Abstract
The promoter of the pepper pathogen-induced membrane protein gene CaPIMP1 was analyzed by an Agrobacterium-mediated transient expression assay in tobacco leaves. Several stress-related cis-acting elements (GT-1, W-box and ABRE) are located within the CaPIMP1 promoter. In tobacco leaf tissues transiently transformed with a CaPIMP1 promoter-β-glucuronidase (GUS) gene fusion, serially 5′-deleted CaPIMP1 promoters were differentially activated by Pseudomonas syringae pv. tabaci, ethylene, methyl jasmonate, abscisic acid, and nitric oxide. The −1,193 bp region of the CaPIMP1 gene promoter sequence exhibited full promoter activity. The −417- and −593 bp promoter regions were sufficient for GUS gene activation by ethylene and methyl jasmonate treatments, respectively. However, CaPIMP1 promoter sequences longer than −793 bp were required for promoter activation by abscisic acid and sodium nitroprusside treatments. CaPIMP1 expression was activated in pepper leaves by treatment with ethylene, methyl jasmonate, abscisic acid, β-amino-n-butyric acid, NaCl, mechanical wounding, and low temperature, but not with salicylic acid. Overexpression of CaPIMP1 in Arabidopsis conferred hypersensitivity to mannitol, NaCl, and ABA during seed germination but not during seedling development. In contrast, transgenic plants overexpressing CaPIMP1 exhibited enhanced tolerance to oxidative stress induced by methyl viologen during germination and early seedling stages. These results suggest that CaPIMP1 expression may alter responsiveness to environmental stress, as well as to pathogen infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Adverse environmental conditions such as pathogen and herbivore attack, or high salinity and drought stresses, limit plant growth and drastically reduce plant productivity. Biotic and abiotic stresses trigger depolarization of the plasma membrane and changes in membrane potentials, and plants respond by transmitting defense signals (Gelli et al. 1997; Pike et al. 1998; Krol et al. 2003). Plasma membrane-associated proteins, which are involved in plant homeostasis and stress responses, comprise a subfamily of membrane proteins localized predominantly in organelles such as vacuoles, the Golgi apparatus, the endoplasmic reticulum, and the nucleus. Structurally distinct transmembrane domains have been found in plasma membrane proteins, and their number and location are highly variable among plant species. For example, two transmembrane domains have been found in Arabidopsis NDR1 (Century et al. 1997), and the barley Mlo protein and the cold-regulated wheat COR-413 protein were found to have seven and nine transmembrane domains, respectively (Breton et al. 2003; Bhat et al. 2005). Only transmembrane helix structures are sufficient to address the biological roles of plasma membrane proteins, including COR-413 and wpi6 (Breton et al. 2003; Imai et al. 2005).
Large proteins localized in plasma membranes are involved in plant stress tolerance in adverse environmental conditions (Shi and Zhu 2002; Véry and Sentenac 2002; Hussain et al. 2004; Takano et al. 2005). The rice Xa21 and tomato Cf-9 resistance proteins are involved in the recognition of bacterial avrXa21 and fungal Avr9 proteins, respectively, in plasma membrane to mediate rapid defense signaling. The plasma membrane-localized K+ and Ca2+ channels play a pivotal role in plant nutrition and cell signaling during abiotic stress (Véry and Sentenac 2002). Na+/H+ antiporters localized in plasma membrane and tonoplast are involved in maintaining cellular homeostasis during high salt stress (Shi and Zhu 2002; Shi et al. 2003; Yokoi et al. 2005), and overexpression of these antiporters confers salt tolerance in Arabidopsis (Aspe et al. 1999; Shi et al. 2003). The Arabidopsis P-type ATPase HMA2 and boron transporter BOR1, which regulate transport of these metals in the plasma membrane, are required for heavy metal homeostasis (Hussain et al. 2004; Takano et al. 2005). HMA2 was also suggested to influence cadmium detoxification (Hussain et al. 2004). More recently, Arabidopsis and rice proteins that belong to the integral and peripheral plasma membrane protein family have been identified by proteomics analyses (Alexandersson et al. 2004; Marmagne et al. 2004; Tanaka et al. 2004; Chen et al. 2007). However, the roles of plasma membrane proteins in cellular adaptation and developmental cues in plants are still poorly understood.
Several cis-acting promoter elements are indispensable for the regulation of defense-related gene expression during biotic and abiotic stress. These elements, such as the W-box, the GCC-box, and CRT/DRE, have been identified in stress- and hormone response-related promoters in several plant species and have been investigated by deletion analysis (Eyal et al. 1993; Eulgem et al. 1999, 2000; Shinozaki et al. 2003). The W-box (TTGACC), the ethylene-responsive GCC-box (AGCCGCC), and the salicylic acid-responsive (SA) as-1 element (TGACG) are also involved in plant disease defense (Eyal et al. 1993; Jupin and Chua 1996; Eulgem et al. 2000). The W-box is a binding site for transcription factors in the WRKY family. Ethylene-inducible pathogenesis-related (PR) gene expression is modulated by ERF transcription factors containing the AP2 domain, which specifically interact with a GCC-box characterized in Arabidopsis and tobacco (Ohta et al. 2000; Oñate-Sánchez and Singh 2002). Recently, several pepper (Capsicum annum) PR promoters were isolated and functionally characterized by Agrobacterium-mediated transient expression in tobacco plants (Hong et al. 2005; Jung et al. 2005; Hong and Hwang 2006). However, there is little information about how defense-related promoters of plasma membrane protein genes are regulated in response to pathogen infection and abiotic elicitors.
We previously demonstrated that the pepper CaPIMP1 gene, which encodes a plasma membrane protein, is differentially expressed in leaf tissues during compatible and incompatible interactions with Xanthomonas campestris pv. vesicatoria (Hong et al. 2008). Overexpression of CaPIMP1 also alters resistance to bacterial and oomycete pathogens. In this study, we analyzed CaPIMP1 expression and promoter activation by biotic and abiotic stimuli. We found that CaPIMP1 transcripts accumulated in pepper leaf tissues upon treatment with abiotic defense elicitors. The CaPIMP1 promoter region was essential for gene expression activated by pathogen infection and defense elicitor treatment. CaPIMP1 overexpression (OX) in transgenic Arabidopsis plants altered osmotic and oxidative stress tolerance.
Materials and methods
Plants and growth conditions
Pepper (Capsicum annuum, cv. Nockkwang) seeds were sown in a soil mix (peat moss/perlite/vermiculite, 5/3/2, v/v/v). Plants were grown in a growth room at 25 ± 1°C and 70 μmol photons/m2 s−1 illumination under a 16-h-light/8-h-dark regime. Plants at the 6-leaf stage were used for treatment with various agents. Tobacco (Nicotiana tabacum, cv. Xanthi-nc) seeds were sown in the same soil mix and grown under the same conditions. Tobacco leaves at the 6-leaf stage were used for Agrobacterium-mediated transient gene expression. Wild-type (Col-0) and CaPIMP1-overexpression (OX) transgenic Arabidopsis thaliana seeds (Hong et al. 2008) vernalized for 4 days at 4°C were sown on a potting soil mix (compost soil/perlite/vermiculite, 3/1/1, v/v/v). Arabidopsis plants were raised in a growth chamber at 24°C/19°C (day/night) with a 12-h photoperiod (100 μmol photons/m2 s−1).
Pathogen inoculation
Tobacco leaves infiltrated with Agrobacterium harboring the binary vector pCAMBIA1381 with CaPIMP1 promoter deletion constructs were inoculated with a suspension (2 × 108 cfu/ml) of Pseudomonas syringae pv. tabaci using a syringe without a needle. Control plants were mock-inoculated by infiltration with 10 mM MgCl2. Mock- and bacteria-infected tobacco plants were maintained in a moist chamber at 26°C for 12 h prior to the GUS activity assay.
Isolation of genomic sequence and promoter of the CaPIMP1 gene
Pepper genomic DNA was extracted from leaf tissues following the method of Hong et al. (2000). A genomic fragment containing the CaPIMP1 gene was amplified by PCR using degenerate primers, primer, 5′-CTATTTTAGTTGAATAGACAAAGTGAA-3′ (forward), and 5′-AAACATAATTTCTCGAAACACTG-3′ (reverse), based on the 5′- and 3′-untranslated regions of the CaPIMP1 cDNA. PCR amplification was performed with initial denaturation at 95°C for 2 min followed by 35 cycles of incubation at 95°C for 1 min, 54°C for 30 s, and 72°C for 2 min, with final extension at 72°C for 10 min. PCR products were cloned into the vector pCR2.1-TOPO (Invitrogen). Genomic DNA sequences were aligned and compared with the CaPIMP1 cDNA nucleotide sequence. The Genome Walker Universal Kit (Clontech Laboratories Inc., Palo Alto, CA, USA) was used to isolate the CaPIMP1 promoter region with antisense CaPIMP1-specific primers, 5′-AGCATAAAAGTCCTTAAACTTGATTTTGA-3′ and 5′-AAATGTTTCTGACAAAATTTCATAGTTT-3′ for primary and secondary nested PCR, respectively, according to the manufacturer’s instructions. The generated PCR product was cloned into pCR2.1-TOPO and sequenced. CaPIMP1 promoter sequences were analyzed by the PLACE Web Signal Scan program (Higo et al. 1999).
Promoter deletion-GUS constructs
A CaPIMP1-GUS construct was generated by fusing a CaPIMP1 promoter fragment (from −1193 to −1 bp, where the first nucleotide of the initiating ATG is designated +1) to the coding region of the GUS reporter gene in pCAMBIA 1381. Serially 5′-deleted CaPIMP1-GUS constructs were created by PCR, using the full-length promoter fragment as a template with the reverse oligonucleotide primer VI (5′-CCATGGTTCACTTTGTCTATTCAACTAAA-3′, with a NcoI restriction site at the 5′-end) with five forward oligonucleotides: primer I (5′-GAATTCACTTGTGAGAAATAGTTTGAGT-3′), primer II (5′-GAATTCCTTATTTCTTTCAAAAGCTTA-3′), primer III (5′-GAATTCTATATTCGATCAATATTCAAGAA-3′), primer IV (5′-GAATTCTTAATAGGATGAAAATACATA-3′) or primer V (5′-GAATTCATTATGTTGTTTGAAACAACG-3′), each containing an EcoRI restriction site at the 5′-ends. Each fragment was digested with EcoRI/NcoI and subcloned into EcoRI/NcoI-digested pCAMBIA 1381 to generate five promoter deletion derivatives. All constructs were verified by nucleotide sequencing. Each promoter-GUS fusion construct was introduced into Agrobacterium tumefaciens strain EHA105 via electroporation.
Agrobacterium-mediated transient expression assay
Assays of the CaPIMP1 promoter-GUS constructs were performed in tobacco leaves using the method of Hong et al. (2005). A. tumefaciens EHA105 harboring each of the serially deleted promoter-GUS constructs was grown on yeast extract peptone medium (10 g yeast extract, 10 g Bacto peptone, 5 g NaCl, 15 g agar/l) supplemented with rifampicin (60 μg/ml) and kanamycin (50 μg/ml). Agrobacterium was cultured at 28°C and harvested by centrifugation for 15 min at 6,000×g, resuspended in infiltration media [0.1× MS salts, 0.1× B5 vitamins, 20 mM MOPS, pH 5.4, 1% (w/v) glucose, 2% (w/v) sucrose, 200 μM acetosyringone (Sigma-Aldrich, St Louis, MO)], and adjusted to an OD600 of 0.7. After infiltration of Agrobacterium suspension into abaxial surfaces of tobacco leaves using a syringe without a needle (Kim et al. 2007), the tobacco plants were maintained in a moist chamber at 26°C for 48 h, followed by P. syringae pv. tabaci inoculation and abiotic elicitor treatments for GUS activity analysis.
GUS activity measurement
GUS activity in Agrobacterium-mediated, transiently expressed tobacco leaves was measured as described by Jefferson et al. (1987). Tobacco leaf tissues were homogenized in 1 ml extraction buffer [50 mM NaH2PO4, pH 7.0, 10 mM EDTA, 0.1% Triton X −100, 0.1% (w/v) sodium laurylsarcosine, 10 mM β-mercaptoethanol]. After centrifuging for 10 min at 12,000×g at 4°C, the supernatant was transferred to a fresh microtube. The fluorogenic reaction was carried out in a 1-ml volume with 1 mM 4-methylumbelliferyl-β-d-glucuronide (Duchefa Biochemie, Haarlem, The Netherlands) in the extraction buffer supplemented with a 0.1-ml aliquot of protein extract supernatants. GUS activity was normalized to protein concentration in each of the crude extracts and was expressed as nmol 4-methylumbelliferone min/mg protein. Total protein in sample extracts was quantified using bovine serum albumin as a standard, according to the method of Bradford (1976). The GUS measurement was repeated at least three times with similar results.
Abiotic elicitor treatments
Ethylene treatment was performed by placing the pepper plants in a tight glass chamber, where 5 μl/l ethylene was applied by injecting the gas, via a syringe, through a rubber septum in the chamber. Pepper plants sprayed with 100 μM methyl jasmonate (MeJA) were packed in a vinyl bag. For abscisic acid (ABA) treatment, the pepper plants were removed from soil. The roots were carefully washed with tap water and then soaked in 100 μM ABA. For salicylic acid (SA) treatment, 5 mM SA was foliar-sprayed onto pepper plants. For β-amino-n-butyric acid (BABA) treatment, 20 mM BABA in water was foliar-sprayed onto pepper plants. For wounding stress, the leaves were pricked with a needle. For low-temperature treatment, pepper plants were placed at 4°C. For NaCl treatment, pepper plants grown in plastic pots containing compost soil mix were gently removed from the soil and their roots were immersed in 400 mM NaCl. At various time points, pepper leaves treated with abiotic elicitors were harvested, frozen in liquid nitrogen, and stored at −70°C until used for RNA blot analyses.
To investigate the activation of the CaPIMP1 promoter by treatment with abiotic elicitors, tobacco leaves infiltrated with Agrobacterium harboring CaPIMP1 promoter-GUS constructs were sprayed with 100 μM ABA or 100 μM sodium nitroprusside (SNP). Tobacco plants were sprayed with water as a mock-treatment. To monitor ethylene responsiveness of the CaPIMP1 promoter, 10 μl/l of ethylene gas was injected into a glass chamber containing tobacco plants. Tobacco plants sprayed with 100 μM MeJA were sealed with a transparent plastic bag. Treated tobacco plants were placed in a growth room for 12 h and then immediately frozen in liquid nitrogen for GUS activity assays.
RNA gel blot analysis
Total RNA was isolated from pepper and Arabidopsis using the guanidium-acid phenol method (Chomczynski and Sacchi 1987; Chung et al. 2007) and Trizol reagent (Invitrogen, Carlsbad, CA, USA), respectively. Ten micrograms of total RNA was separated on 1.2% agarose/formaldehyde gels, blotted onto Tropilon-Plus nylon membranes positively charged (Applied Biosystems, Bedford, MA, USA), and hybridized overnight with 14-dCTP-biotin-labeled CaPIMP1 cDNA (accession no. DQ356278) in the hybridization buffer (1 mM EDTA, 7% SDS, 250 mM Na2HPO4, and 5% dextran sulfate) at 65°C. After hybridization, the nylon membranes were washed as previously described (Hong et al. 2005). Biotin was detected via chemiluminescence with CDP-Star substrate according to manufacturer’s protocol (Applied Biosystems, Bedford, MA, USA). The membranes were exposed to X-ray film. All RNA blot analyses were repeated at least three times.
Evaluation of Arabidopsis responses to abiotic elicitors
Arabidopsis seeds sown on basal MS medium containing 400 mM mannitol, 200 mM NaCl, and 2.5 μM ABA were maintained at 4°C for 4 days, and germination (emergence of radicles) was scored daily. Arabidopsis seedlings were grown in 1× MS agar medium supplemented with 1% sucrose in a growth chamber for 4 days after sowing and transferred to 1× MS agar medium supplemented with mannitol, NaCl, or ABA. Arabidopsis seeds sown on MS medium containing 100 μM methyl viologen (MV) were maintained at 4°C for 2–4 days, and germination (emergence of radicles) was scored daily. To monitor seedling development, Arabidopsis seedlings were grown in 1× MS agar medium supplemented with 1% sucrose in a growth chamber for 7 days after sowing and transferred to 1× MS liquid medium supplemented with MV at different concentrations. Germination and seedling growth assays were repeated at least three times with similar results.
Results
Sequence analysis of the CaPIMP1 gene
The CaPIMP1 genomic sequence was isolated and compared with the CaPIMP1 cDNA sequence, which revealed that it contains three exons and two introns of 698 and 859 bp in length (data not shown). All deduced intron/exon junctions possess the consensus GT/AG splice sites. The nucleotide sequence data in this study appear under the accession number DQ356279 in the DDBJ/EMBL/GenBank nucleotide database.
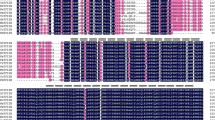
An upstream region including the putative promoter sequence of the CaPIMP1 gene was isolated from pepper genomic DNA, and sequence analysis with the PLACE program revealed several motifs that are found in most eukaryotic promoters for gene expression and regulation (Fig. 1). Potential regulatory elements associated with hormone- and stress-related responses found in other plant promoters were located within the CaPIMP1 promoter: two GT-1 elements, three MYB transcription factor-binding sites, four W-boxes, three ethylene responsive elements (EREs), two ACGT elements, eleven cytokinin-regulated transcription factor ARR1-binding sites, and two gibberellin-responsive elements. The presence of these motifs indicates that CaPIMP1 may be regulated by various cis-acting elements within the promoter as well as corresponding trans-acting factors.
Nucleotide sequence of 5′-flanking promoter regions and putative cis-acting elements of the CaPIMP1 gene. ACGTAterd1, ACGT sequence required for the etiolation-induced expression of erd1 (early responsive to dehydration) in Arabidopsis (Simpson et al. 2003); ARR1At, ARR1-binding element (Sakai et al. 2000; Ross et al. 2004); BP5, OsBP-5 (a MYC protein) binding site in the rice Wx promoter (Zhu et al. 2003); ERE, ethylene-responsive element of the tomato E4 and carnation GST1 genes (Montgomery et al. 1993; Itzhaki et al. 1994); GARE, GA-responsive element (Ogawa et al. 2003); GT1GmSCaM4, GT-1 motif found in the promoter of soybean CaM isoform, SCaM-4 (Park et al. 2004); LeCp, TAAAATAT element in the LeCp (tomato Cys protease) binding cis-element in the LeAcs2 gene (Matarasso et al. 2005); MYB core, binding site for all animal MYBs and the Arabidopsis MYB proteins AtMYB1 and AtMYB2 (Urao et al. 1993); NtBBF1, tobacco Dof protein binding site in the Agrobacterium rhizogenes rolB gene (Baumann et al. 1999); RAV1At, binding consensus sequence for the Arabidopsis transcription factor RAV1 (Kagaya et al. 1999); W-box, binding site for the WRKY transcription factor (Eulgem et al. 2000)
Activation of the CaPIMP1 promoter by bacterial infection and defense signaling molecules
To determine the minimal promoter sequence of the CaPIMP1 gene required for promoter activity, five promoter fragments beginning −1,193, −1,017, −793, −593, and −417 bp upstream of the translational initiation site were fused to the GUS reporter gene (Fig. 2a). Tobacco leaves were infiltrated with Agrobacterium harboring these constructs, and GUS activity expressed in response to bacterial infection and various signal molecules was analyzed by quantitative fluorometry. Twelve hours after inoculation with P. syringae pv. tabaci (Fig. 2b), tobacco leaf tissues harboring the −1,193 bp promoter construct exhibited a threefold higher GUS activity than did mock-inoculated leaves. However, further deletion of the promoter permitted induction of GUS activity in response to P. syringae pv. tabaci.
a Schematic representation of CaPIMP1 promoter constructs for assaying GUS (β-glucuronidase) expression in tobacco leaves. The serially 5′-deleted promoter constructs of the CaPIMP1 gene were fused to the GUS reporter gene in the vector pCAMBIA1381. b CaPIMP1 promoter activation in response to Pseudomonas syringae pv. tabaci infection in tobacco leaf tissues transiently transformed with 5′-CaPIMP1-GUS chimeric constructs. Tobacco leaves were infiltrated with a bacterial suspension of P. syringae pv. tabaci (2 × 108 cfu/ml 10 mM MgCl2) or with 10 mM MgCl2 as a mock-inoculation. GUS activity was analyzed fluorometrically and expressed as nmoles 4-methylumbelliferone (MU)/mg protein min−1. Data are means ± standard deviations from three independent assays of tobacco leaf extracts
Treatments with ethylene, MeJA, ABA, and SNP for 20 h proved sufficient to trigger GUS expression driven by CaPIMP1 promoter constructs (Fig. 3). Ethylene treatment distinctively induced expression driven by all promoter regions between −1,193 and −417 bp. Promoter deletion to −1,017 bp led to a twofold induction of GUS activity. Further deletions to −793 and −593 bp were more effective for ethylene-mediated GUS activation, resulting in 7- and 11-fold increases, respectively. The −417 bp promoter construct showing a 4.5-fold increase also was sufficient for the induction of GUS activity by ethylene. The ethylene-induced GUS activity levels driven by the CaPIMP1 promoter were relatively higher than those by other signal molecules, such as ABA, SNP, and MeJA. All CaPIMP1 promoter fusions except for the −417 bp construct were responsive to MeJA treatment. ABA induced a twofold increase in GUS activity in tobacco leaves harboring the −1,193 bp promoter construct. Deletion to −1,017 and −793 bp regions resulted in roughly threefold increases in GUS activity by ABA treatment. However, significant GUS activity was not observed in ABA-treated tobacco leaves harboring the −593- and −417 bp CaPIMP1 promoter fusions. Treatment with SNP, a nitric oxide donor, also induced GUS expression in tobacco leaves harboring the −1,193, −1,017, and −793 bp regions of the CaPIMP1 promoter. SNP-induced GUS activity gradually decreased following further promoter deletion to −417 bp. GUS activity was abolished by deletion of the CaPIMP1 promoter to −593 and −417 bp.
CaPIMP1 promoter activation in response to ethylene, methyl jasmonate (MeJA), abscisic acid (ABA), and sodium nitroprusside (SNP) applied to tobacco leaf tissues transiently transformed with CaPIMP1-GUS constructs. The numbers over the bars indicate the fold increase in induction of GUS activity after chemical treatment versus mock-treatment. Data are means ± standard deviations from three independent assays of tobacco leaf extracts
CaPIMP1 gene expression in pepper leaves treated with abiotic elicitors
To evaluate the effect of signal molecules on CaPIMP1 expression, ethylene, MeJA, ABA, SA, and BABA were exogenously applied to pepper plants at the 6-leaf stage (Fig. 4). Treatment with ethylene, MeJA, ABA, and BABA activated the CaPIMP1 gene. Transcription began 1 h after ethylene treatment, increased up to 6 h and slightly decreased over 24 h. CaPIMP1 expression was also transiently induced 2–12 h after treatment with MeJA and ABA. CaPIMP1 transcripts were detected 1 h after BABA treatment, with a peak of induction at 2–6 h. To determine whether environmental stresses affect CaPIMP1 expression, pepper plants were exposed to NaCl, wounding, and low temperature. The CaPIMP1 gene was rapidly activated within 1 h after NaCl treatment, and expression drastically declined by 12 h and disappeared by 24 h. The CaPIMP1 gene was markedly expressed within 30 min following mechanical wounding, and thereafter gradually diminished in pepper leaf tissues. In response to cold stress, CaPIMP1 transcripts accumulated in pepper leaves 24 h after low-temperature treatment.
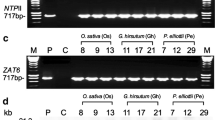
RNA gel blot analysis of CaPIMP1 expression in leaf tissues of pepper plants treated with ethylene, MeJA, ABA, SA, BABA, mechanical wounding, and low temperature. The rRNA in agarose gels was stained with ethidium bromide to show equal loading of RNA. Similar results were obtained in three independent experiments. C healthy controls
Enhanced sensitivity to osmotic stress and ABA of CaPIMP1-OX Arabidopsis
We monitored seed germination and seedling growth of Arabidopsis to examine the responses of transgenic plants to osmotic stress and ABA (Fig. 5). Most (80–90%) wild-type seeds germinated in 400 mM mannitol, 200 mM NaCl, and 2.5 μM ABA within 7 days after sowing, whereas germination of CaPIMP1-OX transgenic seeds was severely reduced under the same conditions (Fig. 5a). In contrast, there was no difference in development for wild-type and transgenic seedlings when they were treated with mannitol, NaCl, or ABA (Fig. 5b).
Enhanced sensitivity to osmotic stresses and abscisic acid (ABA) of CaPIMP1-OX transgenic Arabidopsis plants during germination and early seedling development. a Germination rates of wild-type (WT) and transgenic lines #3, #4 and #5 on 1× MS medium containing 400 mM mannitol, 200 mM NaCl and 2.5 μM ABA 6 days after sowing. Germination was scored when the radicle tips had fully emerged from the seed coats. The data are the mean ± standard deviations of three independent experiments in the evaluation of 100 seeds. b Seedling development of wild-type and transgenic lines on 1× MS agar medium containing 400 mM mannitol, 125 mM NaCl and 10 μM ABA 12 days after sowing
Enhanced tolerance to oxidative stress of CaPIMP1-OX Arabidopsis
Methyl viologen (MV), a redox-cycling herbicide that propagates cellular reactive oxygen species, was used to evaluate the tolerance of CaPIMP1-OX transgenic plants to oxidative stress. Transgenic plants were more resistant than wild-type plants to MV-mediated oxidative stress during the germination and early seedling stages. Following 100 μM MV treatment, the transgenic lines germinated to a higher extent than did the wild-type seeds (Fig. 6a): after 4 days, 75% of the transgenic seeds but only 10% of the wild-type seeds germinated. Lower dosages (0.4–0.8 μM) of MV severely retarded post-germination growth of both wild-type and transgenic plants, and this effect was more pronounced in wild-type plants (Fig. 6b, c). Some wild-type seedlings became bleached and died. Oxidative stress responses of 4-day-old wild-type and transgenic seedlings were not distinctively different after treatment with 0.5 μM MV (Fig. 6d). However, transgenic lines #4 and #5 exhibited slightly higher tolerance to 1.0 μM MV compared to the wild-type plants.
Enhanced tolerance to oxidative stress of CaPIMP1-OX transgenic Arabidopsis plants during germination and early seedling development. a Germination rates of wild-type (WT) and transgenic lines on 1× MS medium containing 100 μM methyl viologen (MV) 6 days after sowing. The data are the means ± standard deviations of three independent experiments in the evaluation of 100 seeds. b Cotyledon formation of wild-type (WT) and transgenic lines on 1× MS medium containing 0.8 μM MV 6 days after sowing. The data are the means ± standard deviations of three independent experiments in the evaluation of 40 seedlings. c Seedling development of wild-type (WT) and transgenic lines on 1× MS medium containing 0.4 μM MV 6 days after sowing. d MV tolerance of seedling plants of transgenic lines. Wild-type and transgenic lines were germinated and grown in 1× MS agar medium in the absence of MV for 4 days. Seedlings were transplanted to liquid medium containing different MV concentrations. Photographs were taken after exposure to MV for 12 days
Discussion
To elucidate the molecular basis of CaPIMP1 gene expression, we analyzed its genomic organization and promoter activity in this study. We further investigated the biological functions of CaPIMP1 during osmotic and oxidative stresses in CaPIMP1-OX transgenic Arabidopsis plants.
CaPIMP1 expression was induced by pathogen and abiotic elicitors. Several putative cis-acting elements, such as the ACGT-box and W-box, were found by computational analysis to reside in the CaPIMP1 promoter, and these elements may be responsible for CaPIMP1 expression by pathogen infection and abiotic elicitors. The −1,193 bp CaPIMP1 promoter was sufficient to drive GUS activity in tobacco leaf tissues infected with P. syringae pv. tabaci. Cis-acting elements essential for activation in response to P. syringae pv. tabaci infection may reside between −1,193 and −1,017 bp. Only a GT-1 element identified in the soybean calmodulin gene promoter activated by pathogen infection and NaCl stress was also found within the CaPIMP1 promoter region from −1,103 to −1,098 bp (Park et al. 2004). The presence of this GT-1 element suggests that it may function in CaPIMP1 promoter activation in response to bacterial infection. Another GT-1 element was also identified at −605 to −599 bp, indicating that this GT-1 element may not be sufficient for CaPIMP1 promoter activation by P. syringae pv. tabaci infection. Three W-boxes and one as-1 element were found within the −1017 CaPIMP1 promoter region. These elements have been suggested to be binding sites for the SA-dependent and pathogen-induced transcription factors WRKY and TGA, respectively (Jupin and Chua 1996; Eulgem et al. 2000). However, promoter constructs containing these cis-acting elements were not activated by P. syringae pv. tabaci infection.
In this study, the minimal promoter region was demonstrated to be differently located for CaPIMP1 activation by abiotic elicitors, ABA, SNP, ethylene, and MeJA. The ABA-responsive, bZIP transcription factor-binding ACGT-box, and EREs were found in the CaPIMP1 promoter region. The −593 bp deletion construct did not respond to ABA treatment, although there are ABA-responsive bZIP and MYB binding sites in this region (Urao et al. 1993, 2000). This observation suggests that the bZIP element identified within the CaPIMP1 promoter may not function in the activation of CaPIMP1 deletion promoters. Three EREs were found in the CaPMIP1 promoter. A 5′-deletion of the CaPMIP1 promoter to −593 bp resulted in a gradual induction of ethylene-responsive promoter activity, indicating that putative cis-acting elements bound by transcriptional repressors may exist between −1,193 and −593 bp. These transcriptional repressors may tightly control CaPIMP1 gene expression by ethylene-mediated signaling. Deletion of the promoter to −417 bp drastically reduced ethylene-responsive promoter activity. An ERE in the −257 bp fragment was sufficient to activate the promoter; however, a lack of putative ERE(s) between −593 and −417 bp responsible for CaPIMP1 promoter activation may reduce promoter activity. Interestingly, the −793 deletion retained the ERE at the −640 bp site, leading to a gradual increase in promoter activity. The GCC-box-like jasmonic acid-responsive element or other jasmonic acid-responsive elements (Menke et al. 1999; Xu and Timko 2004) were not found in the CaPIMP1 promoter region. However, the CaPIMP1 promoter was sufficient for MeJA-induced activation, suggesting that there are unidentified novel jasmonic acid-responsive cis-acting elements in the CaPIMP1 gene promoter region. Synergistic and antagonistic interactions of various cis-acting elements for CaPIMP1 promoter activation remain to be elucidated.
Plasma membrane proteins are involved in the recognition and transduction of endogenous hormonal signals (Blakeslee et al. 2005). CaPIMP1 expression may be dependent on ethylene, MeJA, and ABA. However, SA had no effect on CaPIMP1 gene expression in pepper leaves. Induction of disease resistance-related plasma membrane proteins by plant hormones has not been reported. Cold-regulated plasma membrane protein genes are induced in wheat and rice by ABA treatment (Breton et al. 2003; Imai et al. 2005; Morsy et al. 2005). Inducible CaPIMP1 may be efficient at mediating and enhancing plant defense responses against abiotic stresses.
Environmental stresses, including wounding or exposure to low temperature or high NaCl induced CaPIMP1 expression in pepper plants. Multispanning transmembrane proteins in several plant species have been shown to be regulated by cold stress (Breton et al. 2003). Pathogenesis-related genes isolated from pepper plants were shown to be induced by exogenous hormone treatment and environmental stresses (Jung et al. 2003; Lee and Hwang 2005; Hong and Hwang 2005, 2006). Analysis of transgenic Arabidopsis overexpressing basic PR-1, chitinase, lipid transfer protein, and the Cys2/His2 zinc-finger transcription factor indicates that these pepper pathogenesis-related proteins are involved in environmental stress tolerance (Kim et al. 2004; Hong and Hwang 2005, 2006; Jung et al. 2005; Lee and Hwang 2006). Recently, we found that CaPIMP1 is also rapidly induced by infection with X. campestris pv. vesicatoria, and that CaPIMP1 overexpression in transgenic Arabidopsis alters disease resistance (Hong et al. 2008). These studies support the possibility that the CaPIMP1 protein is also involved in abiotic stress signaling in pepper plants, as well as in disease resistance.
Overexpression of CaPIMP1 in transgenic Arabidopsis results in increased bacterial resistance to P. syringae pv. tomato, but enhanced disease susceptibility to the oomycete biotroph Hyaloperonospora parasitica (Hong et al. 2008), suggesting distinct roles for CaPIMP1 in diverse interactions of pathogens with host plants. Interestingly, ectopic expression of the CaPIMP1 gene in Arabidopsis also caused altered responses to high osmotic stress and oxidative damage during germination and seedling development in this study. CaPIMP1 overexpression in transgenic plants may negatively regulate ABA-related signaling, but positively enhance oxidative stress signaling. Interestingly, negative effect of CaPIMP1-overexpression on ABA-mediated signaling was only shown at the seed germination stage, which may be due to the difference in physiology between germination and seedling growth. It is not evident why overexpression of CaPIMP1 results in increased tolerance to oxidative stress. The CaPIMP1 gene may participate in oxidative burst-mediated disease resistance, which is supported by previous studies of environmental stress perception and of plant antioxidant systems (Foyer and Noctor 2005). Oxidative damage in plants caused by MV may be due to the excess generation of superoxide radicals, which are normally detoxified into oxygen and hydrogen peroxide (H2O2) by superoxide dismutase (Apel and Hirt 2004). Nevertheless, exogenous application of H2O2, causing oxidative stress, did not distinctively affect the germination of CaPIMP1-OX Arabidopsis seeds and early seedling development (data not shown), suggesting that CaPIMP1 may function differently against different sources of reactive oxygen species in plant cells.
In conclusion, we suggest that CaPIMP1 promoter activation by pathogen infection and abiotic elicitor treatment is sufficient to regulate both disease resistance and abiotic stress tolerance in plants. The CaPIMP1 gene may be involved in plant tolerance to a broad spectrum of plant stresses. Further dissection of the CaPIMP1 promoter will reveal the presence of unidentified cis-acting elements for promoter activation by pathogen- and abiotic stimuli. Together with our previous studies of altered disease resistance of CaPIMP1-OX Arabidopsis, these findings emphasize the need to continue elucidating the distinct roles of CaPIMP1 in disease resistance and abiotic stress tolerance.
Abbreviations
- ABA:
-
Abscisic acid
- BABA:
-
β-Amino-n-butyric acid
- ERE:
-
Ethylene-responsive element
- GUS:
-
β-Glucuronidase
- MeJA:
-
Methyl jasmonate
- MV:
-
Methyl viologen
- OX:
-
Overexpression
- PR:
-
Pathogenesis-related
- SA:
-
Salicylic acid
- SNP:
-
Sodium nitroprusside
References
Alexandersson E, Saalbach G, Larsson C, Kjellbom P (2004) Arabidopsis plasma membrane proteomics identifies components of transport, signal transduction and membrane trafficking. Plant Cell Physiol 45:1543–1556
Apel K, Hirt H (2004) Reactive oxygen species: metabolism, oxidative stress, and signal transduction. Annu Rev Plant Biol 55:373–399
Aspe MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by overexpression of a vacuolar Na+/H+ antiporter in Arabidopsis. Science 285:1256–1258
Baumann K, De Paolis A, Costantino P, Gualberti G (1999) The DNA binding site of the Dof protein NtBBF1 is essential for tissue-specific and auxin-regulated expression of the rolB oncogene in plants. Plant Cell 11:323–334
Bhat RA, Miklis M, Schmelzer E, Schulze-Lefert P, Panstruga R (2005) Recruitment and interaction dynamic of plant penetration resistance components in a plasma membrane microdomain. Proc Natl Acad Sci USA 102:3135–3140
Blakeslee JJ, Peer WA, Murphy AS (2005) Auxin transport. Curr Opin Plant Biol 8:494–500
Bradford MM (1976) A rapid sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Breton G, Danyluk J, Charron JBF, Sarhan F (2003) Expression profiling and bioinformatics analyses of a novel stress-regulated multispanning transmembrane protein family from cereals and Arabidopsis. Plant Physiol 132:64–74
Century KS, Shapiro AD, Repetti PP, Dahlbeck D, Holub E, Staskawicz BJ (1997) NDR1, a pathogen-induced component required for Arabidopsis disease resistance. Science 278:1963–1965
Chen F, Yuan Y, Li Q, He Z (2007) Proteomic analysis of rice plasma membrane reveals proteins involved in early defense response to bacterial blight. Proteomics 7:1529–1539
Chomczynski P, Sacchi N (1987) Single-step method of RNA isolation by acid guanidium thiocyanate-phenol-chloroform extraction. Anal Biochem 162:156–159
Chung E, Oh SK, Park JM, Choi D (2007) Expression and promoter analyses of pepper CaCDPK4 (Capsicum annuum calcium dependent protein kinase 4) during plant defense response to incompatible pathogen. Plant Pathol J 23:76–89
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206
Eulgem T, Rushton PJ, Schmelzer E, Hahlbrock K, Somssich IE (1999) Early nuclear events in plant defence signalling: rapid gene activation by WRKY transcription factors. EMBO J 18:4689–4699
Eyal Y, Meller Y, Lev-Yadan S, Fluhr R (1993) A basic-type PR-1 promoter directs ethylene responsiveness, vascular and abscission zone-specific expression. Plant J 4:225–234
Foyer CH, Noctor G (2005) Redox homeostasis and antioxidant signaling: a metabolic interface between stess perception and physiological responses. Plant Cell 17:1866–1875
Gelli A, Higgins VJ, Blumwald E (1997) Activation of plant plasma membrane Ca2+-permeable channels by race-specific fungal elicitors. Plant Physiol 113:269–279
Higo K, Ugawa Y, Iwamoto M, Korenaga T (1999) Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res 27:297–300
Hong JK, Choi DS, Kim SH, Yi SY, Kim YJ, Hwang BK (2008) Distinct roles of the pepper pathogen-induced membrane protein gene CaPIMP1 in bacterial disease resistance and oomycete disease susceptibility. Planta 228:485–497
Hong JK, Hwang BK (2005) Functional characterization of PR-1 protein, β-1, 3-glucanase and chitinase genes during defense response to biotic and abiotic stresses in Capsicum annuum. Plant Pathol J 21:195–206
Hong JK, Hwang BK (2006) Promoter activation of pepper class II basic chitinase gene, CAChi2, and enhanced bacterial disease resistance and osmotic stress tolerance in the CAChi2-overexpressing Arabidopsis. Planta 223:433–448
Hong JK, Jung HW, Kim YJ, Hwang BK (2000) Pepper gene encoding a basic class II chitinase is inducible by pathogen and ethephon. Plant Sci 159:39–49
Hong JK, Lee SC, Hwang BK (2005) Activation of pepper basic PR-1 gene promoter during defense signaling to pathogen, abiotic and environmental stresses. Gene 356:169–180
Hussain D, Haydon MJ, Wang Y, Wong E, Sherson SM, Young J, Camakaris J, Harper JF, Cobbett CS (2004) P-type ATPase heavy metal transporters with roles in essential zinc homeostasis in Arabidopsis. Plant Cell 16:1327–1339
Imai R, Koike M, Sutoh K, Kawakami A, Torada A, Oono K (2005) Molecular characterization of a cold-induced plasma membrane protein gene from wheat. Mol Genet Genomics 274:445–453
Itzhaki H, Maxson JM, Woodson WR (1994) An ethylene-responsive enhancer element is involved in the senescence-related expression of the carnation glutathione-S-transferase (GSTI) gene. Proc Natl Acad Sci USA 91:8925–8929
Jefferson RA, Kavanagh TA, Bevan MW (1987) GUS fusions: β-glucouronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J 13:3901–3907
Jung HW, Kim KD, Hwang BK (2005) Identification of pathogen-responsive regions in the promoter of a pepper lipid transfer protein gene (CALTPI) and the enhanced resistance of the CALTPI transgenic Arabidopsis against pathogen and environmental stresses. Planta 221:361–373
Jung HW, Kim WB, Hwang BK (2003) Three pathogen-inducible genes encoding lipid transfer protein from pepper are differentially activated by pathogens, abiotic, and environmental stresses. Plant Cell Environ 26:915–928
Jupin I, Chua NH (1996) Activation of the CaMV as-1 cis-element by salicylic acid: Differential DNA-binding of a factor related to TGA1a. EMBO J 15:5679–5689
Kagaya Y, Ohmiya K, Hattori T (1999) RAV1, a novel DNA-binding protein, binds to bipartite recognition sequence through two distinct DNA-binding domains uniquely found in higher plants. Nucleic Acids Res 27:470–478
Kim BS, Kim YC, Shin KS, Kim JH (2007) Near-isogenic lines for genes conferring hypersensitive resistance to bacterial spot in chili pepper. Plant Pathol J 23:155–160
Kim SH, Hong JK, Lee SC, Sohn KH, Jung HW, Hwang BK (2004) CAZFP1, Cys2/His2-type zinc-finger transcription factor gene functions as a pathogen-induced early-defense gene in Capsicum annuum. Plant Mol Biol 55:883–904
Krol E, Dziubinska H, Trebacz K (2003) Low-temperature induced transmembrane potential changes in liverwort Conocephalum conicum. Plant Cell Physiol 44:527–533
Lee SC, Hwang BK (2005) Induction of some defense-related genes and oxidative burst is required for the establishment of systemic acquired resistance in Capsicum annuum. Planta 221:790–800
Lee SC, Hwang BK (2006) CASAR82A, a pathogen-induced pepper SAR8.2, exhibits an antifungal activity and its overexpression enhances disease resistance and stress tolerance. Plant Mol Biol 61:95–109
Marmagne A, Roue MA, Ferro M, Rolland N, Alcon C, Joyard J, Garin J, Barbier-Brygoo H, Ephritikhine G (2004) Identification of new intrinsic proteins in Arabidopsis plasma membrane proteome. Mol Cell Proteomics 3:675–691
Matarasso N, Schuster S, Avni A (2005) A novel plant cysteine protease has a dual function as a regulator of 1-aminocyclopropane–1-carboxylic Acid synthase gene expression. Plant Cell 17:1205–1216
Menke FLH, Champion A, Kijne JW, Memelink J (1999) A novel jasmonate- and elicitor-responsive element in the periwinkle secondary metabolite biosynthetic gene Str interacts with a jasmonate- and elicitor-inducible AP2-domain transcription factor, ORCA2. EMBO J 18:4455–4463
Montgomery J, Goldman S, Deikman J, Margossian L, Fischer RL (1993) Identification of an ethylene-responsive region in the promoter of a fruit ripening gene. Proc Natl Acad Sci USA 90:5939–5943
Morsy MR, Almutairi AM, Gibbons J, Yun SJ, de los Reyes BG (2005) The OsLti6 genes encoding low-molecular-weight membrane proteins are differentially expressed in rice cultivars with contrasting sensitivity to low temperature. Gene 344:171–180
Ogawa M, Hanada A, Yamauchi Y, Kuwahara A, Kamiya Y, Yamaguchi S (2003) Gibberellin biosynthesis and response during Arabidopsis seed germination. Plant Cell 15:1591–1604
Ohta M, Ohme-Takagi M, Shinshi H (2000) Three ethylene-responsive transcription factors in tobacco with distinct transactivation functions. Plant J 22:29–38
Oñate-Sánchez L, Singh KB (2002) Identification of Arabidopsis ethylene-responsive element binding factors with distinct induction kinetics after pathogen infection. Plant Physiol 128:1313–1322
Park HC, Kim ML, Kang YH, Jeon JM, Yoo JH, Kim MC, Park CY, Jeong JC, Moon CB, Lee JH, Yoon HW, Lee SH, Chung WS, Lim CO, Lee SY, Hong JC, Cho MJ (2004) Pathogen- and NaCl-induced expression of the SCaM-4 promoter is mediated in part by a GT-1 box that interacts with a GT-1-like transcription factor. Plant Physiol 135:2150–2161
Pike SM, Ádám AL, Pu XA, Hoyos ME, Laby R, Beer SV, Novacky A (1998) Effects of Erwinia amylovora harpin on tobacco leaf cell membranes are related to leaf necrosis and electrolyte leakage and distinct from perturbations caused by inoculated E. amylovora. Physiol Mol Plant Pathol 53:39–60
Ross EJ, Stone JM, Elowsky CG, Arredondo-Peter R, Klucas RV, Sarath G (2004) Activation of the Oryza sativa non-symbiotic haemoglobin-2 promoter by the cytokinin-regulated transcription factor, ARR1. J Exp Bot 55:1721–1731
Sakai H, Aoyama T, Oka A (2000) Arabidopsis ARR1 and ARR2 response regulators operate as transcriptional activators. Plant J 24:703–711
Shi H, Zhu JK (2002) Regulation of expression of the vacuolar Na+/H+ antiporter gene AtNHX1 by salt stress and abscisic acid. Plant Mol Biol 50:543–550
Shi H, Lee B, Wu SJ, Zhu JK (2003) Overexpression of a plasma membrane Na +/H + antiporter gene improves salt tolerance in Arabidopsis thaliana. Nat Biotechnol 21:81–85
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 6:410–417
Simpson SD, Nakashima K, Narusaka Y, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Two different novel cis-acting elements of erd1, a clpA homologous Arabidopsis gene function in induction by dehydration stress and dark-induced senescence. Plant J 33:259–270
Takano J, Miwa K, Yuan L, von Wirén N, Fujiwara T (2005) Endocytosis and degradation of BOR1, a boron transporter of Arabidopsis thaliana, regulated by boron availability. Proc Natl Acad Sci USA 102:12276–12281
Tanaka N, Fujita M, Handa H, Murayama S, Uemura M, Kawamura Y, Mitsui T, Mikami S, Tozawa Y, Yoshinaga T, Komatsu S (2004) Proteomics of the rice cell: systematic identification of the protein populations in subcellular compartments. Mol Gen Genomics 271:566–576
Urao T, Yamaguchi-Shinozaki K, Urao S, Shinozaki K (1993) An Arabidopsis myb homolog is induced by dehydration stress and its gene product binds to the conserved MYB recognition sequence. Plant Cell 5:1529–1539
Urao T, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97:11632–11637
Véry AA, Sentenac H (2002) Cation channels in the Arabidopsis plasma membrane. Trends Plant Sci 7:168–175
Yokoi S, Quintero FJ, Cubero B, Ruiz MT, Bressan RA, Hasegawa PM, Pardo JM (2005) Differential expression and function of Arabidopsis thaliana NHX Na+/H+ antiporters in the salt stress response. Plant J 30:529–539
Zhu Y, Cai XL, Wang ZY, Hong MM (2003) An interaction between a MYC protein and an EREBP protein is involved in transcriptional regulation of the rice Wx gene. J Biol Chem 278:47803–47811
Xu B, Timko M (2004) Methyl jasmonate induced expression of the tobacco putrescine N-methyltransferase genes requires both GCC-box and GCC-motif elements. Plant Mol Biol 55:743–761
Acknowledgments
This research was supported by a grant (CG1133) from the Crop Functional Genomics Center of the 21st Century, Frontier Research Program, funded by the Ministry of Science and Technology, Korea.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequence data reported here has been deposited in the GenBank database under the accession number DQ356279.
Rights and permissions
About this article
Cite this article
Hong, J.K., Hwang, B.K. The promoter of the pepper pathogen-induced membrane protein gene CaPIMP1 mediates environmental stress responses in plants. Planta 229, 249–259 (2009). https://doi.org/10.1007/s00425-008-0824-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-008-0824-z