Abstract
Main conclusion
The ribosome inactivating protein BE27 displays several biological activities in vitro that could result in a broad action against several types of pathogens.
Beetin 27 (BE27), a ribosome-inactivating protein (RIP) from sugar beet (Beta vulgaris L.) leaves, is an antiviral protein induced by virus and signaling compounds such as hydrogen peroxide and salicylic acid. Its role as a defense protein has been attributed to its RNA polynucleotide:adenosine glycosidase activity. Here we tested other putative activities of BE27 that could have a defensive role against pathogens finding that BE27 displays rRNA N-glycosidase activity against yeast and Agrobacterium tumefaciens ribosomes, DNA polynucleotide:adenosine glycosidase activity against herring sperm DNA, and magnesium-dependent endonuclease activity against the supercoiled plasmid PUC19 (nicking activity). The nicking activity could be a consequence of an unusual conformation of the BE27 active site, similar to that of PD-L1, a RIP from Phytolacca dioica L. leaves. Additionally, BE27 possesses superoxide dismutase activity, thus being able to produce the signal compound hydrogen peroxide. BE27 is also toxic to COLO 320 cells, inducing apoptosis in these cells by either activating the caspase pathways and/or inhibiting protein synthesis. The combined effect of these biological activities could result in a broad action against several types of pathogens such as virus, bacteria, fungi or insects.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants produce defense-related proteins upon infection with oomycetes, fungi, bacteria, or viruses, or insect attack (van Loon et al. 2006). Several types of these proteins are common and have been classified into 17 families of pathogenesis-related proteins (PRs). Others such as amylases, cell wall hydroxyproline-rich glycoproteins, glycine-rich proteins, polygalacturonase-inhibiting proteins, lipoxygenases, lipase-like gene products, ribosome-inactivating proteins (RIPs), lectins and cysteine-rich peptides occur more specifically in some plant species (van Loon et al. 2006). Most of them are induced by signaling compounds such as salicylic acid, jasmonic acid or ethylene and display antimicrobial activities in vitro through different enzymatic activities (van Loon et al. 2006).
Some RIPs can be considered as defense-related proteins since they are induced by salt stress (Rippmann et al. 1997), insect elicitors (Engelberth et al. 2012), mechanical wounding (Song et al. 2000; Engelberth et al. 2012), pathogenic fungus (Tan et al. 2013) and temperature and ultraviolet light (Qin et al. 2009). Additionally, some of them are induced by signaling compounds such as jasmonic acid (Dunaeva et al. 1999; Song et al. 2000), salicylic acid (Qin et al. 2009) or abscisic acid (Song et al. 2000). Furthermore, several studies indicate that RIPs protect plants from viral and fungal infection and transgenic plants bearing RIP genes have been constructed that are resistant to virus, fungi and insects (Corrado et al. 2005; Huang et al. 2008; Dowd et al. 2012; Qian et al. 2014). In addition to the defensive role, other functions such as storage proteins, contribution to development or plant senescence have been proposed (Girbes et al. 2004). For example, the role in plant senescence has been attributed to RIPs in Hura crepitans L., Phytolacca americana L. and Triticum aestivum L. (Stirpe et al. 1996; Sawasaki et al. 2008).
RIPs have been extensively researched due to their ability to inhibit protein synthesis and their utility as a toxic moiety for the construction of immunotoxins, conjugates or recombinant chimeras against tumor cells (Ferreras et al. 2011a; Citores et al. 2013).
From a structural point of view, they have been classified into two types: type 1 RIPs (e.g. saporin, PAP, BE27 and PD-L1) consisting of single-chain proteins, and type 2 RIPs (e.g. ricin, abrin, mistletoe lectin) consisting of an A (active) chain with RIP properties covalently linked to a B (binding) chain with lectin properties. The latter RIPs can enter cells more easily because the B chain allows the binding to sugar-containing cell surface receptors, and for this reason they could be potent toxins like ricin, abrin or volkensin (Ferreras et al. 2011a; Citores et al. 2013). Nevertheless, a recent phylogenetic study has revealed the existence of a more complex scenario and three groups of RIPs have been proposed. Group 1: type 1 RIPs from monocots, group 2: type 1 RIPs from dicots and group 3: type 2 RIPs from both monocots and dicots and type 1 RIPs from Euphorbiaceae, Cucurbitaceae, Rosaceae and Iridaceae (Di Maro et al. 2014). Other structures have been found for some RIPs (Girbes et al. 2004; Peumans and Van Damme 2010), however, many authors prefer to maintain the classification of type 1 RIPs, without B chains, and type 2 RIPs, with B chains (Stirpe 2005).
Despite all the work done in the field of RIPs, the mechanism whereby RIPs protect plants from pathogens is not clear. RIPs are protein synthesis inhibitors that irreversibly inactivate mammalian ribosomes (Barbieri et al. 1993; Girbes et al. 2004). They can also inhibit protein synthesis in other animals and yeast (Girbes et al. 2004) and some of them can inactivate plant and bacterial ribosomes although usually these ribosomes are insensitive to the majority of RIPs (Girbes and Ferreras 1998). These proteins are 28S rRNA N-glycosidases (EC 3.2.2.22) that cleave the N-glycosidic bond between the adenine No. 4324 and its ribose in the 60S subunit of rat ribosomes (or the equivalent adenine in sensitive ribosomes from other organisms). This adenine is located in the Sarcin Ricin Loop (SRL) that is crucial for anchoring the elongation factor G or the elongation factor 2 on the ribosome during mRNA–tRNA translocation in prokaryotes and eukaryotes, respectively (Barbieri et al. 1993; Girbes et al. 2004; Puri et al. 2012).
Therefore, the antiviral and antifungal activity was initially attributed to the ability of RIPs to inhibit protein synthesis. Type 1 RIPs (located in the apoplasm) might be part of a general suicide strategy of some higher plants, rendering wounded tissue inefficient for the establishment of an infection by viruses or parasitic fungi (Prestle et al. 1992; Park et al. 2004).
Additionally, some RIPs release adenines from viral genomic RNAs (Barbieri et al. 1994, 1997; Iglesias et al. 2005) and herring sperm DNA (Barbieri et al. 1994, 1997). Thus, Barbieri et al. (1997) have proposed the name of polynucleotide:adenosine glycosidases for these proteins and that extensive deglycosylation of viral RNA and DNA might be responsible for the antiviral action of RIPs.
Nuclease and RNase are enzymatic activities that have also been involved in plant defense (LeBrasseur et al. 2002; van Loon et al. 2006). It has been reported that some RIPs display RNase (Fong et al. 2000) and endonuclease (Aceto et al. 2005) activities. This is a controversial issue because it has been demonstrated that, for some RIPs, this activity is due to RNase or nuclease contaminations (Valbonesi et al. 1999; Barbieri et al. 2000).
On the other hand, some RIPs display superoxide dismutase (SOD) activity (Sharma et al. 2004; Barbieri et al. 2006). SOD activity generates hydrogen peroxide that can be toxic to different types of attackers or could directly or indirectly stimulate plant-defense responses (van Loon et al. 2006).
Cell death induced by RIPs resembles apoptosis instead of necrosis (Das et al. 2012) and initial studies attributed to the rRNA N-glycosidase activity the capacity of RIPs to induce apoptosis. However, the role of depurination activity of the RIPs in apoptosis induction is still controversial and there are three different hypotheses which propose that depurination is essential, essential but not the sole factor, or not essential for apoptosis (Das et al. 2012).
Beetins, BE27 and BE29, are two ribosome-inactivating proteins (RIPs) isolated from sugar beet (Beta vulgaris L.). Both isoforms of beetin represent different levels of glycosylation of the same polypeptide chain (Iglesias et al. 2005). These proteins are synthesized in response to virus infection (Girbes et al. 1996) and are expressed both in infected (hypersensitive response) and non-infected (systemic acquired resistance, SAR) leaves (Iglesias et al. 2005). Furthermore, the signaling compounds hydrogen peroxide and salicylic acid induce the expression of both proteins (Iglesias et al. 2005, 2008). The hypothesis that BE27 is involved in plant defense against viruses is also supported by the fact that the external application of BE27 to sugar leaves prevented infection by artichoke mottled crinkle virus (AMCV) (Iglesias et al. 2005). To gain insights into the protective properties of BE27 against pathogens, we have tested other putative activities of this RIP and we found that BE27 displays rRNA N-glycosidase activity against microbial ribosomes, DNA polynucleotide:adenosine glycosidase activity against eukaryotic DNA, endonuclease activity against supercoiled plasmid DNA and superoxide dismutase activity. BE27 also induces apoptosis in COLO 320 cells by either activating the caspase pathways and/or inhibiting protein synthesis. The combined effect of these biological activities could result in a broad action against several types of pathogens.
Materials and methods
Materials
The sources of the chemicals used in this work have been indicated previously (Ferreras et al. 2011b) and most of them were obtained from Sigma-Aldrich (St Louis, MO, USA). The chromatography media (SP-Sepharose Fast Flow, Superdex 75 HiLoad 26/60 and Red-Sepharose CL-6B) were purchased from GE Healthcare, Spain. Bovine pancreatic ribonuclease A (RNase A), yeast RNA, NBT and WST-1 were purchased from Roche Diagnostics S.L. (Barcelona, Spain). Herring sperm DNA, superoxide dismutase from bovine liver and xanthine oxidase from bovine milk were purchased from Sigma-Aldrich.
Isolation of BE27 from sugar beet leaves
BE27 was isolated from sugar beet (B. vulgaris L.) leaves following a procedure previously described by Iglesias et al. (2005). Briefly, 300 g of sugar beet leaves (collected in September from a crop field at Cobos de Cerrato, Palencia, Spain) were ground in a blender and extracted overnight with 2.4 L of 5 mM sodium phosphate (pH 7.5) buffer containing 140 mM NaCl. The extract was acidified to pH 4 and chromatographed through SP-Sepharose Fast Flow (7 cm height × 5 cm diameter). The bound protein was eluted with 5 mM sodium phosphate (pH 7.5) buffer containing 0.5 M NaCl. The protein solution was further dialyzed and chromatographed through SP-Sepharose Fast Flow (6.5 × 1 cm) using 40 volumes of a linear gradient of 30–200 mM NaCl in 5 mM sodium phosphate (pH 7.5) buffer. The fractions inhibiting protein synthesis were pooled and subjected to chromatography through Superdex 75 HiLoad 26/60 as indicated (Iglesias et al. 2005). The protein peaks showing inhibitory activity on protein synthesis were dialyzed and further chromatographed through Red-Sepharose CL-6B.
Dye-chromatography using Red-Sepharose
Putative nuclease contaminants were removed from BE27 by Red-Sepharose CL-6B chromatography essentially as described before (Valbonesi et al. 1999; Barbieri et al. 2000). Briefly: 0.9 mg of BE27 in 10 mM Tris/HCl, pH 7.8 was applied to a 3.5 × 1 cm column of Red Sepharose CL-6B equilibrated with the same buffer. The unbound protein was washed with buffer and bound protein was eluted with a linear gradient of NaCl (0–1 M) in buffer (total volume 28 mL). Elution of protein was followed by absorbance at 280 nm. Fractions containing BE27 were collected and dialysed extensively against water at 5 °C, lyophilized in aliquots of 25 μg and stored at −20 °C.
rRNA N-glycosidase assay
The depurination assay was conducted as described by Girbes et al. (1993) and Iglesias et al. (2005). Escherichia coli or Agrobacterium tumefaciens ribosomes (200 μg) were incubated with 1 μg of BE27 at 30 °C for 1 h in 50 μL of 40 mM Tris–HCl buffer (pH 7.6) containing 60 mM NH4Cl and 10 mM magnesium acetate. N-glycosidase activity on yeast ribosomes was assayed in 100 μL samples of S-30 lysates from yeast in 10 mM Tris–HCl buffer (pH 7.6) containing 10 mM KCl, 10 mM magnesium acetate and 6 mM 2-mercaptoethanol, which were incubated with 3 μg of BE27 at 30 °C for 1 h. After treatment, the RNA was extracted by phenolization, treated with 1 M aniline acetate (pH 4.5) and precipitated with ethanol. COLO 320 (human colon adenocarcinoma) cells were obtained from the European Collection of Cell Cultures (ECACC) and grown in RPMI 1640 medium (Gibco BRL, Barcelona, Spain) supplemented with 10 % foetal bovine serum under 5 % CO2 at 37 °C. COLO 320 cells (1 × 106/plate) were incubated for 72 h in presence of 1 or 10 μM BE27. After treatment, cells were harvested by centrifugation at 1,000g for 5 min. The pellets were lysed and the RNA was isolated following the instruction of the RNeasy Mini Kit (Qiagen GmbH, Hilden, Germany). RNA was treated with 1 M aniline acetate (pH 4.5) for 10 min at 0 °C and precipitated with ethanol. The RNAs were subjected to electrophoresis at 15 mA for 50 min (bacteria), 1 h 30 min (yeast) or 2 h (COLO 320 cells) in a 7 M urea/5 % (w/v) polyacrylamide gel and stained with ethidium bromide (Sallustio and Stanley 1990).
RNase assay
RNase activity was assayed according to the method by Greiner-Stoeffele et al. (1996). A RNA solution was prepared by dissolving 7 mg yeast RNA in 0.7 mL Mops buffer (0.1 M Mops–HCl, pH 7.5, 2 mM EDTA). Methylene blue buffer was prepared by dissolving 0.1 mg methylene blue in 10 mL Mops buffer. 20 μL RNA solution was mixed with 980 μL methylene blue buffer and 20 μL of either water, BE27 or RNase. The samples were incubated at room temperature and the absorbance decrease at 688 nm was monitored at 15, 30, 45 and 60 min.
Polynucleotide:adenosine glycosidase activity on herring sperm DNA assay
The adenine release was measured according to the method reported by Di Maro et al. (2007) with a few modifications. 10 μg of herring sperm DNA was incubated with 3 μg of RIP in 300 μL of a reaction mixture which contained 100 mM KCl, 50 mM magnesium acetate (pH 4), at 30 °C for 60 min. After incubation, the DNA was precipitated with ethanol at −80 °C for 3 h and centrifugated at 10,000g for 15 min. Adenine released from RIP-treated DNA was determined in the supernatants spectrophotometrically at 260 nm.
DNA cleavage experiments
DNA cleavage experiments were performed as previously reported (Huang et al. 1992). Each reaction contained 1 μg of BE27 and 200 ng of the E. coli plasmid PUC19 (Life Technologies S.A., Madrid, Spain) in a final volume of 10 μL of 10 mM Tris–HCl, 5 mM MgCl2, 50 mM NaCl, 50 mM KCl and 0.1 mM EDTA, pH 7.8. Samples were incubated for 1 h at 37 °C, run on agarose gel (0.8 %) in TAE buffer (0.04 M Tris, 0.04 M acetate, 1 mM EDTA, pH 8.0) and visualized by ethidium bromide staining (0.5 μg/mL). EcoRI linearization was achieved by incubating 250 ng of PUC19 with 1.5 units of EcoRI enzyme according to manufacturer instructions (Roche Diagnostics S.L., Barcelona, Spain).
Superoxide dismutase activity of BE27
Reaction mixtures contained, in a total volume of 1 mL: 0.1 mM xanthine, 0.1 mg NBT (nitro blue tetrazolium), 0.002 units of xanthine oxidase from bovine milk and bovine liver superoxide dismutase as a positive control, or BE27. The reduction of NBT was monitored at A560 for 3 min. The data represent the inhibition of reduction of NBT. Data are the average of six independent experiments in duplicate. Standard deviation never exceeded 8 %.
Cell viability assays
Cell viability was determined by a colorimetric assay based on cleavage of the tetrazolium salt WST-1 to formazan by mitochondrial dehydrogenases in viable cells. COLO 320 cells were seeded in 96-well plates (3 × 103 cells in 0.1 mL) and incubated at 37 °C under 5 % CO2 in the absence or the presence of BE27, as described in the legends to the figures. Next, the cells were incubated for another 2 h with 10 μL/well of the cell proliferation reagent WST-1 at 37 °C under 5 % CO2. Then, the absorbance of the samples was measured using a microtitre plate reader set at 450 nm with 620 nm as reference (ELISA reader Multiskan). The absorbance of wells without cells was subtracted as background.
DNA fragmentation analysis
COLO 320 cells (1 × 106/plate) were incubated for 72 h in presence of BE27 at the concentrations indicated in the text. After treatment, cells were harvested by centrifugation at 1,000g for 5 min. The pellets were lysed and the DNA was isolated following the instruction of the Genomic Prep Cells and Tissue DNA Isolation Kit (GE Healthcare). DNA electrophoresis was carried out in 1.8 % agarose gels using TBE buffer (0.089 M Tris, 0.089 M boric acid, 2 mM EDTA, pH 8.0). DNA was stained with Gel Red (Biotium, Inc., Hayward, CA, USA) and visualized with an ultraviolet lamp.
Caspase-3/7 activity
The caspase-3/7 activity was assessed by the luminescent assay Caspase-GloTM 3/7 (Promega). COLO 320 cells (4 × 103/well) were seeded in 96-well microtiter plates in 80 μL RPMI complete medium and incubated at 37 °C under 5 % CO2 in the absence or the presence of either 1 or 10 μM of BE27. After 48 h of incubation, 70 μL/well of Caspase-Glo™ 3/7 were added. Plates were shaken for 1 min and then incubated for 1 h at room temperature in the dark. The luminescence was measured by SpectroMax L (integration time 10 s) and the values were normalized for the viability.
Molecular modeling of BE27
Complete amino acid sequence of BE27 (accession number AAS67266.1) (Iglesias et al. 2005) was downloaded from the National Center for Biotechnology Information (NCBI) sequence database (http://www.ncbi.nlm.nih.gov/protein/). A three-dimensional structural modeling was carried out on the I-TASSER server (Roy et al. 2010; http://zhanglab.ccmb.med.umich.edu/I-TASSER). Protein structures 2q8w (PAP-S1aci), 3ctk (bouganin), 1qci (PAP), 1wuc (bouganin), 2z4u (PD-L4), 1ift (ricin A-chain) and 1apa (abrin A-chain) were chosen by I-TASSER as the templates in the modeling. Study and graph representations of protein structures were performed with the aid of the Discovery Studio 3.5 suite (http://accelrys.com/).
Other procedures
The relative molecular mass of BE27 was determined using an ESI/Q-TOF mass spectrometer (Waters, Milford, MA, USA) after protein desalting by RP-HPLC as described elsewhere (Di Maro et al. 2009). Preparation of the 30,000 g (S30) supernatants and purified ribosomes from E. coli MRE 600, A. tumefaciens, and yeast were performed as described elsewhere (Girbes et al. 1979; Waters and Blobel 1986; Alegre et al. 1993). Protein synthesis was performed with a coupled transcription–translation in vitro assay using a rabbit reticulocytes lysate system as describe elsewhere (Ferreras et al. 2011b). Protein concentrations were determined using the spectrophotometric method of Kalb and Bernlohr (1977).
Results
rRNA glycosidase activity
BE27 is an rRNA N-glycosidase that splits an adenine from the Sarcin Ricin Loop (SRL) from the mammalian 28S rRNA disabling the ribosomes to bind the Elongation Factor 2 and arresting protein synthesis (Iglesias et al. 2005). Additionally, it acts on the ribosomes from the plant Vicia sativa L. and the enterobacterium E. coli (Iglesias et al. 2005). Because the rRNA N-glycosidase activity might play an antimicrobial role in sugar beet we assayed the effect of BE27 on ribosomes from the parasitic bacterium A. tumefaciens and yeast which might be homologous to several putative plant pathogens. As shown in Fig. 1a–b, BE27 displays rRNA N-glycosidase activity on both types of ribosomes as indicated by the release of the diagnostic fragment (Endo and Tsurugi 1988) upon treatment with acid aniline. The released fragments displayed sizes of 225 ± 5, 245 ± 5 and 360 ± 30 nucleotides for A. tumefaciens, E. coli and yeast, respectively, in accordance with that expected for the SRL deglycosylation (228, 244 and 368 nucleotides for A. tumefaciens, E. coli and yeast, Fig. 1c). Therefore, this RIP might enter into the bacterial and fungal cells and inactivate their ribosomes avoiding the propagation of the pathogen as has been suggested for other RIPs (Park et al. 2004).
rRNA N-glycosidase activity of BE27 on bacterial and yeast ribosomes. a–b N-glycosidase activity was assayed as indicated under "Materials and methods". Each lane contained 3 μg of RNA isolated from either untreated (control) or BE27 treated ribosomes from A. tumefaciens, E. coli (a) or the yeast S. cerevisiae (b). The arrows indicate the RNA fragments released as a consequence of RIP action upon acid aniline treatment (plus). Numbers indicate the size of the standards (M) in nucleotides. c Sarcin Ricin Loop of the large rRNA from A. tumefaciens, E. coli and S. cerevisiae. The sequences (Accession numbers AB102732, AB035926 and J01355) were downloaded from the NCBI sequence database (http://www.ncbi.nlm.nih.gov/nucleotide/). The adenine released by the RIP action (boldfaced), the site of splitting by the acid aniline (arrows) and the size of the generated fragment are also indicated
DNA polynucleotide:adenosine glycosidase activity
Although RIPs were studied for many years as potent inhibitors of protein synthesis, their main enzymatic activity is the depurination of nucleic acids, and for this reason the name of polynucleotide:adenosine glycosidases has been proposed for these proteins. The different RIPs display different abilities to depurinate RNA (Barbieri et al. 1997). BE27 is able to depurinate the RNA from the tobacco mosaic virus (TMV) which might explain its antiviral effect on RNA virus (Iglesias et al. 2005). We have assayed the DNA polynucleotide:adenosine glycosidase activity of BE27 on herring sperm DNA and compared this activity with that of ricin from Ricinus communis L. seeds, which possesses a moderate activity (Barbieri et al. 1997), and PD-L1 from Phytolacca dioica L. leaves, which displays a high activity (Parente et al. 2008; Di Maro et al. 2009; Severino et al. 2010). As shown in Fig. 2, BE27 displays a high DNA N-glycosidase activity on herring sperm DNA, twice and eight-fold higher than PD-L1 and ricin, respectively. This activity might be responsible for the antiviral activity of BE27 against DNA virus.
Polynucleotide:adenosine glycosidase activity of BE27 assayed on herring sperm DNA. Adenine polynucleotide glycosidase activity of 3 μg of either BE27, PD-L1 or ricin was assayed as indicated under “Materials and methods” and the absorbance of the adenine released was measured at 260 nm. The data represent the mean of two duplicate experiments and the bars indicate the standard error of the mean
Ribonuclease activity
It has been published that the RIPs alpha- and beta-momorcharins from Momordica charantia L. seeds possess a weak ribonuclease activity, 100,000- and 6,500-fold lower than the RNase-MC isolated from the same seeds (Fong et al. 2000). Nevertheless, this activity might be responsible for the antiviral and protein synthesis inhibitory activities of these RIPs (Fong et al. 2000). Due to this we investigated the putative RNase activity of BE27 and found that this protein does not display a detectable RNase activity on yeast RNA even at a concentration of 5,200 ng/mL while such activity was apparent for bovine pancreatic ribonuclease A (RNase A) at a concentration of 8 ng/mL, and even slightly detectable at a concentration of 0.08 ng/mL, 65,000-fold lower than that used for BE27 (data not shown). Therefore, it seems that this RIP does not possess RNase activity and that its sole activity on RNA is the reported polynucleotide:adenosine glycosidase (Iglesias et al. 2005).
Endonuclease activity on supercoiled plasmid DNA
It has been reported that some RIPs display endonuclease activity on supercoiled plasmid DNA producing relaxed or even linear plasmids (Aceto et al. 2005; de Virgilio et al. 2010). This capability may be necessary for these proteins to perform different biological roles including resistance to pathogenic micro-organisms or viruses (Aceto et al. 2005; de Virgilio et al. 2010). Therefore, we have tested the endonuclease activity of BE27 on the plasmid PUC19 and found that BE27 promoted the conversion of supercoiled PUC19 DNA into the relaxed and, in less extension, linear forms. This effect was dependent of magnesium ions as reported for PD-L1 (Aceto et al. 2005) and the highest activity was observed at 10 mM of such an ion (data not shown).
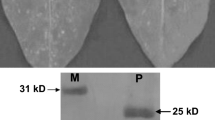
To exclude that the endonuclease activity observed was due to nuclease contamination, BE27 was subjected to Red Sepharose chromatography (Fig. 3a). Most of the protein eluted at a NaCl concentration between 0.36 and 0.54 M. These fractions were pooled and used to assay the nuclease activity. As shown in Fig. 3b mass spectrometry indicated that BE27 is the only protein present in such a preparation with a relative molecular mass of 27,589 corresponding to the molecular mass deduced from its amino acid sequence. Red-Sepharose purified BE27 displays the same protein synthesis inhibiting activity as BE27 prior to purification, both preparations having an IC50 (protein concentration inhibiting 50 % of protein synthesis) of 0.85 ng/mL (31 pM) on a rabbit reticulocytes lysate protein synthesis system (Fig. 3c).
Dye-chromatography on Red-Sepharose of BE27. a 0.9 mg of BE27 was loaded to a Red-Sepharose CL-6B column and eluted with a linear gradient of NaCl (dashed line) as indicated under “Materials and methods”. Shading indicates pooled purified fractions. b Molecular mass of BE27. The relative molecular mass of BE27 was determined using an ESI/Q-TOF mass spectrometer as indicated under “Materials and methods”. c Effect of BE27 either purified (open circle) or non-purified (filled circle) by Red-Sepharose chromatography on protein synthesis. Translation assays were carried out using rabbit reticulocytes lysate as a cell-free system, as indicated in “Materials and methods”. Data represent the percentage of protein synthesis with respect to a control without BE27
Red-Sepharose purified BE27 retained the nicking activity on the supercoiled PUC19 (Fig. 4a). Such activity was blocked by the addition of EDTA, thus indicating that the observed change documented in Fig. 4a was not due to nuclease contamination. It has been reported that high concentrations of some RIPs may alter the electrophoretic mobility of both linear and supercoiled DNA due to a massive binding of protein to the DNA molecules (Barbieri et al. 2000). As shown in Fig. 4b, BE27 does not change the mobility of the linear PUC19 in the conditions used in these assays thus demonstrating that the observed change in Fig. 4a was consequence of an endonuclease activity. On the other hand, little or no linear PUC19 was observed upon treatment with BE27 in our assays, thus indicating that the catalytic action primarily affects one DNA strand.
Nicking activity of BE27 on PUC19 DNA. a 200 ng samples of plasmid DNA were incubated in 10 μL with 1 μg BE27 in the absence or the presence of 25 mM EDTA (plus) as indicated under “Materials and methods”. b PUC19 DNA was previously linearized using EcoRI. R, L, and S indicate relaxed, linear and supercoiled forms of PUC19, respectively and the numbers, the size of the standards (M) in nucleotides
Structure of BE27
Another point that has sparked interest is that not all RIPs possess endonuclease activity. For example, PD-L1 and PD-L4 exhibit very different properties even if they share 82 % amino acid identity and only PD-L1 displays endonuclease activity (Ruggiero et al. 2009). Two differences have been reported concerning the structure of these proteins that may have implications in their enzymatic activities. The first difference is that they exhibit a different distribution of charged residues on their surfaces, in particular, a more pronounced negatively charged patch is found close to the active site cleft of PD-L1, compared to PD-L4 (Ruggiero et al. 2009). These differences may contribute to the different activity/specificity of the two proteins. The second difference is a conformational variation of the loop lining the catalytic site cleft and containing Asp91, a residue which is important for catalysis. This conformational change, induced by the presence of Arg97 (serine in PD-L4) opens the active site cleft allowing the accommodation of supercoiled DNA that has to be cleaved (Ruggiero et al. 2009).
To ascertain the main structural characteristics that may be involved in the different behavior of BE27 with respect to PD-L1 and PD-L4, a 3D-structure was predicted by comparative modeling using several RIP crystal structures as templates. Figure 5 shows such model compared with the X-ray crystal structures of PD-L1 and PD-L4 (Ruggiero et al. 2009). The three RIPs display the characteristic fold of the type 1 RIPs and the A chain of type 2 RIPs in which the confluence of three domains create the active site: the N-terminal domain 1, consisting of β-strands and α-helices; the domain 2, characterized by α-helices; and the C-terminal domain 3, formed by two α-helices and two β-strands (Tahirov et al. 1995; Di Maro et al. 2014). Because of their high amino acid identity values, PD-L1 and PD-L4 share a strong structural similarity (Fig. 5a) displaying a low main chain root mean square deviation (rmsd) of 0.86 Å that contrasts with higher values for the pairs PD-L1/BE27 and PD-L4/BE27 (1.87 and 1.74, respectively). BE27 shares a 40 % sequence identity with PD-L1 and 42 % with PD-L4. Although the models obtained by comparative modeling have a limited value and they should be confirmed by X-ray diffraction studies there are structural characteristics in BE27 that are very evident. The first is that BE27, in contrast with PD-L1, does not display a negatively charged patch close to the active site cleft (Fig. 5b–d). However, it is true that BE27 presents a conformation of the loop lining the catalytic site cleft (from amino acid 84 to amino acid 122) similar to that of PD-L1 (Fig. 5e–h).
Structure of BE27 compared with PD-L1 and PD-L4. The three-dimensional structural modeling was carried out on the I-TASSER server and the figure was generated with DS Visualizer 3.5. a Superposition of α-carbon traces of BE27 (red), PD-L1 (Accession number 3H5K) (blue) and PD-L4 (Accession number 2QES) (green). The balls represent the conserved amino acids in the active site to indicate its position. b–d Electrostatic potential surfaces of BE27 (b), PD-L1 (c) and PD-L4 (d). The circle in c indicates a more pronounced negatively charged patch close to the active site cleft in PD-L1. e Structure of BE27 indicating the position of the loop from amino acid 84 to amino acid 122 (red ribbon). f–h Comparison between the loop from BE27 expanding from amino acid 84 to amino acid 122 (f) and their homologs from PD-L1 (g) and PD-L4 (h). Key amino acids are represented by scaled balls and sticks
In PD-L4, Asp91 is hydrogen bonded to Ser120, a residue involved in adenine binding. This interaction is not conserved in the structure of PD-L1 in which the entire loop is pulled back due to interactions of the backbone atoms of Phe89 and Ile92 with the side chain of Arg97, a serine in PD-L4. The consequence of this conformational change is the opening of the active site cavity. As shown in Fig. 5f the loop in BE27, similarly to PD-L1 (Fig. 5g), is pulled back due to a very much shorter alpha-helix. This catalytic cleft opening may allow the binding of regions of the to-be-cleaved supercoiled DNA whose binding is hampered by the obstructing loop in PD-L4. This loop conformation is also observed in the structure of the pokeweed antiviral protein (PAP), which is able to induce DNA cleavage. In addition, other RIPs exhibiting DNA cleaving activity, like Saporin 6 and Dianthin 30, are characterized by a two-residue shorter loop (Ruggiero et al. 2009).
Apoptotic activity
Although initially the toxicity of RIPs to animal cells was attributed to their ability to inhibit protein synthesis, there is an increasing body of evidence indicating that the main cause of RIP toxicity is their ability to induce apoptosis (Das et al. 2012). Because of this we studied the capacity of BE27 to induce apoptosis in a sensitive cell line. As shown in Fig. 6a, BE27 was toxic to COLO 320 cells in a concentration- and time-dependent manner with IC50s of 2.2 × 10−6 and 4 × 10−7 M after 48 and 72 h exposure, respectively. As expected ricin was cytotoxic at lower concentrations (IC50 of 1.4 × 10−13 M) due to the presence of its B chain, a lectin that allows the protein to go through a more efficient intracellular delivery pathway (Citores et al. 2013). BE27-treated cells exhibited the morphological features characteristic of apoptosis (data not shown) and stimulated the apoptotic pathway(s) as demonstrated by the strong dose-dependent activation of the effector caspase 3/7 (Fig. 6b) and the breakdown of the nuclear DNA into oligonucleosomal fragments in COLO 320 cells (Fig. 6c). Analysis of the ribosomal RNA from BE27-treated cells showed that the ribosomes were depurinated releasing the diagnostic fragment (Fig. 6d), thus indicating that apoptosis induction might be a consequence of protein synthesis inhibition (Das et al. 2012).
Cytotoxicity of BE27 on COLO 320. a Effect of BE27 and ricin on viability of COLO 320 cells. Cells were incubated with different concentrations of ricin (filled triangle) and BE27 (filled circle) for 48 h or BE27 (filled square) for 72 h and cell viability was evaluated by a colorimetric assay as indicated under “Materials and methods”. Data represent the mean ± SD of three experiments performed in triplicate. b Caspase-3/7 activation in COLO 320 cells treated with either 1 or 10 μM of BE27 for 48 h. Activity is expressed as the percentage of control values obtained from cultures grown in the absence of RIPs. Data represent the mean ± SD of two experiments performed in duplicate. c Effect of BE27 on internucleosomal DNA fragmentation. COLO 320 cells were incubated in the absence or presence of 1 and 10 μM of BE27 for 72 h. Then, the DNA was isolated and 4 μg was electrophoresed as indicated in “Materials and methods”. The numbers indicate the corresponding size of the standards (λDNA HindIII/EcoRI) (M) in kb. d rRNA N-glycosidase activity of BE27 on RNA from COLO 320 cells. rRNA N-glycosidase activity was assayed as indicated in "Materials and methods." Each lane contained 2 μg of RNA isolated from either untreated cells or cells incubated with either 1 or 10 μM of BE27 for 72 h. The arrow indicates the RNA fragment released as a consequence of RIP action upon acid aniline treatment (plus). The numbers on the left indicate the corresponding size of the RNA standards (M) in nucleotides
Superoxide dismutase activity
Because superoxide dismutase (SOD) activity stimulate plant-defense responses (van Loon et al. 2006) conferring resistance to viruses (Hao et al. 2011) and fungus (Tertivanidis et al. 2004) and some RIPs have been reported to possess such activity (Sharma et al. 2004; Barbieri et al. 2006) we assayed the putative SOD activity of BE27 (Table 1). The SOD activity of BE27 was checked by an assay consisting of the reduction of NBT by xanthine oxidase and subsequent inhibition of reduction by SOD. We used bovine liver superoxide dismutase as a positive control and SOD activity of BE27 was compared. Two concentrations of BE27 were tested for the ability to inhibit NBT reduction. In our hands, BE27 caused 22 and 26 % inhibition of NBT reduction at 250 and 500 ng/mL respectively, thus indicating that BE27 possesses a considerable SOD activity although smaller than that reported for TRIP isolated from tobacco leaves (Sharma et al. 2004) and the Cucurbita moschata Duchesne RIP (Barbieri et al. 2006).
Discussion
Beetin 27 (BE27) is a defense protein that protects beet leaves against the infection of the ssRNA artichoke mottle crinkle virus (AMCV) (Iglesias et al. 2005) and is expressed in response to virus infection (Girbes et al. 1996) or the signaling compounds hydrogen peroxide and salicylic acid (Iglesias et al. 2005, 2008). This role as a defense protein has been attributed to its RNA polynucleotide:adenosine glycosidase activity on viral RNA since it depurinates the tobacco mosaic virus RNA while beet ribosomes are not sensitive to BE27 (Iglesias et al. 2008). This contrasts with other RIPs such as the pokeweed antiviral protein (PAP) from P. americana L. leaves that inactivates their ‘conspecific’ ribosomes (Prestle et al. 1992) but does not depurinate RNAs from virus (Barbieri et al. 1997). PAP accumulates in the apoplastic space and has been postulated as a part of a general suicide strategy of some higher plants, rendering wounded tissue inefficient for the establishment of an infection by viruses or parasitic fungi (Prestle et al. 1992; Park et al. 2004). Although earlier reports indicated that RIPs were inactive against bacterial ribosomes (Girbes and Ferreras 1998), systematic studies using 31 RIPs and 4 bacteria (Girbes et al. 1993; Ferreras et al. 1994, 1995) revealed that 3 RIPs (crotins 2 and 3 from Croton tiglium L. and momordin I from M. charantia L.) inhibited protein synthesis in all the bacteria tested, 5 RIPs inhibited protein synthesis but not in all the bacteria and 23 RIPs did not inhibit protein synthesis in bacteria (Girbes and Ferreras 1998). The mechanism of inhibition was the release of the above-mentioned adenine in the 23S rRNA SRL (Girbes et al. 1993; Ferreras et al. 1994, 1995). When the gene coding for BE27 was expressed in E. coli the growth of the transformed bacteria was arrested due to the strong activity of BE27 against bacterial ribosomes since the treatment of the corresponding rRNA with acid aniline promoted the release of the RIP diagnostic RNA fragment (Iglesias et al. 2005). This, together with the fact that most RIPs inhibit protein synthesis in yeast might indicate that some RIPs could play an antimicrobial role. BE27 displays rRNA N-glycosidase activity on both bacterial and fungal ribosomes as indicated by the release of the diagnostic fragment (Endo and Tsurugi 1988) upon treatment with acid aniline. Therefore, this RIP might, assisted by other hydrolytic defence proteins, enter into the bacterial and fungal cells and inactivate their ribosomes avoiding the propagation of the pathogen.
The different RIPs display different abilities to depurinate RNA. There are RIPs that possess much more activity than others and even RIPs lacking this activity (Barbieri et al. 1997). However, all the RIPs are able to depurinate herring sperm DNA although this activity may vary by more than 100-fold in some RIPs compared to others (Barbieri et al. 1997). BE27 displays a high DNA polynucleotide:adenosine glycosidase activity on herring sperm DNA, two times and eightfold higher than PD-L1 and ricin, respectively. This activity may be responsible for antiviral activity against DNA virus and antimicrobial activity. However, despite that all the RIPs are active against nucleosomal DNA most of them are inactive against bacterial DNA (Barbieri et al. 2000; Ruggiero et al. 2009). RIP activities other than the polynucleotide:adenosine glycosidase have been the subject of discussion by several authors (Peumans et al. 2001; Robertus and Monzingo 2004). Although some RIPs are active against supercoiled plasmid DNA, it has been reported that this activity is a consequence of contamination by nucleases in the protein preparation (Barbieri et al. 2000). BE27 is active against supercoiled plasmid DNA, a property only shared by some RIPs from P. dioica, P. americana, Saponaria officinalis and Dianthus caryophyllus (Roncuzzi and Gasperi-Campani 1996; Ruggiero et al. 2009). This activity may be the consequence of a particular configuration in the structure surrounding the active site. In particular, we have investigated the loop region between the amino acid residues 84 and 122, homologous to the amino acid residues 82 and 126 in PD-L1 and PD-L4 (Ruggiero et al. 2009). This in silico analysis revealed that structural variation present in this loop region is important for endonuclease activity. Comparison of the loop conformation shows an open cleft in both BE27 and PD-L1 that is not present in PD-L4. On the other hand, BE27 does not display a negatively charged patch in this open cleft as PD-L1 does. However, the open cleft is probably more important for the nicking activity on supercoiled DNA than charge arrangements of amino acid residues. Differences in structural features in this region may contribute to the diverse activity/specificity of ribosome inactivating proteins for endonuclease activity. Two mechanisms have been suggested for the endonuclease activity displayed by some RIPs. A bifunctional role as DNA glycosylase/AP lyases, similar to that displayed by DNA repair enzymes, which produce deadenylation followed by cleavage of apurinic DNA was proposed for gelonin (Nicolas et al. 2000). A second mechanism, spontaneous breakage of phosphodiester bonds after the removal of adenines was proposed for PD-L1, taking into account that phosphodiester bonds in extensively deadenylated regions of supercoiled DNA likely become liable due to the existence of tension in supercoiled DNA (Ruggiero et al. 2009). In this framework, DNA cleavage is a consequence of PD-L1 catalytic action, although not directly catalyzed by the enzyme. In any case, this activity is equivalent to that of nucleases that have been reported as defense proteins (LeBrasseur et al. 2002; van Loon et al. 2006).
It is worthy of mentioning that the classification of type 1 RIPs is too simplistic in saying that they are the same kind of proteins as they may be very different enzymes. In fact type 1 RIPs have been recently classified from a phylogenetic point of view as belonging to three different groups (Di Maro et al. 2014), so they might play different roles. BE27 and PD-L1 cluster in the same group indicating that they may play the same or a similar role (Di Maro et al. 2014).
On the other hand, BE27 like some RIPs displays superoxide dismutase (SOD) activity. SOD activity generates hydrogen peroxide that can be toxic to different types of attackers or could directly or indirectly stimulate plant-defense responses (van Loon et al. 2006). In this respect, it has been reported that SOD and other antioxidant enzymes from rice coordinately participate in the resistance to rice stripe virus (Hao et al. 2011) and that SOD transgenes confer resistance to the fungus Cercospora beticola in sugar beet (Tertivanidis et al. 2004). Additionally, plant tolerance to abiotic stresses may be improved by higher levels of SOD (Gill and Tuteja 2010), thus, the SOD activity of BE27 might be related not only to biotic but also to abiotic stresses.
Although the N-glycosidase activity of RIPs against mammalian, insect, fungal or plant cells has been well-studied, the mechanism of cell death induced by RIPs in these particular cells is not well-understood. It has been shown that cell death induced by RIPs on cells resembles apoptosis instead of necrosis (Das et al. 2012) and initial studies attributed to the rRNA N-glycosidase activity the capacity of RIPs to induce apoptosis. RIPs have been shown to be toxic for a broad spectrum of insects (Dowd et al. 2012; Shahidi-Noghabi et al. 2010) most probably because they induce apoptosis in their gut cells (Shahidi-Noghabi et al. 2010, 2011). BE27 enters COLO 320 cells resulting in protein synthesis inhibition, caspase activation and the induction of apoptosis. Since BE27 displays rRNA N-glycosidase activity on those cells at the same concentrations which activates the effector caspases, it may be assumed that apoptosis is a consequence of protein synthesis inhibition. As has been reported for other RIPs, in addition to the inhibition of protein synthesis BE27 may induce apoptosis via different pathways that could involve interactions with anti-oxidant proteins or the production of reactive oxygen species (Das et al. 2012). BE27 might also induce direct DNA damage if it could reach the nucleus as has been shown for the type 1 RIP saporin. After intoxication of HeLa cells with saporin-S6, DNA gaps resulting from abasic sites and saporin-S6 are found in HeLa nuclei (Bolognesi et al. 2012).
In conclusion, BE27 might favor the resistance of beet against a broad variety of pathogens due to having several enzymatic activities. Further work with different pathogens in vivo and in vitro will be conducted to study the conditions in which the different activities could be more prevalent with respect to the others, since it is probable that the different enzymatic activities have a different significance when the invader pathogen is a virus, a bacterium, a fungus or an insect.
Abbreviations
- NBT:
-
4-Nitro blue tetrazolium
- SOD:
-
Superoxide dismutase
- SRL:
-
Sarcin ricin loop
References
Aceto S, Di Maro A, Conforto B, Siniscalco GG, Parente A, Delli BP, Gaudio L (2005) Nicking activity on pBR322 DNA of ribosome inactivating proteins from Phytolacca dioica L. leaves. Biol Chem 386:307–317
Alegre C, Iglesias R, Ferreras JM, Carbajales ML, Girbes T (1993) Preparation, optimization and characterization of a polyphenylalanine synthetizing system from Agrobacterium tumefaciens. Cell Mol Biol 39:575–581
Barbieri L, Battelli MG, Stirpe F (1993) Ribosome-inactivating proteins from plants. Biochim Biophys Acta 1154:237–282
Barbieri L, Gorini P, Valbonesi P, Castiglioni P, Stirpe F (1994) Unexpected activity of saporins. Nature 372:624
Barbieri L, Valbonesi P, Bonora E, Gorini P, Bolognesi A, Stirpe F (1997) Polynucleotide:adenosine glycosidase activity of ribosome-inactivating proteins: effect on DNA, RNA and poly(A). Nucleic Acids Res 25:518–522
Barbieri L, Valbonesi P, Righi F, Zuccheri G, Monti F, Gorini P, Samori B, Stirpe F (2000) Polynucleotide:adenosine glycosidase is the sole activity of ribosome-inactivating proteins on DNA. J Biochem 128:883–889
Barbieri L, Polito L, Bolognesi A, Ciani M, Pelosi E, Farini V, Jha AK, Sharma N, Vivanco JM, Chambery A, Parente A, Stirpe F (2006) Ribosome-inactivating proteins in edible plants and purification and characterization of a new ribosome-inactivating protein from Cucurbita moschata. Biochim Biophys Acta 1760:783–792
Bolognesi A, Polito L, Scicchitano V, Orrico C, Pasquinelli G, Musiani S, Santi S, Riccio M, Bortolotti M, Battelli MG (2012) Endocytosis and intracellular localisation of type 1 ribosome-inactivating protein saporin-s6. J Biol Regul Homeost Agents 26:97–109
Citores L, Iglesias R, Ferreras JM (2013) Ribosome inactivating proteins from plants: biological properties and their use in experimental therapy. In: Fang EF, Ng TB (eds) Antitumor potential and other emerging medicinal properties of natural compounds. Springer, Dordrecht, pp 127–143
Corrado G, Bovi PD, Ciliento R, Gaudio L, Di Maro A, Aceto S, Lorito M, Rao R (2005) Inducible expression of a Phytolacca heterotepala ribosome-inactivating protein leads to enhanced resistance against major fungal pathogens in tobacco. Phytopathology 95:206–215
Das MK, Sharma RS, Mishra V (2012) Induction of apoptosis by ribosome inactivating proteins: importance of N-glycosidase activity. Appl Biochem Biotechnol 166:1552–1561
de Virgilio M, Lombardi A, Caliandro R, Fabbrini MS (2010) Ribosome-inactivating proteins: from plant defense to tumor attack. Toxins 2:2699–2737
Di Maro A, Chambery A, Daniele A, Casoria P, Parente A (2007) Isolation and characterization of heterotepalins, type 1 ribosome-inactivating proteins from Phytolacca heterotepala leaves. Phytochemistry 68:767–776
Di Maro A, Chambery A, Carafa V, Costantini S, Colonna G, Parente A (2009) Structural characterization and comparative modeling of PD-Ls 1–3, type 1 ribosome-inactivating proteins from summer leaves of Phytolacca dioica L. Biochimie 91:352–363
Di Maro A, Citores L, Russo R, Iglesias R, Ferreras JM (2014) Sequence comparison and phylogenetic analysis by the maximum likelihood method of ribosome-inactivating proteins from angiosperms. Plant Mol Biol 85:577–588
Dowd PF, Johnson ET, Price NP (2012) Enhanced pest resistance of maize leaves expressing monocot crop plant-derived ribosome-inactivating protein and agglutinin. J Agric Food Chem 60:10768–10775
Dunaeva M, Goebel C, Wasternack C, Parthier B, Goerschen E (1999) The jasmonate-induced 60 kDa protein of barley exhibits N-glycosidase activity in vivo. FEBS Lett 452:263–266
Endo Y, Tsurugi K (1988) The RNA N-glycosidase activity of ricin A-chain. The characteristics of the enzymatic activity of ricin A-chain with ribosomes and with rRNA. J Biol Chem 263:8735–8739
Engelberth J, Contreras CF, Viswanathan S (2012) Transcriptional analysis of distant signaling induced by insect elicitors and mechanical wounding in Zea mays. PLoS ONE 7:e34855
Ferreras JM, Alegre C, Iglesias R, Girbes T (1994) Sensitivity of translation by Brevibacterium lactofermentum ribosomes to type 1 and type 2 ribosome-inactivating proteins. Biosci Biotechnol Biochem 58:1458–1462
Ferreras JM, Iglesias R, Barbieri L, Alegre C, Bolognesi A, Rojo MA, Carbajales ML, Escarmis C, Girbes T (1995) Effects and molecular action of ribosome-inactivating proteins on ribosomes from Streptomyces lividans. Biochim Biophys Acta 1243:85–93
Ferreras JM, Citores L, Iglesias R, Jimenez P, Girbes T (2011a) Use of ribosome-inactivating proteins from Sambucus for the construction of immunotoxins and conjugates for cancer therapy. Toxins 3:420–441
Ferreras JM, Citores L, Iglesias R, Souza AM, Jimenez P, Gayoso MJ, Girbes T (2011b) Occurrence of the type two ribosome-inactivating protein nigrin b in elderberry (Sambucus nigra L.) bark. Food Res Int 44:2798–2805
Fong WP, Mock WY, Ng TB (2000) Intrinsic ribonuclease activities in ribonuclease and ribosome-inactivating proteins from the seeds of bitter gourd. Int J Biochem Cell Biol 32:571–577
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Girbes T, Ferreras JM (1998) Ribosome-inactivating proteins from plants. Recent Res Devel Agric Biol Chem 2:1–16
Girbes T, Cabrer B, Modolell J (1979) Preparation and assay of purified Escherichia coli polysomes devoid of free ribosomal subunits and endogenous GTPase activities. Methods Enzymol 59:353–362
Girbes T, Barbieri L, Ferreras M, Arias FJ, Rojo MA, Iglesias R, Alegre C, Escarmis C, Stirpe F (1993) Effects of ribosome-inactivating proteins on Escherichia coli and Agrobacterium tumefaciens translation systems. J Bacteriol 175:6721–6724
Girbes T, de Torre C, Iglesias R, Ferreras JM, Mendez E (1996) RIP for viruses. Nature 379:777–778
Girbes T, Ferreras JM, Arias FJ, Stirpe F (2004) Description, distribution, activity and phylogenetic relationship of ribosome-inactivating proteins in plants, fungi and bacteria. Mini Rev Med Chem 4:461–476
Greiner-Stoeffele T, Grunow M, Hahn U (1996) A general ribonuclease assay using methylene blue. Anal Biochem 240:24–28
Hao Z, Wang L, He Y, Liang J, Tao R (2011) Expression of defense genes and activities of antioxidant enzymes in rice resistance to rice stripe virus and small brown planthopper. Plant Physiol Biochem 49:744–751
Huang PL, Chen HC, Kung HF, Huang PL, Huang P, Huang HI, Lee-Huang S (1992) Anti-HIV plant proteins catalyze topological changes of DNA into inactive forms. BioFactors 4:37–41
Huang M, Hou P, Wei Q, Xu Y, Chen A (2008) A ribosome-inactivating protein (curcin 2) induced from Jatropha curcas can reduce viral and fungal infection in transgenic tobacco. Plant Growth Regul 54:115–123
Iglesias R, Perez Y, de Torre C, Ferreras JM, Antolin P, Jimenez P, Rojo MA, Mendez E, Girbes T (2005) Molecular characterization and systemic induction of single-chain ribosome-inactivating proteins (RIPs) in sugar beet (Beta vulgaris) leaves. J Exp Bot 56:1675–1684
Iglesias R, Perez Y, Citores L, Ferreras JM, Mendez E, Girbes T (2008) Elicitor-dependent expression of the ribosome-inactivating protein beetin is developmentally regulated. J Exp Bot 59:1215–1223
Kalb VF Jr, Bernlohr RW (1977) A new spectrophotometric assay for protein in cell extracts. Anal Biochem 82:362–371
LeBrasseur ND, MacIntosh GC, Perez-Amador MA, Saitoh M, Green PJ (2002) Local and systemic wound-induction of RNase and nuclease activities in Arabidopsis: RNS1 as a marker for a JA-independent systemic signaling pathway. Plant J 29:393–403
Nicolas E, Beggs JM, Taraschi TF (2000) Gelonin is an unusual DNA glycosylase that removes adenine from single-stranded DNA, normal base pairs and mismatches. J Biol Chem 275:31399–31406
Parente A, Conforto B, Di Maro A, Chambery A, De LP, Bolognesi A, Iriti M, Faoro F (2008) Type 1 ribosome-inactivating proteins from Phytolacca dioica L. leaves: differential seasonal and age expression, and cellular localization. Planta 228:963–975
Park SW, Vepachedu R, Sharma N, Vivanco JM (2004) Ribosome-inactivating proteins in plant biology. Planta 219:1093–1096
Peumans WJ, Van Damme EJM (2010) Evolution of plant ribosome-inactivating proteins. In: Lord JM, Hartley MR (eds) Toxic plant proteins—series Plant cell monographs, vol 18. Springer, Heidelberg, pp 1–26
Peumans WJ, Hao O, Van Damme EJ (2001) Ribosome inactivating proteins from plants: more than RNA N-glycosidases? FASEB J 15:11493–11506
Prestle J, Schonfelder M, Adam G, Mundry KW (1992) Type 1 ribosome-inactivating proteins depurinate plant 25S rRNA without species specificity. Nucleic Acids Res 20:3179–3182
Puri M, Kaur I, Perugini MA, Gupta RC (2012) Ribosome-inactivating proteins: current status and biomedical applications. Drug Discov Today 17:774–783
Qian Q, Huang L, Yi R, Wang S, Ding Y (2014) Enhanced resistance to blast fungus in rice (Oryza sativa L.) by expressing the ribosome-inactivating protein alpha-momorcharin. Plant Sci 217–218:1–7
Qin X, Zheng X, Shao C, Gao J, Jiang L, Zhu X, Yan F, Tang L, Xu Y, Chen F (2009) Stress-induced curcin-L promoter in leaves of Jatropha curcas L. and characterization in transgenic tobacco. Planta 230:387–395
Rippmann JF, Michalowski CB, Nelson DE, Bohnert HJ (1997) Induction of a ribosome-inactivating protein upon environmental stress. Plant Mol Biol 35:701–709
Robertus JD, Monzingo AF (2004) The structure of ribosome inactivating proteins. Mini Rev Med Chem 4:477–486
Roncuzzi L, Gasperi-Campani A (1996) DNA-nuclease activity of the single-chain ribosome-inactivating proteins dianthin 30, saporin 6 and gelonin. FEBS Lett 392:16–20
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5:725–738
Ruggiero A, Di Maro A, Severino V, Chambery A, Berisio R (2009) Crystal structure of PD-L1, a ribosome inactivating protein from Phytolacca dioica L. leaves with the property to induce DNA cleavage. Biopolymers 91:1135–1142
Sallustio S, Stanley P (1990) Isolation of Chinese hamster ovary ribosomal mutants differentially resistant to ricin, abrin, and modeccin. J Biol Chem 265:582–588
Sawasaki T, Nishihara M, Endo Y (2008) RIP and RALyase cleave the sarcin/ricin domain, a critical domain for ribosome function, during senescence of wheat coleoptiles. Biochem Biophys Res Commun 370:561–565
Severino V, Chambery A, Di Maro A, Marasco D, Ruggiero A, Berisio R, Giansanti F, Ippoliti R, Parente A (2010) The role of the glycan moiety on the structure-function relationships of PD-L1, type 1 ribosome-inactivating protein from P. dioica leaves. Mol BioSyst 6:570–579
Shahidi-Noghabi S, Van Damme EJ, Mahdian K, Smagghe G (2010) Entomotoxic action of Sambucus nigra agglutinin I in Acyrthosiphon pisum aphids and Spodoptera exigua caterpillars through caspase-3-like-dependent apoptosis. Arch Insect Biochem Physiol 75:207–220
Shahidi-Noghabi S, Van Damme EJ, De Vos WH, Smagghe G (2011) Internalization of Sambucus nigra agglutinins I and II in insect midgut CF-203 cells. Arch Insect Biochem Physiol 76:211–222
Sharma N, Park SW, Vepachedu R, Barbieri L, Ciani M, Stirpe F, Savary BJ, Vivanco JM (2004) Isolation and characterization of an RIP (ribosome-inactivating protein)-like protein from tobacco with dual enzymatic activity. Plant Physiol 134:171–181
Song SK, Choi Y, Moon YH, Kim SG, Choi YD, Lee JS (2000) Systemic induction of a Phytolacca insularis antiviral protein gene by mechanical wounding, jasmonic acid, and abscisic acid. Plant Mol Biol 43:439–450
Stirpe F (2005) Ribosome-inactivating proteins. In: Wiley RG, Lappi DA (eds) Molecular neurosurgery with targeted toxins. Humana Press Inc., Totowa, pp 9–29
Stirpe F, Barbieri L, Gorini P, Valbonesi P, Bolognesi A, Polito L (1996) Activities associated with the presence of ribosome-inactivating proteins increase in senescent and stressed leaves. FEBS Lett 382:309–312
Tahirov TH, Lu TH, Liaw YC, Chen YL, Lin JY (1995) Crystal structure of abrin-a at 2.14 A. J Mol Biol 250:354–367
Tan YC, Yeoh KA, Wong MY, Ho CL (2013) Expression profiles of putative defence-related proteins in oil palm (Elaeis guineensis) colonized by Ganoderma boninense. J Plant Physiol 170:1455–1460
Tertivanidis K, Goudoula C, Vasilikiotis C, Hassiotou E, Perl-Treves R, Tsaftaris A (2004) Superoxide dismutase transgenes in sugarbeets confer resistance to oxidative agents and the fungus C. beticola. Transgenic Res 13:225–233
Valbonesi P, Barbieri L, Bolognesi A, Bonora E, Polito L, Stirpe F (1999) Preparation of highly purified momordin II without ribonuclease activity. Life Sci 65:1485–1491
van Loon LC, Rep M, Pieterse CM (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Waters MG, Blobel G (1986) Secretory protein translocation in a yeast cell-free system can occur posttranslationally and requires ATP hydrolysis. J Cell Biol 102:1543–1550
Acknowledgments
This work was supported by Grants FISPI04/1279 to J. M. F., BIO39/VA42/10 to L. C. and funds from the Second University of Naples. We thank Judy Callaghan (Monash University, Melbourne, Australia) for correcting the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Iglesias, R., Citores, L., Di Maro, A. et al. Biological activities of the antiviral protein BE27 from sugar beet (Beta vulgaris L.). Planta 241, 421–433 (2015). https://doi.org/10.1007/s00425-014-2191-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-014-2191-2