Abstract
Cadmium (Cd) detoxification involves glutathione and phytochelatins biosynthesis: the higher need of nitrogen should require increased nitrate (NO3 −) uptake and metabolism. We investigated inducible high-affinity NO3 − uptake across the plasma membrane (PM) in maize seedlings roots upon short exposure (10 min to 24 h) to low Cd concentrations (0, 1 or 10 μM): the activity and gene transcript abundance of high-affinity NO3 − transporters, NO3 − reductases and PM H+-ATPases were analyzed. Exposure to 1 mM NO3 − led to a peak in high-affinity (0.2 mM) NO3 − uptake rate (induction), which was markedly lowered in Cd-treated roots. Plasma membrane H+-ATPase activity was also strongly limited, while internal NO3 − accumulation and NO3 − reductase activity in extracts of Cd treated roots were only slightly lowered. Kinetics of high- and low-affinity NO3 − uptake showed that Cd rapidly (10 min) blocked the inducible high-affinity transport system; the constitutive high-affinity transport system appeared not vulnerable to Cd and the low-affinity transport system appeared to be less affected and only after a prolonged exposure (12 h). Cd-treatment also modified transcript levels of genes encoding high-affinity NO3 − transporters (ZmNTR2.1, ZmNRT2.2), PM H+-ATPases (ZmMHA3, ZmMHA4) and NO3 − reductases (ZmNR1, ZmNADH:NR). Despite an expectable increase in NO3 − demand, a negative effect of Cd on NO3 − nutrition is reported. Cd effect results in alterations at the physiological and transcriptional levels of NO3 − uptake from the external solution and it is particularly severe on the inducible high-affinity anion transport system. Furthermore, Cd would limit the capacity of the plant to respond to changes in NO3 − availability.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Multiple evidence suggests that coping with cadmium (Cd) toxicity should require increased nitrate (NO3 −) uptake and metabolism. Firstly, Cd detoxification in higher plants mainly occurs through phytochelatins, which are N- and S-containing peptides (Clemens 2006). Their synthesis leads to a depletion of the glutathione pool and to a de-repression of sulfate (SO4 2−) uptake (Nocito et al. 2002), which in turn is known to be linked to NO3 − uptake and metabolism, as described in maize cells or in barley and spinach plants (Clarkson et al. 1989, 1999; Prosser et al. 2001). Secondly, synthesis of phytochelatins requires glutamate, but exposition to Cd inhibits glutamine synthetase and glutamine oxoglutarate aminotransferase so that both leaves and roots of Cd-treated plants tend to accumulate ammonium and deplete their glutamate pool. As a consequence, an increase in glutamate synthesis via glutamate dehydrogenase (GDH) activity has been observed in different plant species such as barley, bean, maize and rice (Boussama et al. 1999a, b; Gouia et al. 2000, 2003; Astolfi et al. 2004; Lee et al. 2010): NO3 − could help avoiding ammonium toxicity and favor glutamate synthesis (Britto and Kronzucker 2002). Finally, the severity of Cd toxicity also depends on the plant N-status and an adequate rate of NO3 − uptake may consequently facilitate Cd detoxification: N-deficient barley plants treated with Cd show different metabolite pools (e.g. lower phytochelatins content), enzymatic activity and gene transcription levels if compared to N-sufficient plants treated with Cd (Finkemeier et al. 2003). It has also been recently suggested that Cd tolerance may involve a preferential accumulation of NO3 − in Arabidopsis roots (Li et al. 2010).
Despite the expectable increase in NO3 − demand, a general negative effect of Cd on NO3 − nutrition in higher plants has been reported (Sanità di Toppi and Gabbrielli 1999). Cd exposure lowers NO3 − accumulation and assimilation both in the root and in the shoot of bean and tomato (Ouariti et al. 1997; Gouia et al. 2000). At the enzymatic level, Cd decreases protein amount and activity of nitrate reductase (NR) in bean and maize, probably as a consequence of a general depression in protein synthesis (Boussama et al. 1999b; Gouia et al. 2003). However, it is often difficult to separate the direct effect of Cd itself from the secondary effect caused by the cellular response to a toxic accumulation of the heavy metal, since experimental conditions frequently include high Cd concentrations (up to the millimolar range) or prolonged exposures of the plants to the heavy metal (some days) as well as additional stress factors, e.g., S-deficiency (Astolfi et al. 2004).
Notwithstanding the contradiction between the expected importance of NO3 − uptake for Cd detoxification and the proved negative effect of Cd on NO3 − nutrition, the effect of Cd exposure on the mechanisms of the anion’s uptake across the plasma membrane (PM) by NO3 − transporters has not been studied in detail. Indeed, a decrease in NO3 − depletion from the external solution, as well as an inhibition of NR at the physiological level, has been described in barley or Pisum sativum (25–50 μM Cd for up to 72 h or 10 day; Hernandez et al. 1997; Boussama et al. 1999a), but the activity of NO3 − transporters has not been monitored. At the molecular level, it has been observed in Arabidopsis that Cd (5 or 50 μM Cd for up to 30 h) can rapidly alter the transcript levels of genes encoding NO3 − transporters (e.g. AtNRT1.1, AtNRT2.1, AtNRT2.2), as well as NO3 − reductases (AtNR1, AtNR2) and PM proton pumps (PM H+-ATPases; e.g. AtAHA2, AtAHA5, AtAHA10, AtAHA11; Herbette et al. 2006).
NO3 − uptake across the root PM represents a complex process with some peculiar characteristics shared by different plant species, such as: (1) a localized rapid induction of high-affinity transporters activity upon supply of the anion, observed at both the transcript and the protein level (Hole et al. 1990; Miller et al. 2007; Wirth et al. 2007), (2) a systemic negative feedback on high-affinity transporters exerted by the intermediate products of NO3 − assimilation (e.g. nitrite, ammonium, glutamine, asparagine, arginine; Fraisier et al. 2000; Vidmar et al. 2000; Loque et al. 2003), and (3) a dependence of NO3 − transport on the electrochemical gradient generated by the activity of the PM H+-ATPases (Miller and Smith 1996).
The induction of high-affinity NO3 − transport is therefore considered one of the first steps of the complex response to external NO3 − with the anion acting not only as a nutrient, but also as a signal eliciting the rapid gene expression of transporters and metabolism enzymes (Krouk et al. 2010). High-affinity NO3 − transporters have also been suggested to play a role in root morphology (Little et al. 2005; Remans et al. 2006) and plant growth (Orsel et al. 2004; Katayama et al. 2009) in Arabidopsis and rice.
Thus, it appears interesting to assess whether Cd might affect the induction of high-affinity NO3 − transport across the PM. In the present work, 5-day-old maize seedlings were exposed to 1 mM NO3 − for up to 24 h (induction) in the presence or absence of low Cd concentrations (1 or 10 μM Cd) and the induction of high-affinity NO3 − uptake was monitored; concomitantly, uptake kinetics, NO3 − accumulation and reduction rate, and PM H+-ATPase activity were measured. The transcript levels of the main genes involved in NO3 − uptake and reduction were also analyzed.
Materials and methods
Plant material and growth conditions
Maize seeds (Zea mays L. cv. Cecilia; Pioneer Hi-Bred Italia Srl, Pieve Delmona, CR, Italy) were germinated over an aerated 0.5 mM CaSO4 solution at 27 °C in the dark. After 3 days, seedlings were transferred into an aerated solution containing 0.5 mM CaSO4 (day/night photoperiod 16/8, light intensity 220 μmol photons m−2 s−1, temperature (day/night) 25/20 °C, RH 70–80 %). After 2 days, seedlings were transferred for NO3 − uptake induction to a nutrient solution (NS) containing (mM) KNO3 1, NH4H2PO4 0.025, CaSO4 0.4, KH2PO4 0.087, MgSO4 0.1, KCl 0.005, FeSO4 0.01, H3BO3 0.0025, MnSO4 0.0002, ZnSO4 0.0002, CuSO4 0.00005, H2MoO4 0.00005 and with 0 (induced), 1 or 10 μM CdSO4 for 0, 4, 8, 12 or 24 h. In the NS for non-induced plants, KNO3 was replaced by K2SO4 0.5 mM.
Measurement of net high-affinity NO3 − uptake and calculation of kinetic parameters
Roots of intact seedlings were immersed in 40 mL of a constantly agitated and aerated solution containing 0.5 mM CaSO4 and 0.2 mM KNO3. Net uptake was measured as NO3 − depletion from the solution per unit of time (Cataldo et al. 1975), removing samples (0.2 mL) for NO3 − determination every 2 min for 10 min, span time during which uptake had a linear trend. Aliquots of 0.2 mL were mixed thoroughly with 0.8 mL of 5 % (w/v) salicylic acid in concentrated H2SO4. After 20 min incubation at room temperature, 19 mL of 2 M NaOH was added. Samples were cooled to room temperature and NO3 − concentration was determined spectrophotometrically by measuring the absorbance at 412 nm.
Kinetic parameters of the high-affinity NO3 − uptake system (V max and K m) were calculated in the 0.15–0.5 concentration range. Uptake rates were measured as described above except that the uptake solution contained 0.125, 0.15, 0.2, 0.3, 0.5, 1, 2, 5 or 10 mM KNO3. Kinetic parameters were calculated after subtracting the linear component of the uptake rate calculated as the slope in the 0.2–0.5 concentration range. The results were obtained using the linearization of Lineweaver–Burk. The linearizations of Hanes–Woolf and Woolf–Augustinsson–Hofstee were used for comparison (Segel 1976) and gave lower absolute values for V max and K m, but confirmed the differences between treatments. These kinetic parameters are not to be attributed to a single transporter, but refer to the overlapping activities of different transporters.
NO3 − reductase
NR was extracted from leaf tissues grinded in a mortar with liquid nitrogen. The extraction buffer (50 mM potassium phosphate buffer, pH 7.5, 1 mM ethylenediaminetetraacetate, 1 mM dithiothreitol (DTT), 1 μM flavin adenine dinucleotide, 10 μM leupeptin and 10 μM chimostatin were then added to the tissue powder (0.04 mL mg−1 FW). The homogenate was centrifuged at 4 °C for 30 min at 12,500g. NR activity was measured immediately in the supernatant. The reaction mixture consisted of 10 mM phosphate buffer, pH 7.5, supplemented with 10 mM KNO3 and 0.1 mM NADH. The reaction was terminated after 15 min at 28 °C in the dark, by addition of an equal volume of sulfanilamide [1 % (w/v) in 1 N HCl] and then naphthylethylene-diamine dihydrochloride [0.01 % (w/v)] to the reaction mixture and the absorbance at 540 nm was measured.
Determination of NO3 − and Cd content
Roots were rinsed three times in distilled water and blotted with paper towels, frozen in liquid nitrogen and stored at −80 °C until use. Leaves were collected and immediately frozen and stored. For NO3 − content, 300 mg tissue was homogenized in ice cold deionised water (10 mL g−1 FW). The homogenate was filtered through four cheesecloth layers and transferred into 2 mL tubes, then centrifuged at 13,000g for 15 min. NO3 − concentration was determined in 200 μL aliquots of the supernatants with the same procedure described for NO3 − uptake assay, except that for each sample a blank was prepared, omitting the salicylic acid from the H2SO4 solution to subtract basal noise.
For Cd content, as described in Zuchi et al. (2009), root and shoot tissues were oven-dried at 80 °C, ashed at 550 °C, dissolved in 1 N HCl and analyzed by inductively coupled plasma atomic emission spectrometry (Varian, Torino, Italy).
Isolation of plasma membranes
Plasma-membrane vesicles were isolated from root samples as described in Tomasi et al. (2009) with slight modifications: 2 g FW root tissue was homogenized with a mortar and pestle in 4 mL freshly prepared ice-cold extraction medium: 250 mM sucrose, 2 mM MgSO4, 2 mM adenosine 5′-triphosphate, 10 % (v/v) glycerol, 10 mM glycerol-1-phosphate, 0.16 % (w/v) BSA, 2 mM ethylene glycol tetraacetic acid, 2 mM DTT, 5.7 % (w/v) choline-iodide, 1 mM phenylmethylsulfonyl fluoride, 20 μg mL−1 chymostatin, 25 mM MES-1,3-bis [tris(hydroxymethyl)-methyloamino] propane (BTP) pH 7.6 and 0.5 g−1 FW polyvinylpolypyrrolidone (PVPP). Homogenates were filtered through four layers of cheesecloth and the suspensions were subjected to differential centrifugation steps in an Eppendorf microcentrifuge at 2 °C: 12,700g for 3 min (pellets discarded), 12,700g for 25 min (pellets recovered). Microsomes, gently resuspended in 400 μL of homogenization medium (extraction medium without PVPP) were loaded onto a discontinuous sucrose gradient made by layering 700 μL sucrose solution (1.13 g cm−3) on a 300 μL sucrose (1.17 g cm−3) cushion and then centrifuged at 12,700g for 1 h. The sucrose solutions were prepared in 5 mM MES-BTP pH 7.4 and contained all of the protectants present in the homogenization medium. Vesicles migrating to the 1.13/1.17 g cm−3 interface were collected, diluted with 1.8 mL homogenization medium and centrifuged at 14,000g for 30 min. Pellets were resuspended in a 100 μL medium containing 250 mM sucrose, 10 % (v/v) glycerol, 1 mM DTT, 50 μg mL−1 chymostatin and 2 mM MES-BTP pH 7.0, immediately frozen in liquid nitrogen and stored at −80 °C.
Measurement of PM H+-ATPase activity
PM H+-ATPase hydrolytic activity was measured at 38 °C in a 0.6 mL reaction medium (50 mM MES-BTP pH 6.5, 5 mM MgSO4, 100 mM KNO3, 600 μM Na2MoO4, 1.5 mM NaN3, 5 mM ATP-BTP pH 6.5, 0.01 % (w/v) polyoxyethylene 20 cetyl ether (Brij 58), with or without 100 μM V2O5). The reaction was started by adding the membrane vesicles containing 0.5 μg of total protein; after 30 min, the reaction was stopped and color developed as previously described by Santi et al. (1995). Inorganic phosphate was quantified spectrophotometrically at 705 nm as described by Forbush (1983). Protein content was determined as described by Bradford (1976), using BSA as standard, after solubilizing membrane vesicles with 0.5 M NaOH (Gogstad and Krutnes 1982). The activity is expressed in μmol P mg protein h−1 subtracting the quantity produced in the enzyme assay in presence of vanadate.
Transcript levels analysis
At harvesting times, root samples were collected, immediately frozen in liquid nitrogen and conserved at −80 °C until further processing. RNA extractions were performed using the Invisorb Spin Plant RNA kit (Stratec Molecular, Berlin, Germany). 1 μg of total RNA (checked for quality and quantity using a spectrophotometer, followed by electrophoresis in agarose gel) of each sample was retro transcribed using 1 pmol of Oligo d(T)23VN (Sigma Aldrich, Milano, Italy), 15 U Prime RNase Inhibitor (Eppendorf, Hamburg, Germany) and 10 U M-MulV RNase H− for 1 h at 42 °C (Finnzymes, Helsinki, Finland). After RNA digestion with 1 U RNase A (USB, Cleveland, OH, USA) for 1 h at 37 °C, transcript levels analyses were performed by adding 0.1 μL of the cDNA to FluoCycleTM sybr green (20 μL final volume; Euroclone, Pero, MI, Italy) in a DNA Engine Opticon Real-Time PCR Detection (Biorad, Hercules, CA, USA).
Primers (Tm = 58 °C) were the following: ZmNRT2.1 (AJ344451), GATCGACGATCACCTATACCTC and GTGCTCCGTTGACATGAG (PCR efficiency 69 %); ZmNRT2.2 (AY659965), CCTACCTTTACGTGTATGCCTTG and GATGTGCCAACGATATTCATC (PCR efficiency 83 %); ZmMHA1 (U09989), CGAGAACAAGACGAGCTTCA and CAGTGGAGATGCTCGACAAA (PCR efficiency 75 %); ZmMHA2 (X85805), TCCGACTGTTGTTTGTCGAG and CACCGACTCCATCCTCATCT (PCR efficiency 71 %); ZmMHA3 (AJ441084), GCCAAGAGACGAGCTGAGAT and CACCGTGTAGTTCTGCTGGA (PCR efficiency 84 %); ZmMHA4 (AJ539534), CGGTGATGTGATTGGAGACA and CGGTGATGTGATTGGAGACA (PCR efficiency 93 %); ZmNR1 (AF153448), CCAGCCGACTTGCCAGCGTAA and GCATGGCCTATGTTATCTGCTGCTC (PCR efficiency 85 %); ZmNADH:NR (M27821), GGTCTTTGGAGGTGGAGGTGCTG and CTCTGGCTGCGTATTCAAACTCTCGT (PCR efficiency 85 %); ZmST1.1 (AF355602), AAGTGGAATCCATGCTTTGG and CTGAGCGGAGCTTCTGGAT (PCR efficiency 74 %). As housekeeping genes, ZmPolyU (polyubiquitin, S94466, GTACCCTCGCCGACTACAAC and ATGGTCTTGCCAGTCAAGGT, PCR efficiency 83 %), and ZmRPL17 (ribosomal protein L17, AF034948, AAAGTCTCGCCACTCCAATG and ACGTCCAAGCCTTTCACATC, PCR efficiency 90 %) were used. Triplicates were performed on three independent experiments; analyses of real-time result were performed using Opticon Monitor 2 software (Biorad) and R (version 2.9.0; http://www.r-project.org/) with the qPCR package (version 1.1-8; http://www.dr-spiess.de/qpcR.html). Efficiencies of amplification were calculated following the authors’ indications (Ritz and Spiess 2008).
Transmembrane topology prediction
Predictions have been carried out at the PSIPRED Protein Structure Prediction Server of the University College of London (http://bioinf.cs.ucl.ac.uk/psipred/) using MEMSAT3 with the default settings and the sequences retrieved from NCBI (http://www.ncbi.nlm.nih.gov/protein) for the following proteins: ZmNRT2.1 (CAC87729.2), ZmNRT2.2 (AY659965.1), ZmNRT1.2 (AAY40798.1) and ZmNAR2.1 (AAY40796.1).
Statistical analysis
Computation of the graphical representation and statistical validation (ANOVA and Student’s t test; P < 0.05) were performed on data belonging to each time point (not between different time points) using SigmaPlot 11.0 (Systat software, Point Richmond, CA, USA). Transcript levels data were illustrated considering the differences in the PCR efficiency of amplification and using the mean transcript level of the housekeeping genes ZmPolyU and ZmRPL17 in roots of control non-induced plants at time zero as reference.
Results
The data presented have been obtained using maize seedlings exposed to 0, 1 or 10 μM Cd during a 24-h induction for NO3 − uptake (1 mM NO3 −). In our experimental conditions Cd exposure did not produce any apparent symptoms of toxicity. As expected (Nocito et al. 2002), a typical detoxification response was activated by all Cd-treated plants, as evidenced by decreased glutathione pools, increased non-protein thiols concentrations and higher SO4 2− uptake capacity (Online Resource 1: Suppl. Figs. S1–S3). At the end of the exposure period, Cd concentration was higher in the roots as compared to the shoots (184 vs. 52 μg g−1 DW and 470 vs. 102 μg g−1 DW, in plants treated with 1 and 10 μM Cd, respectively).
Cd effect on the induction of high-affinity NO3 − transporters’ activity, NO3 − accumulation and reduction
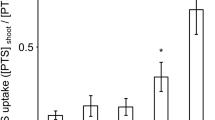
Induced plants, when treated with 1 mM NO3 −, gradually developed a greater net high-affinity NO3 − uptake rate, measured at 0.2 mM NO3 −, with a peak after 12 h of treatment and a subsequent de-induction phase (Pinton et al. 1999; Santi et al. 2003); the increase in net high-affinity NO3 − uptake was not observed in plants not supplied with NO3 − (control non-induced), which maintained their basal uptake rate all along the experimental period (Fig. 1). On the other hand, the presence of Cd in the nutrient solution strongly affected the induction of root high-affinity NO3 − uptake: only a slight induction of the high-affinity net NO3 − uptake rate was observed in plants supplied with both 1 mM NO3 − and 1 μM Cd (induced + 1 μM Cd); the presence of 10 μM Cd strongly impaired induction of the high-affinity NO3 − uptake (Fig. 1). Conversely, no significant alteration was observed in the constitutive net high-affinity NO3 − uptake rate (non-induced + 1 or 10 μM Cd; Fig. 1).
High-affinity net NO3 − uptake rate in roots of maize seedlings supplied with 1 mM NO3 − and 0 (control induced), 1 or 10 μM Cd in the nutrient solution. Control non-induced plants were treated in nutrient solution without NO3 −. Net NO3 − uptake was measured spectrophotometrically as the depletion from a solution containing 0.2 mM NO3 −. Closed circles, control induced; open circles, induced + 1 μM Cd; closed triangles, induced + 10 μM Cd; open triangles, control non-induced; closed squares, non-induced + 1 μM Cd; open squares, non-induced + 10 μM Cd. Data are mean ± SD, letters refer to statistically significant differences within each time point among independent experiments, underlined letters refer to overlapping data not significantly different among treatments (ANOVA, n = 3, P < 0.05)
NO3 − supply also produced a gradual increase in the NR activity of the roots, which was slightly, although significantly, less pronounced in the presence of Cd (Fig. 2a). In induced Cd-treated plants, root NO3 − content was similar to that of the control-induced plants and started to decline after 8 h of treatment with the highest Cd concentration (10 μM Cd) (Fig. 2b). NO3 − accumulation in the roots was faster and more pronounced than in the shoots: NO3 − content was about twofold higher in the roots than in the shoot after 24 h (Fig. 2b). NO3 − content after 24 h exposure to 0, 1 or 10 μM Cd was 100, 94 and 71 %, respectively, in the root, and 100, 84 and 63 %, respectively, in the shoot.
Activity of NR (a) and NO3 − content (b) in shoots (above) and roots (below), measured spectrophotometrically after extraction from root tissues of maize seedlings supplied with 1 mM NO3 − and 0 (control induced), 1 or 10 μM Cd in the nutrient solution. a Closed circles, control induced; open circles, induced + 1 μM Cd; closed triangles, induced + 10 μM Cd. b White bars, control plants at time zero; black bars, induced; light grey bars, induced + 1 μM Cd; dark grey bars, induced + 10 μM Cd. Data are mean ± SD, letters refer to statistically significant differences within each time point among independent experiments (ANOVA, n = 3, P < 0.05)
The activity of the PM H+-ATPase, which is known to increase in maize in response to NO3 − supply (Santi et al. 1995) was measured in vesicles isolated from roots. The time course of ATP hydrolysis rate in the different treatments was similar to that described for NO3 − uptake rate. Figure 3 shows an increase in the ATP hydrolysis rate in the vesicles isolated from roots of control-induced plants, with a peak after 12 h of treatment which matches the time of maximum NO3 − uptake rate measured at 0.2 mM (see Fig. 1). On the other hand, induced Cd-treated plants only showed a slight increase in their ATP hydrolysis rate.
Vanadate-sensitive phospho-hydrolysing activity of the PM H+-ATPase in vesicles isolated from roots of maize seedlings supplied with 1 mM NO3 − and 0 (control induced), 1 or 10 μM Cd in the nutrient solution. Phospho-hydrolysing activity was measured spectrophotometrically on root microsomal fractions. Closed circles, control induced; open circles, induced + 1 μM Cd; closed triangles, induced + 10 μM Cd. Data are mean ± SD, letters refer to statistically significant differences within each time point among independent experiments (ANOVA, n = 3, P < 0.05)
Cd effect on NO3 − uptake kinetics
NO3 − uptake rates as a function of external NO3 − concentration, in the range of 0.125–10 mM, were measured (Figs. 4, 5; Table 1). The kinetic parameters (V max and K m) for the high-affinity transport, which was considered to be operating below 0.5 mM NO3 − (Hogh-Jensen et al. 1997; Siddiqi et al. 1990) were calculated after subtraction of the linear component of the uptake rate, estimated as the slope in the 0.2–0.5 mM concentration range (Table 1).
Net NO3 − uptake kinetics in roots of maize seedlings after prolonged exposure to Cd (12 h), measured in the high-affinity concentration range (0.125–0.5 mM NO3 −, a) and in the low-affinity concentration range (0.5–10 mM NO3 −, b). Control induced plants were supplied with 1 mM NO3 − for 12 h before the uptake assay; control non-induced plants were treated for 12 h in nutrient solution without NO3 −. For Cd-treated plants 1 μM Cd was added to the nutrient solutions for 12 h. Closed circles, control induced; open circles, induced + 1 μM Cd; open triangles, control non-induced; closed squares, non-induced + 1 μM Cd. Net NO3 − uptake was measured spectrophotometrically as depletion from solutions containing different concentrations of NO3 −. Data are mean ± SD, letters refer to statistically significant differences within each time point among independent experiments; underlined letters refer to overlapping data not significantly different among treatments (ANOVA, n = 4, P < 0.05)
Net NO3 − uptake kinetics in roots of maize seedlings after short exposure to Cd (10 min), measured in the high-affinity concentration range (0.125–0.5 mM NO3 −, a) and in the low-affinity concentration range (0.5–10 mM NO3 −, b). Control induced plants were supplied with 1 mM NO3 − for 12 h before the uptake assay; control non-induced plants were treated for 12 h in nutrient solution without NO3 −. For Cd-treated plants 1 μM Cd was added to the assay medium (10 min). Labels, measurements and statistical analyses as described in Fig. 4
In a first set of measurements (long exposure; Table 1; Fig. 4), root NO3 − uptake rates were compared among different treatments: (1) plants supplied for 12 h with a complete nutrient solution containing no NO3 − (non-induced), (2) plants supplied for 12 h with a complete nutrient solution containing no NO3 − and with the addition of Cd (non-induced + 1 μM Cd), (3) plants fed for 12 h with a complete solution containing 1 mM NO3 − (induced), and (4) plants treated for 12 h with both NO3 − and Cd (induced + 1 μM Cd). Induced plants, compared to non-induced ones, showed increased NO3 − uptake rates both in the high- and low-affinity concentration ranges (Fig. 4a, b, respectively). The kinetic parameters calculated for the high-affinity NO3 − uptake (Table 1) showed a decrease in the K m value of induced plants and an increase in the V max. On the other hand, non-induced plants did not show any difference when compared to non-induced ones treated with Cd (Fig. 4a). Finally, induced plants treated with Cd showed an intermediate uptake rate in the low-affinity concentration range (Fig. 4b), while in the high-affinity concentration range the uptake rate remained similar to the constitutive uptake rate of non-induced plants (Fig. 4a). In these latter plants, Cd effect was particularly evident on the K m value, while the V max value was between that of non-induced and control induced plants (Table 1).
A second set of measurements (short exposure, Fig. 5) was performed using plants either non-induced or induced with 1 mM NO3 − for 12 h without any Cd addition, and then exposed to 0 or 1 μM Cd during the 10-min NO3 − uptake assay. Again, when NO3 − uptake was measured in the high-affinity concentration range, the short exposure to Cd did not cause any significant decrease in the constitutive uptake rates of non-induced plants, but strongly depressed uptake rates of induced plants (Fig. 5a). In Cd-treated plants, K m value was similar independent of NO3 − induction and comparable to that of plants not induced for NO3 − uptake and not exposed to Cd during the NO3 − uptake assay (control non-induced; Table 1). V max value was significantly decreased by the short Cd treatment in plants induced for NO3 − uptake (control induced). Different to the prolonged exposure described above, the short exposure to Cd did not affect, either in non-induced or induced plants, the NO3 − uptake rates measured in the low-affinity concentration range, which remained as high as that of plants that were never exposed to the heavy metal (Fig. 5b).
Cd effect on transcript levels of genes related to NO3 − acquisition in root tissues
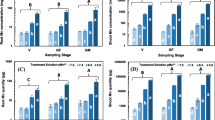
The transcript amount of the genes ZmNRT2.1 and ZmNRT2.2, which encode two putative high-affinity transporters, was analysed. mRNA level of ZmNRT2.1 reached a maximum between 8 and 12 h from the beginning of NO3 − supply in roots of control-induced plants (Fig. 6a). On the other hand, in induced Cd-treated plants no significant change in ZmNRT2.1 transcript accumulation was measured during the experimental time span (Fig. 6a). Consequently, mRNA amount after 12 h from starting NO3 − supply appeared to be significantly higher in control-induced plants than in plants treated with 1 μM Cd. After 24 h ZmNRT2.1 transcripts in induced Cd-treated plants showed values that were not significantly different from those of control-induced plants, where feedback regulation started to occur.
Real-time RT-PCR analyses of gene transcript levels in roots of maize seedlings supplied with 1 mM NO3 − and 0 or 1 μM Cd in the nutrient solution. Analyzed genes encode: a, b high-affinity NO3 − transporters (ZmNRT2.1 and ZmNRT2.2); c, d PM H+-ATPases (ZmMHA3 and ZmMHA4); e, f nitrate reductases (ZmNR1 and ZmNADH:NR). White bars, control non-induced plants at time zero; black bars, control induced; light grey bars, induced +1 μM Cd. Gene mRNA levels were normalized with respect to the mean transcript level of the housekeeping genes ZmPolyU and ZmRPL17; relative changes in gene transcript levels were calculated on the basis of the mean transcript level of housekeeping genes ZmPolyU and ZmRPL17 in roots of control plants at time zero. Data are mean ± SD, letters refer to statistically significant differences within each time point among independent experiments; stars refer to statistically significant differences between each time point and control non-induced plants at time zero (t test, n = 3, P < 0.05)
The time course of ZmNRT2.2 mRNA accumulation showed a peak after 4 h in control-induced plants with a 2.9-fold increase when compared to 0 h (Fig. 6b). Cd-treated plants showed a clear increase in transcript level only after 24 h with values 5.6 times higher than at 0 h.
ZmMHA3 and ZmMHA4 encode two putative PM H+-ATPases. The accumulation of ZmMHA3 transcripts in control-induced plants peaked after 4 h of treatment and later declined (Fig. 6c). In Cd-treated plants the increase in ZmMHA3 mRNA was lower and delayed reaching a peak after 8 h (Fig. 6c).
Transcript amount of ZmMHA4 showed, in control-induced plants, a trend similar to that of ZmMHA3, with a peak of similar extent (8.2 and 7.1-fold increase compared to 0 h, respectively for ZmMHA3 and ZmMHA4) after 4 h. Plants treated with 1 μM Cd did not show any significant accumulation of ZmMHA4 transcript (Fig. 6d). The mRNAs of the genes encoding for two other putative PM H+-ATPases were not detectable (ZmMHA1, ZmMHA2, data not shown), as previously reported for maize roots (Santi et al. 2003).
The transcript levels of ZmNR1 and ZmNADH:NR, encoding two maize NRs (Hyde and Campbell 1990; Campbell 1992; Dwivedi et al. 1994), were also examined. ZmNR1 transcript progressively accumulated in control-induced plants reaching a maximum after 8 h (value 2.5 times higher than 0 h) and then declining (Fig. 6e). In induced plants treated with 1 μM Cd this pattern was delayed: the mRNA amount increased only after 12 h and was significantly higher than in control-induced plants after 24 h (Fig. 6e).
ZmNADH:NR transcript did not show significant accumulation in control-induced plants, but rather a decrease after 8 and 24 h (0.5 times the level measured at 0 h; Fig. 6f). After 12 h, however, the transcript level was comparable to that measured at 0 h and was significantly higher than in induced Cd-treated plants. In induced Cd-treated plants the transcript abundance was significantly lower at any time during the time span of the experiment as compared with values recorded at 0 h (Fig. 6f).
Discussion
Cadmium is known to affect NO3 − nutrition at the physiological and molecular levels in different plant species (Ouariti et al. 1997; Herbette et al. 2006). The Cd concentrations used in this work were low enough to avoid the development of any visible toxicity symptom during the experimental period. Nevertheless, the plants showed a marked and dose-dependent decrease in their capability to develop a higher capacity for NO3 − uptake measured in the high-affinity range (0.2 mM NO3 −; Fig. 1).
As Cd is known to lower the protein and activity levels of NR in maize (Boussama et al. 1999b), the lack of induction of NO3 − uptake in Cd-treated plants could be due to a negative feedback caused by the impairing of the assimilatory pathway, as also reported for barley, tobacco or Arabidopsis (Fraisier et al. 2000; Vidmar et al. 2000; Loque et al. 2003). Indeed, we observed that root NR activity was slightly, but significantly depressed in the presence of Cd (Fig. 2a). However, a decrease rather than an increase of NO3 − accumulation was measured, as compared to induced roots, in the roots of Cd-treated plants, particularly in those treated with 10 μM Cd (Fig. 2b). The highest proportion of NO3 − is generally translocated and undergoes reductive assimilation in the shoot (Lewis et al. 1982). In our experimental conditions, shoot NO3 − content was always lower than in the root and the differences between control NO3 −-induced plants and Cd-treated ones were greater in the shoot as compared to the root (Fig. 2b). Thus, as there was no higher NO3 − accumulation either in the root or in the shoot of Cd-treated plants, the rapid inhibition of NO3 − uptake induction does not appear to be related to a negative feedback due to the impairing of the assimilatory pathway (i.e. lowered NR activity and excess NO3 − accumulation), while it is more compatible with a limited influx of the anion. Interestingly, a preferential accumulation of NO3 − in the roots of Cd-stressed plants has been repeatedly reported in different plant species, suggesting that NO3 − retention in the roots of Cd-treated plants may function as a strategy to protect roots (Hernandez et al. 1997; Chaffei et al. 2004; Li et al. 2010). However, since Cd has a detrimental effect on the development of inducible high-affinity NO3 − uptake, an increase in root NO3 − content through an accelerated uptake rate is unlikely. Coherently, other responses have so far been described such as: (1) reduced root-to-shoot nitrogen translocation due to decreased transpiration in P. sativum (Hernandez et al. 1997), (2) N-recycling (e.g. asparagine synthesis) and shoot-to-root translocation of reduced N-compounds in tomato (Chaffei et al. 2004), and (3) increased activity of AtNRT1.8, an Arabidopsis low-affinity NO3 − transporter localized in the PM of xylem parenchyma cells which unloads NO3 − from the xylem sap and is strongly up-regulated by Cd exposure (Li et al. 2010).
NO3 − uptake is energized by the PM H+-ATPase activity (McClure et al. 1990a, b); it has been shown in maize that the response to NO3 − supply involves an increase both in the transcription and the protein level of the enzyme (Santi et al. 2003). Cd has been reported to alter PM permeability in P. sativum and maize, thus leading to an apparent decrease of the proton pumping activity that, in turn, requires higher ATP hydrolysis rates to sustain the proton electrochemical gradient (Hernandez and Cooke 1997; Astolfi et al. 2005; Nocito et al. 2008). A direct inhibition of PM H+-ATPase has been demonstrated in maize, but only for high Cd concentrations (>50 μM; Nocito et al. 2008). In our experimental conditions there was no direct inhibition of PM H+-ATPase caused by Cd (either on proton pumping or ATP hydrolysis rates, Online Resource 1: Suppl. Fig. S4). On the other hand, the proton pumping rate was lower in vesicles purified from plants induced for NO3 − uptake in the presence of Cd (Online Resource 1: Suppl. Fig. S5), compared to plants induced only in presence of NO3 −. The ATP hydrolysis rate was also markedly lower in vesicles extracted from roots of plants induced for NO3 − uptake in the presence of Cd (Fig. 3), as was high-affinity NO3 − uptake (Fig. 1). Thus, the lack of induction of PM H+-ATPase activity seems to be related to the lack of induction of the high-affinity NO3 − transporter, rather than to an effect of Cd on the PM H+-ATPase itself or on membrane properties (e.g. permeability).
We further studied in detail the Cd effect on the different NO3 − transport systems. The functionality of the constitutive high-affinity NO3 − transport system was not significantly affected by Cd exposure, while the low-affinity transport of NO3 −-induced plants was only affected after a prolonged exposure to Cd (12 h). On the other hand, a 10-min exposure to Cd of NO3 −-induced plants dropped their high-affinity transport rate to the levels of non-induced plants (Fig. 5a). Since a significant Cd accumulation in the root during the 10-min assay can be reasonably excluded, results rather support a direct effect of the heavy metal on the inducible high-affinity NO3 − transporters located at the PM. ZmNRT2.1 and ZmNRT2.2 (putative high-affinity transporters) are predicted to carry one and two cysteine residues, respectively, on the external side of the PM; thus these cysteines would be exposed to Cd present in the external solution and CD readily reacts with thiols. This is not the case for ZmNRT1.2 (putative low-affinity transporter). Moreover, to be fully functional, several NRT2 transporters need to interact with other proteins such as AtNAR2.1 in Arabidopsis (Orsel et al. 2006; Okamoto et al. 2006; Yong et al. 2010; Kotur et al. 2012), OsNAR2.1 in rice (Yan et al. 2011) and HvNAR2.3 in barley (Tong et al. 2005; Ishikawa et al. 2009). A protein with high sequence homology is also found in maize (ZmNAR2.1) and it might be needed for high-affinity NO3 − uptake. Based on the prediction of transmembrane protein topology, ZmNAR2.1 would carry three cysteine residues on the external side of the PM. This might be a possible explanation for the different sensitivity to Cd exhibited by high- and low-affinity NO3 − transporters, although these aspects deserve additional research efforts.
The induction of NO3 − uptake is known to be regulated at the transcriptional level in different plant species (Hole et al. 1990; Wirth et al. 2007). The results of the present work (Figs. 1, 6a) give further support to the idea that ZmNRT2.1 would be the putative main inducible high-affinity transporter for NO3 − uptake from the external solution in maize (Santi et al. 2003; Trevisan et al. 2008). In Cd-treated plants the induction of high-affinity NO3 − uptake was very low (1 μM Cd; Fig. 1) and no significant change in ZmNRT2.1 transcript amount was observed during the experimental period (Fig. 6a). The residual induction of NO3 − uptake in induced plants treated with 1 μM Cd (Fig. 1) might well be due to the operation of low-affinity transporters (Figs. 4b, 5b). By subtracting the linear component due to the low-affinity transporters (Fig. 4a) it became apparent that the only component of the high-affinity transport system still active, after a 12 h Cd treatment, was the constitutive one.
ZmNRT2.2, another putative high-affinity NO3 − transporter, has been found to localize both in the cortex and at higher levels in the central cylinder of maize plants, where it likely plays a role in controlling root-to-shoot translocation of the anion (Trevisan et al. 2008). In control-induced plants it showed an early (4 h) increase in its transcript level, indicating a possible involvement in NO3 − translocation towards the shoot for assimilation (Fig. 6b). In Cd-treated plants ZmNRT2.2 exhibited a marked increase in mRNA accumulation only after 24 h (Fig. 6b). Such a behaviour might be related to the need of balancing the lack of induction of ZmNRT2.1, as it has been suggested for Arabidopsis (Li et al. 2007); alternatively it might be part of a reaction aimed at improving the root-to-shoot translocation of NO3 − notwithstanding the limited influx (Trevisan et al. 2008). Finally, it is interesting to note that both ZmNRT2.1 and ZmNRT2.2 transcripts were detectable before NO3 − supply, supporting the view of their possible involvement in the constitutive high-affinity NO3 − uptake system (Trevisan et al. 2008).
The transcript amounts of ZmMHA3 and ZmMHA4, encoding the putative PM H+-ATPases reported to be up-regulated in response to NO3 − supply (Santi et al. 1995, 2003), showed a peak with similar timing and extent (Fig. 6c, d). However, in Cd-treated plants the response was different for the two genes. ZmMHA4 has been described as more responsive to NO3 − (Santi et al. 2003) and conceivably in Cd-treated plants its mRNA amount remained comparable to that of 0 h throughout the Cd-treatment. Conversely, ZmMHA3 transcript accumulation showed a peak in Cd-treated plants, although delayed and with a reduced amplitude with respect to control-induced plants. These results support the existence of a strict relationship, extending to gene transcription levels, between NO3 − transporters and isoforms of the PM proton pump in both control-induced and induced Cd-treated plants.
In higher plants, NO3 − exposure is known to induce the transcription of genes encoding NR (Gowri et al. 1992; Stitt 1999). A close relationship between NR activity and ZmNR1 transcript level (Figs. 2, 6e) was also observed in our control-induced plants. Moreover, high-affinity NO3 − uptake rate and ZmNRT2.1 transcript level peaked after 12 h from NO3 − supply (Figs. 1, 6a), when ZmNR1 mRNA accumulation was still significantly higher than at 0 h. Therefore, this putative NR isoform might contribute to the assimilation of NO3 − taken up in maize roots. Cd-treatment did not change the expression pattern of the gene; rather it caused a temporal shift with a peak of expression at 12–24 h from the beginning of the experiment (Fig. 6e). This might be due to a reduced influx of NO3 − into the root cells; on the other hand, it might account for the maintenance of a high level of activity during the experimental period even in the presence of Cd. This behaviour might be part of a response to counteract the interference exerted by Cd at the transcriptional and physiological level.
ZmNADH:NR, instead, does not appear to be actively involved, at least at the transcriptional level, in response to incoming NO3 − in maize (Fig. 6f).
Interestingly, we described a delay in gene transcription for ZmNRT2.2, ZmMHA3 and ZmNR1 (Fig. 6b, c, e, respectively) upon exposure to NO3 − in the presence of 1 μM Cd; hence, this response appears to be shared upon Cd stress by several genes coding for proteins involved in the induction of high-affinity NO3 − uptake. Thus, the analysis of gene expression pattern supports the idea that the inhibition of NO3 − uptake by Cd limits the development of a higher uptake capacity (induction) in roots exposed to the anion also through an interference on transcriptional events, and that this interference is exerted with some specificities on the different target genes.
In conclusion, the results of the present work indicate that Cd interference in the process of NO3 − uptake induction involves a direct inhibition of the inducible high-affinity NO3 − transport system. This effect could, in turn, decrease the uptake of the anion and the subsequent induction of physiological and transcriptional processes. These results also show that a Cd concentration as low as 1 μM is able to limit the plant’s ability to respond to fluctuations in the external NO3 − concentration.
Abbreviations
- Cd:
-
Cadmium
- GDH:
-
Glutamate dehydrogenase
- NO3 − :
-
Nitrate
- NR:
-
Nitrate reductase
- PM:
-
Plasma membrane
- SO4 2− :
-
Sulfate
References
Astolfi S, Zuchi S, Passera C (2004) Role of sulphur availability on cadmium-induced changes of nitrogen and sulphur metabolism in maize (Zea mays L.) leaves. J Plant Physiol 161:795–802
Astolfi S, Zuchi S, Passera C (2005) Effect of cadmium on H+ATPase activity of plasma membrane vesicles isolated from roots of different S-supplied maize (Zea mays L.) plants. Plant Sci 169:361–368
Boussama N, Ouariti O, Ghorbal MH (1999a) Changes in growth and nitrogen assimilation in barley seedlings under cadmium stress. J Plant Nutr 22:731–752
Boussama N, Ouariti O, Suzuki A, Ghorbal MH (1999b) Cd-stress on nitrogen assimilation. J Plant Physiol 155:310–317
Bradford MM (1976) Rapid and sensitive method for quantitation of microgram quantities of protein utilizing principle of protein–dye binding. Anal Biochem 72:248–254
Britto DT, Kronzucker HJ (2002) NH4 + toxicity in higher plants: a critical review. J Plant Physiol 159:567–584
Campbell WH (1992) Expression in Escherichia coli of cytochrome-C reductase activity from a maize NADH nitrate reductase complementary-DNA. Plant Physiol 99:693–699
Cataldo DA, Maroon M, Schrader LE, Youngs VL (1975) Rapid colorimetric determination of nitrate in plant tissue by nitration of salicylic acid. Commun Soil Sci Plant Anal 6:71–80
Chaffei C, Pageau K, Suzuki A, Gouia H, Ghorbel MH, Masclaux-Daubresse C (2004) Cadmium toxicity induced changes in nitrogen management in Lycopersicon esculentum leading to a metabolic safeguard through an amino acid storage strategy. Plant Cell Physiol 45:1681–1693
Clarkson DT, Saker LR, Purves JV (1989) Depression of nitrate and ammonium transport in barley plants with diminished sulfate status—evidence of co-regulation of nitrogen and sulfate intake. J Exp Bot 40:953–963
Clarkson DT, Diogo E, Amancio S (1999) Uptake and assimilation of sulphate by sulphur deficient Zea mays cells: the role of O-acetyl-l-serine in the interaction between nitrogen and sulphur assimilatory pathways. Plant Physiol Biochem 37:283–290
Clemens S (2006) Toxic metal accumulation, responses to exposure and mechanisms of tolerance in plants. Biochimie 88:1707–1719
Dwivedi UN, Shiraishi N, Campbell WH (1994) Identification of an essential cysteine of nitrate reductase via mutagenesis of its recombinant cytochrome-B reductase domain. J Biol Chem 269:13785–13791
Finkemeier I, Kluge C, Metwally A, Georgi M, Grotjohann N, Dietz KJ (2003) Alterations in Cd-induced gene expression under nitrogen deficiency in Hordeum vulgare. Plant Cell Environ 26:821–833
Forbush B (1983) Assay of Na, K-ATPase in plasma membrane preparations: increasing the permeability of membrane vesicles using sodium dodecyl sulfate buffered with bovine serum albumin. Anal Biochem 128:159–163
Fraisier V, Gojon A, Tillard P, Daniel-Vedele F (2000) Constitutive expression of a putative high-affinity nitrate transporter in Nicotiana plumbaginifolia: evidence for post-transcriptional regulation by a reduced nitrogen source. Plant J 23:489–496
Gogstad GO, Krutnes MB (1982) Measurement of protein in cell-suspensions using the coomassie brilliant blue dye-binding assay. Anal Biochem 126:355–359
Gouia H, Ghorbal MH, Meyer C (2000) Effects of cadmium on activity of nitrate reductase and on other enzymes of the nitrate assimilation pathway in bean. Plant Physiol Biochem 38:629–638
Gouia H, Suzuki A, Brulfert J, Ghorbal MH (2003) Effects of cadmium on the co-ordination of nitrogen and carbon metabolism in bean seedlings. J Plant Physiol 160:367–376
Gowri G, Ingemarsson B, Redinbaugh MG, Campbell WH (1992) Nitrate reductase transcript is expressed in the primary response of maize to environmental nitrate. Plant Mol Biol 18:55–64
Herbette S, Taconnat L, Hugouvieux V, Piette L, Magniette MLM, Cuine S, Auroy P, Richaud P, Forestier C, Bourguignon J, Renou JP, Vavasseur A, Leonhardt N (2006) Genome-wide transcriptome profiling of the early cadmium response of Arabidopsis roots and shoots. Biochimie 88:1751–1765
Hernandez LE, Cooke DT (1997) Modification of the root plasma membrane lipid composition of cadmium-treated Pisum sativum. J Exp Bot 48:1375–1381
Hernandez LE, Garate A, CarpenaRuiz R (1997) Effects of cadmium on the uptake, distribution and assimilation of nitrate in Pisum sativum. Plant Soil 189:97–106
Hogh-Jensen H, Wollenweber B, Schjoerring JK (1997) Kinetics of nitrate and ammonium absorption and accompanying H+ fluxes in roots of Lolium perenne L. and N2-fixing Trifolium repens L. Plant Cell Environ 20:1184–1192
Hole DJ, Emran AM, Fares Y, Drew MC (1990) Induction of nitrate transport in maize roots, and kinetics of influx, measured with nitrogen-13. Plant Physiol 93:642–647
Hyde GE, Campbell WH (1990) High-level expression in Escherichia coli of the catalytically active flavin domain of corn leaf NADH-nitrate reductase and its comparison to human NADH-cytochrome-B5 reductase. Biochem Biophys Res Commun 168:1285–1291
Ishikawa S, Ito Y, Sato Y, Fukaya Y, Takahashi M, Morikawa H, Ohtake N, Ohyama T, Sueyoshi K (2009) Two-component high-affinity nitrate transport system in barley: membrane localization, protein expression in roots and a direct protein–protein interaction. Plant Biotechnol 26:197–205
Katayama H, Mori M, Kawamura Y, Tanaka T, Mori M, Hasegawa H (2009) Production and characterization of transgenic rice plants carrying a high-affinity nitrate transporter gene (OsNRT2.1). Breeding Sci 59:237–243
Kotur Z, Mackenzie N, Ramesh S, Tyerman SD, Kaiser BN, Glass ADM (2012) Nitrate transport capacity of the Arabidopsis thaliana NRT2 family members and their interactions with AtNAR2.1. New Phytol 194:724–731
Krouk G, Crawford NM, Coruzzi GM, Tsay YF (2010) Nitrate signaling: adaptation to fluctuating environments. Curr Opin Plant Biol 13:266–273
Lee K, Bae DW, Kim SH, Han HJ, Liu X, Park HC, Lim CO, Lee SY, Chung WS (2010) Comparative proteomic analysis of the short-term responses of rice roots and leaves to cadmium. J Plant Physiol 167:161–168
Lewis OAM, Watson EF, Hewitt EJ (1982) Determination of nitrate reductase activity in barley leaves and roots. Ann Bot 49:31–37
Li WB, Wang Y, Okamoto M, Crawford NM, Siddiqi MY, Glass ADM (2007) Dissection of the AtNRT2.1: AtNRT2.2 inducible high-affinity nitrate transporter gene cluster. Plant Physiol 143:425–433
Li JY, Fu YL, Pike SM, Bao J, Tian W, Zhang Y, Chen CZ, Zhang Y, Li HM, Huang J, Li LG, Schroeder JI, Gassmann W, Gong JM (2010) The Arabidopsis nitrate transporter NRT1.8 functions in nitrate removal from the xylem sap and mediates cadmium tolerance. Plant Cell 61:1633–1646
Little DY, Rao HY, Oliva S, Daniel-Vedele F, Krapp A, Malamy JE (2005) The putative high-affinity nitrate transporter NRT2.1 represses lateral root initiation in response to nutritional cues. Proc Nat Acad Sci USA 102:13693–13698
Loque D, Tillard P, Gojon A, Lepetit M (2003) Gene expression of the NO3 − transporter NRT1.1 and the nitrate reductase NIA1 is repressed in Arabidopsis roots by NO2 −, the product of NO3 − reduction. Plant Physiol 132:958–967
McClure PR, Kochian LV, Spanswick RM, Shaff JE (1990a) Evidence for cotransport of nitrate and protons in maize roots: I. effects of nitrate on the membrane potential. Plant Physiol 93:281–289
McClure PR, Kochian LV, Spanswick RM, Shaff JE (1990b) Evidence for cotransport of nitrate and protons in maize roots: II. measurement of NO3 − and H+ fluxes with ion-selective microelectrodes. Plant Physiol 93:290–294
Miller AJ, Smith SJ (1996) Nitrate transport and compartmentation in cereal root cells. J Exp Bot 47:843–854
Miller AJ, Fan XR, Orsel M, Smith SJ, Wells DM (2007) Nitrate transport and signalling. J Exp Bot 58:2297–2306
Nocito FF, Pirovano L, Cocucci M, Sacchi GA (2002) Cadmium-induced sulfate uptake in maize roots. Plant Physiol 129:1872–1879
Nocito FF, Espen L, Crema B, Cocucci M, Sacchi GA (2008) Cadmium induces acidosis in maize root cells. New Phytol 179:700–711
Okamoto M, Kumar A, Li WB, Wang Y, Siddiqi MY, Crawford NM, Glass ADM (2006) High-affinity nitrate transport in roots of Arabidopsis depends on expression of the NAR2-like gene AtNRT3.1. Plant Physiol 140:1036–1046
Orsel M, Eulenburg K, Krapp A, Daniel-Vedele F (2004) Disruption of the nitrate transporter genes AtNRT2.1 and AtNRT2.2 restricts growth at low external nitrate concentration. Planta 219:714–721
Orsel M, Chopin F, Leleu O, Smith SJ, Krapp A, Daniel-Vedele F, Miller AJ (2006) Characterization of a two-component high-affinity nitrate uptake system in Arabidopsis. Physiology and protein–protein interaction. Plant Physiol 142:1304–1317
Ouariti O, Gouia H, Ghorbal MH (1997) Responses of bean and tomato plants to cadmium: growth, mineral nutrition, and nitrate reduction. Plant Physiol Biochem 35:347–354
Pinton R, Cesco S, Iacolettig G, Astolfi S, Varanini Z (1999) Modulation of NO3 − uptake by water-extractable humic substances: involvement of root plasma membrane H+ATPase. Plant Soil 215:155–161
Prosser IM, Purves JV, Saker LR, Clarkson DT (2001) Rapid disruption of nitrogen metabolism and nitrate transport in spinach plants deprived of sulphate. J Exp Bot 52:113–121
Remans T, Nacry P, Pervent M, Girin T, Tillard P, Lepetit M, Gojon A (2006) A central role for the nitrate transporter NRT2.1 in the integrated morphological and physiological responses of the root system to nitrogen limitation in Arabidopsis. Plant Physiol 140:909–921
Ritz C, Spiess AN (2008) qpcR: an R package for sigmoidal model selection in quantitative real-time polymerase chain reaction analysis. Bioinformatics 24:1549–1551
Sanità di Toppi L, Gabbrielli R (1999) Response to cadmium in higher plants. Environ Exp Bot 41:105–130
Santi S, Locci G, Pinton R, Cesco S, Varanini Z (1995) Plasma-membrane H+-ATPase in maize roots induced for NO3 − uptake. Plant Physiol 109:1277–1283
Santi S, Locci G, Monte R, Pinton R, Varanini Z (2003) Induction of nitrate uptake in maize roots: expression of a putative high-affinity nitrate transporter and plasma membrane H+-ATPase isoforms. J Exp Bot 54:1851–1864
Segel IH (1976) Enzymes. Biochemical calculations: how to solve mathematical problems in general biochemistry, 2nd edn. Wiley, Inc., New York, p 208–323
Siddiqi MY, Glass ADM, Ruth TJ, Rufty TW (1990) Studies of the uptake of nitrate in barley: I. kinetics of 13NO3 − influx. Plant Physiol 93:1426–1432
Stitt M (1999) Nitrate regulation of metabolism and growth. Curr Opin Plant Biol 2:178–186
Tomasi N, Kretzschmar T, Espen L, Weisskopf L, Fuglsang AT, Palmgren MG, Neumann G, Varanini Z, Pinton R, Martinoia E, Cesco S (2009) Plasma membrane H+-ATPase-dependent citrate exudation from cluster roots of phosphate-deficient white lupin. Plant Cell Environ 32:465–475
Tong Y, Zhou JJ, Li ZS, Miller AJ (2005) A two-component high-affinity nitrate uptake system in barley. Plant J 41:442–450
Trevisan S, Borsa P, Botton A, Varotto S, Malagoli M, Ruperti B, Quaggiotti S (2008) Expression of two maize putative nitrate transporters in response to nitrate and sugar availability. Plant Biol 10:462–475
Vidmar JJ, Zhuo D, Siddiqi MY, Schjoerring JK, Touraine B, Glass ADM (2000) Regulation of high-affinity nitrate transporter genes and high-affinity nitrate influx by nitrogen pools in roots of barley. Plant Physiol 123:307–318
Wirth J, Chopin F, Santoni V, Viennois G, Tillard P, Krapp A, Lejay L, Daniel-Vedele F, Gojon A (2007) Regulation of root nitrate uptake at the NRT2.1 protein level in Arabidopsis thaliana. J Biol Chem 282:23541–23552
Yan M, Fan XR, Feng HM, Miller AJ, Shen QR, Xu GH (2011) Rice OsNAR2.1 interacts with OsNRT2.1, OsNRT2.2 and OsNRT2.3a nitrate transporters to provide uptake over high and low concentration ranges. Plant Cell Environ 34:1360–1372
Yong Z, Kotur Z, Glass ADM (2010) Characterization of an intact two-component high-affinity nitrate transporter from Arabidopsis roots. Plant J 63:739–748
Zuchi S, Cesco S, Varanini Z, Pinton R, Astolfi S (2009) Sulphur deprivation limits Fe-deficiency responses in tomato plants. Planta 230:85–94
Acknowledgments
This work was supported by Ministero Italiano dell’Istruzione, dell’Università e della Ricerca (MIUR-PRIN), Regione Friuli-Venezia Giulia (LR 26/05), Regione Lombardia (Fondo per la Promozione di Accordi Istituzionali, project BIOGESTECA 15083/RCC). The manuscript greatly benefited from the detailed and constructive criticism of two anonymous reviewers.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rizzardo, C., Tomasi, N., Monte, R. et al. Cadmium inhibits the induction of high-affinity nitrate uptake in maize (Zea mays L.) roots. Planta 236, 1701–1712 (2012). https://doi.org/10.1007/s00425-012-1729-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-012-1729-4