Abstract
Main conclusion
NO acts as the essential signal to enhance poplar tolerance to chilling stress via antioxidant enzyme activities and protein S -nitrosylation modification, NO signal is also strictly controlled by S -nitrosoglutathione reductase and nitrate reductase to avoid the over-accumulation of reactive nitrogen species.
Poplar (Populus trichocarpa) are fast growing woody plants with both ecological and economic value; however, the mechanisms by which poplar adapts to environmental stress are poorly understood. In this study, we used isobaric tags for relative and absolute quantification proteomic approach to characterize the response of poplar exposed to cold stress. We identified 114 proteins that were differentially expressed in plants exposed to cold stress. In particular, some of the proteins are involved in reactive oxygen species (ROS) and reactive nitrogen species (RNS) metabolism. Further physiological analysis showed that nitric oxide (NO) signaling activated a series of downstream defense responses. We further demonstrated that NO activated antioxidant enzyme activities and S-nitrosoglutathione reductase (GSNOR) activities, which would reduce ROS and RNS toxicity and thereby enhance poplar tolerance to cold stress. Suppressing NO accumulation or GSNOR activity aggravated cold damage to poplar leaves. Moreover, our results showed that RNS can suppress the activities of GSNOR and NO nitrate reductase (NR) by S-nitrosylation to fine-tune the NO signal and modulate ROS levels by modulating the S-nitrosylation of ascorbate peroxidase protein. Hence, our data demonstrate that NO signaling activates multiple pathways that enhance poplar tolerances to cold stress, and that NO signaling is strictly controlled through protein post-translational modification by S-nitrosylation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Low-temperature stress, or chilling stress, is one of the major limiting factors of woody plant growth, development, and geographical distribution. Chilling stress results in poor seed germination, stunted seedling growth, yellowing and withering of leaves, delayed crop heading, and increased pollen sterility (Xiong and Zhu 2001; Beck et al. 2004; Minami et al. 2005). Plants have developed multiple strategies to withstand chilling damage. Upon exposure to cold stress, a set of early signals is immediately triggered, including the calcium and reactive oxygen species burst, which activates the MAPK signal cascade, and ultimately initiates the downstream cold-responsive transcriptional cascade, which involves COR (cold-regulated), KIN (cold-induced), LTI (low temperature-induced), and RD (responsive to dehydration) genes. C-repeat (CRT)-binding factors (CBFs), also called dehydration-responsive element-binding proteins (DERBs), can bind directly to the promoter regions of these cold-responsive gene to activate their expression (Zhu et al. 2007). It was recently demonstrated that post-transcriptional regulation or epigenetic modification of vital genes also play a role in cold stress response, demonstrating the intricate nature of plant acclimation to cold stress (Zhu et al. 2007; Chinnusamy et al. 2008; Luo et al. 2012). Woody plants play an essential role in moderating the global carbon cycle and climate (Jackson et al. 2002; Reichstein et al. 2014). However, most studies examining acclimation to cold stress focus on model plants or crops, such as Arabidopsis thaliana (Arabidopsis), Oryza sativa (rice), and Hordeum vulgare (barley). The mechanism by which woody plants respond to chilling stress is poorly understood.
Reactive oxygen species (ROS), particularly H2O2, play an essential role in the plant’s response to multiple physiological processes, including seed germination, growth, senescence, differentiation, and stress responses (Neill et al. 2002a; Desikan et al. 2004). An optimal concentration of ROS is critical for normal plant growth and development, and over-production of ROS may led to oxidative stress and cell death. To avoid the oxidative damage imposed by ROS, plants employ a series of enzymatic and nonezymatic oxidants that detoxify ROS accumulated in cells. These antioxidants include glutathione (GSH), ascorbate (AsA), ascorbate peroxidase (APX), dehydroascorbate reductase (DHAR), and monodehydroascorbate reductase (MDHAR) (Neill et al. 2002a; Foyer and Noctor 2011). Like H2O2, nitric oxide is a gaseous radical molecule that participates in a wide spectrum of physiological processes. Over-accumulated NO can interact with ROS to form reactive nitrogen species (RNS), such as dinitrogen tridoxide (N2O3), nitrogen dioxide (·NO2), peroxynitrite (ONOO−), S-nitrosothiols (SNOs), and S-nitrosoglutathione (GSNO) (Wilson et al. 2008). Among these RNS, the SNOs, GSNO, and peroxynitrite are the most studied. RNS interacts with protein or non-protein sulfhydryl-containing compounds to form SNOs, which also carry out important biological reactions, including NO release, transnitrosation, and redox-based posttranslational modification. For example, the total SNO levels in plants were found to be upregulated by wounding and heavy metal stress (Barroso et al. 2006; Chaki et al. 2011). The S-nitrosylation of NO− with GSH to form GSNO has an important physiological function in plants, as GSNO serves as a mobile reservoir of NO bioactivity. GSNO reductase (GSNOR) modulates GSNO levels in plants by catalyzing the NADH-dependent reduction of GSNO to glutathione disulfite (GSSG) and NH3.(Sakamoto et al. 2002). The activity and expression of GSNOR can be induced by various stimuli, including wounding, low temperature, high temperature, heavy metals, and pathogen infection (Barroso et al. 2006; Corpas et al. 2008; Chaki et al. 2009, 2011), but the roles of RNS and GSNOR in woody plants subjected to chilling stress are unclear.
Quantitative proteomics research identifies and quantifies changes in protein abundance and function with the aim of deciphering deeper biological functions (Jorrin-Novo 2009; Alexandersson et al. 2013). Widely used quantitative proteomics techniques include classical gel-based methods (2DE, DIGE) and liquid chromatography–mass spectrometry-based methods such as isotopic labeling and the relatively new label-free methods. Among these methods, isobaric tags for relative and absolute quantification (iTRAQ) can be used to analyze up to eight different samples within the same experiment (Bindschedler and Cramer 2011; Alexandersson et al. 2013). As fast growing trees, poplars play an important role in forestry and as energy crops. The availability of the full genome sequences of P. trichocarpa makes poplar an ideal model organism for woody plant functional genomics studies (Tuskan et al. 2006). Here, to evaluate the potential role of reactive nitrogen species in poplar acclimation to cold stress, we used physiological and quantitative proteomics to investigate the mechanism underlying poplar acclimation to chilling stress. Our proteomic results showed that a series of proteins associated with ROS and RNS metabolism were markedly differentially expressed during chilling stress in poplar. During the early stages of exposure to chilling stress, NO levels increased quickly after chilling stress; however, over-accumulation of NO interacts with ROS to form RNS, which reduces poplar’s viability after long-term chilling stress and is accompanied by an increase in GSNOR enzyme accumulation and activity. Suppressing GSNOR enzyme activity markedly increased RNS accumulation and aggravated the damage of chilling on leaf viability. Based on our physiological and proteomics results, we propose that feedback regulation of RNS levels by GSNOR plays an essential role during poplar’s response to chilling stress. Our results suggest a strategy to genetically engineer poplar plants with increased tolerance to chilling stress by directly manipulating the levels of small metabolite compounds such as RNS.
Materials and methods
Material and chilling treatment
Stem cuttings (20 cm) taken from poplar (Populus yunnanensis Dode) were rooted and grown in pots filled with 3 L of one-third strength Hoagland’s nutrient solution, pH 6.5, as reported (Gaudet et al. 2011). Cuttings were grown in a controlled greenhouse (50–70 % relative humidity, 25 °C in day time and 18 °C in night time) with a 12-h photoperiod (800 µmol photons m−2 s−1 light intensity). The nutrient solutions were completely replaced every week to prevent depletion of metals, nutrients, and oxygen. Plants were allowed to develop roots and grown for 3 weeks in hydroponic solution. For chilling treatment, 100 healthy plantlets with an average of five nodes were randomly selected and moved to a cold chamber stably maintained at 4 °C with the same humidity and light intensity as above. Three to five replicates were performed per experiment and each replicate consisted of six cutting plantlets. Samples without chilling treatment were used as the controls.
Chemical treatment
The GSNOR inhibitor N6022 purchased from Merck Company (Darmstadt, Germany), the GSNOR inhibitor DA (dodecanoic acid) and cPTIO [2-(4-carboxyphenyl)-4, 4, 5, 5–tetramethylimidazoline-1-oxyl-3-oxide], thioredoxin, and ascorbate were purchased from Sigma Chemical company (St. Louis, MO, USA). For chemical inhibitor treatment, the stock chemical was prepared as 1000-fold, and these chemicals was added to the hydroponic Hoagland’s nutrient solution to the indicated concentration.
Analysis of chlorophyll fluorescence
Chlorophyll fluorescence was analyzed using a pulse amplitude-modulated chlorophyll fluorometer (Heinz Walz, Effeltrich, Germany) as described (Bai et al. 2011b). Briefly, the poplar cuttings were dark adapted for 30 min after treatment and the maximum quantum yield of PSII (Fv/Fm) was determined for each sample by analyzing a whole leaf. Maximal fluorescence (Fm) was recorded using a 0.8-s pulsed light at 4000 µmol s−1 m−2, and the minimal fluorescence (F o) was recorded during the weak measuring pulses.
Protein extraction for iTRAQ analysis
Protein extraction for iTRAQ analysis was performed as described (Kong et al. 2014). The leaves of plantlets exposed to different chilling treatment for different periods were collected and frozen in liquid nitrogen, and proteins were extracted using a previously described phenol extraction procedure (Kong et al. 2014). Briefly, ∼1 g of frozen seedlings was ground with a mortar in liquid nitrogen and 5 mL of extraction buffer containing 100 mM Tris–HCl, pH 7.8, 100 mM KCl, 50 mM l-ascorbic acid, 1 % (v/v) Triton X-100, 1 % (v/v) β-mercaptoethanol, and 1 mM phenylmethylsulfonyl fluoride (PMSF) was added. After centrifugation (12,000g; 15 min; 4 °C), the resulting suspension solution was transferred to a 20-mL tube and Tris–phenol (2 vol; pH 8.0, Amresco, Solon, OH, USA) was added. The mixture was thoroughly vortexed before centrifuging at 12,000g for 15 min at 4 °C. The upper phenolic phases were transferred to a 50-mL tube, and then 5 volumes of 100 mM ammonium acetate/methanol were added. After careful mixing, the mixture was stored at −20 °C overnight. The supernatant was carefully removed after centrifugation at 13,000g for 15 min at 4 °C, and the protein pellets were used immediately or suspended in 25 mL of ice-cooled methanol for 2 h at −20 °C for further analysis.
Protein digestion, iTRAQ labeling, and protein quantification
The protein pellet was dissolved in a solution of 1 % SDS and 100 mM triethylammonium bicarbonate, pH 8.5, and then subjected to reduction, alkylation, trypsin digestion, and labeling using 8-plex iTRAQ reagent kits, according to the manufacturer’s instructions (AB Sciex, Framingham, MA, USA). After labeling, the samples were combined and lyophilized, and the peptide mixture was dissolved in strong cation exchange (SCX) solvent A (25 % (v/v) acetonitrile, 10 mM ammonium formate, pH 2.8). The peptides were fractionated on an Agilent HPLC system 1100 with a polysulfethyl A column (2.1 × 100 mm, 5 μm, 300 A, PolyLC, Columbia, MD, USA). Peptides were eluted at a flow rate of 200 μL/min with a linear gradient of 0–20 % solvent B [25 % (v/v) acetonitrile, 500 mM ammonium formate] over 50 min, followed by ramping up to 100 % solvent B in 5 min and holding for 10 min. The absorbance at 214 nm was monitored, and a total of 12 fractions were collected. Each SCX fraction was lyophilized and dissolved in solvent A [3 % acetonitrile (v/v), 0.1 % formic acid (v/v)] and then analyzed with a Q-Exactive Hybrid Quadrupole-Orbitrap mass spectrometer (Thermo Finnigan Scientific, San Jose, CA, USA). Samples were separated on a Hypersil Gold C18 column (100 × 2.1 mm, 1.9 μm) (Thermo Fisher Scientific, Pittsburgh, PA, USA). Peptides were eluted with a linear gradient of acetonitrile/0.1 % formic acid from 3 to 50 % in 90 min at a flow rate of 250 nL/min. Peptides were then sprayed into the chamber of the Q-Exactive MS/MS system with a spray voltage of 2.2 kV. Full-scan mass spectra were collected over 200–1800 m/z at high resolution (60,000). The four most intense precursor ions were selected for collision-induced fragmentation in a linear ion trap with 50–2000 m/z and 30–2000 ms at a resolution of 7500. Dynamic exclusion was employed within 40 s to prevent repetitive selection of peptides.
The raw LC–MS/MS files were analyzed using Proteome Discoverer 1.3 (Thermo Fisher Scientific). The software was connected to the Mascot Search Engine server, version 2.3 (Matrix Science, Boston, MA, USA). The spectra were searched against the poplar protein database available from NCBI (http://www.ncbi.nlm.nih.gov/protein), which included 40151 sequences (accessed in April 2012, txid3694). Search parameters included iTRAQ 8-plex quantification, carbamidomethylation of cysteine was set as a fixed modification, and oxidation of methionine was set as a variable modification. Trypsin was specified as the proteolytic enzyme, and one missed cleavage was allowed. Peptide mass tolerance was set at 10 ppm and fragment mass tolerance was set at 0.1 Da. An automatic decoy database search was performed as part of the search. False discovery rates (FDRs) for peptide identification of all searches were <1.0 %. The data were prefiltered to exclude MS/MS spectra containing fewer than three peaks or with a total ion count of below 50. Mascot results were filtered with the Mascot Percolator package to improve the accuracy and sensitivity of peptide identification. For differential analyses, all proteins identified and quantified with at least four independent peptides with a high degree of confidence (FDR of 1 %) were selected. The results of all of the measured iTRAQ values were normalized again reporting ratios using Proteome Discoverer1.3 software. Only significant ratios from the replicates were used to calculate the average ratio for the protein. Each P value was generated based on quantitative information derived from at least three independent peptides in each replicate. Cutoffs of 1.5- or 0.5-fold were set to indicate up-regulation or down-regulation of proteins, respectively, and a P value of <0.05 using t test was used to indicate significance.
Immunoblotting and protein S-nitrosylation analysis
Proteins from seedlings subjected to different treatments were extracted in a solution consisting of 50 mM Tris, pH 7.5, 150 mM NaCl, 1 mM EDTA, 0.1 % Triton X-100, 0.5 % β-mercaptoethanol, and 1 mM PMSF with vigorous vortexing at 4 °C. After centrifugation at 20,000g, 4 °C for 15 min, the supernatants were collected. The supernatant protein concentration was determined using the Bio-Rad Quick Start Bradford Kit (cat. 500-0201), according to the manufacturer’s instructions (Bio-Rad Laboratories, Hercules, CA, USA). The extracted proteins (10 μg per sample) were separated on 12 % SDS-PAGE gels and transferred to nitrocellulose membranes using a Mini-PROTEAN system (Bio-Rad). The membranes were blocked for 1 h at room temperature in phosphate-buffered saline (PBS) containing 5 % milk and washed at least three times with PBS buffer containing 0.5 % Tween 20 (PBST), each for 5 min. Primary antibodies were prepared in PBS buffer containing 1 % BSA and then incubated with the membranes overnight at 4 °C. After removing unbound antibodies by washing with PBST, the blots were incubated with rabbit anti-rabbit IgG secondary antibody (horseradish peroxidase conjugated, Promega, Madison, WI, USA) in PBST buffer at a dilution of 1:10,000 and visualized using a chemiluminescent kit (Thermo Fisher Scientific). Anti-APX (1:1000), anti-NR (1:2000), anti-SOD (1:2000), anti-P5CS (1:1000), and anti-tubulin (1:3000) were purchased from Agrisera (Vannas, Sweden). The anti-HSP90 (1:3000), HSP17.6 (1:3000), anti-P5CS (1:1000), and anti-GSNOR (1:2000) antibodies were used as described previously (Bai et al. 2011b; Kong et al. 2014). The biotin switch technique was used to detect protein S-nitrosylation status (Forrester et al. 2009; Bai et al. 2011a). Biotinylated proteins were pulled down with streptavidin agarose CL-6B (Thermo Fisher Scientific) and the S-nitrosylated protein was detected using the corresponding antibody.
Proline and malondialdehyde (MDA) content measurements
Proline content was measured as described (Nakazawa et al. 1982). Approximately, 0.5 g of fresh leaves of each sample was homogenized in 8 mL of 3 % aqueous sulfosalicylic acid, and the homogenate was centrifuged at 2000g for 10 min. Two milliliters of each extract, acidic ninhydrin, and glacial acetic acid were heated for 1 h in a boiling water bath, and the reaction was terminated in an ice bath. The reaction mixture was extracted with 4 mL of toluene, with vigorous mixing using a test tube stirrer for 15–20 s at 25 °C. The chromophore-containing toluene was aspirated from the aqueous phase and warmed to room temperature and its absorbance was read at 520 nm using toluene as a blank. The proline concentration was determined from a standard curve and calculated on a dry weight (DW) basis as follows: (µg mL−1proline × mL toluene) × 5 (g sample)−1 = µg proline g−1DW material.
MDA content was determined as described (Wang et al. 2010). Approximately, 0.5 g of fresh leaves per sample was homogenized in 10 mL of 10 % trichloroacetic acid (TCA) and centrifuged at 12,000g for 10 min. Two milliliters of 0.6 % thiobarbituric acid in 10 % TCA was then added to an aliquot of 2 mL of the supernatant. The mixture was heated in boiling water for 30 min and then quickly cooled in an ice bath. After centrifugation at 10,000g for 10 min, the absorbance of the supernatant at 450, 532, and 600 nm was determined. The MDA concentration, which was expressed as nmol g−1DW, was estimated from the formula: C (nmol mL−1) = 6.45 (A532 − A600) − 0.56A450.
In situ H2O2 and O2 − detection and quantification
In situ detection of H2O2 and O2 − was performed as reported, with some modifications (Able 2003). To detect H2O2, three leaf discs punched out at specific time points during the treatments were vacuum infiltrated in 10 mL of 1 mg mL−1 diaminobenzidine solution for 2 h and then cleared in boiling ethanol (95 %) for 10 min. The samples were subsequently stored and examined in 95 % ethanol. The amount of O2− in leaves was monitored by 10−2 M nitroblue tetrazolium (NBT) reduction at specific time points. Three leaf discs were vacuum infiltrated with 10 mL of NBT for 2 h, cleared in boiling ethanol (95 %) for 10 min, and examined in 95 % ethanol. The contents of H2O2 and O −2 in poplar were measured as described (Bai et al. 2011b).
Determination of ABA and jasmonic acid (JA) contents
The ABA and JA contents were measured as described, with minor modifications (Fragoso et al. 2014). Briefly, ~500 mg of leaf tissue subjected to different treatments was taken and rapidly frozen and ground in liquid nitrogen. Phytohormones were extracted with 1 mL of ethyl acetate spiked with internal standards (100 ng of [2H5]-JA and [2H6]-ABA). After extraction by vortexing for 10 min, the organic phase was obtained by centrifugation at 16,100g for 15 min at 4 °C. Samples were evaporated almost to dryness in a vacuum concentrator (Eppendorf, Hamburg, Germany) under reduced pressure at 30 °C. Leaf samples were then diluted in 600 μL of methanol:water (70:30, v/v). Analysis was performed with a Shimadzu 8040 HPLC–MS/MS system. Chromatography was performed on a Shimadzu FPLC system. Separation was achieved on a Shimadzu pack XR-ODS column (2.0 mm i.d. 75 mm). Formic acid (0.05 %) and ammonium formate (5 mM) in water and methanol were used as the A and B mobile phases, respectively. The elution profile was as follows: 0–0.01 min, 20 % B; 0.01–8.0 min, 20–95 % B; and 8.0–8.1 min, 20 % B. The mobile phase flow rate was 0.3 mL min−1. The column temperature was maintained at 50 °C. An ESI tandem mass spectrometer (Shimadzu) was operated in the negative ionization mode. The instrument parameters were optimized by infusion experiments with pure standards for ABA and JA, where available as requested. The ion spray voltage was maintained at −4500 eV. The heating block temperature was set at 500 °C. The DL temperature was set at 300 °C. Nebulizing gas was set at 3 L min−1, drying gas flow at 15 L min−1, and collision gas at 230 kPa. Sample-derived phytohormones were calculated based on the ratios of their ion intensity and of their respective internal standards. All quantifications were corrected according to the sample dilution, and extractions were performed with five biological replicates. The resulting amount of different phytohormones was then expressed as per grams fresh mass plant material.
Antioxidant enzyme activity assays
Approximately 1 g of leaf tissue from each sample was homogenized in 10 mL of extraction buffer containing 50 mM sodium phosphate (pH 7.0), 1 mM EDTA, 1 mM dithiothreitol (DTT), 1 mM glutathione, 5 mM MgCl2, 1 % PVP-40, and 20 % glycerin. The homogenates were centrifuged at 12,000g for 15 min at 4 °C, and the total soluble protein concentration of supernatants was determined by the Bradford method (Bradford 1976). The activities of MDHAR, DHAR, APX, and GR were determined as described (Chen et al. 2011).
SNO content determination and GSNOR enzyme activity assay
The total SNO content was determined via a chemiluminescence method with minor modifications (Bai et al. 2012). SNO detection was based on the reductive decomposition of nitroso species by an iodine/triiodide mixture that releases NO, and the released NO was subsequently measured by gas-phase chemiluminescence upon reaction with ozone. SNOs are sensitive to mercury-induced decomposition, in contrast to other nitroso species, including nitrosamines (RNNOs) and nitrosyl hemes. Briefly, the treated samples were homogenized in buffer containing 100 mM diethylenetetraminepentaacetic acid (1:5; w/v) and centrifuged at 3000g for 10 min. Then the supernatants were incubated with 10 mM N-ethylmaleimide for 15 min at 48 °C. For each sample, two aliquots were prepared: (1) one treated with 10 mM sulfanilamide for 15 min at 48 °C to eliminate nitrite and (2) one treated with 10 mM sulfanilamide and 7.3 mM HgCl2 for 15 min at 48 °C to eliminate nitrite and SNOs, respectively. These samples were then analyzed in a Nitric Oxide Monitor 410 (2B Technologies, Boulder, CO, USA). The data obtained from both aliquots (1 and 2) represented the total concentration of SNOs. The entire procedure was performed under a red safety light to protect SNOs from light-dependent decomposition.
GSNOR activity was determined spectrophotometrically at 25 °C by monitoring the oxidation of NADH at 340 nm (Bai et al. 2012). The treated samples were quickly homogenized in liquid nitrogen and extracted with an assay mixture containing 20 mM Tris–HCl (pH 8.0), 0.2 mM NADH, and 0.5 mM EDTA at 4 °C and then centrifuged at 3000g for 10 min. The supernatants were subjected to further assays, and the reaction was initiated by adding GSNO (Calbiochem, San Diego, CA, USA) to the supernatants at a final concentration of 400 mM. The activity was expressed as nanomoles NADH consumed per minute per milligram of protein (e340 6.22 mM−1 cm−1).
Electrolyte leakage assay
Electrolyte leakage was assayed according to a described method with some modifications (Wang et al. 2010). Briefly, tubes containing six to eight leaves detached from 20-day-old plants acclimated for varying periods at 4 °C were placed in a low-temperature bath (Grant) set at 0 °C. Ice chips were added to each tube after 1 h of incubation at 0 °C. The bath temperature was reduced at a rate of 2 °C h−1. The tubes were removed at defined temperatures and thawed overnight at 4 °C in the dark and then incubated with 6 mL of deionized water in a shaker with a shaking speed of 200 rpm and at 25 °C for 2 h. Electrical conductivity in the bathing solution was first determined (C1), and thereafter the samples were heated at 100 °C for 30 min and conductivity was determined again in the bathing solution (C2). Relative ion leakage was expressed as a percentage of the total conductivity after heating at 100 °C (i.e., relative ion leakage % = C1/C2 × 100).
Determination of nitrate reductase (NR) and NO synthase (NOS) activities
The NR activity was assayed following a described method with some modifications (Zhao et al. 2009). About 1 g of leaves was ground in liquid nitrogen and then resuspended in extraction buffer containing 100 mM HEPES–KOH (pH 7.5), 1 mM EDTA, 10 % (v/v) glycerol, 5 mM dithiothreitol, 0.1 % Triton X-100, 0.5 mM phenylmethylsulfonyl fluoride, 20 μM FAD, 1 μM leupeptin, 5 μM Na2MoO4, and 1 % polyvinylpyrrolidone. After centrifuging at 10,000g for 20 min at 4 °C, the supernatant was used for NR determination. The NR activity was measured by mixing 1 volume of extract with 5 volumes of pre-warmed (25 °C) assay buffer (100 mM HEPES–KOH, pH 7.5, 5 mM KNO3, and 0.25 mM NADH). The reaction was started by the addition of assay buffer, incubated at 25 °C for 30 min, and then stopped by the addition of 0.1 M zinc acetate. After 15 min, the tubes were centrifuged at 13,000g for 5 min. The nitrite produced was measured colorimetrically at 520 nm by adding 1 mL of 1 % (w/v) sulfanilamide in 3 M HCl plus 1 mL of 0.02 % (v/v) N-(1-naphthyl)-ethylenediamine in distilled water.
NOS activity was determined as described (Zhao et al. 2009). Briefly, approximately 1 g of leaves, together with 50 mg of polyvinylpolypyrrolidone, was ground in liquid N2 and then resuspended in extraction buffer (50 mM Tris–HCl, pH 7.4, 1 mM EDTA, 320 mM sucrose, 1 mM dithiothreitol, 1 μM leupeptin, 1 μM pepstatin, and 1 mM phenylmethylsulfonyl fluoride). After centrifuging at 10,000g for 30 min at 4 °C, the supernatant was used to determine NOS activity by the citrulline assay using the NOS assay kit (Cayman Chemical, Ann Arbor, MI, USA). The reaction mixture (50 μL) contained 25 mM Tris-HCl, pH 7.4, 3 μM tetrahydrobiopterin, 1 μM FAD, 1 μM FMN, 1 mM β-NADPH, 0.6 mM CaCl2, 0.1 μM calmodulin, 0.3 μM (37,000 Bq) [3H]Arg, and 10 μL of enzyme extract. After incubation for 30 min at 37 °C, the reaction was stopped by adding 400 μL of stop buffer (50 mM HEPES, pH 5.5, and 5 mM EDTA). A 100-μL resin slurry was added to the reaction mixture, and the resin was removed by centrifugation. Flowthrough (400 μL) was added to 5 mL of scintillation liquid and radioactivity was counted (LS 6000; Beckman). The protein contents in the supernatant were determined according to the Bradford method (Bradford 1976) with bovine serum albumin as a standard.
Results
Physiological response of poplar leaves to chilling treatment
To characterize the underlying mechanism of chilling stress on poplar growth, we first tested the effects of chilling stress at 4 °C on poplar leaf photosynthesis capability and ion leakage, both of which are important criteria for estimating the degree of damage experienced by a plant in response to chilling stress. As shown in Fig. 1a, prolonging the chilling time markedly suppressed the leaf photosynthesis capability, as measured by Fv/Fm,the parameter to reflect the leave photosynthesis capability. We found that low-temperature treatment gradually reduced the ratio of Fv/Fm, and reduction continued to decrease throughout the observation period at 4 °C (Fig. 1b). The ion leakage is an important index of plasma membrane intactness. We found that ion leakage in leaves gradually increased after chilling treatment at 4 °C (Fig. 1c). These data suggest that chilling stress suppressed leaf viability by suppressing the leaf photosynthesis capability and enhancing leaf electrolyte leakage.
Effects of cold stress on the leaf photosynthesis capability and electrolyte leakage. a Phenotype of poplar leaves subjected to cold stress at 4 °C. Poplar seedlings were cultivated under normal conditions for 3 weeks and then subjected to cold stress at 4 °C. At the indicated time points, images were captured, and the photosynthetic capabilities and ion leakage values were recorded. The photosynthetic capabilities were recorded as Fv/Fm using a PAM chlorophyll fluorometer (right). Images captured under white light were used as controls. The pseudocolor code depicted at the bottom of the images ranges from 0 (red) to 1.0 (purple). The experiment was repeated three times with similar results. One representative leaf is shown. Bars 1.5 cm. b Average Fv/Fm values and ion leakage. Fv/Fm and electrolyte leakage were determined for whole leaves subjected to cold stress at 4 °C for the indicated periods. Data represent the means of five replicate experiments (SE). Means denoted by different letters show significant differences at P < 0.05 according to Tukey’s test
Quantitative protein profiling of poplar in response to chilling
To establish the underlying mechanism of poplar’s response to chilling stress, we conducted an iTRAQ proteome analysis of differential protein changes in poplar leaves after 1, 3, or 7 days of chilling treatments. The sample without cold treatment was used as the control. We repeated the experiments three times and total spectra were generated and the data collected were analyzed with Mascot software. We then determined the ratios of the protein abundance for the following three groups: (1) 1 day of chilling treatment/the control without treatment, (2) 3 days of chilling treatment/the control without treatment, and (3) 7 days of chilling treatment/the control without treatment, with a fold change cutoff of >1.5 for increased expression and <0.5-fold for decreased expression. Our iTRAQ results showed that a total of 114 proteins had significant differences in expression (P < 0.05). Among them, 69 proteins were up-regulated (>1.5-fold) and 45 proteins were down-regulated (<0.5-fold) after chilling treatment, respectively (Tables 1, 2, Suppl. Table S1). The 114 identified protein species were divided into ten groups based on their biological functions (Tables 1, 2; Fig. 2a). Most of these proteins were classified in the group of material and energy metabolism, followed by plant hormone and antioxidant proteins and enzymes (Fig. 2a). Among the 69 up-regulated proteins and 45 down-regulated proteins, a total of 24 proteins were constitutively increased and 8 proteins were constitutively decreased, respectively, at all time points (Fig. 2b). We next conducted a hierarchical cluster analysis to categorize the proteins that showed differential expression profiles during chilling stress (Fig. 3c). We noticed that antioxidant enzymes and proteins, such as catalase (gi|90818818), ascorbate peroxidase (gi|411101112), and glutathione peroxidase (gi|224089376), were up-regulated upon exposure to chilling stress and clustered together (Fig. 2c). In addition, proteins involved in NO metabolism, including NR (gi|743935527) and GSNOR (gi|566159968), were up-regulated in response to chilling stress. Proteins associated with plant hormone biosynthesis and response, such as auxin response factor 2 family protein (gi|566215821), 1-aminocyclopropane-1-carboxylate oxidase (gi|7576446), and allene oxide synthase family protein (gi|222844179), and the defense response, such as heat shock protein 70 (gi|224112795) and delta-1-pyrroline-5-carboxylate synthase (gi|743879886), were differentially accumulated, suggesting their critical roles during poplar exposure to chilling stress.
Characterization of proteins differentially expressed in poplar after exposure to cold stress. a Functional classification of proteins that were differentially expressed after exposure to cold stress. b Venn diagram analysis of proteins that were differentially expressed after cold stress. 1 day/control: the differential protein number between 1 day of cold treatment and the control; 3 days/control: the differential protein number between 3 days of cold treatment and the control; and 7 days/control: the differential protein number between 7 days of cold treatment and the control. c Hierarchical clustering of the poplar leaf protein expression profile following cold stress treatment. Different colors correspond to the differential expression ratios of proteins, depicted in the bar at the bottom of the figure. Three-week-old poplar seedlings were subjected to cold stress at 4 °C for 1, 3, and 7 days, and seedlings not subjected to cold treatment were used as the control. Differences in protein accumulation between the treatment and control sample were compared by iTRAQ, and ratios of >1.5 or <0.5 were selected as cutoffs for differential protein expression
Effects of cold stress on the generation of NO and SNO, and on the activities of enzymes involved in NO metabolism. Three-week-old poplar seedlings were subjected to cold stress at 4 °C for 1, 3, 5, and 7 days, and seedling maintained at 25 °C were used as the control. The generation of NO (a) and SNO (c), the enzyme activities of NR and NOS (b), and GSNOR enzyme activity (d) were measured. Data represent the means of three replicate experiments (±SD). The means denoted by different letters indicate significant differences according to Tukey’s test (P < 0.05)
Chilling stress enhanced the accumulations of NO and RNS, and the activities of NR, NOS, and GSNOR
NO is reported to increase tolerance to cold stress in Arabidopsis (Zhao et al. 2009). Here, our iTRAQ results showed that proteins associated with NO metabolism including NR (gi|743935527) and GSNOR (gi|566159968) were increased after chilling stress, suggesting the potential role of NO in poplar’s response to chilling stress. Thus, we determined NO levels in poplar leaves after exposure to chilling stress. Chilling treatment induced a striking increase in NO levels in poplar leaves after 1, 3, and 7 days of treatment (Fig. 3a). NR and NOS are two major enzymes responsible for NO generation in plants (Neill et al. 2002b). Chilling stress induced both NR and NOS enzyme activity (Fig. 3b), which correlates with the iTRAQ results indicating that the levels of both enzymes increased after chilling stress. Chilling-induced activities of NR and NOS peaked after 3 days of treatment and then slightly declined after 5 and 7 days of treatment. These data suggest that NO is the endogenous signal that regulates poplar’s response to chilling stress.
NO and its derivatives can react with thiol in the tyrosine residues to form S-nitrosothiols (SNOs), which have been shown to alter the activity, localization, or conformation of target proteins (Spadaro et al. 2010; Spoel and Loake 2011). Cellular SNO level are controlled by the conserved cytosolic enzyme GSNO reductase (GSNOR) (Frungillo et al. 2014). Our iTRAQ data demonstrated that chilling stress induced the accumulation of poplar GSNOR (gi|566159968), suggesting a possible role for GSNOR in regulating the cytosolic redox status and SNO levels. To evaluate this possibility, we firstly measured the level of SNOs in poplar leaves. Similar to the pattern observed for NO generation, chilling treatment markedly gradually induced the accumulation of total SNOs in poplar leaves after chilling treatment (Fig. 3c). Chilling stress also induced GSNOR activity, which peaked after 3 days of treatment (Fig. 3d). This result correlates with the iTRAQ data presented above. However, GSNOR activity declined after 5 and 7 days of chilling treatment, although the activity remained higher than that of control leaves not subjected to chilling treatment (Fig. 3d), suggesting the existence of a feedback mechanism that suppressed the increase in GSNOR activity after 5 days of chilling treatment.
Chilling stress induced ROS accumulation and antioxidant enzyme activity
Because the iTRAQ results showed that chilling stress induced the accumulation of a set of antioxidant enzymes, and ROS including H2O2 and O2 − are important signals that mediate the plant’s response to abiotic stress (Neill et al. 2002a), we investigated whether chilling stress-induced ROS accumulation and whether the antioxidant enzymes were involved in ROS scavenging. We first quantified in situ ROS levels, including those of H2O2 and O2 −, in leaves exposed to chilling stress. In the absence of chilling stress, poplar leaves accumulated low levels of H2O2, as indicated by 3,3′-diaminobenzidine (DAB) staining. One day of exposure to chilling stress slightly increased the accumulation of H2O2, but 3 or 5 days of chilling stress resulted in increased H2O2 levels, and browning of the leaves. H2O2 levels increased after 7 days of chilling stress (Fig. 4a). This generation pattern for O2 − is similar to that for superoxide radicals, as indicated by nitroblue tetrazolium (NBT) staining. Compared with control leaves, chilling treatment, especially for 5 or 7 days, markedly increased O2 − accumulation (Fig. 4a). In accordance with this phenotype, the quantitative data of H2O2 (Fig. 4b) and O2 − (Fig. 4b) also demonstrate that chilling treatment substantially induced the increase of H2O2 and O2 − compared with the control leaf not subjected to cold stress. Malondialdehyde (MDA) is the important index to reflect the damage degree of plant plasma membrane under chilling injury. Here, we also found that cold treatment gradually increased the MDA content in the poplar leaves (Fig. 4c).
Effects of cold stress on ROS generation and antioxidant enzymes activities in poplar leaves. Three-week-old poplar seedlings were subjected to cold stress at 4 °C for 1, 3, 5, and 7 days, and seedlings maintained at 25 °C were used as the control. The in situ accumulation of H2O2 and O2 − was detected by DAB and NBT staining (a). The experiment was replicated three times with similar results. One representative leaf is shown. Quantification of H2O2 and O2 − (b), the accumulation of MDA (c), and the antioxidant enzyme activities of MDHAR (d), DHAR (e), APX (f), and GR (g) were analyzed after cold stress treatment for the indicated periods. Data represent the means of three replicate experiments (SE). Means denoted by different letters show significant differences at P < 0.05 according to Tukey’s test
The antioxidant enzyme system could maintain ROS at acceptable levels during environmental stress. Our iTRAQ results showed that chilling stress increased the content of a series of antioxidant enzymes, suggesting their potential role during chilling stress. We thus monitored the antioxidant enzyme activities of MDHAR, DHAR, APX, and GR (Fig. 4d–g) and found that chilling stress gradually induced these enzymes activities to different degrees. These results are in good agreement with our iTRAQ results and indicate the possible role of these antioxidant enzyme in suppressing the over-accumulation of ROS during chilling stress.
Controlling NO metabolism affects the response of poplar leave to chilling stress
Our above results showed that chilling stress induced NO accumulation and GSNOR activity, and increased SNO levels. To further determine the potential roles of NO and GSNOR in modulating poplar’s response to chilling stress, we treated the poplar plantlets with the NO scavenger cPTIO, and the GSNOR inhibitor N6022 or DA, respectively, by adding these inhibitors to the hydroponic solution, before subjecting the plants to chilling treatment. As shown in Fig. 5a, pretreatment with cPTIO, N6022, or DA markedly increased the plant’s sensitivity to chilling stress, as indicated by the lower Fv/Fm fluorescence (Fig. 5b) and higher ion leakage (Fig. 5c) in leaves exposed to 3 days of chilling stress, compared with control leaves. These data suggest that reducing NO accumulation by cPTIO or increasing RNS accumulation by GSNOR inhibitors damages the leaves. We also found that treatment with the GSNOR enzyme inhibitor D6022 or DA markedly increased the accumulation of SNOs and suppressed the activity of GSNOR in poplar leaves after 3 days of chilling treatment (Fig. 6a, b). Treatment with the NO scavenger cPTIO did not significantly affect the activity of GSNOR, but significantly reduced the generation of SNOs (Fig. 6b), possibly because cPTIO completely removed the NO that is required for SNO production in planta. The GSNOR enzyme inhibitor N6022 or DA also reduced the NR and NOS activity in the leaves after 3 days of chilling treatment, but cPTIO did not significantly affect NR or NOS activity (Fig. 6c, d).
Effects of treatment with the NO biosynthesis scavenger or GSNOR inhibitors on the leaf’s photosynthesis ability and electrolyte leakage under cold stress. a Phenotype of poplar leaves subjected to NO biosynthesis inhibitors and GSNOR inhibitor under cold stress at 4 °C. Three-week-old poplar seedlings were treated with the GSNOR enzyme inhibitor N6022 (5-chloro-3-(2-[(4-ethoxyphenyl)ethylamino]-2-oxoethyl)-1H-indole 2-carboxylic acid,2 μM) or DA (dodecanoic acid, 10 mM), or the NO scavenger cPTIO (30 μM), respectively, under cold stress at 4 °C for 3 days, and the photosynthetic capabilities were recorded by Fv/Fm imaging using a PAM chlorophyll fluorometer (right). Images captured under white light were used as controls. The pseudocolor untreated leaves were used as the control. The experiment was repeated three times with similar results. One representative leaf is shown. Bars 1.5 cm. b Average Fv/Fm values and ion leakage. Fv/Fm and electrolyte leakage were determined for whole leaves subjected to the inhibitors described above, and then subjected to cold stress at 4 °C for the indicated periods. Data represent the means of five replicate experiments (SE). Means denoted by different letters show significant differences at P < 0.05 according to Tukey’s test
Effects of different scavengers or inhibitors of RNS generation and RNS metabolism-related enzyme activity of poplar exposed to 3 days of cold stress. Three-week-old poplar seedlings were subjected to different scavenger or inhibitor treatments under cold stress at 4 °C for 3 days, as described above; SNO accumulation (a) and the enzyme activities of NR (b), GSNOR (c), and NOS (d) were measured. Data represent the means of three replicate experiments (SE). Means denoted by different letters show significant differences at P < 0.05 according to Tukey’s test
We then observed the in situ accumulation of H2O2 and O2 − after treatment with the NO scavenger cPTIO or the GSNOR inhibitor N6022 or DA followed by 3 days of cold stress. As shown in Fig. 7a, poplar leaves exposed to 3 days of chilling stress accumulated much higher levels of H2O2 than did control leaves not subjected to chilling stress. Pretreatment with cPTIO, N6022, or DA, respectively, markedly increased the in situ H2O2 accumulation and resulted in leaf browning than only with 3 days of chilling treatment (Fig. 7a). Furthermore, pretreatment with cPTIO, N6022, or DA, respectively, also resulted in more dark blue spots in the leaves, in contrast to leaves subjected only to 3 days of cold stress (Fig. 7a), suggesting that more superoxide radicals accumulated after cPTIO, N6022, or DA pretreatment. Consistent with the observed in situ accumulation of H2O2 and O2 −, the quantitative data demonstrated that pretreatment with cPTIO or N6022 or DA, respectively, strongly enhanced chilling-induced H2O2 or O2 − accumulation (Fig. 7b). Malondialdehyde (MDA) is a useful indicator of plasma membrane lipid oxidation and damage. We found that 3 days of chilling increased the MDA value and pretreatment with cPTIO, N6022, or DA further increased the MDA value (Fig. 7c), suggesting that cPTIO, N6022, or DA pretreatment aggravates chilling-induced plasma membrane damage.
Effects of N6022, DA, and cPTIO on ROS generation and antioxidant enzyme activity in poplar leaves after 3 days of cold stress treatment. Three-week-old poplar seedlings were subjected to the indicated scavengers or inhibitors, followed by cold stress treatment at 4 °C for 3 days. The in situ accumulation of H2O2 and O2 − were detected by 3,3′-diaminobenzidine (DAB) and nitroblue tetrazolium (NBT) staining, respectively (a). The experiment was repeated three times with similar results. One representative leaf is shown. The content of H2O2 and O2 − (b), the accumulation of MDA (c), and the antioxidant enzyme activities of MDHAR (d), DHAR (e), APX (f), and GR (g) were analyzed. Data represent the means of three replicate experiments (SE). Means denoted by different letters show significant differences at P < 0.05 according to Tukey’s test
We further measured the antioxidant enzyme activities of MDHAR, DHAR, APX, and GR after cPTIO, N6022, or DA pretreatment and chilling stress (Fig. 7d–g). Chilling treatment for 3 days markedly increased the activity of these enzymes, and pretreatment with cPTIO, N6022, or DA for 12 h, respectively, partially suppressed these antioxidant enzymes’ activities compared with those in plants exposed to only 3 days of chilling stress, which explains our finding that pretreatment with cPTIO, N6022, or DA caused an increase in H2O2 and O2 − levels (Fig. 7a–c).
Controlling NO metabolism affects defense-related protein accumulation during chilling stress
A previous study demonstrated that proline synthesis enhances Arabidopsis tolerance to cold stress (Zhao et al. 2009). Interestingly, our iTRAQ results showed that chilling induced the accumulation of P5CS (delta-1-pyrroline-5-carboxylate synthase, gi|743879886) and P5CR (pyrroline-5-carboxylate reductase, gi|743844755), both of which are important for proline biosynthesis. Furthermore, the accumulation of proline dehydrogenase (gi|743906676), which catalyzes proline degradation, was reduced after chilling stress, suggesting that proline metabolism is involved in poplar’s response to chilling stress. We thus measured the proline content in polar leaves after chilling stress and found that chilling stress indeed caused proline to accumulate in leaves (Fig. 8a). Treatment with the GSNOR inhibitor DA and N6022 or NO scavenger cPTIO markedly suppressed cold-induced proline biosynthesis in polar leaves (Fig. 8a), suggesting a role for NO metabolism in controlling proline biosynthesis under cold stress.
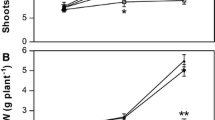
Effects of N6022, DA, and cPTIO on proline and sucrose accumulation and phytohormone biosynthesis in poplar leaves after 3 days of cold stress. Three-week-old poplar seedlings were subjected to the indicated scavenger or inhibitors, followed by cold stress at 4 °C for 3 days. The accumulation of proline (a) and sucrose (b) was measured, and the contents of JA (c) and ABA (d) were analyzed. Data represent the means of three replicate experiments (SE). Means denoted by different letters show significant differences at P < 0.05 according to Tukey’s test
Several lines of evidence indicate that the phytohoromones JA and ABA enhance tolerance to cold or freezing stress in Arabidopsis (Baron et al. 2012; Hu et al. 2013; Ding et al. 2015). Our iTRAQ data also demonstrated that several proteins associated with JA and ABA biosynthesis, such as 9-cis-epoxycarotenoid dioxygenase NCED (gi|566153852), lipoxygenase (gi|227345104), and allene oxide synthase family protein (gi|222844179), were up-regulated after 1 and 3 days of cold stress. In agreement with these findings, the content of JA and ABA in poplar leaves increased after 1 and 3 days of cold stress, and suppressing GSNOR activity by N6022 or DA, or NO accumulation by cPTIO also reduced JA or ABA accumulation after cold treatment (Fig. 8b, c), suggesting that NO signal-mediated JA and ABA biosynthesis has essential roles in poplar’s response to cold stress. Previous research showed that ethylene is also a positive regulator of cold or freezing stress in Arabidopsis (Catala et al. 2014). In agreement with this, our iTRAQ data showed that the levels of 1-aminocyclopropane-1-carboxylate oxidase, (gi|7576446), which is associated with ethylene biosynthesis, increased after cold stress treatment in poplar leaves. Furthermore, our iTRAQ data showed that proteins associated with the auxin and gibberellin biosynthesis and response pathways, such as auxin-responsive protein IAA (gi|62125392) and gibberellin 20-oxidase family protein (gi|566202999), increased after cold stress, suggesting potential roles for auxin and gibberellin in the cold stress response. A previous study showed that the WRKY transcription factor is involved in Glycine max (soybean) or Hordeum vulgare (barley) tolerance to cold stress (Mare et al. 2004; Zhou et al. 2008). Our iTRAQ also provided data that transcription factors, including those belonging to the WRKY transcription factor family, such as WRKY transcription factor 30 (gi|259121425) and WRKY transcription factor 17 (gi|259121399), were differentially regulated by cold stress. What is more, the bZIP transcription factor family protein (gi|566199296) and MYB transcription factor R2R3 type (gi|31980095) were also involved in this process.
Previous studies showed that starch and sucrose signaling are components of the cold stress response in plants (Seo et al. 2011; Kong et al. 2014). Here, our iTRAQ data revealed that glucose-6-phosphate dehydrogenase (gi|566208506) and a series of proteins involved in photosynthesis were differentially regulated after cold stress, suggesting that changes in sucrose accumulation occur during cold stress. Indeed, we measured sucrose accumulation in the leaves of poplar plants subjected or not to various pretreatments and then exposed to cold stress for 3 days, and found that suppressing GSNOR activity with N6022 or DA, or NO accumulation with c PTIO also reduced sucrose accumulation after cold stress (Fig. 8d).
Our iTRAQ data, as well as the physiological results presented above, showed that chilling stress increased the accumulation and activity of a series of enzymes, including those involved in NO metabolism and in antioxidant enzymes. To confirm this finding, we used immunoblot analysis to measure the accumulation of APX, SOD, GSNOR, and NR after 1, 3, and 7 days of chilling stress. As shown in Fig. 9a, we found the chilling treatment for 1 or 3 days increased these proteins’ accumulations compared to the leaves without chilling stress. Treatment for 7 days also induced protein accumulations of GSNOR and NR, but not too strongly (Fig. 9a) Other HSP proteins including HSP90 and HPS17.6 increased after chilling stress as observed by iTRAQ. Western blotting analysis confirmed that these protein accumulation increased after cold stress (Fig. 9a), while the tubulin protein as loading control did not change during different periods of chilling stress. We found that treatment with cPTIO, which reduces NO accumulation, suppressed the chilling-induced accumulation of APX, GR, SOD, HSP90, and HSP17.6, suggesting that NO is an important signal that mediates the chilling-induced accumulation of these proteins (Fig. 9a). Similarly, suppressing GSNOR activity by pretreatment with N6022 or DA also reduced the chilling-induced accumulation of these defense-related proteins (Fig. 9a). It is possible that loss of GSNOR activity caused more accumulation of RNS to damage the leaves by suppressing the accumulation of defense-related proteins.
Effects of different scavengers or inhibitors on cold-responsive protein expression and protein S-nitrosylation modification in poplar leaves subjected to cold stress. a Effects of cold stress on cold-responsive protein expression as determined by immunoblot analysis. Total proteins were extracted from leaves subjected to cold stress for the indicated periods, and the level of protein expression was analyzed using the corresponding antibody. Tubulin is included as a protein loading control. b Effect of the indicated scavenger or inhibitors on protein expression in poplar leaves after 3 days of cold stress. The poplar leaves were treated with cPTIO (30 μM), DA (10 mM), or N6022 (2 μM), respectively, followed by 3 days of cold stress, or only with 3 days of cold stress, and the protein expression level was measured by immunoblot analysis using the corresponding antibody. Tubulin is included as a protein loading control
NO-modulated protein S-nitrosylation during chilling stress
NO is a well-documented modulator of protein function by modulating the status of protein S-nitrosylation (He et al. 2004; Frungillo et al. 2014). We therefore examined if GSNOR is subjected to S-nitrosylation during chilling stress using the biotin switch approach. This technique relies on specific reduction of SNO groups by ascorbate followed by their labeling with biotin. Using this technique, we found that GSNOR is S-nitrosylated after 1 day of chilling treatment, while no obvious S-nitrosylation of GSNOR was detected in leaves not exposed to chilling treatment (Fig. 10a). Similarly, we found that chilling treatment increased the protein S-nitrosylation of APX and NR (Fig. 10a). Previous research reported that protein S-nitrosylation suppressed the activity of GSNOR in Arabidopsis (Frungillo et al. 2014). It is possible that S-nitrosylation also suppressed the activity of GSNOR1, APX, and NR after prolonged chilling treatment, which may explain our finding that long-term chilling (i.e., a 7-day chilling treatment) reduced the activities of NR, GSNOR, or APX in poplar leaves.
Modification of protein S-nitrosylation status by cold stress. a Cold induced the S-nitrosylation of NR, APX, and GSNOR. Poplar leaves were treated with N6022 (2 μM), respectively, followed by 1 day of cold stress, or with only 1 day of cold stress, and the degree of protein S-nitrosylation was measured by immunoblot analysis using the biotin switch technique. Total NR, APX, or GSNOR proteins were detected by the corresponding antibody before biotin switch assay. Ponceau staining was used to confirm the equal addition among different samples. During biotin switch assay, the sample without Asc (20 mM ascorbate) treatment was used as the negative control. The signal ratio means the protein S-nitrosylation degree in contrast to the total protein. b Modification of protein S-nitrosylation status by thioredoxin. Poplar leaves were treated with 3 days of cold stress, and the total proteins were extracted for biotin switch assay. For thioredoxin treatment, thioredoxin (TRX, 50 μM) was incubated with the total protein before detecting SNO–GSNOR by biotin switch assay. For denitrosylation (De-nitro) treatment, DTT (5 mM) was incubated with the total protein before protein S-nitrosylation detection. The total GSNOR protein content was detected by a GSNOR antibody. Ponceau staining ensures equal protein loading. The signal ratio shows the S-nitrosylation degree of GSNOR in contrast to the total GSNOR protein
Treatment with the GSNOR inhibitor N6022 markedly increased the protein S-nitrosylation degree of APX, NR, and GSNOR in poplar leaves after 1 day of chilling stress (Fig. 10a), probably because treatment with GSNOR inhibitor induced the over-production of SNOs in leaves. This result explains why treatment with N6022 aggravated leaf damage in plants subjected to chilling stress. Our iTRAQ data indicated that thioredoxin m (gi|222843925) was induced by cold treatment, and Kneeshaw et al. (2014) reported that the Arabidopsis oxidoreductase thioredoxin-h5 (TRXh5) showed a selective protein denitrosylation activity to modulate plant immunity. We also detected the effect of thioredoxin on the cold-induced protein S-nitrosylation status. As shown in Fig. 10b, thioredoxin treatment obviously reduced the S-nitrosylation status of GSNOR after 3 days of cold stress; as the negative control, we treated the sample after 3-day cold stress with the reducing reagent dithiothreitol (DTT) to denitrosylate the protein. DTT treatment completely abolished the protein nitrosylation status for GSNOR, indicating a dynamic regulating of protein S-nitrosylation and enzyme activity by the NO signal and reducing the thioredoxin system in the response of poplar to cold stress.
Discussion
Poplar applies multiple strategies to enhance its tolerance to chilling stress
Low-temperature stress is a major factor limiting the geographical distribution of plants and their growth and development (He et al. 2004; Wilson et al. 2008; Yu et al. 2014). Many plants exposed to low temperatures have evolved mechanisms to enhance their tolerance to cold stress (Zhao et al. 2009). Identifying and characterizing the genes involved in cold acclimation have advanced our knowledge of the molecular mechanisms underlying cold acclimation and freezing tolerance in plants. Previous studies of cold acclimation mainly focused on crop plants, such as rice and barley, or model plants, such as Arabidopsis (He et al. 2004). The molecular mechanism underlying the woody plant’s response to cold stress is poorly understood. Here, we applied a quantitative proteomic approach to investigate the molecular mechanism of poplar’s response to chilling stress. Our iTRAQ results showed that a series of proteins exhibited dramatic changes after chilling stress. These proteins can be classified into ten categories. Among these, we identified proteins associated with NO metabolism, including NR and GSNOR, which suggests the importance of the NO signal during the response to chilling stress in poplar. In agreement with this notion, we found that the main enzyme activities responsible for NO biosynthesis, including NR and NOS, increased in activity after chilling stress, and that reducing NO accumulation using the NO scavenger cPTIO reduced the tolerance to chilling stress. Thus, NO is an important factor in the response to chilling stress in poplar.
Previously, it was reported that NO enhanced the accumulation of proline by modulating the expressions of P5CS and ProDH to increase tolerance to cold stress in Arabidopsis, and that this tolerance could be impaired by treatment with an NR inhibitor or NO scavenger (Zhao et al. 2009). We found that cold stress induced the rapid accumulation of oxygen species, including H2O2 and O2 −, and increased both the level and activity of antioxidant proteins, which suggests that cold stress increases the activities of antioxidant enzymes to reduce ROS damage (Figs. 4, 7), as a reduction in antioxidant enzyme activity markedly increased the over-accumulation of ROS and reduced the photosynthesis capability during cold stress. Our iTRAQ and immunoblot analyses showed that cold stress increased the accumulation of P5CS, P5CR, and proline. Such an effect could be inhibited by pretreatment with the NO scavenger cPTIO (Figs. 8a, 9a), suggesting that NO-mediated proline biosynthesis has a similar function in poplar as in Arabidopsis under cold stress. Furthermore, cold also induced sucrose accumulation, and this effect could be suppressed by pretreatment with a GSNOR enzyme inhibitor or the NO scavenger cPTIO, which confirmed the previous suggestion that starch and sucrose signaling modulates cold stress in plants (Seo et al. 2011; Kong et al. 2014).
Previous research showed that protein stability or protein degradation, such as ICE1 degradation in Arabidopsis, was involved in the plant’s response to cold stress (Dong et al. 2006). In plants, two major proteolytic systems degrade damaged or oxidized proteins under stress, the ubiquitin–proteasome system and vacuolar autophagy. In plant cells, the ubiquitin–proteasome system and a specific type of vacuolar autophagy, chaperone-mediated autophagy, are involved in the degradation of proteins during oxidative stress (Vierstra 1996). Autophagy is involved in the degradation of oxidized proteins under oxidative stress conditions in Arabidopsis (Xiong et al. 2007). In the ubiquitin–proteasome system, ubiquitin is covalently attached to target proteins through the action of three enzymes known as E1, E2, and E3, and ubiquitination results in the degradation of the target protein. The iTRAQ data presented in the present work demonstrated that cold stress induced the accumulation of proteins involved in protein degradation (Table 1). In addition, the iTRAQ and immunoblotting analyses revealed that autophagy 4b and autophagy 8 were induced after cold stress (Table 1; Fig. 9a), suggesting the potential and novel role of autophagy in degrading oxidative proteins during cold stress. Given that NO has been reported to modulate protein degradation in response to light stimulation (Lozano-Juste and Leon 2011), it is possible that NO is also involved in protein degradation during cold stress. Among the differentially expressed proteins identified by iTRAQ, we identified several proteins associated with epigenetic modification, including histone acetylation, such as histone deacetylase (gi|224131666) and SET domain protein (gi|224117488). These data suggest that epigenetic regulation may contribute to the cold stress response in poplar. In agreement with our data, a previous study demonstrated that HDACs (histone deacetylases), including HDA6 and HDA19, mediate histone deacetylation in response to ABA or salt stress in Arabidopsis (Chen and Wu 2010). The histone acetyltransferase (HAT) GCN5 interacts with the transcriptional adaptor (ADA) to regulate cold tolerance (Stockinger et al. 2001). It is possible that cold-induced epigenetic changes might be one of the mechanisms underlying adaptation to cold stress in poplar.
It is reported that the plant hormones JA, ABA, and ethylene positively regulate tolerance to cold stress in Arabidopsis (Shi et al. 2012; Hu et al. 2013; Catala et al. 2014; Ding et al. 2015). Our iTRAQ data showed that many proteins associated with JA, ABA, and ethylene biosynthesis were induced after cold stress. Consistent with this observation, we found that cold stress induced increases in JA and ABA contents (Fig. 8c, d). Furthermore, proteins involved in gibberellin (GA) and auxin signaling were also differentially accumulated after cold stress. The GA signal has been reported to participate in the plant’s response to abiotic stress through DELLA, which interacts with EIN3 or JAZs, components of the JA signal transduction pathway, to modulate the response to environmental stress or development in Arabidopsis (An et al. 2012). Furthermore, it is possible that the GA signal interacts with the JA or ethylene signal to control the response to cold stress in poplar. In our study, we also found that several proteins associated with cell division and the cell cycle were less abundant after cold stress, which might explain the slow growth observed during cold conditions.
Based on our iTRAQ data and physiological analysis, we propose that poplar adopts multiple strategies, including activating antioxidant enzyme activity, inducing proline, phytohormones, and sucrose accumulation, controlling cell division, triggering degradation at the post-translation level, and even imitating epigenetic modification, to adapt to cold environments.
GSNOR-mediated control of RNS levels and the NO signal affect poplar defenses against chilling stress.
In plants, a burst of reactive oxygen and nitrogen species can mediate the cellular response to the environment by modifying cysteine residues in signaling proteins (Spadaro et al. 2010). Cysteine reactivity toward NO leads to the formation of SNOs, which play important roles in pathogenesis and immunity (Liu et al. 2001). Arabidopsis knockout mutants deficient in GSNO reductase 1 (GSNOR1) fail to remove cellular SNOs and consequently accumulate high levels of protein-SNO; these mutants are also impaired in SA-dependent immune signaling and are highly susceptible to disease (Feechan et al. 2005), demonstrating that GSNOR1 plays a critical role in governing protein-SNO levels during the plant immune responses. Our iTRAQ data revealed that cold stress induced the accumulation of GSNOR, suggesting that this enzyme functions in the response to cold stress in poplar. Further enzyme activity analysis showed that GSNOR activity was markedly increased after exposure to cold stress. Cold stress also induced an increase in RNS, whereas suppressing GSNOR activity reduced RNS content and further impaired leaf vigor, suggesting that GNSOR adjusts RNS levels in poplar during cold stress. In agreement with our result, a previous study showed that GSNOR regulates NO homeostasis under high temperature stress, and that loss of GSNOR activity rendered the Arabidopsis GSNO reductase 1 (gsnor1) mutant more sensitive to high temperatures (Lee et al. 2008).
Genetic manipulation of the NO signal in different NO metabolism mutants also affects the plant’s response to abiotic stress. Our iTRAQ results and physiological results demonstrated that cold stress induced the NR and NOS enzyme activities responsible for NO generation. Suppressing NR and NOS activities reduced plant tolerance to cold stress, inhibited antioxidant enzyme activities, and increased ROS over-accumulation, confirming that the NO signal enhances tolerance to cold stress in poplar by scavenging ROS. Suppressing NO generation also reduced proline and sucrose accumulation, and influenced JA and ABA biosynthesis, suggesting that the NO signal has broad functions in triggering tolerance to cold stress in poplar. Previously, feedback regulation of NR activity by the NO signal during nitrate uptake was suggested (Frungillo et al. 2014). Our iTRAQ results and enzyme activity analysis showed that long-term cold stress (i.e., for 5 or 7 days) partially reduced NR and NOS activity compared with the 3-day treatment, suggesting that a similar feedback regulatory mechanism exists for NR activity in poplar under long-term cold stress. NR activity has been coupled to GSNOR1 activity through S-nitrosylation modification in Arabidopsis (Frungillo et al. 2014), because the over-generation of NO by NR activity caused the over-accumulation of RNS, which is strictly controlled by GSNOR activity, while RNS often regulates enzyme activity through S-nitrosylation. Our immunoblot analysis revealed that levels of S-nitrosylation increased sharply after 5 days of cold stress, and that promoting RNS accumulation by inhibiting GSNOR activity further increased the S-nitrosylation degree of NR protein, suggesting that NR activity is controlled by a novel post-transcriptional modification mechanism under cold stress. We also observed that APX is modified by S-nitrosylation. The GSNOR enzyme itself could be S-nitrosylated after long-term exposure to cold stress (Fig. 9c), which could explain why long-term cold stress reduces GSNOR activity. Recently, Kneeshaw et al. (2014) reported that the Arabidopsis oxidoreductase thioredoxin-h5 (TRXh5) reverses SNO modifications by acting as a selective protein-SNO reductase. Thioredoxin is a class of small redox proteins and plays a role in many important biological processes including redox signaling, and it acts as antioxidant by facilitating the reduction of other proteins by cysteine thiol–disulfide exchange. Our iTRAQ data found that thioredoxin m (gi|222843925) was induced by cold treatment, suggesting the potential role of thioredoxin in reverse S-nitrosylation modification to antagonize the damage of S-nitrosylation on protein activity. Our iTRAQ data also show that other proteins, such as L-ascorbate peroxidase family protein (gi|224103203), ascorbate peroxidase (gi|411101112), glutathione peroxidase (gi|224089376), and ascorbate peroxidase (gi|411101112) could be induced after cold stress; these enzymes could modulate the cytosolic redox status that possibly reverse S-nitrosylation damage during cold stress. Thus, GSNOR activity and protein S-nitrosylation status may be regulated by a feedback mechanism in response to cold stress.
Based on our quantitative proteomics and physiological findings, we conclude that poplar has multiple strategies to enhance its adaptation to cold stress and that homeostasis plays a central role in this process. We propose a model that illustrates the essential function of the NO signal during adaptation to cold stress in poplar (Fig. 11). Upon cold stress, the poplar seedling quickly induces the generation of NO through enhancing NR and NOS enzyme activity. NO is the central activator of antioxidant enzymes that scavenge the over-accumulated ROS in response to cold stress. Furthermore, NO also activates GSNOR to reduce the over-accumulation of RNS after cold stress. NO subsequently acts as the signal that activates the downstream defense response, including proline and sucrose biosynthesis, stimulates phytohormone biosynthesis, and controls cell division. Protein degradation and epigenetic modification also participate in poplar’s response to cold stress. Furthermore, NO and RNS homeostases are strictly controlled by NR and GSNOR enzymes, respectively, and the activities of NR, GSNOR, and antioxidant enzymes are reduced by NO or RNS through a feedback mechanism involving S-nitrosylation. In conclusion, we show that the NO signal and GSNOR enzyme activity have important roles in the plant’s response to cold stress in poplar. These findings may be used to genetically engineer poplar with enhanced resistance to cold stress.
Schematic representation of a proposed model for enhancing poplar leaf tolerance to cold stress. Upon cold stress, NO production is rapidly induced in the poplar leaves through NR or NOS enzyme activity. NO plays the central role in enhancing poplar tolerance to cold stress: NO can trigger the downstream defense response by increasing proline and sucrose accumulation, induce phytohormone biosynthesis, modulate cell division, and trigger various protein degradation and epigenetic modification mechanisms. NO also activates antioxidant enzymes, which scavenge the over-accumulated ROS, and activate GSNOR activity to avoid the over-accumulation of RNS subjected to cold stress. Furthermore, the NO signal fine-tunes by feedback regulating NR, GSNOR, and APX enzyme activity through protein S-nitrosylation
Author contribution statement
X.H. and J.S. designed the research. T.C., J.C., P.W. and G.W. performed the research. X.H., J.S. and A.E. analyzed the data. X.H and J.S wrote the article.
Abbreviations
- ABA:
-
Abscisic acid
- APX:
-
Ascorbate peroxidase
- cPTIO:
-
2-(4-Carboxyphenyl)-4, 4, 5, 5–tetramethylimidazoline-1-oxyl-3-oxide
- DA:
-
Dodecanoic acid
- DHAR:
-
Dehydroascorbate reductase
- GR:
-
Glutathione reductase
- GSNO:
-
S-nitrosoglutathione
- GSNOR:
-
GSNO reductase
- MDA:
-
Malondialdehyde
- NO:
-
Nitric oxide
- NOS:
-
NO synthase
- NR:
-
Nitrate reductase
- JA:
-
Jasmonic acid
- MDHAR:
-
Monodehydroascorbate reductase
- RNS:
-
Reactive nitrogen species
- ROS:
-
Reactive oxygen species
- SNO:
-
S-nitrosothiol
References
Able AJ (2003) Role of reactive oxygen species in the response of barley to necrotrophic pathogens. Protoplasma 221:137–143
Alexandersson E, Ali A, Resjo S, Andreasson E (2013) Plant secretome proteomics. Front. Plant Sci 4:9
An F, Zhang X, Zhu Z, Ji Y, He W, Jiang Z, Li M, Guo H (2012) Coordinated regulation of apical hook development by gibberellins and ethylene in etiolated Arabidopsis seedlings. Cell Res 22:915–927
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Bai X, Yang L, Tian M, Chen J, Shi J, Yang Y, Hu X (2011a) Nitric oxide enhances desiccation tolerance of recalcitrant Antiaris toxicaria seeds via protein S-nitrosylation and carbonylation. PLoS One 6:e20714
Bai X, Yang L, Yang Y, Ahmad P, Yang Y, Hu X (2011b) Deciphering the protective role of nitric oxide against salt stress at the physiological and proteomic levels in maize. J Proteome Res 10:4349–4364
Bai XG, Chen JH, Kong XX, Todd CD, Yang YP, Hu XY, Li DZ (2012) Carbon monoxide enhances the chilling tolerance of recalcitrant Baccaurea ramiflora seeds via nitric oxide-mediated glutathione homeostasis. Free Radic Biol Med 53:710–720
Baron KN, Schroeder DF, Stasolla C (2012) Transcriptional response of abscisic acid (ABA) metabolism and transport to cold and heat stress applied at the reproductive stage of development in Arabidopsis thaliana. Plant Sci 188–189:48–59
Barroso JB, Corpas FJ, Carreras A, Rodriguez-Serrano M, Esteban FJ, Fernandez-Ocana A, Chaki M, Romero-Puertas MC, Valderrama R, Sandalio LM, del Rio LA (2006) Localization of S-nitrosoglutathione and expression of S-nitrosoglutathione reductase in pea plants under cadmium stress. J Exp Bot 57:1785–1793
Beck EH, Heim R, Hansen J (2004) Plant resistance to cold stress: mechanisms and environmental signals triggering frost hardening and dehardening. J Biosci 29:449–459
Bindschedler LV, Cramer R (2011) Quantitative plant proteomics. Proteomics 11:756–775
Catala R, Lopez-Cobollo R, Mar Castellano M, Angosto T, Alonso JM, Ecker JR, Salinas J (2014) The Arabidopsis 14-3-3 protein RARE COLD INDUCIBLE 1A links low-temperature response and ethylene biosynthesis to regulate freezing tolerance and cold acclimation. Plant Cell 26:3326–3342
Chaki M, Fernandez-Ocana AM, Valderrama R, Carreras A, Esteban FJ, Luque F, Gomez-Rodriguez MV, Begara-Morales JC, Corpas FJ, Barroso JB (2009) Involvement of reactive nitrogen and oxygen species (RNS and ROS) in sunflower-mildew interaction. Plant Cell Physiol 50:265–279
Chaki M, Valderrama R, Fernandez-Ocana AM, Carreras A, Gomez-Rodriguez MV, Pedrajas JR, Begara-Morales JC, Sanchez-Calvo B, Luque F, Leterrier M, Corpas FJ, Barroso JB (2011) Mechanical wounding induces a nitrosative stress by down-regulation of GSNO reductase and an increase in S-nitrosothiols in sunflower (Helianthus annuus) seedlings. J Exp Bot 62:1803–1813
Chen LT, Wu K (2010) Role of histone deacetylases HDA6 and HDA19 in ABA and abiotic stress response. Plant Signal Behav 5:1318–1320
Chen Q, Yang L, Ahmad P, Wan X, Hu X (2011) Proteomic profiling and redox status alteration of recalcitrant tea (Camellia sinensis) seed in response to desiccation. Planta 233:583–592
Chinnusamy V, Gong Z, Zhu JK (2008) Abscisic acid-mediated epigenetic processes in plant development and stress responses. J Integr Plant Biol 50:1187–1195
Corpas FJ, Chaki M, Fernandez-Ocana A, Valderrama R, Palma JM, Carreras A, Begara-Morales JC, Airaki M, del Rio LA, Barroso JB (2008) Metabolism of reactive nitrogen species in pea plants under abiotic stress conditions. Plant Cell Physiol 49:1711–1722
Desikan R, Cheung MK, Bright J, Henson D, Hancock JT, Neill SJ (2004) ABA, hydrogen peroxide and nitric oxide signalling in stomatal guard cells. J Exp Bot 55:205–212
Ding Y, Li H, Zhang X, Xie Q, Gong Z, Yang S (2015) OST1 kinase modulates freezing tolerance by enhancing ICE1 stability in Arabidopsis. Dev Cell 32:278–289
Dong CH, Agarwal M, Zhang Y, Xie Q, Zhu JK (2006) The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degradation of ICE1. Proc Natl Acad Sci USA 103:8281–8286
Feechan A, Kwon E, Yun BW, Wang Y, Pallas JA, Loake GJ (2005) A central role for S-nitrosothiols in plant disease resistance. Proc Natl Acad Sci USA 102:8054–8059
Forrester MT, Foster MW, Benhar M, Stamler JS (2009) Detection of protein S-nitrosylation with the biotin-switch technique. Free Radic Biol Med 46:119–126
Foyer CH, Noctor G (2011) Ascorbate and glutathione: the heart of the redox hub. Plant Physiol 155:2–18
Fragoso V, Rothe E, Baldwin IT, Kim SG (2014) Root jasmonic acid synthesis and perception regulate folivore-induced shoot metabolites and increase Nicotiana attenuata resistance. New Phytol 202:1335–1345
Frungillo L, Skelly MJ, Loake GJ, Spoel SH, Salgado I (2014) S-nitrosothiols regulate nitric oxide production and storage in plants through the nitrogen assimilation pathway. Nat Commun 5:5401
Gaudet M, Pietrini F, Beritognolo I, Iori V, Zacchini M, Massacci A, Mugnozza GS, Sabatti M (2011) Intraspecific variation of physiological and molecular response to cadmium stress in Populus nigra L. Tree Physiol 31:1309–1318
He Y, Tang RH, Hao Y, Stevens RD, Cook CW, Ahn SM, Jing L, Yang Z, Chen L, Guo F, Fiorani F, Jackson RB, Crawford NM, Pei ZM (2004) Nitric oxide represses the Arabidopsis floral transition. Science 305:1968–1971
Hu Y, Jiang L, Wang F, Yu D (2013) Jasmonate regulates the inducer OF CBF expression-c-repeat binding factor/DRE binding factor1 cascade and freezing tolerance in Arabidopsis. Plant Cell 25:2907–2924
Jackson RB, Banner JL, Jobbagy EG, Pockman WT, Wall DH (2002) Ecosystem carbon loss with woody plant invasion of grasslands. Nature 418:623–626
Jorrin-Novo JV (2009) Plant proteomics. J Proteomics 72:283–284
Kneeshaw S, Gelineau S, Tada Y, Loake GJ, Spoel SH (2014) Selective protein denitrosylation activity of thioredoxin-h5 modulates plant immunity. Mol Cell 56:153–162
Kong X, Ma L, Yang L, Chen Q, Xiang N, Yang Y, Hu X (2014) Quantitative proteomics analysis reveals that the nuclear cap-binding complex proteins Arabidopsis CBP20 and CBP80 modulate the salt stress response. J Proteome Res 13:2495–2510
Lee U, Wie C, Fernandez BO, Feelisch M, Vierling E (2008) Modulation of nitrosative stress by S-nitrosoglutathione reductase is critical for thermotolerance and plant growth in Arabidopsis. Plant Cell 20:786–802
Liu L, Hausladen A, Zeng M, Que L, Heitman J, Stamler JS (2001) A metabolic enzyme for S-nitrosothiol conserved from bacteria to humans. Nature 410:490–494
Lozano-Juste J, Leon J (2011) Nitric oxide regulates DELLA content and PIF expression to promote photomorphogenesis in Arabidopsis. Plant Physiol 156:1410–1423
Luo M, Liu X, Singh P, Cui Y, Zimmerli L, Wu K (2012) Chromatin modifications and remodeling in plant abiotic stress responses. Biochim Biophys Acta 1819:129–136
Mare C, Mazzucotelli E, Crosatti C, Francia E, Stanca AM, Cattivelli L (2004) Hv-WRKY38: a new transcription factor involved in cold- and drought-response in barley. Plant Mol Biol 55:399–416
Minami A, Nagao M, Ikegami K, Koshiba T, Arakawa K, Fujikawa S, Takezawa D (2005) Cold acclimation in bryophytes: low-temperature-induced freezing tolerance in Physcomitrella patens is associated with increases in expression levels of stress-related genes but not with increase in level of endogenous abscisic acid. Planta 220:414–423
Nakazawa K, Tanaka H, Arima M (1982) Rapid, simultaneous and sensitive determination of free hydroxyproline and proline in human serum by high-performance liquid chromatography. J Chromatogr 233:313–316
Neill S, Desikan R, Hancock J (2002a) Hydrogen peroxide signalling. Curr Opin Plant Biol 5:388–395
Neill SJ, Desikan R, Clarke A, Hurst RD, Hancock JT (2002b) Hydrogen peroxide and nitric oxide as signalling molecules in plants. J Exp Bot 53:1237–1247
Reichstein M, Bahn M, Mahecha MD, Kattge J, Baldocchi DD (2014) Linking plant and ecosystem functional biogeography. Proc Natl Acad Sci USA 111:13697–13702
Sakamoto A, Ueda M, Morikawa H (2002) Arabidopsis glutathione-dependent formaldehyde dehydrogenase is an S-nitrosoglutathione reductase. FEBS Lett 515:20–24
Seo PJ, Kim MJ, Ryu JY, Jeong EY, Park CM (2011) Two splice variants of the IDD14 transcription factor competitively form nonfunctional heterodimers which may regulate starch metabolism. Nat Commun 2:303
Shi Y, Tian S, Hou L, Huang X, Zhang X, Guo H, Yang S (2012) Ethylene signaling negatively regulates freezing tolerance by repressing expression of CBF and type-A ARR genes in Arabidopsis. Plant Cell 24:2578–2595
Spadaro D, Yun BW, Spoel SH, Chu C, Wang YQ, Loake GJ (2010) The redox switch: dynamic regulation of protein function by cysteine modifications. Physiol Plant 138:360–371
Spoel SH, Loake GJ (2011) Redox-based protein modifications: the missing link in plant immune signalling. Curr Opin Plant Biol 14:358–364
Stockinger EJ, Mao Y, Regier MK, Triezenberg SJ, Thomashow MF (2001) Transcriptional adaptor and histone acetyltransferase proteins in Arabidopsis and their interactions with CBF1, a transcriptional activator involved in cold-regulated gene expression. Nucleic Acids Res 29:1524–1533
Tuskan GA, Difazio S, Jansson S, Bohlmann J, Grigoriev I, Hellsten U, Putnam N, Ralph S, Rombauts S, Salamov A et al (2006) The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313:1596–1604
Vierstra RD (1996) Proteolysis in plants: mechanisms and functions. Plant Mol Biol 32:275–302
Wang L, Yang L, Yang F, Li X, Song Y, Wang X, Hu X (2010) Involvements of H2O2 and metallothionein in NO-mediated tomato tolerance to copper toxicity. J Plant Physiol 167:1298–1306
Wilson ID, Neill SJ, Hancock JT (2008) Nitric oxide synthesis and signalling in plants. Plant Cell Environ 31:622–631
Xiong L, Zhu JK (2001) Abiotic stress signal transduction in plants: molecular and genetic perspectives. Physiol Plant 112:152–166
Xiong Y, Contento AL, Nguyen PQ, Bassham DC (2007) Degradation of oxidized proteins by autophagy during oxidative stress in Arabidopsis. Plant Physiol 143:291–299
Yu M, Lamattina L, Spoel SH, Loake GJ (2014) Nitric oxide function in plant biology: a redox cue in deconvolution. New Phytol 202:1142–1156
Zhao MG, Chen L, Zhang LL, Zhang WH (2009) Nitric reductase-dependent nitric oxide production is involved in cold acclimation and freezing tolerance in Arabidopsis. Plant Physiol 151:755–767
Zhou QY, Tian AG, Zou HF, Xie ZM, Lei G, Huang J, Wang CM, Wang HW, Zhang JS, Chen SY (2008) Soybean WRKY-type transcription factor genes, GmWRKY13, GmWRKY21, and GmWRKY54, confer differential tolerance to abiotic stresses in transgenic Arabidopsis plants. Plant Biotechnol J 6:486–503
Zhu JH, Dong CH, Zhu JK (2007) Interplay between cold-responsive gene regulation, metabolism and RNA processing during plant cold acclimation. Curr Opin Plant Biol 10:290–295
Acknowledgments
The authors would like to extend their sincere appreciation to the Deanship of Scientific Research at King Saud University for funding this research (Research Group NO. RG 1435-014).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Cheng, T., Chen, J., EF, A. et al. Quantitative proteomics analysis reveals that S-nitrosoglutathione reductase (GSNOR) and nitric oxide signaling enhance poplar defense against chilling stress. Planta 242, 1361–1390 (2015). https://doi.org/10.1007/s00425-015-2374-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-015-2374-5