Abstract
ASCL1 is one of the master transcription factors of small cell lung carcinoma (SCLC). To investigate the significance of ASCL1 in pulmonary neuroendocrine carcinoma, we performed 2 comparative RNA-seq studies between H69 (ASCL1-positive, classical type SCLC) and H69AR (ASCL1-negative, variant type SCLC) and between ASCL1-transfected A549 adenocarcinoma cell lines (A549(ASCL1+) cell lines) and A549(control) cell lines. RNA-seq analyses revealed that 940 genes were significantly different between the H69 and H69AR cell lines, and 728 between the A549(ASCL1+) and A549(control) cell lines. In total, 120 common genes between these analyses were selected as candidate ASCL1-related genes, and included genes with various cellular functions, such as neural development, secretion, growth, and morphology. Their expression degrees in three classical and two variant SCLC cell lines, two A549(ASCL1+) and two A549(control) cell lines were subjected to quantitative PCR analyses. Since the candidate ASCL1-related genes were strongly expressed in the classical SCLC and A549(ASCL1+) cell lines and more weakly expressed in the variant SCLC and A549(control) cell lines, the ASCL1-related 7 molecules INSM1, ISL1, SYT4, KCTD16, SEZ6, MS4A8, and COBL were further selected. These molecules suggested diverse functions for A549(ASCL1+): INSM1 and ISL1 are transcription factors associated with neuroendocrine differentiation, while SYT4, KTCD16, and SEZ6 may be related to neurosecretory functions and MS4A8 and COBL to cell growth and morphology. An immunohistochemistry of these seven molecules was performed on lung carcinoma tissues and the xenotransplanted tumors of A549(ASCL1+), and they were preferentially and positively stained in ASCL1-postive tumor tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lung cancer includes various histological types, among which neuroendocrine (NE) tumors are a distinct family of lung cancers with shared morphological, ultrastructural, immunohistochemical, and molecular characteristics (Brambilla et al. 2015). Small cell lung cancer (SCLC) is the highest malignant type among this lung cancer family, and is characterized by rapid growth and metastasis as well as NE differentiation (Travis et al. 2015; Pietanza et al. 2015; Bunn et al. 2016). SCLC is characterized by the expression of NE molecules, which are regulated by signal pathway-transcription factor networks, such as the Notch signaling pathway and achaete-scute complex homologue 1 (ASCL1) (Ito et al. 2003, 2017; Ball 2004; Meder et al. 2016).
ASCL1 is a proneuronal basic helix-loop-helix transcription factor that plays an important role in cell fate decisions during neurogenesis (Guillemot et al. 1993). The significance of ASCL1 in the NE differentiation of the mouse fetal lung epithelium and human SCLC was previously demonstrated (Borges et al. 1997). ASCL1 expression is regulated by Notch signaling (Ito et al. 2000; Morimoto et al. 2012), and mouse pulmonary NE cells are positive for ASCL1, but negative for Notch1-3 and hairy and enhancer of split 1 (Hes1), and many pulmonary NE cells appear in Hes1 gene-deficient mice (Ito et al. 2000). Linnoila et al. reported that the overexpression of ASCL1 and SV40 large T antigen in lung epithelial cells using a club cell secretory protein gene promoter induced non-SCLC with NE features in mice (Linnoila et al. 2000). Transfection studies have been performed on the ASCL1 gene in various human lung adenocarcinoma (ADC) cell lines, and NE differentiation was confirmed in ASCL1-transfected ADC cells (Osada et al. 2008; Fujino et al. 2015; Meder et al. 2016). After transfection of the ASCL1 gene, A549 ADC cells showed the expression of SYP, CHGB, and SCG2 (Osada et al. 2008), H1975 and H358 ADC cells showed the expression of CHGA, SYP, and NCAM (Fujino et al. 2015), and PC9 ADC cells showed the expression of NCAM (Meder et al. 2016). Besides the induction of NE differentiation by the ASCL1 gene, ASCL1 may play roles in growth, cell survival, migration, epithelial-mesenchymal transition, and the modulation of tumor-initiating cells (Sriuranpong et al. 2001; Osada et al. 2005, 2008; Jiang et al. 2009; Demelash et al. 2012; Li and Linnoila 2012; Ito et al. 2017; Tenjin et al. 2019). The ASCL1 gene is considered to be a master regulator of NE differentiation (Borges et al. 1997) and a lineage-specific oncogene of SCLC (Gazdar et al. 2017). SCLC may be subdivided into four main subtypes based on the expression of lineage-specific transcription factors, with SCLC having ASCL1 being the most common subtype according to a recent review (Rudin et al. 2019). Transcriptional targets of ASCL1 have been examined using chromatin immunoprecipitation sequencing (ChIP-seq) (Augustyn et al. 2014; Borromeo et al. 2016). ChIP-seq studies identified many target molecules of ASCL1, including genes related to cell survival (Augustyn et al. 2014) and tumorigenesis (Borromeo et al. 2016) as well as NE differentiation, which supports ASCL1 functioning as a lineage-specific oncogene. A ChIP-seq analysis of the ASCL1 gene revealed neural and NE differentiation-related targets molecules, such as GRP, SYT1, INSM1, SNAP25, and potassium voltage-gated channel subfamily members (Augustyn et al. 2014; Borromeo et al. 2016). Zhang et al. recently examined SCLC tumors and cell lines using a 50-gene expression-based NE score, which was developed by expression arrays and RNA sequence (RNA-seq) data (Zhang et al. 2018a). SCLC cell lines have been subdivided into two groups, classical and variant SCLCs (Gazdar et al. 1985; Zhang et al. 2018a). Classical SCLC cell lines are characterized by floating cell growth in medium and distinct NE differentiation, and variant SCLC cell lines by adhesive growth and poor NE differentiation (Gazdar et al. 1985; Zhang et al. 2018a). Classical cell lines belong to NE high SCLC, and variant cell lines to NE low SCLC, in which Notch activity and the loss of ASCL1 expression have been demonstrated (Zhang et al. 2018a).
The purpose of this study is to identify common biomarkers or therapeutic targets in ASCL1-associated PNECs, as ASCL1 is the most common cell-lineage-specific transcription factor in SCLC (Rudin et al. 2019) and it is important to know the ASCL1-associated molecules for further understanding of PNECs including SCLC. Some studies have clarified the genes associated with ASCL1 with use of DNA microarray (Osada et al. 2008; Zhang et al. 2018a), RNA-seq (Zhang et al. 2018a) and ChIP-seq analyses (Augustyn et al. 2014; Borromeo et al. 2016). While referring to these previous studies, to find the candidate common molecules associated with ASCL1, we performed RNA-seq analyses using 2 sets of lung cancer cell lines; (1) H69, ASCL1-positive, classical subtype SCLC cell line comparing with H69AR, ASCL1-negative, variant subtype SCLC cell line, and (2) ASCL1-transfected A549 (ASCL1-induced pulmonary NE carcinoma; A549(ASCL1+)) cell line comparing with ASCL1-negative mock-A549 (A549 (control)) cell line. 120 ASCL1-associated molecules were detected through these studies, and to improve the accuracy of them as the ASCL1-associated molecule, we applied them with quantitated PCR analyses in 3 ASCL1-positive and 2 ASCL1-negative SCLC cell lines, 2 ASCL1+ and 2 control-A549 cell lines, and we have selected the authentic ASCL1-associated molecules which are highly expressed in the ASCL1-positive SCLC and ASCL1 + A549 cell lines, and lowly in the ASCL1-negative SCLC and A549(control) cell lines. We finally selected 7 genes, 5 are NE-related, and the other 2 are related to proliferation and morphology. Our observation could provide valuable information for future studies of ASCL1-positive PNECs including SCLC.
Materials and methods
Cell lines
Five SCLC cell lines (H69, H889, SBC1, H69AR, and SBC3) and one ADC cell line, A549 were used in the present study. H69, H889, H69AR, and A549 were purchased from ATCC (Manassas, Virginia), and SBC1 and SBC3 from the Japan Collection of Research Bioresources Cell Bank (Osaka, Japan).
Plasmid construction
To construct pCAG-IRES-Puro-ASCL1, we replicated the ASCL1 gene of a human ASCL1 cDNA ORF clone (OriGene, Rockville, Maryland) and replaced it with ASCL1. We generated pCAG-IRES-Puro mock from a human ASCL1 cDNA ORF clone by cleaving out the ASCL1 gene. These plasmids were transfected into A549 cells with FuGENE6 (Roche, Risch-Rotkreua, Switzerland) as described in the manufacturer’s instructions. After 48 h, 5 μg/mL puromycin (SIGMA, St. Louis, Missouri) was added to cells for 4 weeks, with a medium change every 3 days. Stably transfected cells with resistance to puromycin were cloned for each transfectant.
Reverse transcription-polymerase chain reaction (RT-PCR) and qRT-PCR
Total RNA was isolated using an RNeasy Mini Kit (Qiagen, Hilden, Germany) and cDNA was produced using the SuperScriptII First-Strand RT-PCR kit (Invitrogen, Osaka, Japan). RT-PCR was conducted in accordance with the standard protocol for TAKARA Ex taq (TAKARA Bio, Shiga, Japan), and DNA amplification was performed using a Mastercycler® nexus (Eppendorf, Hamburg, Germany). Cycling conditions were as follows: one cycle at 95 °C for 2 min, followed by 32 cycles at 95 °C for 30 s, 57 °C for 30 s, and 72 °C for 30 s. Amplification products were separated on a 1.5% agarose gel, and identified after ethidium bromide staining. qRT-PCR was conducted according to the standard protocol of SYBR fast qPCR Mix (TAKARA Bio) on a LiteCycler Nano (Roche). Data were obtained from triplicate reactions. The means and standard deviation (SD) of the copy number were normalized to the value for glyceraldehyde-3-phosphate dehydrogenase (GAPDH) mRNA. A list of the specific primers is shown in Table 1.
RNA sequence analysis
Total RNA was isolated from cultured cells, including A549(control) cells (pCAG-IRES-Puro Flag HA), A549(ASCL1+) cells (pCAG-IRES-Puro Flag HA ASCL1), H69 cells, and H69AR cells using an RNeasy Mini Kit (Qiagen). A 2100 Bioanalyzer (Agilent, Santa Clara, California) was used to detect the concentration and purity of total RNA. All samples with an RNA integrity number (RIN) > 7.5 were used for sequencing. Nextseq 500 (Illumina, San Diego, California) was used to analyze data, which were converted to FastQ files. The quality control of data was performed by FastQC. Filtered reads were used to map to the UCSC hg19 genome reference genome using HISAT2 2.1.0. Fragments per kilobase of exon per million mapped reads (FPKM) values were calculated using Cufflinks. Significant genes were extracted by cuffdiff (p < 0.05).
Gene ontology (GO) analysis
GO annotation and classification were based on three categories: biological process, molecular function, and cellular component. The Database for Annotation, Visualization, and Integrated Discovery 6.7 (DAVID 6.7, https://www.david.niaid.nih.gov) was used for the GO analysis (Huang et al. 2009). The gene list contained significant genes in the RNA-seq analysis. The DAVID online database was used to visualize key molecular functions and biological processes. p < 0.05 was regarded as the cut-off criterion with a significant difference.
Western blotting
Cultured cells were prepared for the WB analysis, as described previously (Motooka et al. 2017). A list of the primary antibodies used is shown in Table 2. The membrane was washed and incubated with the respective secondary antibodies conjugated with horseradish peroxidase (Cell Signaling, Danvers, MA) for 1 h, and the immunocomplex was visualized with chemiluminescence substrate (Amersham Pharmacia Biotech, Buckinghamshire, United Kingdom).
Tissue samples
Tissue samples of SCLC (n = 10), ADC (n = 10), and SCC (n = 10) resected at the Department of Thoracic Surgery of Kumamoto University Hospital were obtained from 30 patients for the following studies. A histological diagnosis of the samples was made in accordance with World Health Organization criteria (Brambilla et al. 2015). These sections were used for immunohistochemistry. The present study followed the guidelines of the Ethics Committee of Kumamoto University.
Transcriptome data of SCLC tissues
The transcriptome data of SCLC tissue samples from the GSE60052 (n = 79) RNA-seq dataset (Jiang et al. 2016) were used to examine the relationship between the expression levels of ASCL1 and the other mRNAs studied.
IHC
Formalin-fixed, paraffin-embedded specimens were cut into 3-μm-thick sections and mounted onto MAS-GP-coated slides (Matsunami Glass Ind., Osaka, Japan). After being deparaffinized and rehydrated, the sections were heated using an autoclave in 0.01 mol/L citrate buffer (pH 6.0 or 7.0) for antigen retrieval. Sections were incubated with 0.3% H2O2 in absolute methanol for 20 min to block endogenous peroxidase activity. Sections were then incubated with Protein Block Serum-Free (Dako, Glostrup, Denmark) for 10 min to block non-specific staining. After this blocking step, sections were incubated with the primary antibodies (Table 2) at 4 °C overnight. This was followed by sequential 1-h incubations with the secondary antibodies (En Vision + System-HRP-labeled polymer; Dako) and visualization with liquid DAB + substrate Chromogen System (Dako). All slides were counterstained with hematoxylin for 30 s before being dehydrated and mounted. The specificity of the immunolabeling of each antibody was tested using normal mouse IgG (Santa Cruz Biotechnology) and normal rabbit IgG (Santa Cruz Biotechnology). We discerned that the tissue samples were positive, when more than 10% of tumor cells were positively stained for each antigen.
Xenotransplantation
A total of 1.0 × 106 cells each of A549(control) cells and A549(ASCL1+) cells were injected subcutaneously into the backs of Rag2−/−:Jak3−/− mice (a generous gift from Prof. Seiji Okada Kumamoto University). Twenty days after the injection, tumors were removed. Samples were fixed with phosphate-buffered 4% paraformaldehyde solution and embedded in paraffin. Tissue sections were stained with hematoxylin and eosin (HE), and additional sections were used for immunohistochemical analyses. All animal experiments were conducted in accordance with the guidelines of the Animal Care and Use Committee of Kumamoto University.
Statistical analysis
All data were obtained from independent experiments, and were expressed as the means ± SDs of triplicate measurements. Differences in mean values between the two groups were statistically analyzed using the t test. Prism v.7 software (GraphPad Software, San Diego, California) was used for statistical analyses. p < 0.05 was considered to be significant.
Results
ASCL1 is associated with NE differentiation in SCLC cell lines
SCLC belongs to the category of high-grade PNEC (Brambilla et al. 2015). To confirm the expression relationship between ASCL1 and NE markers, we performed a WB analysis on the 5 SCLC cell lines. As well as ASCL1, we examined the other lineage-specific transcription factors such as NEUROD1, POU2F3 and YAP1. WB revealed that ASCL1 and NE markers including CHGA, SYP, and NCAM were simultaneously expressed in 3 (H69, H889, and SBC1) out of the 5 SCLC cell lines examined. On the other hand, in accordance with the weaker expression of ASCL1, H69AR cells lost NE markers. The SBC3 cell line is a variant subtype of SCLC (Gazdar et al. 1985; Wael et al. 2014), in which ASCL1 and NE markers were negligibly expressed, but YAP1 positively expressed (Fig. 1a). These results suggest that ASCL1 is closely related to NE differentiation in SCLC cell lines, such as H69, H889, and SBC1. The H69AR cell line is a drug (Adriamycin)-resistant form of the H69 cell line (Mirski et al. 1987), and lose ASCL1 expression, but showed YAP1 expression. H69 cells were the floating type, while H69AR cells were the adherent type, as shown in phase contrast images (Fig. 1b).
a H69, H889, and SBC1 cell lines express ASCL1 and NE markers, including CHGA, SYP, and NCAM, and only SBC1 expresses NEUROD1. However, H69AR and SBC3 do not show the expression of ASCL1, NEUROD1, POU2F3, or NE markers, but show the expression of YAP1. b H69 and H69AR cells are shown by phase contrast images. H69 cells are floating, while H69AR cells are adherent
ASCL1 plays critical roles in NE differentiation, cell adhesion, and proliferation
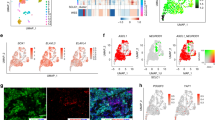
To examine the biological significance of ASCL1 in the lung cancer cell line, we performed ASCL1 gene transfection into A549 cells, which is an ADC cell that expresses neither ASCL1 nor any other NE markers. We established A549(control) cells using an empty vector as the control. As the expression of the NE markers such as CHGA and SYP, in the ASCL1-transfected A549 (A549(ASCL1+)) cells was detected, the A549(ASCL1+) cells can be confirmed as NE carcinoma cells. The expression of phosphorylated-histone H3, a mitotic marker, was also stronger in A549(ASCL1+) cells than in A549(control) cells. In contrast, the expression of E-CADHERIN was significantly reduced after the forced expression of ASCL1 (Fig. 2a). It is important to note that A549(control) cells tightly adhered to the dish, whereas A549(ASCL1+) cells showed a round-shaped morphology with weaker adhesion (Fig. 2b). We then performed xenotransplantation experiments on these cells using immunocompromised mice. A549(control) transplanted tumor cells had an ADC morphology, whereas A549(ASCL1+) cells showed undifferentiated features along with prominent nucleoli and increased mitosis by HE staining. IHC staining revealed that A549(ASCL1+) cells had increased expression levels of CHGA and Ki67, but decreased levels of E-CADHERIN (Fig. 2c). A RNA-seq study revealed that 728 molecules were significantly different between the A549 cell lines with or without ASCL1. We extracted molecules associated with NE differentiation from RNA-Seq data and generated heat maps (Fig. 2d). In the comparison between A549(ASCL1+) cells and A549(control) cells, we identified up-regulated genes, such as INSM1, CHGB, SYP, and CPLX2. We also identified down-regulated genes, such as SYTL3, NTRK3, and NCF2. In the other functional categories, for example, “Cell Proliferation”, we identified up-regulated genes, such as GAS2, CCND2, and INHBB, in a similar manner. In contrast, we also identified down-regulated genes, such as CREG1, PLK2, and CCND3. We then attempted to gain insights into the category “Cell Adhesion”. We identified up-regulated genes, such as CLDN4, BCAM, and JUP. Meanwhile, CDH1, CD44, and CLMP were identified as down-regulated genes. These results suggest that ASCL1 plays important roles in the regulation of NE differentiation, cell adhesion, and proliferation in this cell line.
a A WB analysis was performed using A549(no treatment), A549(control), and ASCL1-transfected A549 (A549(ASCL1+)) cell lines. CHGA and SYP were induced by the forced expression of ASCL1. pHH3 expression was stronger in A549(ASCL1+) cells than in A549(control) cells. On the other hand, E-CADHERIN expression levels were lower in A549(ASCL1+) cells than in A549(control) cells. β-actin serves as an internal control. b A549(control) cells show a rhomboid-shaped morphology, whereas A549(ASCL1+) cells show a round-shaped morphology in phase contrast images. Scale bar = 100 μm. c Xenotransplantation experiments using immunocompromised mice. A549(ASCL1+) transplanted tumor cells appear undifferentiated, while A549(control) transplanted tumor cells show adenocarcinoma features with tubular formation by hematoxylin and eosin staining (HE). IHC staining confirmed that A549(ASCL1+) transplanted tumors had higher expression levels of CHGA and Ki67, but lower expression levels of E-CADHERIN. Scale bar = 50 μm. d Heatmap of the FPKM of significant genes obtained by A549(control) and A549(ASCL1+) RNA-seq analysis. We annotated the genes to the ontologies which related to neuroendocrine, proliferation and adhesion. In the significant up-regulated genes, INSM1, CHGB, SYP and CPLX2 appear in neuroendocrine category. GAS2, CCND2 and INHBB appear in proliferation category. BCAM, JUP and CLDN4 appear in adhesion category
RNA-sequencing analysis showed that ASCL1 drives characteristic downstream target genes
To identify the genes regulated by ASCL1, we performed the two sets of RNA-seq studies: (1) a comparison between H69 and H69AR cells, and (2) a comparison between A549(ASCL1+) and A549(control) cells. Through these 2 sets of RNA-seq studies, we identified 120 common genes that showed significantly different mRNA expression in the 2 groups (Fig. 3a). The DAVID online bioinformatics tool was used for a GO functional analysis of the 120 shared genes. The top 20 significantly enriched terms (p < 0.05) in the molecular function and biological process category are shown in Fig. 3b. Biological process contains the terms involved in neuron-related matters, for example, neuronal development and differentiation or the regulation of synaptic transmission. Molecular function contains the terms involved in endocrine or cell metabolic processes, such as hormone activity, growth factor activity, or enzyme activity. We screened the 120 genes by qRT-PCR to extract target genes that show high expression levels in all ASCL1-positive SCLC cell lines and the A549(ASCL1+) cell lines (supplementary Table S1). We ultimately selected 7 out of 120 common genes, which included NE-associated molecules, such as INSM1 (Fujino et al. 2015), ISL1 (Agaimy et al. 2013), SYT4 (Zhang et al. 2018a), KCTD16 (van Coevorden-Hameete et al. 2019), and SEZ6 (Zhang et al. 2018a). Moreover, MS4A8 and COBL, which function in cell proliferation (Ye et al. 2014) and morphogenesis (Ahuja et al. 2007), were selected. We then examined the expression profiles of all 7 genes in SCLC cell lines, A549(control) cell lines, and A549(ASCL1+) cell lines using RT-PCR (Fig. 3c). We confirmed that ASCL1-positive SCLC cell lines and A549(ASCL1+) cell lines strongly expressed these 7 genes. These results suggest that ASCL1 potently drove the expression of NE-related molecules, such as INSM1, ISL1, SYT4, KCTD16, and SEZ6, and also correlated with the non-NE-related gene expression of MS4A8 and COBL. Based on the result showing that 7 representative ASCL1 and ASCL1-regulated genes, INSM1, ISL1, SYT4, KCTD16, SEZ6, MS4A8, and COBL, were more likely to be mainly co-expressed in the classical SCLC cell lines and A549(ASCL1+) cell lines, we surveyed public datasets that used the RNA-seq method (Jiang et al. 2016) of gene expression profiling in human SCLC tumor tissue samples and their correlations were evaluated. The RNA-seq dataset using the human tumor tissue samples of 79 SCLC patients confirmed the coordinated expression of 7 representative ASCL1 and ASCL1-regulated genes, INSM1 (GSE60052: ρ = 0.3579), ISL1 (ρ = 0.3929), SYT4 (ρ = 0.6167), KCTD16 (ρ = 0.3126), SEZ6 (ρ = 0.3360), MS4A8 (ρ = 0.5970), and COBL (ρ = 0.3240) in human SCLC tumor tissue samples (Fig. 4). These results support INSM1, ISL1, SYT4, KCTD16, SEZ6, MS4A8, and COBL expression being positively regulated by ASCL1.
a Two sets of RNA sequence analyses were performed. In total, 940 significant genes in the comparison of H69 and H69AR cell lines were extracted. In the same manner, 728 significant genes in the comparison of A549(ASCL1+) and A549(control) cell lines were extracted. In total, 120 genes were shared in the 2 groups. b Gene Ontology (GO) terms of biological processes or molecular functions assessed by the 120 shared genes were sorted by combined scores. The top 20 categories in each of the terms are presented. Values indicate − log10 (p value). c The expression patterns of ASCL1, INSM1, ISL1, SYT4, KCTD16, SEZ6, MS4A8, and COBL mRNA are shown by RT-PCR in 5 SCLC, A549(no treatment), A549(control), and A549(ASCL1+) cell lines. GAPDH serves as an internal control
Expression levels of ASCL1 and INSM1, ISL1, SYT4, KCTD16, SEZ6, MS4A8 and COBL in RNA-seq dataset of SCLC tissues. GSE 60052 (n = 79) dataset were analyzed. Positive correlation of ASCL1 and the other 7 genes mRNA expression were seen respectively. Spearman correlation coefficients (ρ) and p values are indicated
Expression of ASCL1-regulated genes positively correlated in SCLC tumor tissue
We examined the expression of ASCL1, INSM1, ISL1, SYT4, KCTD16, SEZ6, MS4A8, and COBL in xenotransplanted A549(ASCL1+) cells and A549(control) cells tumor tissues from immunocompromised mice and in surgically resected SCLC and NSCLC tissues. The immunohistochemical analysis showed that ASCL1, INSM1, and ISL1 were strongly expressed in the nuclei of A549(ASCL1+) tumor cells and SCLC tumor cells. As expected, ASCL1, INSM1, and ISL1 were not expressed in A549(control) transplanted tumor cells or NSCLC tumor cells (Fig. 5). Similarly, SYT4, KCTD16, SEZ6, MS4A8, and COBL were expressed in A549(ASCL1+) cells and SCLC cells, but not in A549(control) transplanted tumor cells or NSCLC tumor cells (Fig. 5). ASCL1 was strongly expressed in the nuclei of A549(ASCL1+) tumors cells and 70% of SCLC tumor samples, and not in A549(control) tumor cells or NSCLC tumor samples. The expression of ISL1 was similar to that of ASCL1, whereas INSM1 was positive in ASCL1-negative SCLC tumor samples (Table 3). SYT4, KCTD16, SEZ6, MS4A8, and COBL were strongly expressed by A549(ASCL1+) tumor cells and the majority of ASCL1-positive SCLC tumor samples. However, some NSCLC tumor samples were also positive for SYT4, KCTD16, SEZ6, MS4A8, and COBL (Table 3).
ASCL1, INSM1, and ISL1 are positively immunostained in nuclei of A549(ASCL1+) transplanted tumor cells and SCLC tissues. SYT4, SEZ6, and COBL were expressed in both the cell membrane and cytoplasm, while KCTD16 and MS4A8 were expressed in the cell membranes of A549(ASCL1+) and SCLC tumor cells. All of these markers were mostly negative in A549(control) transplanted tumor cells and ADC and SCC tissues. Representative images are shown. Scale bar = 100 μm
The difference of gene expressions between H69 and H69AR cell lines.
We identified 940 genes with expression levels that were significantly different between H69 cells and H69AR cells in RNA-seq (Fig. 3a). These genes were then statistically tested in a GO annotation enrichment analysis using DAVID. The top 3 GO terms in biological processes were neuron differentiation, neuron development, and neuron projection development (Fig. 6). These 3 GO terms related to neurons included BDNF, ISL1, NFASC, DMD, GDNF, and VCAN. On the other hand, drug resistance is generally associated with ATP-binding cassette transporters and solute carrier proteins (Li and Shu 2014; Kathawala et al. 2015; Domenichini et al. 2019). Therefore, the GO term analysis on molecular function within the top 10 revealed passive transmembrane transporter activity and metal ion transmembrane transporter activity, which included ABCC4 and ABCC8, SLC30A3, SLC39A6 and SLC40A1. Not only NE differentiation but also drug resistance or cell adhesive properties could be differed between these cell lines.
Discussion
In the present study, we used H69, H889, and SBC1 as classical SCLC subtypes, and they were positive for ASCL1 with NE markers, such as CHGA, SYP, and NCAM, and showed floating growth. We used H69AR and SBC3 as the variant SCLC subtypes, and they were negative for ASCL1 and the NE markers and showed adhesive growth. RNA-seq analyses comparing these cell lines suggest important molecules related to biological differences in classical and variant SCLC cell lines, including NE differentiation. In the present study, we focused on the significance of ASCL1, the master transcription factor of SCLC (Gazdar et al. 2017), in SCLC, and selected ASCL1-associated candidate molecules with combination RNA-seq studies using A549(ASCL1+).
We investigated the pathological features of A549(ASCL1+), which are generated by ASCL1 transfection in the human lung ADC cell line, A549, To clarify the biological significance of ASCL1. A549(ASCL1+) showed NE differentiation similar to other ASCL1-transfected lung ADC cells previously reported (Osada et al. 2008; Fujino et al. 2015; Meder et al. 2016), and the present study supports the crucial role of ASCL1 in NE differentiation in lung cancer cells. Moreover, A549(ASCL1+) showed a floating morphology in the culture medium, which mimicked the classical SCLC subtype, and we previously reported that ASCL1 modified epithelial-mesenchymal transition (Ito et al. 2017). Western blotting and histological studies using tumors grown in immunocompromised mouse subcutaneous tissue revealed that ASCL1 increased cell proliferation activity and apoptosis. Thus, ASCL1 is considered to play roles in various important cell biological functions other than NE differentiation in pulmonary NE tumors.
We selected a further 120 common ASCL1-associated molecules from RNA-seq analyses of the two sets of lung cancer cell lines. The DAVID analysis revealed the 120 molecules have various biological functions, including neuronal differentiation and development, neuronal secretory activity, cell growth, stem cell biology, enzyme activity, and others, which supports ASCL1 being one of the master genes of SCLC (Gazdar et al. 2017; Rudin et al. 2019). We performed qRT-PCR analyses of these molecules in 3 classical and 2 variant SCLC cell lines, A549(ASCL1+) and A549(control) cell lines and we further selected seven molecules that were strongly expressed in ASCL1-positive cell lines, but weakly or not expressed in the ASCL1-negative cell lines. We performed immunohistochemistry for these molecules in human lung carcinoma samples, including SCLC and A549(ASCL1+), to confirm whether these molecules are useful as biomarkers of SCLC. All molecules functioned as a positive marker for SCLC and A549(ASCL1+). INSM1 and ISL1 were transcription factors. Before the ChIP-seq study for ASCL1 (Borromeo et al. 2016), and our RNA-seq studies, we confirmed that INSM1 played important roles in the NE differentiation of lung neoplasms in cooperation with ASCL1 (Fujino et al. 2015). INSM1 was recently identified as a valuable marker for confirming the NE differentiation of neoplasms from the lung (Rooper et al. 2017; Mukhopadhyay et al. 2019) and other organs (Rosenbaum et al. 2015; Fujino et al. 2017). Among SCLC samples, INSM1 was positive expression in a few ASCL1-negative samples, and INSM1 expression in the ASCL1-negative samples could be explained by regulation by lineage-specific transcription factor such as NeuroD1 (Borromeo et al. 2016). ISL1 was previously reported to be a preferential immunostain for pancreatic NE neoplasms (Schmitt et al. 2008; Agaimy et al. 2013), and may be a marker for NE neoplasms of the lung. In our pilot ISL1 RNA knockdown study, the cell proliferation activity and NE differentiation of an ISL1-positive SCLC cell line were not affected, and ISL1 was not essential for the NE differentiation of pulmonary NE neoplasms. Synaptotagmins play roles in neurotransmitter release or hormone secretion in nervous and NE cells, and SYT4 is expressed by many NE cells (Moghadam and Jackson 2013). SYT4 was shown to be expressed in rat pheochromocytoma (Vician et al. 1995), and the present study demonstrated that SYT4 has potential as a good protein marker of pulmonary NE neoplasms and has an important function in the secretory activity of tumors. Regarding neuronal function, in addition to SYT4, we found that SEZ6 and KCTD16 may be common ASCL1-target molecules. SEZ6 was previously reported to be localized in the brain (Kim et al. 2002), functioned in neuronal development (Gunnersen et al. 2007), and was involved in β-cell function in pancreas islets (Stützer et al. 2013). KCTD16 was shown to play roles in the regulation of GABAB receptors (Zuo et al. 2019), and regulated membrane channel activities via the modulation of GTPases (Teng et al. 2019). The biological significance of SEZ6 and KCTD16 has not yet been investigated in malignant neoplasms, including SCLC, but may play roles in the neuronal aspects of SCLC cells. The other 2 molecules we selected in the present study, MS4A8 and COBL, are not always related to the neuronal phenotype of SCLC. MS4A family proteins are expressed on the cell surface and play roles in growth and development (Eon et al. 2016). Previous studies reported that MS4A8 was expressed in carcinomas of the stomach, colon, and prostate (Michel et al. 2013; Ye et al. 2014; Sun et al. 2018) as well as in SCLC (Bangur et al. 2004). The biological roles of MS4A8 vary among cancer types. In the gastrointestinal tract, MS4A8 may function in epithelial differentiation, and its forced expression suppressed cell proliferation and motility in colon cancer cells (Michel et al. 2013; Eon et al. 2016). In contrast, in prostate cancer, MS4A8 promoted prostatic cancer cells (Carney et al. 1985). Using cDNA subtraction and DNA microarrays, MS4A8 was found to be strongly expressed in SCLC, and was detected in 3 out of 11 SCLC cell lines and 1 out of 2 SCLC samples (Bangur et al. 2004). Although the biological significance of MS4A8 remains unclear, MS4A8 may be one of the representative molecules of ASCL1-expressing SCLC, and has potential utility as a biomarker and therapeutic target of SCLC. One of the selected ASCL1-target molecules in PNEC is COBL. COBL functions as an actin nucleation factor and modulates cell morphology because it is localized at sites of high actin dynamics (Ahuja et al. 2007). However, the significance of COBL has not yet been examined with a focus on cell biological functions in cell morphology and motility in carcinomas, except for a study on prognostic genes for squamous cell carcinoma of the lung (Zhang et al. 2018b).
In summary, in the present study, we performed comparative RNA-seq studies on two sets of samples including H69, an ASCL1-positive classical SCLC cell line with H69AR, an ASCL1-negative variant SCLC cell line, and an A549(ASCL1+) cell line with a A549(control) cell line to clarify ASCL1-related molecules. Based on previous ChIP-seq studies (Augustyn et al. 2014; Borromeo et al. 2016) and NE score data (Zhang et al. 2018a), 120 molecules were confirmed to be common ASCL1-associated molecules totally, and further 7 molecules were carefully selected after qRT-PCR analyses of the 120 molecules in 5 ASCL1-positive and 4 ASCL1-negative cell lines. Their putative functions are related to neuronal development and secretion, proliferation, and morphology, and positive immunohistochemical staining was confirmed in formalin-fixed and paraffin-embedded samples.
Abbreviations
- ASCL1:
-
Achaete-scute complex homologue 1
- HES1:
-
Hairy and enhancer of split 1
- CHGA:
-
Chromogranin A
- CHGB:
-
Chromogranin B
- SCG2:
-
Secretogranin 2
- SYP:
-
Synaptophysin
- NCAM:
-
Neural cell adhesion molecule
- GRP:
-
Gastrin releasing peptide,
- SYT1:
-
Synaptotagmin1
- INSM1:
-
Insulinoma associated protein 1
- SNAP25:
-
Synaptosome associated protein 25
- YAP1:
-
Yes associated protein 1
- NEUROD1:
-
Neurogenic differentiation 1
- POU2F3:
-
POU class 2 homeobox 3
- CPLX2:
-
Complexin 2
- SYTL3:
-
Synaptotagmin like 3
- NTRK3:
-
Neurotrophic receptor tyrosine kinase 3
- NCF2:
-
Neutrophil cytosolic factor 2
- GAS2:
-
Growth arrest specific 2
- CCND2:
-
Cyclin D2
- INHBB:
-
Inhibin subunit beta B
- CREG1:
-
Cellular repressor of E1A stimulated genes 1
- PLK2:
-
Polo-like kinase 2
- CCND3:
-
Cyclin D3
- CLDN4:
-
Claudin 4
- BCAM:
-
Basal cell adhesion molecule
- JUP:
-
Junction plakoglobin
- CDH1:
-
Cadherin 1
- CD44:
-
CD44 molecule
- CLMP:
-
CXADR like membrane protein
- ISL1:
-
Islet1
- SYT4:
-
Synaptotagmin4
- KCTD16:
-
Potassium channel tetramerization domain containing 16
- SEZ6:
-
Seizure-related 6 homolog
- MS4A8:
-
Membrane spanning 4 domains A8
- COBL:
-
Cordon-bleu WH2 repeat protein
- BDNF:
-
Brain derived neurotrophic factor
- NFASC:
-
Neurofascin
- DMD:
-
Dystrophin
- GDNF:
-
Glial cell derived neurotrophic factor
- VCAN:
-
Versican
- ABCC4:
-
ATP binding cassette subfamily C member 4
- ABCC8:
-
ATP binding cassette subfamily C member 8
- SLC30A3:
-
Solute carrier family 30 member 3
- SLC39A6:
-
Solute carrier family 39 member 6
- SLC40A1:
-
Solute carrier family 40 member 1
References
Agaimy A, Erlenbach-Wünsch K, Konukiewitz B, Schmitt AM, Rieker RJ, Vieth M, Kiesewetter F, Hartmann A, Zamboni G, Perren A, Klöppel G (2013) ISL1 expression is not restricted to pancreatic well-differentiated neuroendocrine neoplasms, but is also commonly found in well and poorly differentiated neuroendocrine neoplasms of extrapancreatic origin. Mod Pathol 26(7):995–1003
Ahuja R, Pinyol R, Reichenbach N, Custer L, Klingensmith J, Kessels MM, Qualmann B (2007) Cordon-bleu is an actin nucleation factor and controls neuronal morphology. Cell 131(2):337–350
Augustyn A, Borromeo M, Wang T, Fujimoto J, Shao C, Dospoy PD, Lee V, Tan C, Sullivan JP, Larsen JE, Girard L, Behrens C, Wistuba II, Xie Y, Cobb MH, Gazdar AF, Johnson JE, Minna JD (2014) ASCL1 is a lineage oncogene providing therapeutic targets for high-grade neuroendocrine lung cancers. Proc Natl Acad Sci USA 111(41):14788–14793
Ball DW (2004) Achaete-scute homolog-1 and notch in lung neuroendocrine development and cancer. Cancer Lett 204(2):159–169
Bangur CS, Johnson JC, Switzer A, Wang YH, Hill B, Fanger GR, Wang T, Retter MW (2004) Identification and characterization of L985P, a CD20 related family member over-expressed in small cell lung carcinoma. Int J Oncol 25(6):1583–1590
Borges M, Linnoila RI, van de Velde HJ, Chen H, Nelkin BD, Mabry M, Baylin SB, Ball DW (1997) An achaete-scute homologue essential for neuroendocrine differentiation in the lung. Nature 386(6627):852–855
Borromeo MD, Savage TK, Kollipara RK, He M, Augustyn A, Osborne JK, Girard L, Minna JD, Gazdar AF, Cobb MH, Johnson JE (2016) ASCL1 and NEUROD1 reveal heterogeneity in pulmonary neuroendocrine tumors and regulate distinct genetic programs. Cell Rep 16(5):1259–1272
Brambilla E, Beasley MB, Austin JHM, Capelozzi VL, Chirieac LR, Devesa SS, Frank GA, Gazder A, Ishikawa Y, Jen J, Jett J, Marchevsky AM, Nicholson S, Pelosi G, Powell CA, Rami- Porta R, Scagliotti G, Thunnissen E, Travis WD, van Schil P, Yang P (2015) Neuroendocrine tumours small cell carcinoma. In: Travis WD, Brambilla E, Burke AP, Marx A, Nicholson AG (eds) WHO classification of tumours of the Lung, Pleura, Thymus and Heart, 4th edn. IARC, Lyon, pp 63–68
Bunn PA Jr, Minna JD, Augustyn A, Gazdar AF, Ouadah Y, Krasnow MA, Berns A, Brambilla E, Rekhtman N, Massion PP, Niederst M, Peifer M, Yokota J, Govindan R, Poirier JT, Byers LA, Wynes MW, McFadden DG, MacPherson D, Hann CL, Farago AF, Dive C, Teicher BA, Peacock CD, Johnson JE, Cobb MH, Wendel HG, Spigel D, Sage J, Yang P, Pietanza MC, Krug LM, Heymach J, Ujhazy P, Zhou C, Goto K, Dowlati A, Christensen CL, Park K, Einhorn LH, Edelman MJ, Giaccone G, Gerber DE, Salgia R, Owonikoko T, Malik S, Karachaliou N, Gandara DR, Slotman BJ, Blackhall F, Goss G, Thomas R, Rudin CM, Hirsch FR (2016) Small cell lung cancer: can recent advances in biology and molecular biology be translated into improved outcomes? J Thorac Oncol 11(4):453–474
Carney DN, Gazdar AF, Bepler G, Guccion JG, Marangos PJ, Moody TW, Zweig MH, Minna JD (1985) Establishment and identification of small cell lung cancer cell lines having classic and variant features. Cancer Res 45(6):2913–2923
Demelash A, Rudrabhatla P, Pant HC, Wang X, Amin ND, McWhite CD, Naizhen X, Linnoila RI (2012) Achaete-scute homologue-1 (ASH1) stimulates migration of lung cancer cells through Cdk5/p35 pathway. Mol Biol Cell 23(15):2856–2866
Domenichini A, Adamska A, Falasca M (2019) ABC transporters as cancer drivers: potential functions in cancer development. Biochim Biophys Acta 1:52–60
Eon Kuek L, Leffler M, Mackay GA, Hulett MD (2016) The MS4A family: counting past 1, 2 and 3. Immunol Cell Biol 94(1):11–23
Fujino K, Motooka Y, Hassan WA, Ali Abdalla MO, Sato Y, Kudoh S, Hasegawa K, Niimori-Kita K, Kobayashi H, Kubota I, Wakimoto J, Suzuki M, Ito T (2015) Insulinoma-associated protein 1 is a crucial regulator of neuroendocrine differentiation in lung cancer. Am J Pathol 185(12):3164–3177
Fujino K, Yasufuku K, Kudoh S, Motooka Y, Sato Y, Wakimoto J, Kubota I, Suzuki M, Ito T (2017) INSM1 is the best marker for the diagnosis of neuroendocrine tumors: comparison with CGA, SYP andCD56. Int J Clin Exp Pathol 10(5):5393–5405
Gazdar AF, Carney DN, Nau MM, Minna JD (1985) Characterization of variant subclasses of cell lines derived from small cell lung cancer having distinctive biochemical, morphological, and growth properties. Cancer Res 45(6):2924–2930
Gazdar AF, Bunn PA, Minna JD (2017) Small-cell lung cancer: what we know, what we need to know and the path forward. Nat Rev Cancer 17(12):765
Guillemot F, Lo LC, Johnson JE, Auerbach A, Anderson DJ, Joyner AL (1993) Mammalian achaete-scute homolog 1 is required for the early development of olfactory and autonomic neurons. Cell 75(3):463–476
Gunnersen JM, Kim MH, Fuller SJ, De Silva M, Britto JM, Hammond VE, Davies PJ, Petrou S, Faber ES, Sah P, Tan SS (2007) Sez-6 proteins affect dendritic arborization patterns and excitability of cortical pyramidal neurons. Neuron 56(4):621–639
Huang DW, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 4:44–57
Ito T, Udaka N, Yazawa T, Okudela K, Hayashi H, Sudo T, Guillemot F, Kageyama R, Kitamura H (2000) Basic helix-loop-helix transcription factors regulate the neuroendocrine differentiation of fetal mouse pulmonary epithelium. Development 127(18):3913–3921
Ito T, Udaka N, Okudela K, Yazawa T, Kitamura H (2003) Mechanisms of neuroendocrine differentiation in pulmonary neuroendocrine cells and small cell carcinoma. Endocr Pathol 14(2):133–139
Ito T, Kudoh S, Ichimura T, Fujino K, Hassan WA, Udaka N (2017) Small cell lung cancer, an epithelial to mesenchymal transition (EMT)-like cancer: significance of inactive Notch signaling and expression of achaete-scute complex homologue 1. Hum Cell 30(1):1–10
Jiang L, Huang J, Higgs BW, Hu Z, Xiao Z, Yao X, Conley S, Zhong H, Liu Z, Brohawn P, Shen D, Wu S, Ge X, Jiang Y, Zhao Y, Lou Y, Morehouse C, Zhu W, Sebastian Y, Czapiga M, Oganesyan V, Fu H, Niu Y, Zhang W, Streicher K, Tice D, Zhao H, Zhu M, Xu L, Herbst R, Su X, Gu Y, Li S, Huang L, Gu J, Han B, Jallal B, Shen H, Yao Y (2016) Genomic landscape survey identifies SRSF1 as a key oncodriver in small cell lung cancer. PLoS Genet. 12(4):1005895
Jiang T, Collins BJ, Jin N, Watkins DN, Brock MV, Matsui W, Nelkin BD, Ball DW (2009) Achaete-scute complex homologue 1 regulates tumor-initiating capacity in human small cell lung cancer. Cancer Res 69(3):845–854
Kathawala RJ, Gupta P, Ashby CR Jr, Chen ZS (2015) The modulation of ABC transporter-mediated multidrug resistance in cancer: a review of the past decade. Drug Resist Updat 18:1–17
Kim MH, Gunnersen JM, Tan SS (2002) Localized expression of the seizure-related gene SEZ-6 in developing and adult forebrains. Mech Dev 118(1–2):171–174
Li Q, Shu Y (2014) Role of solute carriers in response to anticancer drugs. Mol Cell Ther 27(2):15
Li Y, Linnoila RI (2012) Multidirectional differentiation of achaete-scute homologue-1-defined progenitors in lung development and injury repair. Am J Respir Cell Mol Biol 47(6):768–775
Linnoila RI, Zhao B, DeMayo JL, Nelkin BD, Baylin SB, DeMayo FJ, Ball DW (2000) Constitutive achaete-scute homologue-1 promotes airway dysplasia and lung neuroendocrine tumors in transgenic mice. Cancer Res 60(15):4005–4009
Meder L, König K, Ozretić L, Schultheis AM, Ueckeroth F, Ade CP, Albus K, Boehm D, Rommerscheidt-Fuss U, Florin A, Buhl T, Hartmann W, Wolf J, Merkelbach-Bruse S, Eilers M, Perner S, Heukamp LC, Buettner R (2016) NOTCH, ASCL1, p53 and RB alterations define an alternative pathway driving neuroendocrine and small cell lung carcinomas. Int J Cancer 138(4):927–938
Michel J, Schönhaar K, Schledzewski K, Gkaniatsou C, Sticht C, Kellert B, Lasitschka F, Géraud C, Goerdt S, Schmieder A (2013) Identification of the novel differentiation marker MS4A8B and its murine homolog MS4A8A in colonic epithelial cells lost during neoplastic transformation in human colon. Cell Death Dis 4:e469. https://doi.org/10.1038/cddis.2012.215
Mirski SE, Gerlach JH, Cole SP (1987) Multidrug resistance in a human small cell lung cancer cell line selected in adriamycin. Cancer Res 47(10):2594–2598
Moghadam PK, Jackson MB (2013) The functional significance of synaptotagmin diversity in neuroendocrine secretion. Front Endocrinol (Lausanne) 18(4):124
Morimoto M, Nishinakamura R, Saga Y, Kopan R (2012) Different assemblies of Notch receptors coordinate the distribution of the major bronchial Clara, ciliated and neuroendocrine cells. Development 139(23):4365–4373
Motooka Y, Fujino K, Sato Y, Kudoh S, Suzuki M, Ito T (2017) Pathology of Notch2 in lung cancer. Pathology 49:486–493
Mukhopadhyay S, Dermawan JK, Lanigan CP, Farver CF (2019) Insulinoma-associated protein 1 (INSM1) is a sensitive and highly specific marker of neuroendocrine differentiation in primary lung neoplasms: an immunohistochemical study of 345 cases, including 292 whole-tissue sections. Mod Pathol 32(1):100–109
Osada H, Tatematsu Y, Yatabe Y, Horio Y, Takahashi T (2005) ASH1 gene is a specific therapeutic target for lung cancers with neuroendocrine features. Cancer Res 65(23):10680–10685
Osada H, Tomida S, Yatabe Y, Tatematsu Y, Takeuchi T, Murakami H, Kondo Y, Sekido Y, Takahashi T (2008) Roles of achaete-scute homologue 1 in DKK1 and E-cadherin repression and neuroendocrine differentiation in lung cancer. Cancer Res 68(6):1647–1655
Pietanza MC, Byers LA, Minna JD, Rudin CM (2015) Small cell lung cancer: will recent progress lead to improved outcomes? Clin Cancer Res 21(10):2244–2255
Rooper LM, Sharma R, Li QK, Illei PB, Westra WH (2017) INSM1 demonstrates superior performance to the individual and combined use of synaptophysin, chromogranin and CD56 for diagnosing neuroendocrine tumors of the thoracic cavity. Am J Surg Pathol 41(11):1561–1569
Rosenbaum JN, Guo Z, Baus RM, Werner H, Rehrauer WM, Lloyd RV (2015) INSM1: a novel immunohistochemical and molecular marker for neuroendocrine and neuroepithelial neoplasms. Am J Clin Pathol. 144(4):579–591
Rudin CM, Poirier JT, Byers LA, Dive C, Dowlati A, George J, Heymach JV, Johnson JE, Lehman JM, MacPherson D, Massion PP, Minna JD, Oliver TG, Quaranta V, Sage J, Thomas RK, Vakoc CR, Gazdar AF (2019) Molecular subtypes of small cell lung cancer: a synthesis of human and mouse model data. Nat Rev Cancer 19(5):289–297
Schmitt AM, Riniker F, Anlauf M, Schmid S, Soltermann A, Moch H, Heitz PU, Klöppel G, Komminoth P, Perren A (2008) Islet 1 (Isl1) expression is a reliable marker for pancreatic endocrine tumors and their metastases. Am J Surg Pathol 32(3):420–425
Sriuranpong V, Borges MW, Ravi RK, Arnold DR, Nelkin BD, Baylin SB, Ball DW (2001) Notch signaling induces cell cycle arrest in small cell lung cancer cells. Cancer Res 61(7):3200–3205
Stützer I, Selevsek N, Esterházy D, Schmidt A, Aebersold R, Stoffel M (2013) Systematic proteomic analysis identifies β-site amyloid precursor protein cleaving enzyme 2 and 1 (BACE2 and BACE1) substrates in pancreatic β-cells. J Biol Chem 288(15):10536–10547
Sun L, Zhang Y, Zhang C (2018) Distinct expression and prognostic value of MS4A in gastric cancer. Open Med (Wars) 13:178–188
Teng X, Aouacheria A, Lionnard L, Metz KA, Soane L, Kamiya A, Hardwick JM (2019) KCTD: a new gene family involved in neurodevelopmental and neuropsychiatric disorders. CNS Neurosci Ther 25(7):887–902
Tenjin Y, Kudoh S, Kubota S, Yamada T, Matsuo A, Sato Y, Ichimura T, Kohrogi H, Sashida G, Sakagami T, Ito T (2019) Ascl1-induced Wnt11 regulates neuroendocrine differentiation, cell proliferation, and E-cadherin expression in small-cell lung cancer and Wnt11 regulates small-cell lung cancer biology. Lab Invest. https://doi.org/10.1038/s41374-019-0277-y
Travis WD, Brambilla E, Burke AP et al (2015) WHO Classification of tumours of the lung, pleura thymus and heart. International Agency for Research on Cancer, Lyon
van Coevorden-Hameete MH, de Bruijn MAAM, de Graaff E, Bastiaansen DAEM, Schreurs MWJ, Demmers JAA, Ramberger M, Hulsenboom ESP, Nagtzaam MMP, Boukhrissi S, Veldink JH, Verschuuren JJGM, Hoogenraad CC, Sillevis Smitt PAE, Titulaer MJ (2019) The expanded clinical spectrum of anti-GABABR encephalitis and added value of KCTD16 autoantibodies. Brain 142(6):1631–1643
Vician L, Lim IK, Ferguson G, Tocco G, Baudry M, Herschman HR (1995) Synaptotagmin IV is an immediate early gene induced by depolarization in PC12 cells and in brain. Proc Natl Acad Sci USA 92(6):2164–2168
Wael H, Yoshida R, Kudoh S, Hasegawa K, Niimori-Kita K, Ito T (2014) Notch1 signaling controls cell proliferation, apoptosis and differentiation in lung carcinoma. Lung Cancer 85(2):131–140
Ye L, Yao XD, Wan FN, Qu YY, Liu ZY, Shen XX, Li S, Liu XJ, Yue F, Wang N, Dai B, Ye DW (2014) MS4A8B promotes cell proliferation in prostate cancer. Prostate 74(9):911–922
Zhang W, Girard L, Zhang YA, Haruki T, Papari-Zareei M, Stastny V, Ghayee HK, Pacak K, Oliver TG, Minna JD, Gazdar AF (2018a) Small cell lung cancer tumors and preclinical models display heterogeneity of neuroendocrine phenotypes. Transl Lung Cancer Res 7(1):32–49
Zhang W, Cui Q, Qu W, Ding X, Jiang D, Liu H (2018b) TRIM58/cg26157385 methylation is associated with eight prognostic genes in lung squamous cell carcinoma. Oncol Rep 40(1):206–216
Zuo H, Glaaser I, Zhao Y, Kurinov I, Mosyak L, Wang H, Liu J, Park J, Frangaj A, Sturchler E, Zhou M, McDonald P, Geng Y, Slesinger PA, Fan QR (2019) Structural basis for auxiliary subunit KCTD16 regulation of the GABAB receptor. Proc Natl Acad Sci USA 116(17):8370–8379
Acknowledgments
We thank Ms. Takako Maeda for her technical assistance and Mr. Shingo Usuki and the staff of the LILA of the Institute of Molecular Embryology and Genetics, Kumamoto University for their technical support of the RNA-seq analysis. This study was supported in part by the program of the Joint Usage/Research Center for Developmental Medicine, Institute of Molecular Embryology and Genetics, Kumamoto University, by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan (18K19480), by a Grant from the Smoking Research Foundation, and by an endowment from Dr. Yuko Aihara of Aihara Allergy and Pediatric Clinic.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest:
All authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kudoh, S., Tenjin, Y., Kameyama, H. et al. Significance of achaete-scute complex homologue 1 (ASCL1) in pulmonary neuroendocrine carcinomas; RNA sequence analyses using small cell lung cancer cells and Ascl1-induced pulmonary neuroendocrine carcinoma cells. Histochem Cell Biol 153, 443–456 (2020). https://doi.org/10.1007/s00418-020-01863-z
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-020-01863-z