Abstract
Evidence of genetic heterogeneity in ALS has been found, with at least 31 genes being identified to date as causing ALS, and other genes being suggested as risk factors for susceptibility to the disease and for phenotype modifications. In recent years, new molecular genetic methodologies, especially GWAS and exome sequencing, have contributed to the identification of new ALS genes. Some of these genes (SOD1, TARDBP, FUS, and C9orf72) have homogenous frequencies in different populations. However, a few genes are rare in populations other than those in which they were first identified. Here we investigate the frequency of the PFN1 gene in a Catalan ALS population. A mutational analysis of the PFN1 gene was carried out on a Catalan cohort of 42 ALS families (FALS) and 423 sporadic ALS patients (SALS). The screening included 600 healthy controls. No PFN1 mutations were identified in either the FALS or SALS group. We also found no mutations in the control group. Our results are consistent with those described in other populations with very low frequencies, suggesting that PFN1 is a very rare cause of ALS worldwide. Together with the absence of a distinctive phenotype associated with ALS18, these results mean that this gene should be a second or third line for inclusion in screening in patients requesting genetic counseling.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disorder predominantly affecting motor neurons, leading to generalized muscle paralysis and death from respiratory failure. Mean survival subsequent to clinical onset is 3–5 years, although patients may survive longer if they agree to mechanical ventilation and other invasive life support procedures. There is no effective treatment for the disease apart from these measures, except for riluzole, which is the only approved drug for the treatment of ALS, and which prolongs survival by 3 months.
Although its etiopathogenesis is still unclear, ALS is considered a multifactorial disease with interactive multiple pathogenic mechanisms. A number of genetic studies suggest that genetic defects in some ALS patients make them susceptible to the disease. This is true of both the familial forms of ALS (FALS) and some of the condition’s sporadic forms (SALS) [1–3].
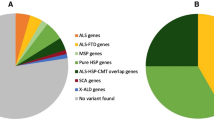
The genetics of ALS are complex. At least 31 genes and 8 different genetic loci with dominant, recessive and X-linked patterns of inheritance have been identified for FALS, and an increasing number of susceptible and modifier loci have also been suggested for sporadic and familial ALS cases. Gene–gene and gene–environment interactions also seem to play an important role in the appearance and phenotype of the disease [1–8].
Most of these genes have been described in the last 3–4 years, thanks to new gene screening methodologies, especially genome-wide association (GWAS) studies, genomic structural variation studies, and whole exome sequencing (WES), which have revolutionized gene testing. However, the role of many genes has yet to be verified in large cohorts other than those in which they have been reported, as doubts remain as to the credibility of the analysis of these putative causing genes identified using bioinformatics [1–3, 9, 10]. It is therefore important to investigate the prevalence of genetic defects in different populations worldwide.
Missense mutations in the profilin 1 gene (PFN1) have recently been identified in two large ALS families of Caucasian and Sephardic Jewish origin using exome sequencing [8]. In the same paper, the authors identified 4 mutations in 7 of the 274 FALS cases. PFN-1 has subsequently been reported in FALS, SALS, FTLD and healthy populations in mutational analyses of other populations [11–23]. Profilin 1 is a small 140 amino acid protein that is essential for the polymerization of monomeric G-actin to filamentous-actin [24, 25]. Although it mainly acts by stimulating actin polymerization, Profilin 1 and its ligands are involved in other processes, including vesicle trafficking, axonal transport, glutamate neuromodulation, endocytosis and nuclear export [25–28]. All these target mechanisms have been proposed as hypotheses of neuronal damage in ALS.
Despite the lack of consensus on unified criteria regarding how to numerate the various genetic forms of ALS, in contrast to other neuromuscular diseases, there is a tendency to numerate them, as can be seen on websites containing gene-specific content, which define this form of ALS caused by PFN1 mutations as ALS18 (#614808) [29, 30]. Genetic characterization of ALS18 families should provide information on the distribution of PFN1 mutations in different ethnic groups. The prevalence of ALS18 in Catalonia, a region of approximately 7,000,000 inhabitants in northeastern Spain, has not been studied. Here we report on the results from 42 FALS index cases and 423 apparently sporadic ALS cases.
Materials and methods
Study population
This study included a cohort of 42 FALS index cases and 423 SALS patients. The patients had been diagnosed with definite, probable, and probable—laboratory-supported ALS according to the El Escorial revised criteria at the ALS Unit in the Neurology Department of Hospital Universitari Vall d’Hebron in Barcelona. The criteria used by Byrne et al. [31] were adopted to identify FALS cases, and individuals presenting no family history of ALS were considered sporadic ALS cases. The study population consisted of patients originating in Catalonia whose four grandparents were born in Catalonia, except for 16 patients of Hispanic, African and Asian origin.
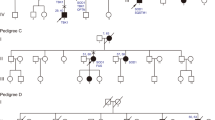
During the study of familial forms, the patients were asked about relatives with similar symptoms and other family members diagnosed with dementia. Whenever possible, both patients with dementia as their initial symptom and those with frontotemporal signs appearing after motor neuron signs were evaluated by the ALS Unit’s neuropsychologist, who sought findings pointing to FTLD.
All the patients had previously been screened for SOD1, TARDBP, FUS/TLS, and C9orf72 mutations. Eleven FALS and four SALS cases were found to have a SOD1 mutation [32]; two FALS and one SALS patients had an FUS/TLS mutation [33]; six FALS and three SALS patients had an expansion for C9orf72 (unpublished data). These ALS patients with identified mutations other than PFN1 were included in the study to investigate the co-occurrence of more than one genetic defect among FALS patients, a phenomenon recently described as occurring in 1.6 % of patients [9].
We included blood samples from 600 healthy subjects of Spanish origin extracted from the patients’ spouses and blood donors as a control population.
Informed written consent was provided by patients or family members (when the patients had writing difficulties) and from the control group, in accordance with a protocol approved by our institution’s Clinical Research Ethics Committee, and with the Declaration of Helsinki [34].
Mutational analysis
Genomic DNA was extracted from peripheral whole blood using a QIAamp DNA Blood Minikit (Qiagen, Valencia, CA, USA). All three coding exons of the PFN1 gene and the intron/exon boundaries were amplified by PCR as previously described [8], purified, and then forward and reverse sequenced using sequencing primers, the Cycle Sequencing Kit and the Big-Dye Terminator v3.1 kit (Applied Biosystems, Foster City, CA, USA). After purification, the products were run on an ABI 3730xl Genetic Analyzer (Applied Biosystems). Sequence analysis was performed using Sequencher 4.8 software (Gene Codes, Ann Arbor, MI, USA). The sequences obtained were compared with the PFN1 reference sequence (GenBank entry NM_005022).
Results
Clinical data
The mean age at onset in the FALS group was 51.4 ± 13.7 years (range 21–92). In the SALS group it was 59.6 ± 12.9 years (range 23–89). The demographic and clinical characteristics are presented in Table 1.
There was no significant difference between ALS patients and the control group in terms of the age and gender of the patients. The mean survival time among the ALS patients who had died was 36.7 ± 27.5 months (range 6–244 months).
Mutation analysis of the PFN1 gene
PFN1 sequencing analysis failed to identify any pathogenic mutations or synonymous variants in any of the three groups (FALS, SALS, and control population).
Discussion
In this study, we present a mutational analysis for the PFN1 gene in a Catalan cohort of 42 FALS and 423 SALS patients. The prevalence of PFN1 in ALS has not previously been studied in Spain. We did not identify any new mutations, previously reported pathogenic mutations, or silent mutations. These results indicate that mutations in the PFN1 gene are an improbable cause of FALS and SALS in our population. Our results contribute to an improved knowledge of the true prevalence of defects in the PFN1 gene among the ALS population.
These negative findings are consistent with those described in ALS patients of European and Chinese ancestry. At least 13 studies have investigated the prevalence of PFN1 gene mutations in their populations since it was first identified in two large ALS families of Caucasian and Sephardic Jewish origin. To date, the prevalence of PFN1 gene mutations has been investigated in ALS patients in China, France, Canada, Belgium, Bulgaria, the United States, Germany, Scandinavia, Italy, and Australia. Clearly negative results (i.e. no pathogenic mutations) have been reported in Chinese, French Canadian, Belgian and Bulgarian patients (see Table 2). In short, PFN1 mutations have to date been checked in just over 10,000 ALS patients worldwide, with an incidence of 0.8 % for FALS, 0.25 % for SALS, and 0.1 % for the control population. For most authors, the incidence of PFN1 mutations in ALS is similar to those reported in public SNP databases.
These data suggest that in comparison with C9orf72, SOD1, FUS/TLS and TARDBP, PFN1 is not a major gene causing ALS. There are doubts as to the role of PFN1 in ALS, especially bearing in mind that: (1) most of the variants that have been reported (e.g. p.T16T, p.D19D, p.T35T, p.G38G, p.Q69Q, p.L88L, p.V98V, p.L112L, p.S133S, and p.Q139Q) are polymorphisms (silent mutations), which do not modify the encoded amino acid, (2) proving the pathogenicity of the most prevalent non-synonymous variant (p.E117G) has been extremely difficult, and (3) for this mutation the prevalence was not significantly increased in ALS patients versus healthy control subjects (see Table 2). These doubts are reinforced by the fact that the prevalence of all the PFN1 variants reported to date among ALS patients is similar to the prevalence among the almost 20,000 healthy controls analyzed. Furthermore, there is no distinctive phenotype for ALS18, suggesting that it is a possible profilinopathy. The phenotype of ALS18 is similar to that of ALS1, i.e. a variable age at onset (between 27 and over 70 years old), predominant lower motor neuron signs, exceptional signs of dementia and a third of patients experiencing bulbar onset. Obligate asymptomatic carriers have also been described in ALS18.
In short, our study provides information on the rarity of profilin 1 mutations in our ALS population, in both sporadic and familial forms. These data do not replicate the results of the seminal study in 2012. Together with the absence of a distinctive phenotype associated with ALS18, these results mean that this gene should be a second or third line for inclusion in screening in patients requesting genetic counseling.
References
Andersen PM, Al-Chalabi A (2011) Clinical genetics of amyotrophic lateral sclerosis: what do we really know? Nat Rev Neurol 7(11):603–615
Al-Chalabi A, Hardiman O (2013) The epidemiology of ALS: a conspiracy of genes, environment and time. Nat Rev Neurol 9(11):617–628
Renton AE, Chiò A, Traynor BJ (2014) State of play in amyotrophic lateral sclerosis genetics. Nat Neurosci 17(1):17–23
DeJesus-Hernandez M, Mackenzie IR, Boeve BF, Boxer AL, Baker M, Rutherford NJ et al (2011) Expanded GGGGCC hexanucleotide repeat in noncoding region of C9ORF72 causes chromosome 9p-linked FTD and ALS. Neuron 72(2):245–256
Kabashi E, Valdmanis PN, Dion P, Spiegelman D, McConkey BJ, VandeVelde C et al (2008) TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nat Genet 40(5):572–574
Leblond CS, Kaneb HM, Dion PA, Rouleau GA (2014) Dissection of genetic factors associated with amyotrophic lateral sclerosis. Exp Neurol. pii: S0014-4886(14)00115-0
Vance C, Rogelj B, Hortobágyi T, De Vos KJ, Nishimura AL, Sreedharan J et al (2009) Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323(5918):1208–1211
Wu CH, Fallini C, Ticozzi N, Keagle PJ, Sapp PC, Piotrowska K et al (2012) Mutations in the profilin 1 gene cause familial amyotrophic lateral sclerosis. Nature 488(7412):499–503
Van Blitterswijk M, van Es MA, Hennekam EA, Dooijes D, van Rheenen W, Medic J et al (2012) Evidence for an oligogenic basis of amyotrophic lateral sclerosis. Hum Mol Genet 21(17):3776–3784
Abel O, Powell JF, Andersen PM, Al-Chalabi A (2013) Credibility analysis of putative disease-causing genes using bioinformatics. PLoS ONE 8(6):e64899
Calvo A, Restangno G, Brunetti M, Barberis M, Ossola I, Moglia C et al (2013) PFN1 mutations are an uncommon cause of familial amyotrophic lateral sclerosis. Abstract P174, 24th international symposium on ALS/MND, Milan
Chen Y, Zheng ZZ, Huang R, Chen K, Song W, Zhao B et al (2013) PFN1 mutations are rare in Han Chinese populations with amyotrophic lateral sclerosis. Neurobiol Aging 34(7):1922.e1–5
Daoud H, Dobrzeniecka S, Camu W, Meininger V, Dupré N, Dion PA, Rouleau GA (2013) Mutation analysis of PFN1 in familial amyotrophic lateral sclerosis patients. Neurobiol Aging 34(4):1311.e1–2
Fratta P, Charnock J, Collins T, Devoy A, Howard R, Malaspina A et al (2014) Profilin1 E117G is a moderate risk factor for amyotrophic lateral sclerosis. J Neurol Neurosurg Psychiatry 85(5):506–508
Ingre C, Landers J, Rizik N, Volk A, Akimoto C, Birve A et al (2013) Mechanisms of a novel phosphorylation site mutation in profilin 1. Abstract P172, 24th international symposium on ALS/MND, Milan
Ingre C, Landers JE, Rizik N, Volk AE, Akimoto C, Birve A et al (2013) A novel phosphorylation site mutation in profilin 1 revealed in a large screen of US, Nordic, and German amyotrophic lateral sclerosis/frontotemporal dementia cohorts. Neurobiol Aging 34(6):1708.e1–6
Lattante S, Le Ber I, Camuzat A, Brice A, Kabashi E (2013) Mutations in the PFN1 gene are not a common cause in patients with amyotrophic lateral sclerosis and frontotemporal lobar degeneration in France. Neurobiol Aging 34(6):1709.e1–2
Soong BW, Lin KP, Guo YC, Lin CC, Tsai PC, Liao YC et al (2014) Extensive molecular genetic survey of Taiwanese patients with amyotrophic lateral sclerosis. Neurobiol Aging. doi:10.1016/j.neurobiolaging.2014.05.008.
Tiloca C, Ticozzi N, Pensato V, Bagarotti A, Del Bo R, Gagliardi S et al (2013) Screening of the PFN1 gene in sporadic amyotrophic lateral sclerosis and in frontotemporal dementia. Abstract P173, 24th international symposium on ALS/MND, Milan
Tiloca C, Ticozzi N, Pensato V, Corrado L, Del Bo R, Bertolin C et al (2013) Screening of the PFN1 gene in sporadic amyotrophic lateral sclerosis and in frontotemporal dementia. Neurobiol Aging 34(5):1517.e9–10
Van Blitterswijk M, Baker MC, Bieniek KF, Knopman DS, Josephs KA, Boeve B et al (2013) Profilin-1 mutations are rare in patients with amyotrophic lateral sclerosis and frontotemporal dementia. Amyotroph Lateral Scler Frontotemporal Degener 14(5–6):463–469
Yang S, Fifita JA, Williams KL, Warraich ST, Pamphlett R, Nicholson GA et al (2013) Mutation analysis and immunopathological studies of PFN1 in familial and sporadic amyotrophic lateral sclerosis. Neurobiol Aging 34(9):2235.e7–10
Zou ZY, Sun Q, Liu MS, Li XG, Cui LY (2013) Mutations in the profilin 1 gene are not common in amyotrophic lateral sclerosis of Chinese origin. Neurobiol Aging 34(6):1713.e5–6
Witke W, Podtelejnikov AV, Di Nardo A, Sutherland JD, Gurniak CB, Dotti C et al (1998) In mouse brain profilin I and profilin II associate with regulators of the endocytic pathway and actin assembly. EMBO J 17(4):967–976
Witke W (2004) The role of profilin complexes in cell motility and other cellular processes. Trends Cell Biol 14(8):461–469
Munsie LN, Truant R (2012) The role of the cofilin-actin rod stress response in neurodegenerative diseases uncovers potential new drug targets. Bioarchitecture 2(6):204–208
Kullmann JA, Neumeyer A, Wickertsheim I, Böttcher RT, Costell M, Deitmer JW et al (2012) Purkinje cell loss and motor coordination defects in profilin1 mutant mice. Neuroscience 25(223):355–364
Caraballo-Miralles V, Cardona-Rossinyol A, Garcera A, Villalonga P, Soler RM, Olmos G et al (2012) SMN deficiency attenuates migration of U87MG astroglioma cells through the activation of RhoA. Mol Cell Neurosci 49(3):282–289
http://omim.org/entry/614808?search=614808&highlight=614808. Retrieved 8 Sep 2014
http://www.genecards.org/cgi-bin/carddisp.pl?gene=PFN1&search=f688fbbdb89dec910cf355b001d57020. Retrieved 8 Sep 2014
Byrne S, Bede P, Elamin M, Kenna K, Lynch C, McLaughlin R et al (2011) Proposed criteria for familial amyotrophic lateral sclerosis. Amyotroph Lateral Scler 12(3):157–159
Gamez J, Corbera-Bellalta M, Nogales G, Raguer N, García-Arumí E, Badia-Canto M et al (2006) Mutational analysis of the Cu/Zn superoxide dismutase gene in a Catalan ALS population: should all sporadic ALS cases also be screened for SOD1? J Neurol Sci 247(1):21–28
Syriani E, Morales M, Gamez J (2011) FUS/TLS gene mutations are the second most frequent cause of familial ALS in the Spanish population. Amyotroph Lateral Scler 12(2):118–123
World Medical Association Declaration of Helsinki. Ethical principles for medical research involving human subjects. Clinical review and education. http://jama.jamanetwork.com. Published online: 19 Oct 2013. doi:10.1001/jama.2013.281053.
Dillen L, Van Langenhove T, Engelborghs S, Vandenbulcke M, Sarafov S, Tournev I et al (2013) Explorative genetic study of UBQLN2 and PFN1 in an extended Flanders-Belgian cohort of frontotemporal lobar degeneration patients. Neurobiol Aging 34(6):1711.e1–5
Acknowledgments
This study has been supported by Spanish Fondo de Investigaciones Sanitarias grants (PI10-01070-FEDER and FIS PI13-01272) and by a Marató de TV3 grant. The authors are indebted to the patients and their relatives for their cooperation.
Conflicts of interest
None.
Ethical standard
The authors declare that this study was approved by our hospital's ethics committee and was therefore been performed in accordance with the ethical standards laid down in the 1964 Declaration of Helsinki.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Syriani, E., Salvans, C., Salvadó, M. et al. PFN1 mutations are also rare in the Catalan population with amyotrophic lateral sclerosis. J Neurol 261, 2387–2392 (2014). https://doi.org/10.1007/s00415-014-7501-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00415-014-7501-x