Abstract
Plant class III peroxidases are involved in numerous responses related to pathogen resistance including controlling hydrogen peroxide (H2O2) levels and lignin formation. Peroxidases catalyze the oxidation of organic compounds using H2O2 as an oxidant. We examined the mechanisms of disease resistance in a transgenic carrot line (P23) which constitutively over-expresses the rice cationic peroxidase OsPrx114 (previously known as PO-C1) and which exhibits enhanced resistance to necrotrophic foliar pathogens. OsPrx114 over-expression led to a slight enhancement of constitutive transcript levels of pathogenesis-related (PR) genes. These transcript levels were dramatically increased in line P23 compared to controls [GUS construct under the control of 35S promoter (35S::GUS)] when tissues were treated with cell wall fragments of the fungal pathogen Sclerotinia sclerotiorum (SS-walls), and to a lesser extent with 2,6-dichloroisonicotinic acid. There was no basal increase in basal H2O2 levels in tissues of the line P23. However, during an oxidative burst response elicited by SS-walls, H2O2 accumulation was reduced in line P23 despite, typical media alkalinization associated with oxidative burst responses was observed, suggesting that OsPrx114 was involved in rapid H2O2 consumption during the oxidative burst response. Tap roots of line P23 had increased lignin formation in the outer periderm tissues, which was further increased during challenge inoculation with Alternaria radicina. Plant susceptibility to a biotrophic pathogen, Erysiphe heraclei, was not affected. Disease resistance to necrotrophic pathogens in carrot as a result of OsPrx114 over-expression is manifested through increased PR transcript accumulation, rapid removal of H2O2 during oxidative burst response and enhanced lignin formation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plant class III peroxidases (Prx) are present in all higher plants and are encoded by a large gene family which can comprise of over 70 genes (Passardi et al. 2005). A large number of peroxidase genes are induced during pathogen challenge. For example, up to ten peroxidase genes were upregulated in Magnaporthe-infected rice (Sasaki et al. 2004). In Arabidopsis plants, seven and three peroxidase genes were upregulated when challenged with Pseudomonas spp. (Mohr and Cahill 2007) and Botrytis cinerea (Chassot et al. 2007), respectively. Plant peroxidases catalyze the oxidation of a range of organic substrates using hydrogen peroxide (H2O2) as an oxidant (Hiraga et al. 2001). As well, many different biochemical functions in plant pathogen defense responses are associated with peroxidases. These include lignin formation (El Mansouri et al. 1999; Lagrimini 1991; Quiroga et al. 2000), xylem wall thickening (Hilaire et al. 2001) and phytoalexin biosynthesis (Kristensen et al. 1999). Peroxidases have also been shown to be involved in the generation of reactive oxygen species (Bestwick et al. 1997; Bindschedler et al. 2006) and scavenging of H2O2 (Baker et al. 2000; Kawaoka et al. 2003). These reports indicate that peroxidases have multiple functions in plant defenses against both biotic and abiotic stresses (Passardi et al. 2005). Since the functions of peroxidases are broad and the physiological reactions they are involved with are complex, it has been difficult to definitely demonstrate the role of peroxidases in plant–pathogen interactions.
In previous reports, a reduction of peroxidase activity in Arabidopsis (Bindschedler et al. 2006) and bell pepper plants (Choi et al. 2007) resulted in reduced H2O2 accumulation and enhanced susceptibility to biotrophic fungal and bacterial pathogens. Similarly, over-expression of peroxidase resulted in increased H2O2 accumulation and enhanced tolerance to biotrophic pathogens in Arabidopsis (Choi et al. 2007), wheat (Schweizer 2008) and tobacco plants (Kim et al. 2008). In contrast, antisense suppression of a tomato peroxidase LePrx06 (previously Ep5C) resulted in increased H2O2 accumulation and enhanced resistance to Pseudomonas syringe pv. tomato (Coego et al. 2005), indicating that LePrx06 may be involved in reduction of H2O2 levels. These reports indicate that different peroxidases function in distinct manners, with certain peroxidases involved in defense against biotrophic pathogens while others may function in necrotrophic interactions.
We previously generated transgenic carrot plants that constitutively over-express the rice OsPrx114 (previously referred to as PO-C1) gene (Wally et al. 2009a), which encodes for a pathogen-inducible cationic class III (PR-9) peroxidase. In rice, this gene was induced to high levels following inoculation with avirulent strains of Xanthomonas oryzae (Young et al. 1995). OsPrx114 encodes a 311 amino acid peptide with a putative N-terminus extracellular localization signal. Extracellular transport was confirmed by immune-localization of inoculated rice (Hilaire et al. 2001) as well as enhanced cell wall bound enzyme fractions in transgenic carrots (Wally et al. 2009a). Currently, there are only short carrot peroxidase protein sequences of 10–17 amino acids which have only limited similarity. The highest degree of amino acid similarity to OsPRx114 found in wheat peroxidase TaPrx103 (Altpeter et al. 2005; Schweizer 2008) and barley HvPrx08 (Johrde and Schweizer 2008). In the transgenic plant transformed with the construct ubi::OsPrx114 referred to as line P23, peroxidase enzyme activity was increased by 3.5-fold in the cell wall ionically bound protein fraction (Wally et al. 2009a). The OsPrx114 gene plays an important role in the early defense responses in resistant rice cultivars and was proposed to be a putative lignin-forming enzyme (Hilaire et al. 2001). In our previous work, the OsPrx114 expressing carrot lines were highly resistant to the foliar necrotrophic pathogens Sclerotinia sclerotiorum and Botrytis cinerea, without showing any visible phenotypic abnormalities (Wally et al. 2009a). This was the first report of peroxidase over-expression in plants which has resulted in disease resistance against necrotrophic pathogens.
In the present study, we explored the possible mechanisms by which OsPrx114 expression could enhance resistance to necrotrophic fungal pathogens in root and foliar tissues by quantifying pathogenesis-related (PR) protein gene expression, as well as H2O2, lignin and phenolic production, in transgenic line compared to control carrot lines transformed with 35S::GUS construct.
Materials and methods
Transgenic carrot plants
Primary transformants of transgenic ‘Nantes Coreless’ carrots constitutively over-expressing the rice cationic peroxidase gene OsPrx114 under the control of the maize ubiquitin promoter, or 35S::GUS control plants containing the 35S::GUS chimera construct developed in previous studies (Wally et al. 2008, 2009a), were grown under greenhouse conditions or maintained in suspension cultures in MS media supplemented with 0.5 mg l−1 2,4-dichlorophenoxyacetic acid (Chen and Punja 2002). Several lines expressing OsPrx114 were resistant to B. cinerea and S. sclerotiorum, line P23 was selected for analysis since this line exhibited the highest levels of pathogen resistance (Wally et al. 2009a).
Chemicals
All chemicals were obtained from Sigma (Oakville, ON, Canada) unless otherwise noted.
Effect of OsPrx114 on PR transcript levels
Total RNA was extracted from lyophilized carrot suspension cultures (~250 mg fresh weight) from the OsPrx114 line P23 and the 35S::GUS line using the monophasic RNA extraction method (Chomczynski and Sacchi 1987). The suspension cultures were treated with 2,6-dichloroisonicotinic acid (INA) (500 μM), methyl-jasmonic acid (JA) (100 μM), H2O2 (500 μM) or S. sclerotiorum cell walls (SS-walls) [100 μg ml−1 dry weight (dw)]. The SS-walls were prepared from 7-day-old S. sclerotiorum cultures grown on potato dextrose broth at room temperature according to published procedures (Tweddell et al. 1994). The lyophilized SS-walls were suspended in H2O (10 mg ml−1 dw) and sterilized by autoclaving. Ten micrograms of total RNA was separated on a formaldehyde-agarose gel, blotted and bound to Hybond-XL membrane (GE healthcare, Buckinghamshire, UK), and hybridized overnight at 65°C with various 32P labeled probes. Defense genes from carrot including: phenylalanine ammonia lyase (DcPAL), DcPR1, DcPR2, DcPR3, DcPR5, as well as DcAct and Dc18S, were generated using RT-PCR on non-transformed carrot cDNA using specific primers (Wally et al. 2009b). Random primers were used to label the specific carrot RT-PCR products using [α-32P] dCTP and Prime-A-gene kit (Promega) following the manufacturers protocols and used as radioactive DNA probes. Blots were washed several times for 20 min each; twice with 2× SSC at 60°C, twice with 1× SSC at 65°C and once with 0.25× SSC at 65°C, all containing 0.1% SDS. The blots were either exposed to X-ray film at −80°C for 3–7 days with an intensifying screen or for quantitative expression of carrot PR genes was assessed by exposing filters to phospho-storage screens (GE) and by subsequently scanning with the phospho-imaging system Si 445 (Molecular Dynamics, CA, USA). The signal was taken as the average of signal intensities normalized against both actin and 18S rRNA and standardized to the time zero points for the 35S::GUS lines. For quantitative analysis, RNA extraction and blotting were repeated a minimum of three times.
Detection of hydrogen peroxide
Qualitative in vivo assessment of H2O2 production in carrot leaves was conducted by placing samples in 3′,3-diaminobenzidine (DAB) solution (1 mg ml−1) overnight (Thordal-Christensen et al. 1997). The chlorophyll was cleared by boiling the leaves in solution of ethanol (95% v/v):glycerol:glacial acetic acid (3:1:1) for up to 45 min.
Elicitation for measurement of oxidative burst, 25 ml of 10-day-old carrot cell cultures were transferred to sterile 100 ml Erlenmeyer flask and agitated on a rotating shaker at 150 rpm. Xylenol orange was used to specifically monitor the production of H2O2 during the oxidative burst response in carrot cell suspension cultures. The hydroperoxides are reduced by ferrous ions in the acid solution to form a ferric–xylenol orange complex that was detected spectrophotometrically at 560 nm (Bindschedler et al. 2001; Gay et al. 1999). The H2O2 measurements were conducted at early time points up until 3 h after elicitation. Inhibitors of the oxidative burst including KCN (1 mM), NaN3 (1 mM), l-cysteine (2.5 mM) (Sariri et al. 2006) and diphenylene iodonium (DPI 50 μM), were added 30 min prior to elicitation. Scavenging potential was measured by spiking the cultures with exogenous H2O2 up to 500 μM, and monitoring the rate of H2O2 reduction.
Quantification of lignin and soluble phenolics
Total lignin was extracted from carrot tissues from the alcohol-insoluble residue (AIR) and preferential solubilization through derivatization of lignin with a modified thiogylcolic acid method (Doster and Bostock 1988). Tap roots were examined for lignin content in the outer 2 mm peel comprised mainly of periderm and phloem tissues, using 0.5 g fresh weight of each tissue type. For suspension cultures, lignin was extracted from 50 mg (dw) of cells. All tissues were homogenized with a tissue polytron in absolute methanol; the cell material was pelleted and resuspended with five washes of methanol. The AIR was dried under vacuum overnight. Approximately 50 mg of AIR was incubated in 5 ml of 10% thiogylcolic acid in 2 N HCl at 95°C for 6 h. The soluble ligno-thioglycolic acid derivative (LTGA) product was precipitated by centrifugation for 10 min in a clinical centrifuge at 800g, and washed with water. The LTGA was suspended in 2 ml of 0.5 N NaOH through gentle shaking for 16 h at room temperature, the extract was cleared by centrifugation and the supernatant was acidified with 0.5 ml of concentrated HCl. The LTGA was precipitated following 4 h incubation at 4°C followed by 10 min centrifugation, and the pellet was washed twice with water before being resuspended in 0.5 ml of 0.5 N NaOH. The LTGA was quantified as A280 readings relative to the reading for the 35S::GUS plants (Doster and Bostock 1988).The experiments were conducted three times with nine replicates for each of the treatments described.
Total soluble phenolics were measured from 100 μl of the methanol-soluble fraction. The extract was incubated with 1 ml of freshly prepared 2% (w/v) NaCO3 for 5 min, to which 25 μl of undiluted Folin–Ciocalteu reagent was added (Vermerris and Nicholson 2006). The sample was mixed thoroughly and incubated for 30 min at 25°C. The absorbance at 750 nm was measured spectrophotometrically. Guaiacol was used to generate a standard curve.
Assessment of disease resistance
Greenhouse-grown carrot roots (20–24 weeks of age) were washed with water and placed in sealed plastic trays lined with moistened paper towels. Mycelial plugs of the fungal pathogen Alternaria radicina (provided by Dr. Barry M. Pryor, University of Arizona, Tucson, USA) from 2-week-old V8 agar cultures were placed evenly along the root (2–3 plugs per root). Lesion area was measured 10 days after inoculation (dai) as mm2 and compared to 35S::GUS lines. Three replicates consisting of four roots each were assayed. The experiment was conducted three times.
For assessment of powdery mildew resistance (Erysiphe heraclei), heavily infected leaves from non-transgenic plants were used as inoculum. Leaves from transgenic and non-transgenic 35S::GUS control plants were harvested, washed in water, and cut into 3–4 cm segments and placed adaxial side up on water agar (6 g l−1) containing benzimidazole (0.1 g l−1) in 100 × 15 mm petri dishes, until 90% of the agar surface was covered. Spores were blown into a settling tower, allowed to settle over the tissues for 5 min, and the dishes were sealed and placed on the laboratory bench 22–25º C for 10 days. Newly formed sporulating colonies were counted per dish at 7 and 10 days, using a stereo dissecting microscope. The experiment was conducted three times, consisting of ten plates for each replicate.
Statistical analysis
Treatments were analyzed for significant differences using one-way ANOVA, followed by Tukey–Kramer HSD test and subsequently compared to the 35S::GUS controls using Dunnett’s control test using the JMP version 7 software (SAS institute 2008). LSD values at α <0.05 were used to determine significance.
Results
Effect of OsPrx114 on defense gene transcript levels
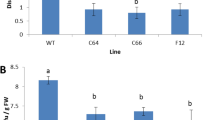
Constitutive transcript levels of genes DcPR1, DcPR2, DcPR3 and DcPR5 in line P23 were elevated compared to the 35S::GUS control levels (Table 1). There was a marked increase in the transcript levels of DcPR1, DcPR2, DcPR3 and DcPR5 in cell cultures elicited with purified cell wall extracts of S. sclerotiorum (SS-walls), with no significant increase in DcPAL levels, after elicitation (Fig. 1); the maximum expression was observed 12–24 h after elicitation (Fig. 2). The PR genes were induced to a greater extent in the P23 cultures, with up to 30-fold increase in DcPR1(Fig. 2a), 45-fold increase in DcPR2 (Fig. 2b), DcPR3 (Fig. 2c) and a 20-fold increase with DcPR5 (Fig. 2d) in comparison to 18-, 8-, 15- and 4.5-fold maximal induction for DcPR1–DcPR5, respectively in the 35S::GUS controls (Fig. 2a–d). Application of the functional salicylic acid analog 2,6-dichloroisonicotinic acid (INA) (500 μM) or the phytohormone jasmonic acid (JA) (100 μM) enhanced defense gene expression in both P23 and 35S::GUS lines, although the P23 line was more responsive, with more rapid expression of DcPR1, DcPR2, DcPR3 and DcPR5 compared to the 35S::GUS. However, the degree of induction due to INA and JA was much lower than with the SS-walls treatment (Supplementary Figs. S1, S2).
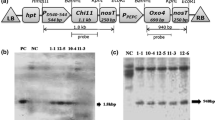
Time-course induction of PR genes in transgenic carrot lines elicited by S. sclerotiorum cell wall fragments. Total RNA from transgenic OsPrx114 expressing line P23 and control 35S::GUS suspension cultures was extracted at 0, 1, 3, 6, 12, 24, 48 and 72 h after treatment with S. sclerotiorum cell walls (100 μg ml−1 dw). Northern blots were probed with the radio-labeled cDNA probes listed. Shown is a representative blot, from five separate replicates
Induction of various carrot pathogenesis-related (DcPR) genes following induction of suspension cultures using purified S. sclerotiorum cell wall fragments (SS-walls) at various time points, comparing OsPrx114 over-expressing P23 line against 35S::GUS control line. Induction was quantified from Northern blots relative to 0 h transcript level for the control lines for a DcPR1, b DcPR2, c DcPR3 and d DcPR5. Transcripts were standardized to both 18S and Actin levels. The experiment was repeated 5 times, and the vertical error bars represent the standard error of the mean
Two non-specific peroxidase inhibitors, NaN3 (1 mM) or KCN (1 mM), were added to the P23 cell cultures to determine if over-expression of peroxidase was responsible for the increase in defense gene transcript levels in transgenic tissues. Overall, defense gene transcript levels after addition of SS-walls were greatly reduced by adding either NaN3 or KCN (Fig. 3).
Time-course induction of PR genes in transgenic P23 carrot lines elicited by S. sclerotiorum cell wall fragments, comparing effect of inhibitors. Total RNA from suspension cultures of transgenic OsPrx114 expressing line P23, with or without 1 mM NaN3 or 1 mM KCN was extracted at 0, 1, 3, 6, 12 and 24 h after treatment with S. sclerotiorum cell walls (100 μg ml−1 dw). Northern blots were probed with the radio-labeled cDNA probes listed
To determine if changes in gene transcript levels were due to increased levels of extracellular H2O2, we investigated the effect of adding exogenous H2O2. 35S::GUS control lines responded weakly to 250 μM, with a slight increase in DcPR1, DcPR2 and DcPR5 expression. Much larger increases in transcript levels of these genes were observed after H2O2 addition in line P23 (Table 2). This increase, however, was less than that observed following SS-walls elicitation (Table 2).
Detection of H2O2
Extracellular peroxidases can generate H2O2 through the hydroxylic cycle, which is an alternative to the normal peroxidative cycle (Passardi et al. 2005). Previous over-expression of a sweet potato peroxidase (IbPrx04) in transgenic tobacco showed high levels of constitutive H2O2 accumulation in leaves following staining with 3′,3-diaminobenzidine (DAB) (Kim et al. 2008). We therefore attempted to determine if H2O2 was enhanced in the P23 transgenic carrot leaves. There was no noticeable difference in the accumulation of H2O2 in leaves (Supplementary Fig. S3) or roots (not shown) of line P23 compared to the 35S::GUS plants. A similar low staining of polymerized DAB was observed near the wound sites in both 35S::GUS and P23 lines.
Control 35S::GUS cells treated with SS-walls at a concentration of 100 mg l−1 exhibited a rapid and reproducible oxidative burst response when assayed with xylenol orange (Fig. 4a). In contrast, when the P23 cells were treated with SS-walls, the oxidative burst response was negligible and there was no apparent increase in H2O2 levels (Fig. 4a).
Biochemical responses in suspension cultures of 35S::GUS controls and transgenic P23 carrot lines. a Treatment with 100 μg ml−1 glucose equivalents of S. sclerotiorum cell wall elicitor. Production of H2O2 was measured using a xylenol orange assay. b Rate of H2O2 consumption following addition of 100 μM H2O2, and 1 mM NaN3 pre-applied to line P23. c Inhibitor treatment with 1 mM NaN3 and diphenylene iodonium (DPI) 50 μM. d Changes in pH of culture media following elicitation with S. sclerotiorum cell wall fragments (100 μg ml−1). Vertical error bars represent the standard error of the mean from 5 replicates from a minimum of independent 4 experiments
To determine if the H2O2 generated through the oxidative burst response was due to activity of peroxidases or NADH-oxidases, inhibitors were applied to the cell cultures. NAD(P)H oxidases are highly sensitive to inhibition by low concentrations of DPI, whereas DPI concentrations greater than 100 μM are required for peroxidase inhibition (Davies et al. 2006). Peroxidase activity was inhibited in P23 by addition of NaN3 and KCN, which do not affect NADH-oxidases. The oxidative burst response was lowered slightly following DPI treatment, with the 35S::GUS cells showing slightly lower extracellular H2O2 across all time points (Fig. 4b). The addition of either NaN3 or KCN resulted in a near abolition of the oxidative burst response, with the H2O2 levels in both 35S::GUS controls and P23 lines detected at basal levels (Fig. 4b). Since both KCN and NaN3 are highly oxidized and non-specific inhibitors of peroxidase, we also examined l-cysteine as an alternative inhibitor (Sariri et al. 2006). l-Cysteine (2.5 mM) reduced the oxidative burst by nearly 80% in the 35S::GUS lines and did not alter the levels found in line P23 (not shown). These findings indicate that the extracellular oxidative burst response in carrot cells was due mainly to peroxidase activity and not NADH-oxidases.
H2O2 scavenging potential
When suspension cultures were initially spiked with 25 μM H2O2, levels of H2O2 were undetectable after 30 s in the P23 line, indicating extremely rapid H2O2 consumption (data not shown). Additional H2O2 experiments were conducted with higher concentrations of H2O2. When 100 μM of H2O2 was applied to the P23 line, the levels were reduced by 80% within 2 min and were at basal levels after 5 min (Fig. 4c). In contrast, the 35S::GUS controls exhibited a more gradual reduction in H2O2, with an 80% reduction after 40 min and a return to basal levels after 50 min. NaN3 was applied to the cells prior to H2O2 application to suppress peroxidase activity and to determine if heightened peroxidase levels were responsible for the increased scavenging. This resulted in the P23 lines removing H2O2 at levels comparable to the 35S::GUS (Fig. 4c), and similar results were obtained using KCN (not shown). The ability of 35S::GUS lines to consume H2O2 was largely unaffected by the presence of either NaN3 or KCN.
To determine if the oxidative burst response was reduced in the P23 cells and was not a consequence of the heightened scavenging ability, we examined the effect on extracellular akalinization in suspension cultures when elicited with SS-walls. The response of line P23 was identical to the 35S::GUS line during the initial 30 min of the oxidative burst response, with a rapid increase in the pH of the growth medium from 5.3 to over 5.8 (Fig. 4d). After the initial phase, the rate of akalinization slowed in the 35S::GUS, peaking at 60 min before gradually decreasing. In contrast, the pH continued to increase in the P23 suspension cultures, peaking at a pH of 6.3 at 120 min and maintaining a pH higher than 6.0 past 200 min (Fig. 4d).
Assessment of disease resistance
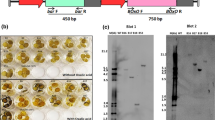
The OsPrx114 over-expressing line P23 was previously found to be highly resistant to the foliar necrotrophic pathogens B. cinerea and S. sclerotiorum (Wally et al. 2009a). Resistance to the root pathogen A. radicina and the biotrophic foliar pathogen E. heraclei was further investigated in this study. The tap roots of line P23 were highly resistant to infection by A. radicina (Fig. 5a), and the total lesion area was reduced by up to 80% compared to the 35S::GUS roots (Fig. 6a). Furthermore, the lesions were largely superficial on the P23 roots, with A. radicina producing mycelia only on the surface of the root (Fig. 5a, b). The degree of fungal penetration was also greatly reduced in the P23 line (Fig. 5b). These differences were very apparent when examined microscopically in roots stained for the presence of lignin (Fig. 5c, d). The 35S::GUS roots had very low levels of lignin (detected with phloroglucinol) and tissue degradation and infection proceeded beyond the periderm (Fig. 5c). The P23 tap roots produced a thick lignin layer directly below the infected area, preventing further penetration and colonization of the tissues (Fig. 5d). There was no fungal mycelium seen below the lignin layer in P23, while a high degree of tissue necrosis with visible mycelia was present beneath the thin lignin layer in the 35S::GUS roots. Interestingly, the peridermal zone of high lignin did not appear to be constitutively formed, since areas away from the lesion did not have detectable lignin using phloroglucinol (not shown). The P23 leaves and petioles exhibited comparable susceptibility towards the biotrophic pathogen E. heraclei as the 35S::GUS tissues (Fig. 5b).
Resistance of transgenic P23 carrot roots to A. radicina 10 days after inoculation, compared to 35S::GUS roots. a Typical large developing lesions in 35S::GUS (upper root), compared to relatively lesion free P23 root (bottom root). b Cross-section through the lesion area. The deep necrotic lesion is apparent in the 35S::GUS (top), while the lesion is fairly superficial on P23 root (bottom). Low magnification micrograph of the 35S::GUS (c) and P23 root (d), taken from the margin of the infected area and stained with phloroglucinol to detect lignin (violet red zone of cells)
Response of transgenic OsPrx114 expressing carrots line P23 to black rot and powdery mildew infection. a Resistance of harvested tap roots to black rot (A. radicina) 10 dai measured as the average total area of each lesion 10 days after inoculation. b Response of the P23 line compared to 35S::GUS on excised leaflets inoculated with E. heraclei spores on water agar plates, based on the number of newly formed sporulating colonies at 7 and 10 days after inoculation. Three replicates of 10 plates were counted for each line. Error bars represent standard error of the means, LSD <0.05
Determination of lignin levels
Lignin levels were measured in the tap roots in the outermost (~2 mm) peel, consisting of periderm and inner layer of vascular tissue comprised mainly of the phloem parenchyma. Additionally, lignin levels were measured in suspension cultures under non-induced and SS-walls treated conditions. The P23 line had 20% greater total lignin levels in the peel under non-induced conditions than the 35S::GUS control line, with no increase in lignin in the phloem tissues (Fig. 7). Lignin levels were increased further in the P23 roots when challenged with A. radicina, in both the peel (by 30%) and phloem tissues (by 50%) (Fig. 7). The 35S::GUS roots had no significant increase in lignin due to infection (Fig. 7). Suspension cells of line P23 had lignin levels similar to that of 35S::GUS cells, both when non-induced or treated with SS-walls after 5 days (data not shown).
Total derivatized lignin-thioglycolic acid complex content in roots of 35S::GUS control and peroxidase over-expressing carrot line P23. Samples were taken from the outer 2 mm peel or from the secondary phloem in uninoculated and inoculated roots 3 days after inoculation with A. radicina. Vertical error bars indicate standard error of the mean from 9 replicate samples for each treatment, LSD <0.05
Since the level of lignification was enhanced in the presence of a pathogen, we also examined the level of soluble phenolic compounds to determine if these were increased or if the levels were limited to lignin deposition during infection. Total soluble phenolics were measured in both 35S::GUS and P23 taproots and leaf tissue, in the absence and presence of the pathogen S. sclerotiorum. There was a slight increase in the phenolic levels during infection; however, the variation was very high and no significant differences were detected between the 35S::GUS and P23 lines (data not shown). Using the suspension culture cells, with defense response elicited by SS-walls, there was also no increase in the phenolic levels in P23 cells compared to 35S::GUS during the first 24 h. However, there was a significant increase in the production of phenolics after 48 h in line P23, which continued to increase to 96 h (Fig. 8).
Discussion
Peroxidases have been reported to have important roles in tolerance to abiotic and biotic stresses, as well in various aspects of plant growth and development, including generating and utilization of H2O2 (Passardi et al. 2005; Johrde and Schweizer 2008; Cosio and Dunand 2009). H2O2 is involved in multiple ways in plant defense responses to pathogens. At low concentrations, H2O2 can function as a signaling molecule, involved in gene regulation and activation of secondary defense pathways (Bolwell 1999; Neill et al. 2002a, b). At higher concentrations, H2O2 has been associated with cell wall modification, the hypersensitive response, or even directly inhibiting pathogens (Lamb and Dixon 1997). However, the role of H2O2 in pathogen defense is complex, since there is evidence to suggest that H2O2 accumulation can be beneficial for colonization of plant tissue by necrotrophic pathogens (Govrin and Levine 2002), while being suppressive to biotrophic pathogens. In contrast, the local production of H2O2 has been associated with enhanced resistance towards the necrotrophic fungus B. cinerea in tomato plants (Asselbergh et al. 2007). Tobacco plants over-expressing a sweet potato peroxidase IbPrx04 had up to fivefold the basal H2O2 levels in leaves (Kim et al. 2008). Similarly, over-expression of a sweet pepper peroxidase CaPrx02 in Arabidopsis was associated with increased H2O2 levels (Choi et al. 2007). In contrast, OsPrx114 over-expressing carrot organs had no detectable increase in H2O2 (Supplemental Fig S3). Apoplastic oxidative burst response is typically controlled either through cell wall bound peroxidases as in French bean (Bindschedler et al. 2001) and Arabidopsis (Bindschedler et al. 2006; Davies et al. 2006) or through the function of NADH-oxidases as in rose cells (Bolwell et al. 1998). The oxidative burst response in carrot tissues elicited by SS-walls appears to be controlled mainly by cell wall bound peroxidases rather than NADH-oxidases, since the burst was effectively eliminated following addition of NaN3, KCN or l-cysteine, and only marginally reduced by adding DPI (Fig. 4). Interestingly, the P23 line had no detectable accumulation of H2O2 in culture media. Extracellular alkalization is a key component of the oxidative burst, independent of H2O2 levels (Bolwell et al. 1998). In tissue-cultured cells of line P23, alkalinization of the medium continued at a longer and slightly stronger rate than in the 35S::GUS line, indicating the oxidative burst response was not inhibited rather there was no accumulation of H2O2 (Fig. 4d). Additionally, cells of P23 were able to rapidly remove high levels of exogenous H2O2, and this ability was effectively eliminated by adding a peroxidase inhibitor (Fig. 4c). These findings may indicate that OsPrx114 over-expression in the P23 line caused a rapid utilization or scavenging of H2O2 generated during the oxidative burst response. OsPrx114 appears to function by rapidly removing the available H2O2 in the apoplastic space through oxidation of available phenolic metabolites rather than as a H2O2 generating peroxidase, operating in a similar fashion to the LePrx06 peroxidase from tomato (Coego et al. 2005).
The signal transduction pathway downstream of H2O2 may be modulated directly or indirectly through the activity of peroxidases, potentially through interactions with extracellular signaling recognition via protein kinases (Lamb and Dixon 1997). Enhanced constitutive expression of many defense-related genes was observed in transgenic tobacco over-expressing a peroxidase gene (Kim et al. 2008). The level of expression was reduced significantly by application of either NaN3 or KCN, which was attributed to a reduction in the overall level of H2O2 in the tobacco cells (Kim et al. 2008). There was a slight constitutive enhancement of defense gene expression in P23 carrot cells, but the level of expression was dramatically increased when the cells were elicited with SS-walls, indicating that these tissues were primed for a more rapid and intense response to pathogens (Table 1). While the addition of NaN3 or KCN lowered the enhanced induction of PR gene transcripts, the transcript levels were still significantly higher than the induced 35S::GUS control levels (Fig. 3). The application of exogenous phytohormones INA, JA (Supplemental figures S1, S2) or H2O2 (Table 2) resulted in marginal increases in transcripts in both 35S::GUS and P23 lines; however, the extent of transcript induction was a fraction of that observed with the SS-walls. The use of complex oligosaccharides found in fungal cell walls has been shown to induce multi-factorial signaling events in plants, generally resulting in much stronger levels of gene induction than a single hormone (Leitner et al. 2008; Jayaraj et al. 2009) and ultimately providing an induced resistance response.
Lignin is a strong structural polymer which is very difficult for pathogens to penetrate or degrade (Quiroga et al. 2000). OsPrx114 has been identified as a putative lignin-forming peroxidase (Hilaire et al. 2001). Previously, we found that OsPrx114 expressing carrot petioles accumulated 40% more lignin than 35S::GUS controls without pathogen challenge, which further increased to 70% in the presence of S. sclerotiorum (Wally et al. 2009a). Since excess lignin can potentially reduce the palatability of root tissue, the lignin levels of mature tap roots in line P23 were measured and compared to 35S::GUS roots. Lignin levels in the peel of the transgenic root (consisting mainly of periderm) were enhanced similar to that observed in the petioles, both constitutively and following pathogen challenge. However, there was no constitutive increase in the lignin levels in the phloem tissues, with increases only observed during pathogen challenge. Since the peel is generally removed before carrot consumption, the increased lignin in the peel or petioles should not be overly detrimental to the palatability (Fig. 7). There was also no increase in the lignin levels in P23 suspension cultures constitutively or induced with SS-walls, possibly indicating a lack of lignin precursors, which limits the lignin levels in fleshy tissues. Increases in soluble phenolic compounds have been associated with peroxidase over-expression in tobacco (Lagrimini 1991; Kim et al. 2008); however, the mechanism of this induction is unknown. When quantified, the OsPrx114 over-expressing carrot line did not exhibit elevated phenolic content in roots, leaves or suspension cultures. There was a gradual increase in the total phenolic content in P23 suspension cultures after elicitation with SS-walls after 24 h and continuing past 96 h compared to 35S::GUS (Fig. 8). This increase in phenolic levels suggests that peroxidase over-expression may allow for more rapid responses of the phenylpropanoid pathway, rather than constitutive induction.
The taproots of P23 carrots had greatly enhanced resistance to the necrotrophic pathogen A. radicina, showing up to 80% reduction in lesion area (Figs. 5, 6). This level of resistance is high, comparable to foliar resistance to S. sclerotiorum or B. cinerea (Wally et al. 2009a). The susceptibility of line P23 to powdery mildew differs from what was reported in wheat (Altpeter et al. 2005; Johrde and Schweizer 2008), barley (Johrde and Schweizer 2008) and tobacco (Kim et al. 2008), where over-expression of a peroxidase gene led to enhanced resistance to powdery or downy mildews. The P23 line did not show a constitutive increase in H2O2 nor the high levels of PR gene expression reported in transgenic tobacco (Kim et al. 2008). The level of peroxidase expression was also greater in barley and tobacco, in excess of 40-fold (Johrde and Schweizer 2008; Kim et al. 2008) compared to carrot, which may have led to reduced mildew infection, as reported in wheat (Altpeter et al. 2005).
In summary, we have shown that line P23 expressing OsPrx114 had reduced accumulation of H2O2 when undergoing an oxidative burst response. In addition, heightened induced expression of defense-related gene transcripts and taproot lignin levels were observed. These findings suggest that constitutive OsPrx114 expression is involved in multiple signaling and defense responses in carrot tissues.
Abbreviations
- PR:
-
Pathogenesis related
- Prx:
-
Peroxidase
- GUS:
-
β-Glucuronidase
- JA:
-
Methyl-jasmonic acid
- INA:
-
2,6-Dichloroisonicotinic acid
- SS-walls:
-
Sclerotinia sclerotiorum cell wall fragments
- DAB:
-
3′,3-Diaminobenzidine
- AIR:
-
Alcohol-insoluble residue
- LTGA:
-
Ligno-thioglycolic acid derivative
References
Altpeter F, Varshney A, Abderhalden O, Douchkov D, Sautter C, Kumlehn J, Dudler R, Schweizer P (2005) Stable expression of a defense-related gene in wheat epidermis under transcriptional control of a novel promoter confers pathogen resistance. Plant Mol Biol 57:271–283
Asselbergh B, Curvers K, Franca SC, Audenaert K, Vuylsteke M, Van Breusegem F, Hofte M (2007) Resistance to Botrytis cinerea in sitiens, an abscisic acid-deficient tomato mutant, involves timely production of hydrogen peroxide and cell wall modifications in the epidermis. Plant Physiol 144:1863–1877
Baker CJ, Deahl K, Domek J, Orlandi EW (2000) Scavenging of H2O2 and production of oxygen by horseradish peroxidase. Arch Biochem Biophys 382:232–237
Bestwick CS, Brown IR, Bennett MHR, Mansfield JW (1997) Localization of hydrogen peroxide accumulation during the hypersensitive reaction of lettuce cells to Pseudomonas syringae pv. phaseolicola. Plant Cell 9:209–221
Bindschedler LV, Minibayeva F, Gardner SL, Gerrish C, Davies DR, Bolwell GP (2001) Early signalling events in the apoplastic oxidative burst in suspension cultured French bean cells involve cAMP and Ca2+. New Phytol 151:185–194
Bindschedler LV, Dewdney J, Blee KA, Stone JM, Asai T, Plotnikov J, Denoux C et al (2006) Peroxidase-dependent apoplastic oxidative burst in Arabidopsis required for pathogen resistance. Plant J 47:851–863
Bolwell GP (1999) Role of active oxygen species and NO in plant defence responses. Curr Opin Plant Biol 2:287–294
Bolwell GP, Davies DR, Gerrish C, Auh CK, Murphy TM (1998) Comparative biochemistry of the oxidative burst produced by rose and French bean cells reveals two distinct mechanisms. Plant Physiol 116:1379–1385
Chassot C, Nawrath C, Metraux JP (2007) Cuticular defects lead to full immunity to a major plant pathogen. Plant J 49:972–980
Chen WP, Punja ZK (2002) Transgenic herbicide- and disease-tolerant carrot (Daucus carota L.) plants obtained through Agrobacterium-mediated transformation. Plant Cell Rep 20:929–935
Choi HW, Kim YJ, Lee SC, Hong JK, Hwang BK (2007) Hydrogen peroxide generation by the pepper extracellular peroxidase CaPO2 activates local and systemic cell death and defense response to bacterial pathogens. Plant Physiol 145:890–904
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidinium thiocyanate phenol chloroform extraction. Anal Biochem 162:156–159
Coego A, Ramirez V, Ellul P, Mayda E, Vera P (2005) The H2O2-regulated Ep5C gene encodes a peroxidase required for bacterial speck susceptibility in tomato. Plant J 42:283–293
Cosio C, Dunand C (2009) Specific functions of individual class III peroxidase genes. J Exp Bot 60:391–408
Davies DR, Bindschedler LV, Strickland TS, Bolwell GP (2006) Production of reactive oxygen species in Arabidopsis thaliana cell suspension cultures in response to an elicitor from Fusarium oxysporum: implications for basal resistance. J Exp Bot 57:1817–1827
Doster MA, Bostock RM (1988) Quantification of lignin formation in almond bark in response to wounding and infection by Phytophthora species. Phytopathology 78:473–477
El Mansouri I, Mercado JA, Santiago-Domenech N, Pliego-Alfaro F, Valpuesta V, Quesada MA (1999) Biochemical and phenotypical characterization of transgenic tomato plants overexpressing a basic peroxidase. Physiol Plant 106:355–362
Gay C, Collins J, Gebicki JM (1999) Hydroperoxide assay with the ferric-xylenol orange complex. Anal Biochem 273:149–155
Govrin EM, Levine A (2002) Infection of Arabidopsis with a necrotrophic pathogen, Botrytis cinerea, elicits various defense responses but does not induce systemic acquired resistance (SAR). Plant Mol Biol 48:267–276
Hilaire E, Young SA, Willard LH, Mcgee JD, Sweat T, Chittoor JM, Guikema JA, Leach JE (2001) Vascular defense responses in rice: peroxidase accumulation in xylem parenchyma cells and xylem wall thickening. Mol Plant Microbe Interact 14:1411–1419
Hiraga S, Sasaki K, Ito H, Ohashi Y, Matsui H (2001) A large family of class III plant peroxidases. Plant Cell Physiol 42:462–468
Jayaraj J, Rahman M, Wan A, Punja ZK (2009) Enhanced resistance to foliar fungal pathogens in carrot by application of elicitors. Ann Appl Biol 155:71–80
Johrde A, Schweizer P (2008) A class III peroxidase specifically expressed in pathogen-attacked barley epidermis contributes to basal resistance. Mol Plant Pathol 9:687–696
Kawaoka A, Matsunaga E, Endo S, Kondo S, Yoshida K, Shinmyo A, Ebinuma H (2003) Ectopic expression of a horseradish peroxidase enhances growth rate and increases oxidative stress resistance in hybrid aspen. Plant Physiol 132:1177–1185
Kim YH, Kim CY, Song WK, Park DS, Kwon SY, Lee HS, Bang JW, Kwak SS (2008) Overexpression of sweetpotato swpa4 peroxidase results in increased hydrogen peroxide production and enhances stress tolerance in tobacco. Planta 227:867–881
Kristensen BK, Bloch H, Rasmussen SK (1999) Barley coleoptile peroxidases. Purification, molecular cloning, and induction by pathogens. Plant Physiol 120:501–512
Lagrimini LM (1991) Wound-induced deposition of polyphenols in transgenic plants overexpressing peroxidase. Plant Physiol 96:577–583
Lamb C, Dixon RA (1997) The oxidative burst in plant disease resistance. Annu Rev Plant Physiol Plant Mol Biol 48:251–275
Leitner M, Kaiser R, Rasmussen MO, Driguez H, Boland W, Mithofer A (2008) Microbial oligosaccharides differentially induce volatiles and signalling components in Medicago truncatula. Phytochemistry 69:2029–2040
Mohr PG, Cahill DM (2007) Suppression by ABA of salicylic acid and lignin accumulation and the expression of multiple genes, in Arabidopsis infected with Pseudomonas syringae pv. tomato. Funct Integr Genomics 7:181–191
Neill S, Desikan R, Hancock J (2002a) Hydrogen peroxide signalling. Curr Opin Plant Biol 5:388–395
Neill S, Desikan R, Clarke A, Hurst RD, Hancock JT (2002b) Hydrogen peroxide and nitric oxide as signalling molecules in plants. J Exp Bot 53:1237–1247
Passardi F, Cosio C, Penel C, Dunand C (2005) Peroxidases have more functions than a Swiss army knife. Plant Cell Rep 24:255–265
Quiroga M, Guerrero C, Botella MA, Barcelo A, Amaya I, Medina MI, Alonso FJ (2000) A tomato peroxidase involved in the synthesis of lignin and suberin. Plant Physiol 122:1119–1127
Sariri R, Sajedi RH, Jafarian V (2006) Inhibition of horseradish activity by thiol inhibitors. J Mol Liq 123:20–23
Sasaki K, Iwai T, Hiraga S, Kuroda K, Seo S, Mitsuhara I, Miyasaka A et al (2004) Ten rice peroxidases redundantly respond to multiple stresses including infection with rice blast fungus. Plant Cell Physiol 45:1442–1452
Schweizer P (2008) Tissue-specific expression of a defence-related peroxidase in transgenic wheat potentiates cell death in pathogen-attacked leaf epidermis. Mol Plant Pathol 9:45–57
Thordal-Christensen H, Zhang ZG, Wei YD, Collinge DB (1997) Subcellular localization of H2O2 in plants. H2O2 accumulation in papillae and hypersensitive response during the barley-powdery mildew interaction. Plant J 11:1187–1194
Tweddell RJ, Jabaji-hare SH, Charest PM (1994) Production of chitinases and beta-1,3-glucanases by Stachybotrys elegans, a mycoparasite of Rhizoctonia solani. Appl Environ Microbiol 60:489–495
Vermerris W, Nicholson R (2006) Isolation and identification of phenolic compounds, a practical guide. In: Vermerris W, Nicholson R (eds) Phenolic compound biochemistry. Springer, Amsterdan, pp 151–196
Wally O, Jayaraj J, Punja ZK (2008) Comparative expression of beta-glucuronidase with five different promoters in transgenic carrot (Daucus carota L.) root and leaf tissues. Plant Cell Rep 27:279–287
Wally O, Jayaraj J, Punja ZK (2009a) Comparative resistance to foliar fungal pathogens in transgenic carrot plants expressing genes encoding for chitinase, beta-1,3-glucanase and peroxidase. Eur J Plant Pathol 123:331–342
Wally O, Jayaraj J, Punja ZK (2009b) Broad spectrum disease resistance to necrotrophic and biotrophic pathogens in transgenic carrots (Daucus carota L.) expressing an Arabidopsis NPR1 gene. Planta 231:131–141
Young SA, Guo A, Guikema JA, White FF, Leach JE (1995) Rice cationic peroxidase accumulates in xylem vessels during incompatible interactions with Xanthomonas oryzae pv oryzae. Plant Physiol 107:1333–1341
Acknowledgments
Funding for this research was provided by the Natural Sciences and Engineering Research Council of Canada, Discovery Grants program. We thank Terry Holmes, Pacific Forestry Center, Victoria, BC, Canada, for performing the microscopy work and Dr. Jayaraj Jayaraman for designing primers and subcloning cDNA fragments.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wally, O., Punja, Z.K. Enhanced disease resistance in transgenic carrot (Daucus carota L.) plants over-expressing a rice cationic peroxidase. Planta 232, 1229–1239 (2010). https://doi.org/10.1007/s00425-010-1252-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-010-1252-4