Abstract
MicroRNAs (miRNAs) are a class of non-coding small endogenous RNAs with lengths of ~22 nucleotides (nt) that have been shown to regulate gene expression at the post-transcriptional levels by targeting mRNAs for degradation or by inhibiting protein translation. Although thousands of miRNAs have been identified in many species, miRNAs have not yet been identified in switchgrass (Panicum virgatum), one of the most important bioenergy crops in the United States and around the world. In this study, we identified 121 potential switchgrass miRNAs, belonging to 44 families, using a well-defined comparative genome-based computational approach. We also identified miRNA clusters and antisense miRNAs in switchgrass expressed sequences tags. These identified miRNAs potentially target 839 protein-coding genes, which can act as transcription factors, and take part in multiple biological and metabolic processes including sucrose and fat metabolism, signal transduction, stress response, and plant development. Gene ontology (GO) analysis, based on these targets, showed that 527 biological processes were involved. Twenty-five of these processes were demonstrated to participate in the metabolism of carbon, glucose, starch, fatty acid, and lignin and in xylem formation. According to pathway enrichment analysis based on Kyoto Encyclopedia of Genes and Genomes (KEGG), 118 metabolism networks were found. These networks are involved in sucrose metabolism, fat metabolism, carbon fixation, hormone regulation, oxidative stress response, and the processing of other secondary metabolites.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
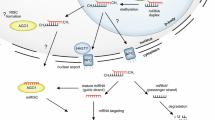
MicroRNAs (miRNAs) are an extensive class of non-coding small endogenous RNAs with ~22 nt in length that are derived from self-complementary foldback structures of longer precursor sequences (pre-miRNAs) and are generated by Dicer-like 1 (DCL1) in plants (Bartel 2004). Mature miRNAs inhibit gene expression at the post-transcriptional levels by either targeting mRNAs for degradation or inhibiting protein translation. Both processes are accomplished by the complementary base pairing of miRNAs to their target mRNA sequences (Ambros 2004). In plants, for a majority of cases, miRNAs interact with their targets through perfect or near-perfect base pairing and lead to target mRNA degradation (Jones-Rhoades et al. 2006). Increasing evidences have revealed that miRNAs play an important role in a wide range of development processes in plants including cell proliferation, stress response, metabolism, inflammation, and signal transduction (Ambros 2004; Jones-Rhoades et al. 2006; Zhang et al. 2007a).
To date, more than 10,000 miRNAs have been identified from 115 species and deposited in the publicly available database miRBase (Release 14) (Griffiths-Jones et al. 2008). The majority of plant miRNAs have been found in species with fully sequenced genomes including 190 from Arabidopsis thaliana, 234 from Populus trichocarpa, 414 from Oryza sativa, 109 from Zea mays, 140 from Sorghum bicolor, and 108 from Medicago truncatula (Griffiths-Jones et al. 2008). miRNA-related research is continuously growing and miRNAs, along with their functions, are being identified and elucidated using a wide variety of computational tools and experimental methods including direct cloning, deep sequencing, and other approaches. Comparison of miRNAs across multiple plant species has demonstrated that some miRNAs are highly evolutionary conserved from species to species, such as from mosses to higher flowering eudicots in the plant kingdom (Floyd and Bowman 2004; Zhang et al. 2006b). Conservation of miRNA sequences has provided a powerful strategy for identifying miRNAs in other species (Pan et al. 2007; Zhang et al. 2005). Currently, comparative genome-based homolog searches have been used to identify conserved miRNAs in many plant species, including cotton (Zhang et al. 2007b), mustard (Xie et al. 2007), soybean (Zhang et al. 2008), wheat (Jin et al. 2008), corn (Zhang et al. 2006a), tomato (Pilcher et al. 2007), potato (Zhang et al. 2009), citrus (Song et al. 2009), and apple (Gleave et al. 2008).
Switchgrass (Panicum virgatum) is a warm-season perennial grass commonly grown in the midwest and grassland areas in the United States. Switchgrass has recently received extensive attention due to its huge potential in the development of cellulosic biofuels (Bouton 2007). Actually, switchgrass has become one of the most important dedicated bioenergy crops for North America. Its use in the production of liquid fuel, such as ethanol, is thought of being one of the important potential energy alternatives to replace dependence on fossil fuels (Chen and Schnoor 2009; Uppugundla et al. 2009). Switchgrass has an important economic and environmental value. To our knowledge, although progress has been made on switchgrass, there is little knowledge about miRNAs in switchgrass. In this study, we employed a well-defined comparative genome-based homolog search to identify switchgrass miRNAs. We also investigated the potential functions of predicted switchgrass miRNAs, particularly in the formation of switchgrass biomass and in biofuel-related biological and metabolic processes.
Materials and methods
Sequence databases
A total of 1,699 known plant miRNAs were downloaded and used as a reference miRNA set for identifying conserved miRNAs in switchgrass. These miRNAs are all currently available miRNAs deposited in miRBase database (http://www.mirbase.org/, Release 14: Sept 2009) (Griffiths-Jones et al. 2008); these miRNAs come from 29 plant species, including A. thaliana, O. sativa (rice), P. trichocarpa, Brassica napus, soybean (Glycine max), M. truncatula, Physcomitrella patens, Saccharum officinarum, S. bicolor, and Z. mays.
Switchgrass expressed sequences tags (ESTs) and protein databases were obtained from the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/, NCBI). Currently, a total of 436, 535 ESTs and 52 protein sequences are available for switchgrass in the NCBI database. All EST sequences were used for predicting conserved miRNAs as well as for identifying potential miRNA targets.
The GO database was downloaded from the Gene Ontology website (http://www.geneontology.org/GO.downloads.shtml) (Ashburner and Bergman 2005). The KEGG database was obtained from the KEGG website (ftp://ftp.genome.jp/pub/kegg/pathway/) (Kanehisa and Goto 2000).
Software
The alignment tool WATER was employed to identify potential conserved miRNAs and their targets and was downloaded as the EMBOSS package (EMBOSS 6.1.0.1) from the public EMBOSS website (http://emboss.sourceforge.net/) (Smith and Waterman 1981). BLAST 2.2.19 was downloaded from NCBI (ftp://ftp.ncbi.nlm.nih.gov/blast/) and used for removing repeated sequences and protein-coding genes. RNAfold was obtained from Vienna RNA Package 1.8.4 (http://www.tbi.univie.ac.at/~ivo/RNA/index.html) (Zuker and Stiegler 1981; Hofacker 2003). MySQL was used for managing all data and was downloaded from the website (http://www.mysql.com/). Perl scripts were developed for data mining in the identification of miRNAs and their targets.
Identification of conserved miRNAs in switchgrass from EST by homologs search
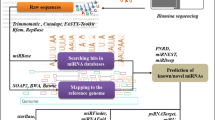
Comparative genome-based EST analysis is a well-established approach to identify conserved miRNAs in one species using already known miRNAs in another species. Since it was developed, EST analysis has been widely used to identify conserved miRNAs in many plant species, including cotton, soybean, oilseed, tomato, and apple. One big issue for current EST analysis is that traditional Blastn searches overlook a lot of potential miRNAs because it is difficult for Blastn searches to align two sequences with deletions, insertions, and gaps within such a small query sequence. Thus, Blastn searches are not an ideal tool to identify conserved small RNA sequences, including miRNAs. To avoid this issue, we adopted WATER to identify potential conserved miRNAs in the bioenergy crop switchgrass.
All of the databases and software were downloaded from the previously mentioned websites. miRNA predication was performed locally using a high performance computer as described in our previous reports. Briefly, we used WATER to align all known mature miRNA sequences to all switchgrass EST sequences in order to identify potential homologs with no >2 nt substitutions, including deletion, and insertion mutations. After removing the repeated and protein-coding sequences and considering proper secondary structure, the sequences only fitting the following criteria were considered potential miRNAs in switchgrass: (1) there were no >2 nt substituted between the EST sequence and the query miRNA sequence; (2) the minimum length of the pre-miRNA was 45 nt; (3) the pre-miRNA could be folded into a perfect stem-loop hairpin secondary structure with the miRNA sitting in one arm of the stem at either the 5′ or 3′ end; (4) there were no more than six nucleotides mismatched between the predicted mature miRNA sequence and its opposite miRNA* sequence in the secondary structure; (5) there were no loops or breaks in the miRNA: miRNA* complex; and (6) the predicted pre-miRNA sequences had a high minimal folding energy (MFE) and MFE index (MFEI). By using these criteria, we could significantly reduce the total number of sequences for subsequent analyses, ultimately saving time and increasing work efficiency. More importantly, the application of these criteria significantly reduced the total number of false miRNA predictions.
Prediction of miRNA targets in switchgrass
Growing evidences have shown that most plant miRNAs function by either perfectly or near-perfectly binding to complementary sites on their target mRNA sequences (Schwab et al. 2005). This provides a powerful way to identify potential miRNA targets simply by aligning and comparing miRNAs with potential target sequences. The criteria for prediction of potential miRNA targets in switchgrass was similar to that described by Schwab and her co-workers but with some modifications (Schwab et al. 2005; Zhang et al. 2008). WATER was employed as an alignment tool to predict miRNA target sequences. Because only a small number of protein-coding genes were reported in switchgrass, all switchgrass ESTs were also used to predict potential miRNA targets. Switchgrass ESTs were Blastx searched against the Arabidopsis protein database in order to identify potential protein-coding homolog genes in switchgrass. Repeated protein-coding sequences with an E value of 1e−25 were removed by additional Blastx searches against the switchgrass protein/EST database and the non-redundant database on NCBI. The predicted target genes with unknown function were discarded. In this research, the following criteria were used for identifying potential miRNA targets: (1) no more than four mismatches were allowed between the mature miRNA and its potential target site; (2) no more than one mismatch was allowed at nucleotide positions 1–9; (3) no more than two consecutive mismatches were allowed; and (4) no mismatches were allowed at positions 10 and 11. These criteria significantly reduced the total number of false-positive targets.
Analysis of GO and KEGG pathway
GO and KEGG pathway analyses were employed to further investigate the biological processes and corresponding metabolic networks regulated by potential miRNAs. We constructed database structures on MySQL, which was used for managing and mining KEGG information. All predicted targets with an E value of 1e−30 were identified by Blastx searching against the GO protein database. GO and pathway analyses were performed by using a combined query search against the GO and KEGG databases. In order to investigate the relationship between the miRNAs and the KEGG pathway, we adopted a formula 1 to convert E values into a relation, which is expressed as an integer from 0 to 16. A large relation number indicates a close relationship. The formula is listed as follows:
where abs is the absolute value of the function, ln is the natural logarithm function having base e in the set, Max is the function of retrieving maximum value of a set, Min is the function of retrieving minimum value of a set, and Int is the function for returning integer of a parameter.
Results
Identifying potential miRNAs in switchgrass
There, currently, are 1,669 miRNAs from 29 plant species deposited in the miRBase database. After removal of the repeated miRNAs, a total of 755 unique miRNA sequences were collected. These 755 unique miRNAs belong to 321 families. In order to identify as many miRNAs as possible in the bioenergy crop switchgrass, we used all currently available 755 unique plant miRNA sequences as queries. After aligning the 755 plant miRNAs with 436,535 switchgrass ESTs, removing the repeated sequences, and considering proper secondary structure, we were able to identify 121 conserved miRNAs in switchgrass, which belong to 44 miRNA families (Table 1). This indicates that miRNAs widely exist in switchgrass and further demonstrates that many miRNAs are highly evolutionary conserved among species in the plant kingdom.
Characterization of microRNAs in switchgrass
The 121 identified switchgrass miRNAs belong to 44 families with an average of about 3 miRNA members per family. The size of a miRNA family varies from family to family; some miRNA families have a larger number of members, but for a majority of families, only one member has been identified (Fig. 1). The miR-444 family has the largest number (13) of members followed by the miR-414 family with 11 members. Both the miR-169 and the miR-2102 families have seven members. There are three miRNA families (miR-156, miR-167, and miR-531) that each contain six members. In this study, for 19 out of the 44 miRNA families, we only identified one member. The remaining 18 miRNA families have 2–4 members, respectively.
miR-444 was originally identified in rice by a direct cloning approach (Sunkar et al. 2005). Currently, miR-444 has only been identified in two other species, wheat (Yao et al. 2007) and Brachypodium distachyon (Unver and Budak 2009), and each species contains only one family member. This suggests that miR-444 only exists in a limited number of plant species and that it is potentially a miRNA specific to monocots. In this study, we identified 13 members of the miR-444 family in switchgrass, which is much larger than the number found in other plant species. Another interesting miRNA is miR-414. Although miR-414 has been identified in Arabidopsis (Wang et al. 2004) and rice (Wang et al. 2004), for more than 6 years miR-414 had not been identified in another plant species except moss (P. patens) (Fattash et al. 2007). Because Arabidopsis and rice belong to two different plant groups (dicots and monocots) and miR-414 was identified in both of these groups as well as in moss, this suggests that miR-414 existed in the plant kingdom several million years ago before the divergence of plants into dicots and monocots. Therefore, miR-414 should be found in almost all plant species. However, it seems that miR-414 is only limited to several specific plant species. In this study, we identified miR-414 in a fourth plant species, switchgrass. Interestingly, miR-414 has more members in switchgrass than in any other species in which miR-414 has been identified. It is unclear what has caused this evolution pattern.
Mature miRNA sequences have been shown to be located on either arm of the secondary stem-loop hairpin structure of the potential pre-miRNA. Of the 121 identified switchgrass miRNAs, 53 (43.8%) were found to be located on the 5′ arm of the stem-loop hairpin structure while 68 (56.2%) resided on the 3′ arm.
The length of switchgrass miRNAs varies from 18 to 24 nt with an average of 20.6 ± 1.0 nt (Fig. 2a). A majority (62 out of 121 or 51.2%) of miRNAs are 21 nucleotides in length. The length of switchgrass pre-miRNA also varies from 46 to 707 nt with an average of 181 ± 135 nt. However, a majority of the pre-miRNAs are 60–139 nt in length (Fig. 2b). The length distribution of miRNAs and their precursor sequences are similar to previous reports in other plant species (Zhang et al. 2006c, 2007b, 2008).
MFE is very important for RNAs forming their secondary structures. Generally speaking, the lower the MFE, the more stable the secondary structure of a RNA sequence. The average value of MFEs was −78.04 ± 60.37 kcal/mol with a range of −10.3 to −395.4 kcal/mol (Fig. 2c). MFEI is a criterion for distinguishing miRNAs from other RNAs. Previous studies have shown that it is more likely to be a potential miRNA if a sequence has a MFEI value >0.85 (Zhang et al. 2006c). For the newly identified 121 switchgrass miRNAs, the average MFEI was 0.86 ± 0.25 with a range of 0.39-1.73 (Fig. 2d).
miRNA clusters in switchgrass
In animals, many miRNAs have been shown to cluster together and have been speculated to have similar expression profiles and functions (Altuvia et al. 2005; Seitz et al. 2004; Tanzer et al. 2005; Tanzer and Stadler 2004; Yu et al. 2006). However, miRNA clusters have rarely been observed in plants. Currently, there are only several clusters have been identified in plants (Jones-Rhoades and Bartel 2004; Talmor-Neiman et al. 2006; Zhang et al. 2006b, 2007b). In this study, due to limited resources, we identified only one miRNA cluster in switchgrass. This cluster includes two miRNAs (miR-2118a and miR-2118b) within the same EST sequence with a distance of 139 nt separating the two (Fig. 3). Based on our best knowledge, this is the first time a cluster involving miR-2118ab has been identified in plants.
miRNA-2118a–2118b cluster in switchgrass EST FL967393. a A schematic diagram of the organization of the cluster. b The EST sequence containing the miRNAs encoded within the cluster. Shadowed sequences represent pre-miRNAs; underlined sequences represent the mature miRNAs. c The predicted secondary structures of miR-2118a and miR-2118b
Antisense miRNAs in switchgrass
Antisense miRNAs are another class of miRNAs that were identified for the first time in invertebrates and vertebrates, including fruit flies and humans (Bender 2008; Stark et al. 2008; Tyler et al. 2008). In this case, miRNAs are transcribed and processed from both sense and antisense transcripts derived from the same genomic loci. In our previous study, we identified five pairs of sense and antisense miRNAs in soybeans and these five pairs belonged to three miRNA families (miR-157, miR-162 and miR-396) (Zhang et al. 2008). In this study, we found one more miRNA, miR-164, with an antisense miRNA in switchgrass (Fig. 4). Since the mature sequence and precursor sequences of sense/antisense miRNAs are different, we propose that they might have different targets or implement their functions through different mechanisms in plants (Zhang et al. 2008).
Target prediction
We adopted more stringent criterion (Schwab et al. 2005) to predict the potential targets of the 121 identified miRNAs in switchgrass. After carefully considering the aligned results, we identified at least one target for each miRNA family and a total of 839 potential targets were predicted. In this study, we identified many miRNA targets that are conserved across several plant species, including Arabidopsis, rice, poplar, cotton, soybean, and corn.
Many studies have demonstrated, by experimental and/or computational approaches, that miRNAs target many transcription factors that help control plant development. We also found this class of targets in switchgrass. MYB transcription factors represent a family of proteins with a conserved MYB DNA-binding domain. This domain was considered to be involved in regulation of secondary metabolism, control of cellular morphogenesis, and regulation of meristem formation and the cell cycle (Jin and Martin 1999). Our results show that MYB proteins might be the target of miR-156, miR-166, miR-414, and miR-2102 in switchgrass. The general transcription factor TFIID, comprised of the TATA-binding protein (TBP) and a set of 13–14 TBP-associated factors, is employed to the promoters of active, and possibly repressed, genes (Cler et al. 2009). In switchgrass, we predict that miR-2102 directs the regulation of TFIID. Aside from WRKY, MYB, and TFIID, there are several transcription factor groups that have been detected to be targets of miRNAs, like GATA transcription factor (miR-414, miR-2102), BHLH transcription factor (miR-2102), and Zinc finger protein (miR-531, miR-2102, miR-2911).
Current studies suggest that miRNAs are involved in plant response to environmental stresses, such as drought, salinity, low temperature, and nutrient deficiency (Chiou 2007; Jagadeeswaran et al. 2009; Jones-Rhoades and Bartel 2004; Jung and Kang 2007; Lu et al. 2005; Sunkar et al. 2006, 2007; Sunkar and Zhu 2004; Zhang et al. 2005). Several miRNAs also target stress-inducible genes. In this study, we identified that 63 stress-related genes were potential targets of 18 miRNA families, including miR-156, miR-167, miR-169, miR-171, miR-397, miR-408, miR-414, miR-477, miR-531, miR-831, miR-854, miR-1132, miR-1436, miR-1535, miR-1858, miR-2102, miR-2118, and miR-2911. The target genes identified are translated in response to such environmental stresses as heavy metal, high-salinity, pathogen infection, endogenous oxidation stress, and apoptosis. Once a plant suffers stress, several physiological regulation mechanisms are enacted in order to promote homeostasis, like hormone-related signal transduction (Jia et al. 2009). Much evidence has shown that miRNAs play a crucial role in hormone signal transduction. For instance, miR-398 has been shown to be involved in ABA-meditated salt resistance response in Arabidopsis (Jia et al. 2009). Our results show that six miRNAs might target 18 switchgrass genes that are involved in signal transduction of auxin, ethylene, brassinosteroid, salicylic acid, gibberellins, and zeatin.
Because switchgrass is one of the most important bioenergy crops for cellulosic ethanol biofuels, we also asked whether miRNAs target traits relating to biofuel production in addition to those necessary for plant development and the creation of biomass. In this study, we did find that 14 miRNAs potentially target 39 genes that are related to the biofuel regulation network. These miRNA targets genes include those for cellulose and sucrose biosynthesis as well as the genes for glycosyltransferase, xylanase, plygalacturonase, and a sucrose transporter. All of these genes play an important role in cellulose and carbohydrate biosynthesis.
Discussion
Identification of conserved miRNAs in switchgrass
In this study, we identified 121 potential switchgrass miRNAs, belonging to 44 families from a total of 436,535 currently available switchgrass ESTs. This suggests that ~0.0277% of switchgrass ESTs contain potential miRNAs. The ratio is much higher than the previously reported 0.0175% for soybean and 0.010% for other plant species (Zhang et al. 2006b, 2008). Several reasons may contribute to this high ratio for identifying conserved miRNAs from switchgrass EST sequences. One reason is that we employed WATER alignment tool for the first time instead of Blastn alignment tool to identify potential conserved miRNAs. In previous studies, researchers have employed Blastn searches in order to identify potential conserved miRNAs. This is because the Blastn search method is easy to use and people can directly search homologs using the NCBI website. Also, it saved time to use Blastn searches because many problems, such as word-size limitation and point mutants, were avoided when analyzing sequences. Blastn search, however, is more favorable to homolog searches with long sequences as search results with shorter sequences are more time consuming to analyze and the results tend to exclude insertion, deletion, and point mutations. All of the shortcomings of the Blastn search tool can be overcome by the WATER program. WATER uses Smith-Water algorithm tool, which ensures optimal local alignment by exploring all possible alignments and selecting the best (Smith and Waterman 1981). This means that deletion, insertion, and substitution of base alignment between known mature miRNAs and EST sequences can be considered, ultimately enhancing identification efficiency (Zhang et al. 2008). One issue with using the Smith-Water algorithm for alignment is that it is time-consuming with a low program-running speed. The WATER program, however, was able to identify a larger number of potential miRNAs; this could be due to the optimal local alignment feature, which enhances the efficiency of miRNA identification. Another reason for an increase in switchgrass miRNA identification is that we used all known plant miRNAs as a reference set; this would allow for the prediction of more conserved miRNAs using our method. New technologies, such as high through-put sequencing, are gradually becoming more and more used for identifying novel and conserved miRNAs in plants and over the next few years, the total number of plant miRNAs in the miRBase is expected to rise. The WATER program provides an inexpensive and reliable method that can continue to identify potentially conserved miRNAs in other plant species.
Analysis of potential miRNA targets in switchgrass
Switchgrass is considered to be a main biofuel feedstock and its cellulosic material is normally digested with industrial enzymes in order to facilitate production into ethanol. (Uppugundla et al. 2009). The synthesis of cellulose is a complicated process that depends on carbon fixation, sugar metabolism and transit, and fat metabolism (Peng et al. 2002; Zhong et al. 2005). In this study, we found that 39 potential switchgrass targets were involved in the biological synthesis of cellulose. 14 miRNAs in switchgrass are predicted to have an effect on the regulation of cellulose biosynthesis (Table 2). For example, our results showed that miR-164 in switchgrass potentially targets five different genes including sucrose synthase 2 and C3 phosphoenolpyruvate carboxylase. This suggests that miR-164 might play a central role in sucrose metabolism and carbon fixation. Among the targets predicted to be involved in cellulose biosynthesis, some important proteins such as cellulose synthase, glycosyltransferase, polygalacturonase, sucrose transporter, and fiber protein Fb34 were detected to be targeted by miR-166, miR-414, miR-531, and miR-2102. Cellulose synthase was predicted to be a target of miR-397 in maize (Zhang et al. 2006a). However, we assumed that miR-166 might regulate cellulose synthase in switchgrass.
To better understand miRNA function, the identified target gene set was subjected to analysis by Gene Ontology (GO), which is viewed as a promising method for uncovering the miRNA-gene regulatory network on the basis of biological process and molecular function (Ashburner and Bergman 2005). In this study, miRNAs were detected in 527 biological processes in switchgrass. Our GO biological process enrichment results (Table 3) demonstrate that a total of 19 miRNA families might be involved in 25 biofuel-related metabolic pathways. Ultimately, this data can be used for improving biofuel production from switchgrass. The data also indicates that we can employ miRNA regulation mechanisms to interfere with cellulose biosynthesis genes ranging from carbon fixation to sucrose metabolism. Following GO analysis, we used KEGG to create a pathway enrichment of predicted miRNA target genes. 118 metabolism networks were found to be involved (data not shown). The results reveal that many miRNA family groups of switchgrass take part in the same pathways including amino acid metabolism, carbon metabolism, fatty acid metabolism, sucrose metabolism, sulfur metabolism, and others (Fig. 5). It suggests that several miRNA families co-participate in the same pathways and may play an important role in complicated metabolic pathways by interacting with each other or other components. In this pathway analysis, miR-414, miR-444, miR-531, miR-1535, miR-1848, miR-2102, and miR-2911 might regulate starch and sucrose metabolism in the same pathway network. Carbon fixation in photosynthesis might be collectively regulated by miR-164, miR-414, miR-1535, miR-2102, and miR-2911. These data suggest that biomass yields and photosynthetic efficiency can be improved by regulation of one or all of the miRNAs in a group. Furthermore, there were also several miRNA regulatory groups believed to be involved in fatty acid metabolism, gluconeogenesis, and nitrogen metabolism. Obviously, these results provide a novel idea of utilizing miRNA regulator groups for controlling biomass yields for biofuel production in switchgrass.
KEGG pathways among partial predicted targets of the identified miRNAs in switchgrass. Each column corresponds to a pathway and each row refers to a group of genes targeted by a specific miRNA. Red was classified into 17 kinds. The color content class is calculated by a formula with one parameter of E value from doing Blastx with protein database (“Methods”). Dark red means closer relation between target and pathway as well as a closer relationship with miRNA and pathway
Abbreviations
- DCL1:
-
Dicer-like 1
- EST:
-
Expressed sequence tag
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- miRNA:
-
MicroRNA
- MFE:
-
Minimal folding free energy
- MFEI:
-
Minimal folding free energy index
- nt:
-
Nucleotide
- pre-miRNAs:
-
miRNA precursor
- NCBI:
-
National Center for Biotechnology Information
- RISC:
-
RNA-induced silencing complex
- SBP:
-
Squamosa-promoter-binding protein-like protein
- TAF:
-
TBP-associated factors
- TBP:
-
TATA-binding protein
References
Altuvia Y, Landgraf P, Lithwick G, Elefant N, Pfeffer S, Aravin A, Brownstein MJ, Tuschl T, Margalit H (2005) Clustering and conservation patterns of human microRNAs. Nucleic Acids Res 33:2697–2706
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355
Ashburner M, Bergman CM (2005) Drosophila melanogaster: a case study of a model genomic sequence and its consequences. Genome Res 15:1661–1667
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Bender W (2008) MicroRNAs in the Drosophila bithorax complex. Genes Dev 22:14–19
Bouton JH (2007) Molecular breeding of switchgrass for use as a biofuel crop. Curr Opin Genet Dev 17:553–558
Chen B, Schnoor JL (2009) Role of suberin, suberan, and hemicellulose in phenanthrene sorption by root tissue fractions of switchgrass (Panicum virgatum) seedlings. Environ Sci Technol 43:4130–4136
Chiou TJ (2007) The role of microRNAs in sensing nutrient stress. Plant Cell Environ 30:323–332
Cler E, Papai G, Schultz P, Davidson I (2009) Recent advances in understanding the structure and function of general transcription factor TFIID. Cell Mol Life Sci 66:2123–2134
Fattash I, Voss B, Reski R, Hess WR, Frank W (2007) Evidence for the rapid expansion of microRNA-mediated regulation in early land plant evolution. BMC Plant Biol 7:13
Floyd SK, Bowman JL (2004) Gene regulation: ancient microRNA target sequences in plants. Nature 428:485–486
Gleave AP, Ampomah-Dwamena C, Berthold S, Dejnoprat S, Karunairetnam S, Nain B, Wang Y-Y, Crowhurst RN, MacDiarmid RM (2008) Identification and characterization of primary microRNAs from apple (Malus domestica cv. Royal Gala) expressed sequence tags. Tree Genet Genomes 4:343–358
Griffiths-Jones S, Saini HK, van Dongen S, Enright AJ (2008) miRBase: tools for microRNA genomics. Nucleic Acids Res 36:D154–D158
Hofacker IL (2003) Vienna RNA secondary structure server. Nucleic Acids Res 31:3429–3431
Jagadeeswaran G, Saini A, Sunkar R (2009) Biotic and abiotic stress down-regulate miR398 expression in Arabidopsis. Planta 229:1009–1014
Jia X, Wang WX, Ren L, Chen QJ, Mendu V, Willcut B, Dinkins R, Tang X, Tang G (2009) Differential and dynamic regulation of miR398 in response to ABA and salt stress in Populus tremula and Arabidopsis thaliana. Plant Mol Biol 71:51–59
Jin H, Martin C (1999) Multifunctionality and diversity within the plant MYB-gene family. Plant Mol Biol 41:577–585
Jin WB, Li NN, Zhang B, Wu FL, Li WJ, Guo AG, Deng ZY (2008) Identification and verification of microRNA in wheat (Triticum aestivum). J Plant Res 121:351–355
Jones-Rhoades MW, Bartel DP (2004) Computational identification of plant microRNAs and their targets, including a stress-induced miRNA. Mol Cell 14:787–799
Jones-Rhoades MW, Bartel DP, Bartel B (2006) MicroRNAs and their regulatory roles in plants. Annu Rev Plant Biol 57:19–53
Jung HJ, Kang H (2007) Expression and functional analyses of microRNA417 in Arabidopsis thaliana under stress conditions. Plant Physiol Biochem 45:805–811
Kanehisa M, Goto S (2000) KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Lu SF, Sun YH, Shi R, Clark C, Li LG, Chiang VL (2005) Novel and mechanical stress-responsive microRNAs in Populus trichocarpa that are absent from Arabidopsis. Plant Cell 17:2186–2203
Pan XP, Zhang BH, SanFrancisco M, Cobb GP (2007) Characterizing viral microRNAs and its application on identifying new microRNAs in viruses. J Cell Physiol 211:10–18
Peng L, Kawagoe Y, Hogan P, Delmer D (2002) Sitosterol-beta-glucoside as primer for cellulose synthesis in plants. Science 295:147–150
Pilcher RLR, Moxon S, Pakseresht N, Moulton V, Manning K, Seymour G, Dalmay T (2007) Identification of novel small RNAs in tomato (Solanum lycopersicum). Planta 226:709–717
Schwab R, Palatnik JF, Riester M, Schommer C, Schmid M, Weigel D (2005) Specific effects of microRNAs on the plant transcriptome. Dev Cell 8:517–527
Seitz H, Royo H, Bortolin ML, Lin SP, Ferguson-Smith AC, Cavaille J (2004) A large imprinted microRNA gene cluster at the mouse Dlkl-Gtl2 domain. Genome Res 14:1741–1748
Smith TF, Waterman MS (1981) Identification of common molecular subsequences. J Mol Biol 147:195–197
Song CN, Fang JG, Li XY, Liu H, Chao CT (2009) Identification and characterization of 27 conserved microRNAs in citrus. Planta 230:671–685
Stark A, Bushati N, Jan CH, Kheradpour P, Hodges E, Brennecke J, Bartel DP, Cohen SM, Kellis M (2008) A single Hox locus in Drosophila produces functional microRNAs from opposite DNA strands. Genes Develop 22:8–13
Sunkar R, Zhu JK (2004) Novel and stress-regulated microRNAs and other small RNAs from Arabidopsis. Plant Cell 16:2001–2019
Sunkar R, Girke T, Jain PK, Zhu JK (2005) Cloning and characterization of microRNAs from rice. Plant Cell 17:1397–1411
Sunkar R, Kapoor A, Zhu JK (2006) Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 18:2051–2065
Sunkar R, Chinnusamy V, Zhu JH, Zhu JK (2007) Small RNAs as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci 12:301–309
Talmor-Neiman M, Stav R, Frank W, Voss B, Arazi T (2006) Novel micro-RNAs and intermediates of micro-RNA biogenesis from moss. Plant J 47:25–37
Tanzer A, Stadler PF (2004) Molecular evolution of a microRNA cluster. J Mol Biol 339:327–335
Tanzer A, Amemiya CT, Kim CB, Stadler PF (2005) Evolution of microRNAs located within Hox gene clusters. J Exp Zool B Mol Dev Evol 304B:75–85
Tyler DM, Okamura K, Chung WJ, Hagen JW, Berezikov E, Hannon GJ, Lai EC (2008) Functionally distinct regulatory RNAs generated by bidirectional transcription and processing of microRNA loci. Genes Develop 22:26–36
Unver T, Budak H (2009) Conserved microRNAs and their targets in model grass species Brachypodium distachyon. Planta 230:659–669
Uppugundla N, Engelberth A, Vandhana Ravindranath S, Clausen EC, Lay JO, Gidden J, Carrier DJ (2009) Switchgrass water extracts: extraction, separation and biological activity of rutin and quercitrin. J Agric Food Chem 57:7763–7770
Wang XJ, Reyes JL, Chua NH, Gaasterland T (2004) Prediction and identification of Arabidopsis thaliana microRNAs and their mRNA targets. Genome Biol 5:R65
Xie FL, Huang SQ, Guo K, Xiang AL, Zhu YY, Nie L, Yang ZM (2007) Computational identification of novel microRNAs and targets in Brassica napus. FEBS Lett 581:1464–1474
Yao YY, Guo GG, Ni ZF, Sunkar R, Du JK, Zhu JK, Sun QX (2007) Cloning and characterization of microRNAs from wheat (Triticum aestivum L.). Genome Biol 8:R96
Yu J, Wang F, Yang GH, Wang FL, Ma YN, Du ZW, Zhang JW (2006) Human microRNA clusters: genomic organization and expression profile in leukemia cell lines. Biochem Biophys Res Commun 349:59–68
Zhang BH, Pan XP, Wang QL, Cobb GP, Anderson TA (2005) Identification and characterization of new plant microRNAs using EST analysis. Cell Res 15:336–360
Zhang BH, Pan XP, Anderson TA (2006a) Identification of 188 conserved maize microRNAs and their targets. FEBS Lett 580:3753–3762
Zhang BH, Pan XP, Cannon CH, Cobb GP, Anderson TA (2006b) Conservation and divergence of plant microRNA genes. Plant J 46:243–259
Zhang BH, Pan XP, Cox SB, Cobb GP, Anderson TA (2006c) Evidence that miRNAs are different from other RNAs. Cell Mol Life Sci 63:246–254
Zhang BH, Wang QL, Pan XP (2007a) MicroRNAs and their regulatory roles in animals and plants. J Cell Physiol 210:279–289
Zhang BH, Wang QL, Wang KB, Pan XP, Liu F, Guo TL, Cobb GP, Anderson TA (2007b) Identification of cotton microRNAs and their targets. Gene 397:26–37
Zhang BH, Pan XP, Stellwag EJ (2008) Identification of soybean microRNAs and their targets. Planta 229:161–182
Zhang WW, Luo YP, Gong X, Zeng WH, Li SG (2009) Computational identification of 48 potato microRNAs and their targets. Comput Biol Chem 33:84–93
Zhong R, Pena MJ, Zhou GK, Nairn CJ, Wood-Jones A, Richardson EA, Morrison WH 3rd, Darvill AG, York WS, Ye ZH (2005) Arabidopsis fragile fiber8, which encodes a putative glucuronyltransferase, is essential for normal secondary wall synthesis. Plant Cell 17:3390–3408
Zuker M, Stiegler P (1981) Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res 9:133–148
Acknowledgments
This work was partially supported by a grant from the North Carolina Biotechnology Center (2009-BRG-1202) and a grant from DuPont Science & Engineering.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Xie, F., Frazier, T.P. & Zhang, B. Identification and characterization of microRNAs and their targets in the bioenergy plant switchgrass (Panicum virgatum). Planta 232, 417–434 (2010). https://doi.org/10.1007/s00425-010-1182-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-010-1182-1