybenzaldehyde was proposed (French et al. 1976). Similar event was found in carrot, where 4-hydroxybenzoic acid was produced via formation of 4-hydroxybenzaldehyde with a non-CoA-dependent non-β-oxidative reaction (Schnitzler et al. 1992). In an earlier report, 4-hydroxybenzoic acid formation was described as a non-β-oxidative, non-CoA-dependent reaction mechanism via 4-hydroxybenzaldehyde as an intermediate in cell cultures of Lithospermum erythrorhizon (Yazaki et al. 1991). Later on, in the same plant culture, it was described that a CoA-dependent β-oxidative pathway is the major biosynthetic route to 4-hydroxybenzoate, where 4-hydroxybenzaldehyde was not produced as intermediate (Fig. 3a). In that report, the rate of CoA-dependent β-oxidative reaction was described as faster than CoA-independent non-β-oxidative reaction. Formation of 4-hydroxybenzaldehyde as intermediate was described as a result of a possible retro-aldol reaction (Löscher and Heide 1994) (Fig. 3b). Thereafter, similar report was found for benzoic acid and salicylic acid biosynthesis in Cucumis sativus and Nicotiana attenuate, where benzaldehyde is not an intermediate (Jarvis et al. 2000). On the contrary, benzaldehyde was found to be an intermediate in benzoic acid formation in cell cultures of Hypericum androsaemum L. (Ahmed et al. 2002).
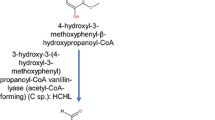
In case of vanillin biosynthesis, similar divergence was observed, where both the CoA-dependent β-oxidative and non-CoA-dependent non-β-oxidative reactions were found (Zenk 1965; Podstolski et al. 2002). After finding of vanillic acid and vanillin biosynthesis from ferulic acid through feruloyl-CoA and vanilloyl-CoA formation by a CoA-dependent β-oxidative cleavage, another model of hydroxycinnamate chain shortening was discovered. An enzyme, 4-hydroxycinnamoyl-CoA hydratase/lyase (HCHL), catalyzed this reaction (Fig. 3c; Table 1). This enzyme and its gene were isolated from Pseudomonas fluorescens strain AN103, grown in a medium containing ferulic acid as a sole carbon source (Narbad and Gasson 1998; Gasson et al. 1998; Mitra et al. 1999). This enzyme was found to be a crotonase homologue, which hydrated 4-hydroxycinnamoyl-CoA thioesters and then catalyzed a retro-aldol cleavage to produce the corresponding 4-hydroxybenzaldehyde. Therefore, this observation supported the previous finding in cell cultures of Lithospermum erythrorhizon (Löscher and Heide 1994). In addition, HCHL was also found to catalyze the hydration of caffeoyl-CoA and feruloyl-CoA to produce their aldehyde, protocatechuic aldehyde, and vanillin, respectively, along with the other cleavage product such as acetyl-CoA. The catalytic centre activity of HCHL was found to be low (2.3 molecules s−1 at 30 °C) with 4-coumaroyl-CoA as substrate. The interesting point in this enzymatic reaction was that it was found to be a non-β-oxidative yet CoA-dependent reaction, which was not found in wild-type plants until Ahmed et al. (2002) observed this kind of activity in Hypericum androsaemum. Thereafter, Gallage et al. (2014) identified the gene VpVAN that encoded vanillin synthase in V. planifolia pod (discussed in the following section). The similar enzymatic activity and gene were also discovered in other Pseudomonas strains (Priefert et al. 1997; Venturi et al. 1998). Its operon was characterized in strain HR199, a strain grown on eugenol (Overhage et al. 1999). This strain converted eugenol to vanillin via ferulic acid formation (Fig. 2). A functional homologue of HCHL was expressed in Amycolatopsis strain HR166, a gram-positive bacterium that synthesised high yields of vanillin from ferulic acid (Achterholt et al. 2000). Later on, similar hydratase/lyase activity was observed in cell cultures of Hypericum androsaemum, which produced benzaldehyde and acetyl-CoA from cinnamoyl-CoA (Ahmed et al. 2002) (Fig. 3b). Another enzyme benzaldehyde dehydrogenase finally converted benzaldehyde to benzoic acid.
The concept of non-CoA-dependent non-β-oxidative formation of vanillin was corroborated when a hydroxycinnamate chain shortening enzyme was partially purified and characterized from V. planifolia cell cultures; the enzyme was designated as hydroxybenzaldehyde synthase (HBS) (Podstolski et al. 2002) (Table 1). This enzyme activity supported the previous radio-labelled precursor feeding experiments that showed benzoic or hydroxybenzoic acid formation via benzaldehyde or hydroxybenzaldehyde by single-step side-chain shortening of 4-coumaric acid (Fig. 3e). HBS showed high specificity to 4-coumaric acid (no activity was found with o-coumaric acid) as substrate and only 2% activity (of that observed with 4-coumaric acid) was found with ferulic acid, which produced 3-methoxy-4-hydroxybenzaldehyde. Roots, leaves, pods, and embryo cultures of V. planifolia also exhibited HBS activity. This enzyme activity exhibited CoA dependency but that was non-specific, and CoA played a role as a ‘thiol reagent’, which was found to be replaced more efficiently by dithiothreitol or dithioerythritol. After partial purification of the enzyme, the main fraction containing HBS activity was shown to have a protein of Mr 28 kDa, along with other proteins of Mr 31–45 kDa. Kinetic analysis of this partially purified fraction did not show saturation with the substrate (4-coumaric acid), even when used as high as 100 mM.
Similar side-chain shortening enzymatic activity was found in the cell-free extract of elicited hairy root cultures of Daucus carota (Sircar and Mitra 2008) and fragrant roots of Hemidesmus indicus (Kundu et al. 2012). In both the cases, the enzyme activity resulted in formation of 4-hydroxybenzaldehyde from 4-coumaric acid. In root cultures of D. Carota, 4-hydroxybenzaldehyde was then converted to 4-hydroxybenzoic acid by an enzyme, 4-hydroxybenzaldehyde dehydrogenase, while in H. indicus roots, 4-hydroxybenzaldehyde was hypothesized as an intermediate of biosynthetic pathway of 2-hydroxy-4-methoxybenzaldehyde, a homologous fragrant molecule of vanillin.
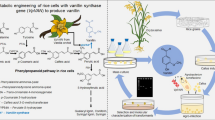
Recently, another side-chain shortening enzyme activity was reported in V. planifolia pod that converted ferulic acid and its glucoside to vanillin and its glucoside through two stepped reaction (Gallage et al. 2014) (Fig. 3d). The enzyme was designated as vanillin synthase (Table 1). The gene of that enzyme was cloned and its heterologous expression was carried out in Nicotiana benthamiana, Hordeum vulgare, and Saccharomyces cerevisiae. This report demonstrated de novo biosynthesis of vanillin by administration of radio-labelled precursor of ferulic acid, such as phenylalanine and cinnamic acid in the slices of developing V. planifolia pod, which resulted in the formation of radio-labelled vanillin glucoside only in the inner part of the pod. The in vivo localization of vanillin biosynthesis was confirmed by in situ studies of co-occurrence of vanillin synthase transcripts and proteins in the inner part of the pod. When radio-labelled 4-hydroxybenzaldehyde was fed to the slices, surprisingly, no vanillin glucoside was found to be produced, which contradicted the previous report, where hydroxybenzaldehyde was demonstrated as a major intermediate of vanillin biosynthesis, produced by HBS enzyme activity through C2 side-chain shortening mechanism (Podstolski et al. 2002). Therefore, the question regarding the true intermediate of vanillin biosynthesis remained unanswered, as both ferulic acid and 4-coumaric acid were found to be used for enzymatic side-chain shortening towards vanillin.
4-Coumaric acid or ferulic acid: selection as substrate by the same enzyme
Several putative radio-labelled compounds were administrated for feeding in V. planifolia to identify the nearest precursor of vanillin or its glucoside. In a report, ferulic acid was found to be the nearest precursor as maximum (29%) radio-activities were incorporated into vanillin glucoside after feeding of 14C-labelled ferulic acid, while only 15% incorporation was observed on 14C-labelled 4-coumaric acid feeding (Negishi et al. 2009). Therefore, it was necessary to identify the enzymes those were involved in incorporations of radio-labelled precursors. Identification of vanillin synthase gene (VpVAN) revealed that it was the same gene that was proposed to encode the HBS enzyme in V. planifolia cell culture, which executed a C2 chain shortening of 4-coumaric acid to produce 4-hydroxybenzaldehyde (Podstolski et al. 2002). In subsequent experiments, the enzyme was transiently expressed in yeast, tobacco, and barley, and was verified through coupled in vitro transcription/translation assay using a range of plausible substrates including 4-coumaric acid. All these assays were done in planta, and none of the assays exhibited conversion of 4-coumaric acid to 4-hydroxybenzaldehyde or its glucoside (Gallage et al. 2014). From the LC–MS monitoring of the assays, it was suggested that the presence of one free or glycosilated ‘-hydroxyl’ group at 4′ position and one ‘-methoxy’ group at 3′ position was required for vanillin synthase activity. Therefore, ferulic acid fitted better as substrate for the VpVAN encoded enzyme. Nevertheless, side-chain shortening vanillin synthase assay with cell-free extract from V. planifolia pod was not succeeded due the presence of high amount of endogenously produced 4-hydroxybenzaldehyde and vanillin glucoside; the presence of these compounds interrupted the detection of diminutive amount of product. Therefore, the investigators concluded that, in the work of Podstolski et al. (2002), a different isoform of this enzyme was partially purified which showed activity with 4-coumaric acid. The other possible interpretations were the presence of impurities in the reaction or simultaneous background reaction or endogenously bound 4-hydroxybenzaldehyde, which resulted in detection of 4-hydroxybenzaldehyde as product of HBS activity.
The reaction mechanism of vanillin synthase enzyme was to be similar to the HCHL enzyme from P. fluorescens, which has been discussed in the third section. The reaction was demonstrated to be occurred in two steps including an initial hydration addition reaction followed by a retro-aldol elimination reaction (Gallage et al. 2014) (Fig. 2d). A similar retro-aldol elimination reaction was previously proposed by some investigators (Löscher and Heide 1994) to demonstrate benzoic or 4-hydroxybenzoic acid biosynthesis (discussed in the third section), but in those reports, 4-coumaric acid was substrate for the enzymatic conversion instead of ferulic acid, and benzaldehyde or 4-hydroxybenzaldehyde was produced via 4-coumaroyl-CoA by retro-aldol elimination and releasing of acetyl-CoA. Furthermore, insignificant sequence similarity was found on sequence alignment of VpVAN with HCHL sequence from P. fluorescens. Only 11 similarly positioned amino-acid residues were observed among 55 conserved residues scattered over the protein sequence belonging to crotonase superfamily from selected bacteria (Achterholt et al. 2000). On the other hand, this enzyme showed a higher amino acid similarity with the enzymes of cysteine protease family. There are more than 150 cysteine proteases in the plants. Two putative protease cleavage sites in VpVAN were identified and it also contained a pro-peptide sequence, which may help in intracellular targeting, proper folding of the mature enzyme and keeping the enzyme in inactive form to balance its function according to the physiological demand of the cell. It was also demonstrated that replacement of this pro-peptide sequence with ER-targeting putative signal peptide and putative pro-peptide protease cleavage site from tobacco cysteine protease, enhanced the enzyme activity when expressed in tobacco cell (Gallage et al. 2014). The most plausible reason is the proper compartmentalization of the protein due to this peptide replacement, which helped the enzyme to be folded more accurately with proper post-translational modification that resulted in increased enzyme activity (Fig. 4). Conversely, according to the patent application publication by Havkin-Frenkel et al. (2003), sequencing of HBS exhibited high sequence identity and many of the characteristic of cathepsin-like cysteine proteases, which suggested the possibility that HBS was evolved from a cysteine protease. Glechoma hederacea, another vanillin producing plant, was reported to contain an enzyme showing 71% sequence similarity with the VpVAN and demonstrated to be a vanillin synthase by transient expressions in tobacco (Gallage et al. 2014).
The identification and characterization of HBS, HCHL, and VpVAN and their sequence similarity with cysteine protease family suggested that they have evolved from the same evolutionary origin and are either encoded by the same gene or three different isoforms. They are shown to use similar chain shortening mechanism on either 4-coumaric acid or ferulic acid, and have different products that still lead to vanillin biosynthesis.
O-methyltransferases in vanillin biosynthesis
A number of O-methyltransferases are present in plants, which are involved in various secondary metabolites biosynthesis using S-adenosyl methionine as methyl group donor. Specially, in lignin monomer biosynthesis, O-methyltransferases play a key role. It has been accepted that methylation for lignin monomer biosynthesis starts at caffeic acid level by caffeic acid O-methyltransferase (COMT), which was characterized from several species (Ibrahim et al. 1998). This COMT was found to have multi-substrate specificity. Among the putative substrates, caffeic acid showed least specificity while 5-hydroxyconiferaldehyde, the key substrate for S-lignin production, was found to be primary substrate (Osakabe et al. 1999; Li et al. 2000; Parvathi et al. 2001). Formation of ferulic acid from caffeic acid is also conducted by caffeic acid/5-hydroxyferulic acid O-methyltransferase (Lam et al. 2007). Therefore, in vanillin biosynthesis via ferulic acid, COMT plays a vital role. According to Gallage et al. (2014), VpVAN used ferulic acid for two stepped conversion to vanillin; this ferulic acid biosynthesis followed the general COMT-mediated pathway which started from caffeoyl-CoA. In another report, an REF-1 gene was identified in aspen, which encoded an enzyme that converted coniferaldehyde to ferulic acid (Nair et al. 2004). However, this enzyme has not been reported in V. planifolia, but there is scope of investigation to find out the alternative route to ferulic acid production which may leads to vanillin biosynthesis.
In non-CoA-dependent non-β-oxidative pathway-mediated vanillin biosynthesis, the penultimate step to vanillin is mediated by a multifunction O-methyltransferase. This enzyme was identified and characterized from V. planifolia cell cultures having 365 amino-acid residues and a predicted molecular weight of 40.66 kDa (Pak et al. 2004). The enzyme was demonstrated to have a broad range of substrate specificity, which also converted 3,4-dihydroxybenzaldehyde to vanillin. However, the tissue distribution and substrate preferences of this enzyme suggested that it might have primary function in lignin biosynthesis. Phylogenetic analysis of this O-methyltransferase through comparing 18 similar methyltransferase sequences demonstrated the relationship of this O-methyltransferase sequence to methyltransferases reported from other species.
Both these O-methyltransferases are shown to have primary function in lignin biosynthesis; therefore, there is a possibility of the presence of O-methyltransferases particularly devoted to vanillin biosynthesis. This finding is necessary to make the choice in selection of pathway in between ferulic acid and hydroxybenzaldehyde-mediated vanillin biosynthesis in V. planifolia.
Conclusion
Investigation in vanillin biosynthetic pathway is necessary to improve the commercial production and to fulfill the worldwide demand of vanillin as flavouring and pharmaceutical molecule. Though the enzymatic pathway leading to vanillin biosynthesis has been explored to a comprehensible point, the entire biochemical and molecular characterization of the all involved enzymes and genes in V. planifolia or other vanillin producing plants will provide a clearer picture. The relation between vanillin and benzoic acid biosynthetic pathway revealed the similarity in the C2 side-chain shortening mechanism, which is the key step of the whole pathway. Though some divergences in reported enzymes are there, but a clear aspect of the key biosynthetic route of vanillin now can be comprehended. From the reports, which revealed the sequential similarity among the side-chain shortening enzymes, mainly VpVAN and 4-HBS, it can be concluded that they have evolved from the same origin and are either encoded by the same gene or different isoforms. In addition, it is also necessary to shed more light on the characteristic distinctions among CoA-dependent β-oxidative pathway, non-CoA-dependent non-β-oxidative pathway, and CoA-dependent yet non-β-oxidative pathway enzymes. However, the current theory of non-CoA-dependent non-β-oxidation of ferulic acid for vanillin synthesis through VpVAN enzyme with inevitable explanations is decidedly accepted. This report exhibited encouraging progress in elucidating the enzymes for vanillin biosynthesis that will offer new opportunities in metabolic engineering-based production in vanilla pod industries.
Author contribution statement
AKundu planned and wrote the review manuscript, designed the figures and table, and corresponded for publication.
Abbreviations
- HBS:
-
Hydroxybenzaldehyde synthase
- HCHL:
-
4-Hydroxycinnamoyl-CoA hydratase/lyase
- COMT:
-
Caffeic acid O-methyltransferase
References
Achterholt S, Priefert H, Steinbüchel A (2000) Identification of Amycolatopsis sp. strain HR167 genes, involved in the bioconversionof ferulic acid to vanillin. Appl Microbiol Biotechnol 54:799–807
Ahmed MA, El-Mawla A, Beerhues L (2002) Benzoic acid biosynthesisin cell cultures of Hypericum androsaemum. Planta 214:727–733
Anwar MH (1963) Paper chromatography of monohydroxyphenols in vanilla extract. Anal Chem 35:1974–1976
Dignum MJW, Kerler J, Verpoorte R (2001) Vanilla production: technological, chemical, and biosynthetic aspects. Food Rev Int 17:199–219
French CJ, Vance CP, Towers GHN (1976) Conversion of p-coumaric acid to p-hydroxybenzoic acid by cell free extracts of potato tubers and Polyporushispidus. Phytochemistry 15:564–566
Fritz RR, Hodgins DS, Abell CW (1976) Phenylalanine ammonia-lyase. Induction and purification from yeast and clearance in mammals. J Biol Chem 251:4646–4650
Funk C, Brodelius PE (1990a) Phenylpropanoid metabolism in suspension cultures of Vanilla planifolia Andr. II Effects of precursor feeding and metabolic inhibitors. Plant Physiol 94:95–101
Funk C, Brodelius PE (1990b) Phenylpropanoid metabolism in suspension cultures of Vanilla planifoliaAndr. III Conversion of 4-methoxycinnamic acids into 4-hydroxybenzoic acids. Plant Physiol 94:102–108
Gallage NJ, Møller BL (2015) Vanillin–bioconversion and bioengineering of the most popular plant flavor and its de novo biosynthesis in the vanilla orchid. Mol Plant 8:40–57
Gallage NJ, Hansen EH, Kannangara R, Olsen CE, Motawia MS, Jørgensen K, Holme I, Hebelstrup K, Grisoni M, Møller BL (2014) Vanillin formation from ferulic acid in Vanilla planifolia is catalysed by a single enzyme. Nat Commun 5:4037. doi:10.1038/ncomms5037
Gasson MJ, Kitamura Y, McLauchlan WR, Narbad A, Parr AJ, Parsons ELH, Payne J, Rhodes MJC, Walton NJ (1998) Metabolism of ferulic acid to vanillin: a bacterial gene of the enoyl-SCoA hydratase/isomerase superfamily encodes an enzyme for the hydration and cleavage of a hydroxycinnamic acid SCoA thioester. J Biol Chem 273:4163–4170
Havkin-Frenkel D, Podstolski A, Dixon R (2003) Vanillin biosynthetic pathway enzyme from Vanilla planifolia. United States patent application publication. https://docs.google.com/viewer?url=patentimages.storage.googleapis.com/pdfs/US20030070188.pdf
Ibrahim RK, Bruneau A, Bantignies B (1998) Plant O-methyltransferases: molecular analysis, common signature and classification. Plant Mol Biol 36:1–10
Jarvis AP, Schaaf O, Oldham NJ (2000) 3-Hydroxy-3-phenylpropanoic acid is an intermediate in the biosynthesis of benzoic acidand salicylic acid but benzaldehyde is not. Planta 212:119–126
Kanisawa T, Tokoro K, Kawahara S (1994) In: Kurihara K, Suzuki N, Ogawa H (eds) Olfaction taste XI (Proceeding of the International Symposium). Springer, Tokyo, p 268
Kaur B, Chakraborty D (2013) Biotechnological and molecular approaches for vanillin production: a review. Appl Biochem Biotechnol 169(4):1353–1372
Kundu A, Jawali N, Mitra A (2012) Shikimate pathway modulates the elicitor-stimulated accumulation of fragrant 2-hydroxy-4-methoxybenzaldehyde in Hemidesmusindicusroots. Plant PhysiolBiochem 56:104–108
Lam KC, Ibrahim RK, Behdad B, Dayanandan S (2007) Structure, function, and evolution of plant O-methyltransferases. Genome 50:1001–1013
Li L, Popko JL, Umezawa T, Chiang VL (2000) 5-Hydroxyconiferylaldehyde modulates enzymatic methylation for syringyl monolignol formation, a new view of monolignol biosynthesis in angiosperms. J Biol Chem 275:6537–6545
Löscher R, Heide L (1994) Biosynthesis of p-hydroxybenzoate fromp-coumarate and p-coumaroyl-coenzyme A in cell-free extracts of Lithospermum erythrorhizon cell cultures. Plant Physiol 106:271–279
Makkar HPS, Beeker K (1994) Isolation of tannins from leaves of some trees and shrubs and their properties. J Agric Food Chem 42:731–734
Mitra A, Kitamura Y, Gasson MJ, Narbad A, Parr AJ, Payne J, Rhodes MJC, Sewter C, Walton NJ (1999) 4-Hydroxycinnamoyl-CoA hydratase/lyase (HCHL)-an enzyme of phenylpropanoid chain cleavage from Pseudomonas. Arch Biochem Biophys 365:10–16
Nair RB, Bastress KL, Ruegger MO, Denault JW, Chapple C (2004) The Arabidopsis thaliana REDUCED EPIDERMAL FLUORESCENCE1 gene encodes an aldehyde dehydrogenase involved in ferulic acid and sinapic acid biosynthesis. Plant Cell 16:544–554
Narbad A, Gasson MJ (1998) Metabolism of ferulic acid via vanillinusing a novel CoA-dependent pathway in a newly-isolated strain of Pseudomonas fluorescens. Microbiology 144:1397–1404
Negishi O, Sugiura K, Negishi Y (2009) Biosynthesis of vanillin via ferulic acid in Vanilla planifolia. J Agric Food Chem 57:9956–9961
Ni J, Tao F, Du H, Xu P (2015) Mimicking a natural pathwayfor de novo biosynthesis: natural vanillin production from accessible carbon sources. Sci Rep. doi:10.1038/srep13670
Osakabe K, Tsao CC, Li L, Popko JL, Umezawa T, Carraway DT (1999) Coniferyl aldehyde 5-hydroxylation and methylation direct syringyl lignin biosynthesis in angiosperms. Proc Natl Acad Sci USA 96:8955–8960
Overhage J, Priefert H, Steinbüchel A (1999) Biochemical andgenetic analyses of ferulic acid catabolism in Pseudomonas sp. Strain HR199. Appl Environ Microbiol 65:4837–4847
Pak FE, Gropper S, Dai WD, Havkin-Frenkel D, Belanger FC (2004) Characterization of a multifunctional methyltransferases from the orchid Vanilla planifolia. Plant Cell Rep 22:959–966
Parvathi K, Chen F, Guo D, Blount JW, Dixon RA (2001) Substrate preferences of O-methyltransferases in alfalfa suggest new pathways for 3-O-methylation of monolignols. Plant J 25:193–202
Podstolski A, Havkin-Frenkel D, Malinowski J, Blount JW, Kourteva G, Dixon RA (2002) Unusual 4-hydroxybenzaldehyde synthase activity from tissue cultures of the vanilla orchid Vanilla planifolia. Phytochemistry 61:611–620
Priefert H, Rabenhorst J, Steinbüchel A (1997) Molecular characterization of genes of Pseudomonas sp. strain HR199 involved inbioconversion of vanillin to protocatechuate. J Bacteriol 179:2595–2607
Ro DK, Mah N, Ellis BE, Douglas CJ (2001) Functional characterization and subcellular localization of poplar (Populustrichocarpa × Populusdeltoides) cinnamate 4-hydroxylase. Plant Physiol 126:317–329
Schnitzler J-P, Madlung J, Rose A, Seitz HU (1992) Biosynthesis of p-hydroxybenzoic acid in elicitor-treated carrot cell cultures. Planta 188:594–600
Schoch G, Goepfert S, Morant M, Hehn A, Meyer D, Ullmann P, Werck-Reichhart D (2001) CYP98A3 from Arabidopsis thaliana is a 30-hydroxylase of phenolic esters, a missing link in the phenylpropanoid pathway. J Biol Chem 276:36566–36574
Sinha AK, Sharma UK, Sharma N (2008) A comprehensive review on vanilla flavor: extraction, isolation and quantification of vanillin and others constituents. Int J Food Sci Nutr 59(4):299–326
Sircar D, Mitra A (2008) Evidence for p-hydroxybenzoate formation involving phenylpropanoid chain-cleavage in hairy roots of Daucus carota. J Plant Physiol 165:407–414
Tokoro K, Kawahara S, Amano A, Kanisawa T, Indo M (1990). In: Bessiere Y, Thomas AF (eds) Flavour science and technology, vol 73. Wiley, Chichester, p 73
Venturi V, Zennaro F, Degrassi G, Okeke BC, Bruschi CV (1998) Genetics of ferulic acid bioconversion to protocatechuic acidin plant-growth-promoting Pseudomonas putida WCS358. Microbiology 144:965–973
Walton NJ, Mayer MJ, Narbad A (2003) Vanillin. Phytochemistry 63:505–515
Yazaki K, Heide L, Tabata M (1991) Formation of p-hydroxybenzoic acid from p-coumaric acid by cell free extract of Lithospermum erythrorhizon cell cultures. Phytochemistry 30:2233–2236
Zamzuri NA, Abd-Aziz S (2012) Biovanillin from agro wastes as an alternative food flavour. J Sci Food Agric 93:429–438
Zenk MH (1965) Biosynthese von vanillin in Vanilla planifolia. Andr Z Pflanzenphysiol 53:404
Acknowledgements
The author acknowledges the Indian Institute of Technology Kharagpur for permitting to access the digital library and for providing ‘Institutional Assistantship’ during preparing the manuscript. Author also acknowledges Ms. Shruti Mishra, M.Sc. for helping in linguistic editing of the article.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kundu, A. Vanillin biosynthetic pathways in plants. Planta 245, 1069–1078 (2017). https://doi.org/10.1007/s00425-017-2684-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2684-x