Abstract
Mutations in MYBPC3 gene, encoding cardiac myosin-binding protein C (cMyBP-C), frequently cause hypertrophic cardiomyopathy (HCM), which affects 0.2 % of the general population. This myocardial autosomal-dominant disorder is the leading cause of sudden cardiac death particularly in young athletes. The current pharmacological and surgical treatments of HCM focus on symptoms relief, but do not address the cause of the disease. With the development of novel strategies targeting the endogenous mutation, causal HCM therapy is now possible. This review will discuss the current knowledge on HCM from the identification of MYBPC3 gene mutations to potential RNA-based correction.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
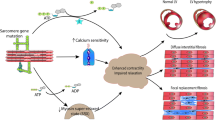
Hypertrophic cardiomyopathy (HCM) is a myocardial disease characterized by left ventricular hypertrophy, mainly involving the interventricular septum, diastolic dysfunction, and myocardial disarray [15]. The prevalence of HCM has been estimated to be 1:500 in the general population [38]. It is therefore the most frequent inherited cardiac disease and the leading cause of sudden cardiac death (SCD), particularly in young athletes [37]. The clinical outcome of HCM varies from a benign asymptomatic course to atrial fibrillation, heart failure, and SCD due to arrhythmias (for detailed reviews, see [14, 21]).
HCM is transmitted in an autosomal-dominant fashion and is caused by more than 1,000 individual mutations in at least 10 genes encoding components of the sarcomere (for detailed recent reviews, see [18, 55]). The MYBPC3 gene encoding cardiac myosin-binding protein C (cMyBP-C) has been assigned on chromosome 11p11.2 by the group of Labeit [19]. As a component of the sarcomere interacting with myosin, titin, and actin, MYBPC3 was the ideal candidate gene for the fourth HCM chromosomal locus (CMH4), which was identified on chromosome 11, using the segregation analysis of microsatellite markers in a large French family, by the group of Schwartz 20 years ago [7]. By screening for MYBPC3 mutations in HCM families, the groups of Schwartz and Seidman simultaneously identified the first three disease mutations [5, 63]. Unexpectedly, these mutations disrupted the reading frame and produced C-terminal truncated cMyBP-C. During the last 2 decades, a body of evidence revealed MYBPC3 as the most frequently mutated HCM gene, representing about 40–50 % of all HCM mutations, and nonsense or frameshift mutations as the most common in MYBPC3 [46, 49, 55].
MYBPC3 mutations and molecular mechanisms
The complete structure and sequence of the human MYBPC3 gene was established in 1997 [6]. It contains more than 21 kbp and is composed of 35 exons of which 34 are coding (Table 1 and Fig. 1a). Exons 10 and 14 are very small (3 bp) and were often not considered, although several pathogenic mutations were found in the flanking introns [16]. Amplifying complementary DNA (cDNA) from human myocardial samples always revealed these small exons.
Structure of human MYBPC3 exons and cMyBP-C protein domains. a Structure of the MYBPC3 exons. MYBPC3 gene consists of 35 exons, of which 34 are coding. The untranslated regions (gray) and the coding regions (light and dark blue) are indicated. Exons in light blue could be skipped without inducing a frameshift, whereas exons in dark blue could not be skipped without introducing a frameshift. The red numbers indicate the sizes (in base pairs) of corresponding exons and introns. Structure was modified from Ensembl. b Structure of cMyBP-C protein and MYBPC3 cDNA. The cMyBP-C protein is composed of 11 domains (C0–C10). Eight are I class immunoglobulin domains (rectangles), and three are fibronectin type III domains (ellipses). The domains of interaction with other sarcomeric proteins (solid black lines) as well as the specific four phosphorylation sites located in the MyBP-C motif (yellow rectangle) are indicated. Correspondence between domains and exons is indicated by dashed lines. Indicated in the cDNA structure are the exons that could be skipped alone (red) and the exons that could be skipped only together to maintain the reading frame (blue). Abbreviation used: PPPP, cMyBP-C motif containing the four phosphorylation sites
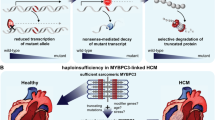
More than 350 HCM-associated mutations (460 in HGMD professional 2013.2; http://www.hgmd.org/) were identified in MYBPC3 (Table 2). Although several missense mutations have been described, which should result in stable mutant proteins that could be incorporated into the sarcomere, 64 % of the known MYBPC3 mutations are frameshift or nonsense (reviewed in [2, 8, 36, 50]). These mutations result in a premature termination codon (PTC) in the transcribed messenger RNA (mRNA) and are expected to produce C-terminal truncated cMyBP-C lacking myosin-binding and/or titin-binding sites [6]. A 25-bp deletion, including the branch point of intron 32, leads to a frameshift due to exon 33 skipping and has been associated with a higher risk of heart failure in 4 % of the population in South Asia [13].
The molecular mechanisms by which MYBPC3 mutations lead to HCM remain elusive. Findings in humans support the view that cMyBP-C haploinsufficiency is the main molecular mechanism, even for patients with missense mutations [39, 40, 44, 53, 59, 60]. Haploinsufficiency is also involved in cat and mouse models of HCM that carry either a missense or a frameshift mutation [43, 61]. Haploinsufficiency has been shown to result from regulations by the nonsense-mediated mRNA decay and/or the ubiquitin–proteasome system (UPS) in mice [61]. Recent findings in HCM mouse models suggest that adrenergic stress or aging induces UPS impairment and potential accumulation of truncated proteins that may act as poison polypeptides [54, 56]. This suggests that the “poison peptide” mechanism could contribute in worsening the HCM phenotype when the UPS is impaired.
RNA-based therapy
The current therapy of HCM focuses on symptoms relief by pharmacological and/or surgical treatments but does not address the cause of the disease (see detailed reviews [17, 58]). With the development of new strategies that target the endogenous gene or pre-mRNA, it is now possible to envision causal therapy for HCM. RNA-based therapeutic approaches such as exon skipping, exon inclusion, and spliceosome-mediated RNA trans-splicing (SMaRT) were developed for neuromuscular disorders during the last decade (see detailed reviews [25, 27, 34]). The development of these approaches for cardiac genetic diseases is very new.
Exon skipping
Exon skipping can be achieved by using antisense oligonucleotides (AONs). AONs are designed to mask exonic splicing enhancer (ESE) motifs and therefore inhibit binding of trans-acting regulatory splicing factors that mediate inclusion of specific exons into the mature mRNA [64]. This is expected to result in the skipping of the targeted exons [25, 64]. An important prerequisite for removing one or more exons is that it should be in-frame and result in an internally deleted (shortened) protein with normal or at least near-normal function, in order to rescue the phenotype.
AONs can be delivered as chemically modified molecules or packaged into adeno-associated virus (AAV). A big hurdle of chemically modified AONs is their delivery into the heart [1, 23]. For example, 2′-O-methyl phosphothioates have shown relatively poor exon skipping efficiency in the heart of dystrophin-deficient mice [3, 22, 28]. On the other hand, the phosphorodiamidate morpholino oligomer (PMO) chemistry has a promising potential for the heart [30]. To improve heart delivery, AONs can also be delivered to the spliceosome machinery by using AAV encoding modified U7 or U1 small nuclear RNA (snRNA) and the appropriate AAV serotype for the best cardiotropism [20, 24, 25].
Out of the 34 coding exons of MYBPC3, 14 exons (2–4, 8–11, 14, 20, 22, and 24–27) are in-frame and therefore may be skipped to remove the mutations (Fig. 1a). If we discard exons involved in an important protein structure or function (such as phosphorylation or protein binding sites), then the exons 2, 22, 24, 25, 26, and 27 are ideal targets for skipping (Fig. 1b). Finally, skipping of the single exon 25 will remove a total of 34 mutations, which represents about 11 % of exonic MYBPC3 mutations (Table 2). It is also possible to skip two or more exons at once to keep the reading frame and the function of the protein (Fig. 1b). We recently published the first proof-of-principle study demonstrating exon skipping in a Mybpc3-targeted knock-in (KI) mouse model of HCM [20]. The Mybpc3-KI mouse carries one of the most frequent human mutations, the G > A transition on the last nucleotide of exon 6 (c.772G > A), which is part of the consensus 5′ splice donor site sequence [61]. This results in three aberrant mRNAs. In addition, we revealed the presence of an alternative splice variant deleted of the exons 5 and 6 (Var-4), which was also expressed at a low level in wild-type mice and very stable after gene transfer in cardiac myocytes [20]. To enhance Var-4 expression, we designed two AONs that mask predicted ESEs in exons 5 and 6 (Fig. 2a). AONs were inserted into U7snRNA and packaged in AAV. Transduction of cardiac myocytes or systemic administration of AAV-U7-AON-5–6 increased Var-4 mRNA/protein levels and reduced aberrant mRNAs. Importantly, injection of newborn KI mice abolished cardiac dysfunction and prevented left ventricular hypertrophy [20].
RNA-based approaches to remove a G > A transition in the last nucleotide of exon 6 of mouse MYBPC3. a Exon skipping strategy to remove the mutated exon 6 together with exon 5 to maintain the reading frame in MYBPC3 mRNA using antisense oligonucleotides (AONs) complementary to exonic splicing enhancer (ESE) motifs (green stripes) in exon 5 (AON-5) and exon 6 (AON-6). b Schematic representation of 5′-trans-splicing bypassing the G > A transition. A pre-trans-splicing molecule (PTM), containing the 5′-splicing site (5′SS) and a specific binding domain (BD) complementary to intron 6, was generated to interfere with the endogenous cis-splicing and to produce full-length repaired MYBPC3 mRNA. Figure adapted from [20, 41]
Exon inclusion
Besides exon skipping, the spliceosome could also be forced to include an exon into the mRNA. This process is called exon inclusion and could be used when the mutation itself is expected to result in exon skipping and frameshift. In this case, AONs are directed against exonic splicing silencer (ESS) or intronic splicing silencer (ISS) motifs, which are normally recognized by splicing repressor proteins such as heterogeneous nuclear ribonucleoprotein A1 [29]. Masking the splicing silencers allows exon recognition by the spliceosome and therefore inclusion of the exon into the mRNA. The efficiency of exon inclusion can be increased by using bifunctional AONs containing an antisense sequence complementary to the silencer plus a potent enhancer sequence to support the binding of splicing enhancer proteins [9].
Exon inclusion application was used successfully in different spinal muscular atrophy (SMA) mouse models as well as in SMA patient-derived fibroblasts [29, 47, 57]. To date, no studies have been published using exon inclusion to treat HCM. Splicing mutations (and in general, intronic mutations leading to exon skipping) are targets for exon inclusion. Out of the 54 described intronic MYBPC3 mutations (Table 2), 40 (11 % of the total) are located in splice sites and are therefore candidates for exon inclusion. The limitation of exon inclusion is that a specific AON has to be generated for every single mutation, whereas for exon skipping, a single AON can be used for patients with different mutations in the targeted exon. Since every single AON is considered as a new drug and has to go through all clinical trials individually, exon skipping is more attractive.
Spliceosome-mediated RNA trans-splicing
Another approach to target mutant pre-mRNA is the spliceosome-mediated RNA trans-splicing (SMaRT) in which two independently transcribed RNA molecules, a target mutant pre-mRNA, and a therapeutic pre-trans-splicing molecule (PTM) are spliced together (for review, see [62]). As a result, a full-length repaired mRNA is formed. Trans-splicing is useful to treat dominant diseases, and it is a suitable alternative to repair a mutant pre-mRNA when exon skipping and/or exon inclusion is not feasible. Trans-splicing is carried out by the endogenous splicing machinery and is restricted to those cells expressing the target pre-mRNA, minimizing the risk of ectopic expression of the repaired mRNA. This approach allows the replacement of endogenous mutations at pre-mRNA level with wild-type coding sequences enclosed in the PTMs. In addition, PTMs should also carry conserved splicing sequences together with a binding domain complementary to the target intronic region. The binding domain is crucial for the specificity of the PTM, and it is assumed that its length positively correlates with its specificity. The feasibility of trans-splicing in vitro has been shown in several studies, most of them using the 3′-trans-splicing approach [45, 51, 52]. Replacement of an internally mutated exon is the most elegant trans-splicing procedure, and its feasibility to repair Duchenne dystrophin transcripts in vitro was recently shown [35]. In the last decade, the therapeutic potentiality of trans-splicing was successfully applied to correct in vivo genetic diseases such as SMA [11, 12], tauopathies [4], and hemophilia A [10], all of them being 3′-trans-splicing approaches. Moreover, in a mouse model of SMA, it was demonstrated that the combination of trans-splicing and AONs to reprogram the mutant SMN2 pre-mRNA is able to lessen the severity of the SMA phenotype, therefore extending the survival of the mice [11].
Recently, our group published the first in vitro and in vivo applications of 5′-trans-splicing for HCM (Fig. 2b; [41]). In this study, the feasibility and efficacy of 5′-trans-splicing were tested in neonatal cardiac myocytes and in the heart of Mybpc3-targeted KI mice. The therapeutic PTM was designed to have a specific cardiac myocyte promoter (human TNNT2 promoter), and the SV40 polyadenylation (polyA) signal was deleted to maintain the PTM transcripts in the nucleus and to prevent its translation. The packaging in cardiac-specific AAV serotypes allowed efficient transduction of cardiac myocytes ex vivo or in the heart in vivo. However, the translation into a corrected cMyBP-C protein was poor. Even though the efficiency of this process needs to be optimized, it is conceivable to generate two PTM molecules, targeting mutations located in either the 5′ or 3′ regions of MYBPC3 pre-mRNA in order to treat all HCM-associated MYBPC3 mutations.
Conventional gene therapy
In larger animal models, conventional cardiac gene therapy has been tested for heart failure (HF), a nongenetic cardiac disease, targeting proteins involved in calcium handling such as phospholamban [32] and S100A1 [48]. Recently, the successful completion of phase II trials for gene therapy of SERCA2a demonstrates the feasibility and safety of AAV1-mediated gene transfer and improvement of the symptoms, and exercise capacity of the patients with advanced HF [31].
Haploinsufficiency of cMyBP-C likely plays a major role in HCM pathogenesis [39, 40, 55]. Insufficient amount of cMyBP-C could produce an imbalance in the stoichiometry of the thick filaments and alter sarcomeric structure and function. Therefore, conventional gene therapy could be also used to treat or prevent the disease. One should note, however, that the presence of poison peptide could hinder the presumed positive effect of MYBPC3 gene transfer. Spontaneous larger animal models for genetic diseases are rare; however, a cat model carrying an MYBPC3 mutation associated with cardiac hypertrophy has been described [43]. Last year, it has been shown that direct myocardial injection of a lentivirus-encoding mouse Mybpc3 cDNA improved myofilament contractile functions in Mybpc3-targeted deficient mice [42]. However, the use of lentivirus and the arduous delivery method in this study limit the direct application of this method to HCM patients.
Conclusion and future directions
MYBPC3 gene mutations are the most frequent cause in HCM and are associated with variable clinical manifestations. There is a need for novel therapies, which will be curative, particularly for the severe forms of the disease, leading to a very bad prognosis. The cat model of HCM, which carries a missense MYBPC3 mutation leading to both haploinsufficiency and poison peptide, represents a good model to evaluate different gene-based therapeutic strategies. Furthermore, developing new models through HCM human-induced pluripotent stem cells (hIPSC; [33]), such as cardiac myocytes or engineered heart tissue [26], should enable high-throughput investigation of molecular mechanisms and molecular therapy for MYBPC3-related HCM. Both the cat and hIPSC-derived cardiac myocytes and engineered heart tissue are required steps before development of clinical trials.
References
Aartsma-Rus A, van Ommen GJ (2009) Less is more: therapeutic exon skipping for Duchenne muscular dystrophy. Lancet Neurol 8:873–875
Alcalai R, Seidman JG, Seidman CE (2008) Genetic basis of hypertrophic cardiomyopathy: from bench to the clinics. J Cardiovasc Electrophysiol 19:104–110
Alter J, Lou F, Rabinowitz A, Yin H, Rosenfeld J, Wilton SD, Partridge TA, Lu QL (2006) Systemic delivery of morpholino oligonucleotide restores dystrophin expression bodywide and improves dystrophic pathology. Nat Med 12:175–177
Avale ME, Rodriguez-Martin T, Gallo JM (2013) Trans-splicing correction of tau isoform imbalance in a mouse model of tau mis-splicing. Hum Mol Genet 22:2603–2611
Bonne G, Carrier L, Bercovici J, Cruaud C, Richard P, Hainque B, Gautel M, Labeit S, James M, Beckmann J, Weissenbach J, Vosberg HP, Fiszman M, Komajda M, Schwartz K (1995) Cardiac myosin binding protein-C gene splice acceptor site mutation is associated with familial hypertrophic cardiomyopathy. Nat Genet 11:438–440
Carrier L, Bonne G, Bahrend E, Yu B, Richard P, Niel F, Hainque B, Cruaud C, Gary F, Labeit S, Bouhour JB, Dubourg O, Desnos M, Hagege AA, Trent RJ, Komajda M, Fiszman M, Schwartz K (1997) Organization and sequence of human cardiac myosin binding protein C gene (MYBPC3) and identification of mutations predicted to produce truncated proteins in familial hypertrophic cardiomyopathy. Circ Res 80:427–434
Carrier L, Hengstenberg C, Beckmann JS, Guicheney P, Dufour C, Bercovici J, Dausse E, Berebbi-Bertrand I, Wisnewsky C, Pulvenis D, Fetler L, Vignal A, Weissenbach J, Hillaire D, Feingold J, Bouhour JB, Hagege A, Desnos M, Isnard R, Dubourg O, Komajda M, Schwartz K (1993) Mapping of a novel gene for familial hypertrophic cardiomyopathy to chromosome 11. Nat Genet 4:311–313
Carrier L, Schlossarek S, Willis MS, Eschenhagen T (2010) The ubiquitin-proteasome system and nonsense-mediated mRNA decay in hypertrophic cardiomyopathy. Cardiovasc Res 85:330–338
Cartegni L, Wang J, Zhu Z, Zhang MQ, Krainer AR (2003) ESEfinder: A web resource to identify exonic splicing enhancers. Nucleic Acids Res 31:3568–3571
Chao H, Mansfield SG, Bartel RC, Hiriyanna S, Mitchell LG, Garcia-Blanco MA, Walsh CE (2003) Phenotype correction of hemophilia A mice by spliceosome-mediated RNA trans-splicing. Nat Med 9:1015–1019
Coady TH, Baughan TD, Shababi M, Passini MA, Lorson CL (2008) Development of a single vector system that enhances trans-splicing of SMN2 transcripts. PLoS ONE 3:e3468
Coady TH, Lorson CL (2010) Trans-splicing-mediated improvement in a severe mouse model of spinal muscular atrophy. J Neurosci 30:126–130
Dhandapany PS, Sadayappan S, Xue Y, Powell GT, Rani DS, Nallari P, Rai TS, Khullar M, Soares P, Bahl A, Tharkan JM, Vaideeswar P, Rathinavel A, Narasimhan C, Ayapati DR, Ayub Q, Mehdi SQ, Oppenheimer S, Richards MB, Price AL, Patterson N, Reich D, Singh L, Tyler-Smith C, Thangaraj K (2009) A common MYBPC3 (cardiac myosin binding protein C) variant associated with cardiomyopathies in South Asia. Nat Genet 41:187–191
Elliott P, Andersson B, Arbustini E, Bilinska Z, Cecchi F, Charron P, Dubourg O, Kuhl U, Maisch B, McKenna WJ, Monserrat L, Pankuweit S, Rapezzi C, Seferovic P, Tavazzi L, Keren A (2008) Classification of the cardiomyopathies: a position statement from the European Society Of Cardiology Working Group on Myocardial and Pericardial Diseases. Eur Heart J 29:270–276
Elliott P, McKenna WJ (2004) Hypertrophic cardiomyopathy. Lancet 363:1881–1891
Frank-Hansen R, Page SP, Syrris P, McKenna WJ, Christiansen M, Andersen PS (2008) Micro-exons of the cardiac myosin binding protein C gene: flanking introns contain a disproportionately large number of hypertrophic cardiomyopathy mutations. Eur J Hum Genet 16:1062–1069
Frey N, Luedde M, Katus HA (2012) Mechanisms of disease: hypertrophic cardiomyopathy. Nat Rev Cardiol 9:91–100
Friedrich FW, Carrier L (2012) Genetics of hypertrophic and dilated cardiomyopathy. Curr Pharm Biotechnol 13:2467–2476
Gautel M, Zuffardi O, Freiburg A, Labeit S (1995) Phosphorylation switches specific for the cardiac isoform of myosin binding protein-C: a modulator of cardiac contraction? Embo J 14:1952–1960
Gedicke-Hornung C, Behrens-Gawlik V, Reischmann S, Geertz B, Stimpel D, Weinberger F, Schlossarek S, Precigout G, Braren I, Eschenhagen T, Mearini G, Lorain S, Voit T, Dreyfus PA, Garcia L, Carrier L (2013) Rescue of cardiomyopathy through U7snRNA-mediated exon skipping in Mybpc3-targeted knock-in mice. EMBO Mol Med 5:1128–1145
Gersh BJ, Maron BJ, Bonow RO, Dearani JA, Fifer MA, Link MS, Naidu SS, Nishimura RA, Ommen SR, Rakowski H, Seidman CE, Towbin JA, Udelson JE, Yancy CW (2011) 2011 ACCF/AHA guideline for the diagnosis and treatment of hypertrophic cardiomyopathy: a report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines. J Thorac Cardiovasc Surg 142:e153–e203
Goyenvalle A, Babbs A, Powell D, Kole R, Fletcher S, Wilton SD, Davies KE (2010) Prevention of dystrophic pathology in severely affected dystrophin/utrophin-deficient mice by morpholino-oligomer-mediated exon-skipping. Mol Ther 18:198–205
Goyenvalle A, Babbs A, van Ommen GJ, Garcia L, Davies KE (2009) Enhanced exon-skipping induced by U7 snRNA carrying a splicing silencer sequence: Promising tool for DMD therapy. Mol Ther 17:1234–1240
Goyenvalle A, Vulin A, Fougerousse F, Leturcq F, Kaplan JC, Garcia L, Danos O (2004) Rescue of dystrophic muscle through U7 snRNA-mediated exon skipping. Science 306:1796–1799
Hammond SM, Wood MJ (2011) Genetic therapies for RNA mis-splicing diseases. Trends Genet 27:196–205
Hansen A, Eder A, Bonstrup M, Flato M, Mewe M, Schaaf S, Aksehirlioglu B, Schworer A, Uebeler J, Eschenhagen T (2010) Development of a drug screening platform based on engineered heart tissue. Circ Res 107:35–44
Havens MA, Duelli DM, Hastings ML (2013) Targeting RNA splicing for disease therapy. Wiley Interdiscip Rev RNA 4:247–266
Heemskerk HA, de Winter CL, de Kimpe SJ, van Kuik-Romeijn P, Heuvelmans N, Platenburg GJ, van Ommen GJ, van Deutekom JC, Aartsma-Rus A (2009) In vivo comparison of 2′-O-methyl phosphorothioate and morpholino antisense oligonucleotides for Duchenne muscular dystrophy exon skipping. J Gene Med 11:257–266
Hua Y, Krainer AR (2012) Antisense-mediated exon inclusion. Methods Mol Biol 867:307–323
Jearawiriyapaisarn N, Moulton HM, Buckley B, Roberts J, Sazani P, Fucharoen S, Iversen PL, Kole R (2008) Sustained dystrophin expression induced by peptide-conjugated morpholino oligomers in the muscles of mdx mice. Mol Ther 16:1624–1629
Jessup M, Greenberg B, Mancini D, Cappola T, Pauly DF, Jaski B, Yaroshinsky A, Zsebo KM, Dittrich H, Hajjar RJ (2011) Calcium Upregulation by Percutaneous Administration of Gene Therapy in Cardiac Disease (CUPID): a phase 2 trial of intracoronary gene therapy of sarcoplasmic reticulum Ca2+-ATPase in patients with advanced heart failure. Circulation 124:304–313
Kaye DM, Preovolos A, Marshall T, Byrne M, Hoshijima M, Hajjar R, Mariani JA, Pepe S, Chien KR, Power JM (2007) Percutaneous cardiac recirculation-mediated gene transfer of an inhibitory phospholamban peptide reverses advanced heart failure in large animals. J Am Coll Cardiol 50:253–260
Lan F, Lee AS, Liang P, Sanchez-Freire V, Nguyen PK, Wang L, Han L, Yen M, Wang Y, Sun N, Abilez OJ, Hu S, Ebert AD, Navarrete EG, Simmons CS, Wheeler M, Pruitt B, Lewis R, Yamaguchi Y, Ashley EA, Bers DM, Robbins RC, Longaker MT, Wu JC (2013) Abnormal calcium handling properties underlie familial hypertrophic cardiomyopathy pathology in patient-specific induced pluripotent stem cells. Cell Stem Cell 12:101–113
Le Roy F, Charton K, Lorson CL, Richard I (2009) RNA-targeting approaches for neuromuscular diseases. Trends Mol Med 15:580–591
Lorain S, Peccate C, Le Hir M, Garcia L (2010) Exon exchange approach to repair Duchenne dystrophin transcripts. PLoS ONE 5:e10894
Marian AJ (2010) Hypertrophic cardiomyopathy: from genetics to treatment. Eur J Clin Invest 40:360–369
Maron BJ, Gardin JM, Flack JM, Gidding SS, Kurosaki TT, Bild DE (1995) Prevalence of hypertrophic cardiomyopathy in a general population of young adults. Echocardiographic analysis of 4111 subjects in the CARDIA Study. Coronary Artery Risk Development in (Young) Adults. Circulation 92:785–789
Maron BJ, Pelliccia A, Spirito P (1995) Cardiac disease in young trained athletes. Insights into methods for distinguishing athlete’s heart from structural heart disease, with particular emphasis on hypertrophic cardiomyopathy. Circulation 91:1596–1601
Marston S, Copeland O, Gehmlich K, Schlossarek S, Carrier L (2012) How do MYBPC3 mutations cause hypertrophic cardiomyopathy? J Muscle Res Cell Motil 33:75–80
Marston S, Copeland ON, Jacques A, Livesey K, Tsang V, McKenna WJ, Jalilzadeh S, Carballo S, Redwood C, Watkins H (2009) Evidence from human myectomy samples that MYBPC3 mutations cause hypertrophic cardiomyopathy through haploinsufficiency. Circ Res 105:219–222
Mearini G, Stimpel D, Kramer E, Geertz B, Braren I, Gedicke-Hornung C, Precigout G, Muller OJ, Katus HA, Eschenhagen T, Voit T, Garcia L, Lorain S, Carrier L (2013) Repair of Mybpc3 mRNA by 5′-trans-splicing in a mouse model of hypertrophic cardiomyopathy. Mol Ther Nucleic Acids 2:e102
Merkulov S, Chen X, Chandler MP, Stelzer JE (2012) In vivo cardiac Myosin binding protein C gene transfer rescues myofilament contractile dysfunction in cardiac Myosin binding protein C null mice. Circ Heart Fail 5:635–644
Meurs KM, Sanchez X, David RM, Bowles NE, Towbin JA, Reiser PJ, Kittleson JA, Munro MJ, Dryburgh K, Macdonald KA, Kittleson MD (2005) A cardiac myosin binding protein C mutation in the Maine Coon cat with familial hypertrophic cardiomyopathy. Hum Mol Genet 14:3587–3593
Moolman JA, Reith S, Uhl K, Bailey S, Gautel M, Jeschke B, Fischer C, Ochs J, McKenna WJ, Klues H, Vosberg HP (2000) A newly created splice donor site in exon 25 of the MyBP-C gene is responsible for inherited hypertrophic cardiomyopathy with incomplete disease penetrance. Circulation 101:1396–1402
Murauer EM, Gache Y, Gratz IK, Klausegger A, Muss W, Gruber C, Meneguzzi G, Hintner H, Bauer JW (2011) Functional correction of type VII collagen expression in dystrophic epidermolysis bullosa. J Invest Dermatol 131:74–83
Olivotto I, Girolami F, Ackerman MJ, Nistri S, Bos JM, Zachara E, Ommen SR, Theis JL, Vaubel RA, Re F, Armentano C, Poggesi C, Torricelli F, Cecchi F (2008) Myofilament protein gene mutation screening and outcome of patients with hypertrophic cardiomyopathy. Mayo Clin Proc 83:630–638
Osman EY, Yen PF, Lorson CL (2012) Bifunctional RNAs targeting the intronic splicing silencer N1 increase SMN levels and reduce disease severity in an animal model of spinal muscular atrophy. Mol Ther 20:119–126
Pleger ST, Shan C, Ksienzyk J, Bekeredjian R, Boekstegers P, Hinkel R, Schinkel S, Leuchs B, Ludwig J, Qiu G, Weber C, Raake P, Koch WJ, Katus HA, Muller OJ, Most P (2011) Cardiac AAV9-S100A1 gene therapy rescues post-ischemic heart failure in a preclinical large animal model. Sci Transl Med 3:92ra64
Richard P, Charron P, Carrier L, Ledeuil C, Cheav T, Pichereau C, Benaiche A, Isnard R, Dubourg O, Burban M, Gueffet JP, Millaire A, Desnos M, Schwartz K, Hainque B, Komajda M (2003) Hypertrophic cardiomyopathy: distribution of disease genes, spectrum of mutations and implications for molecular diagnosis strategy. Circulation 107:2227–2232
Richard P, Villard E, Charron P, Isnard R (2006) The genetic bases of cardiomyopathies. J Am Coll Cardiol 48:A79–A89
Rodriguez-Martin T, Anthony K, Garcia-Blanco MA, Mansfield SG, Anderton BH, Gallo JM (2009) Correction of tau mis-splicing caused by FTDP-17 MAPT mutations by spliceosome-mediated RNA trans-splicing. Hum Mol Genet 18:3266–3273
Rodriguez-Martin T, Garcia-Blanco MA, Mansfield SG, Grover AC, Hutton M, Yu Q, Zhou J, Anderton BH, Gallo JM (2005) Reprogramming of tau alternative splicing by spliceosome-mediated RNA trans-splicing: implications for tauopathies. Proc Natl Acad Sci U S A 102:15659–15664
Rottbauer W, Gautel M, Zehelein J, Labeit S, Franz WM, Fischer C, Vollrath B, Mall G, Dietz R, Kubler W, Katus HA (1997) Novel splice donor site mutation in the cardiac myosin-binding protein-C gene in familial hypertrophic cardiomyopathy. Characterization of cardiac transcript and protein. J Clin Invest 100:475–482
Schlossarek S, Englmann DR, Sultan KR, Sauer M, Eschenhagen T, Carrier L (2012) Defective proteolytic systems in Mybpc3-targeted mice with cardiac hypertrophy. Basic Res Cardiol 107:1–13
Schlossarek S, Mearini G, Carrier L (2011) Cardiac myosin-binding protein C in hypertrophic cardiomyopathy: mechanisms and therapeutic opportunities. J Mol Cell Cardiol 50:613–620
Schlossarek S, Schuermann F, Geertz B, Mearini G, Eschenhagen T, Carrier L (2012) Adrenergic stress reveals septal hypertrophy and proteasome impairment in heterozygous Mybpc3-targeted knock-in mice. J Muscle Res Cell Motil 33:5–15
Skordis LA, Dunckley MG, Yue B, Eperon IC, Muntoni F (2003) Bifunctional antisense oligonucleotides provide a trans-acting splicing enhancer that stimulates SMN2 gene expression in patient fibroblasts. Proc Natl Acad Sci U S A 100:4114–4119
Spoladore R, Maron MS, D’Amato R, Camici PG, Olivotto I (2012) Pharmacological treatment options for hypertrophic cardiomyopathy: high time for evidence. Eur Heart J 33:1724–1733
van Dijk SJ, Dooijes D, Dos Remedios C, Michels M, Lamers JM, Winegrad S, Schlossarek S, Carrier L, Ten Cate FJ, Stienen GJ, van der Velden J (2009) Cardiac Myosin-Binding Protein C Mutations and Hypertrophic Cardiomyopathy. Haploinsufficiency, Deranged Phosphorylation, and Cardiomyocyte Dysfunction. Circulation 119:1473–1483
van Dijk SJ, Paalberends ER, Najafi A, Michels M, Sadayappan S, Carrier L, Boontje NM, Kuster DW, van Slegtenhorst M, Dooijes D, Dos Remedios C, Ten Cate FJ, Stienen GJ, van der Velden J (2012) Contractile dysfunction irrespective of the mutant protein in human hypertrophic cardiomyopathy with normal systolic function. Circ Heart Fail 5:36–46
Vignier N, Schlossarek S, Fraysse B, Mearini G, Kramer E, Pointu H, Mougenot N, Guiard J, Reimer R, Hohenberg H, Schwartz K, Vernet M, Eschenhagen T, Carrier L (2009) Nonsense-Mediated mRNA decay and ubiquitin-proteasome system regulate cardiac myosin-binding protein C mutant levels in cardiomyopathic mice. Circ Res 105:239–248
Wally V, Murauer EM, Bauer JW (2012), Spliceosome-Mediated trans-splicing: the therapeutic cut and paste. J Invest Dermatol
Watkins H, Conner D, Thierfelder L, Jarcho JA, MacRae C, McKenna WJ, Maron BJ, Seidman JG, Seidman CE (1995) Mutations in the cardiac myosin binding protein-C gene on chromosome 11 cause familial hypertrophic cardiomyopathy. Nat Genet 11:434–437
Woodley L, Valcarcel J (2002) Regulation of alternative pre-mRNA splicing. Brief Funct Genomic Proteomic 1:266–277
Acknowledgments
This work was supported by the Fritz Thyssen Stiftung (Az. 10.09.1.139), the seventh Framework Program of the European Union (Health-F2-2009-241577; BIG-Heart project).
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published as part of the Special Issue on MYBPC3.
Rights and permissions
About this article
Cite this article
Behrens-Gawlik, V., Mearini, G., Gedicke-Hornung, C. et al. MYBPC3 in hypertrophic cardiomyopathy: from mutation identification to RNA-based correction. Pflugers Arch - Eur J Physiol 466, 215–223 (2014). https://doi.org/10.1007/s00424-013-1409-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00424-013-1409-7