Abstract
Centromeres are specialized chromosome domain that serve as the site for kinetochore assembly and microtubule attachment during cell division, to ensure proper segregation of chromosomes. In higher eukaryotes, the identity of active centromeres is marked by the presence of CENP-A (centromeric protein-A), a histone H3 variant. CENP-A forms a centromere-specific nucleosome that acts as a foundation for centromere assembly and function. The posttranslational modification (PTM) of histone proteins is a major mechanism regulating the function of chromatin. While a few CENP-A site-specific modifications are shared with histone H3, the majority are specific to CENP-A-containing nucleosomes, indicating that modification of these residues contribute to centromere-specific function. CENP-A undergoes posttranslational modifications including phosphorylation, acetylation, methylation, and ubiquitylation. Work from many laboratories have uncovered the importance of these CENP-A modifications in its deposition at centromeres, protein stability, and recruitment of the CCAN (constitutive centromere-associated network). Here, we discuss the PTMs of CENP-A and their biological relevance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Equal distribution of chromosomes during mitosis is critical for normal cellular functioning and organism development. Errors in chromosome segregation can lead to a state of aneuploidy, which is defined by the presence of extra or fewer chromosomes than normal. Cancers are frequently aneuploid, and the specific loss or gain of tumor suppressor and oncogenes associated with changes in chromosome number may contribute to the prevalence of aneuploidy in cancer (Giam and Rancati 2015; Holland and Cleveland 2009; Pfau and Amon 2012; Sen 2000).
The centromere is a chromosomal locus that orchestrates chromosome movement during mitosis and meiosis. The kinetochore assembles at the site of the centromere and provides the physical attachment site between the chromosome and microtubule spindle. Likewise, the mitotic checkpoint signaling apparatus assembles at the centromere and delays mitosis until chromosome alignment is achieved (Cleveland et al. 2003; Fukagawa and Earnshaw 2014; Westhorpe and Straight 2014).
Although the centromere is classically viewed as the central constriction in mitotic chromosomes, the centromere persists throughout the cell cycle (Allshire and Karpen 2008). Centromeric DNA in humans ranges from 0.3 to 5 M base pairs and is comprised of higher order repeats (HOR) of 171 base pairs α-satellites (Fukagawa and Earnshaw 2014). While many organisms show this pattern of restricted centromere assembly embedded in unique repetitive DNA sequences, several organisms have distributed their centromeres across the entire length of the chromosome known as holocentromeres. (Drinnenberg et al. 2014; Neumann et al. 2015). Holocentromeres may exist in diffused pattern or may occupy several discrete sites throughout the chromosome (Melters et al. 2012).
The location of the centromeres in many yeasts and most-higher eukaryotes is determined epigenetically by the centromere protein-A (CENP-A), independent of DNA sequence (Black and Cleveland 2011; Choo 2000; Henikoff and Dalal 2005; Henikoff and Furuyama 2010; Perpelescu and Fukagawa 2011). At centromeres, CENP-A replaces the canonical histone H3 and forms nucleosome with H2A, H2B, and H4 (Black and Cleveland 2011). The centromere requires CENP-A for the recruitment of the constitutive centromere-associated network (CCAN) and kinetochore proteins in order to facilitate proper chromosome segregation (Carroll et al. 2010; Carroll et al. 2009; Cheeseman and Desai 2008; Fachinetti et al. 2013; Foltz et al. 2006; Hori et al. 2008; Izuta et al. 2006; Kato et al. 2013; Okada et al. 2006). Neocentromeres are the new centromeres that arise naturally and are independent of underlying DNA sequence and recruit CENP-A (Lo et al. 2001; Saffery et al. 2000; Warburton 2004). Assembling CENP-A containing chromatin at non-centromeric loci is sufficient to establish the centromere and assembles a kinetochore capable of microtubule attachments, demonstrating the primary role of CENP-A nucleosomes in forming the centromere (Barnhart et al. 2011; Bergmann et al. 2011; Guse et al. 2011; Mendiburo et al. 2011). Notably, CENP-A-containing nucleosomes are not arranged in a continuous fashion, but are instead interspersed with H3-containing nucleosome that contain a distinctive set of posttranslational modifications important for centromere function (Blower et al. 2002; Fukagawa 2017; Garcia Del Arco and Erhardt 2017).
Posttranslational modifications of centromeric chromatin components have emerged as an important regulator of overall structure and function of centromeres. Compared with canonical histones, which are subjected to a large combinatorial array of posttranslational modifications that directly or indirectly regulate their function, the degree and the information about CENP-A posttranslational modifications are more limited. Nonetheless, CENP-A undergoes a variety of posttranslational modifications including phosphorylation, ubiquitylation, methylation, and acetylation on its amino terminus and histone-fold domain and this review will summarize the known posttranslational modifications of CENP-A and their roles in the context of centromere biology.
An overview of CENP-A structure and posttranslational modifications
The importance of CENP-A nucleosomes for centromere function and inheritance generates an obvious question: what special features does CENP-A possess that confer its centromere-specific function? Among H3 variants, CENP-A is the most diverse variant with just 48% overall similarity to canonical histone H3. It has a highly divergent N–terminus (Sullivan et al. 1994; Tachiwana et al. 2012), and a C–terminus-histone-fold domain (HFD), which is 62% identical to that of H3 (Fig. 1). Within the HFD, the loop1 (L1) along with flanking regions of α1 and α2 helices (aa 75-114) form the CENP-A targeting domain (CATD), which is necessary and sufficient for its centromere localization (Fig. 1) (Black et al. 2004). Sequences in the amino and carboxyl termini are not required for CENP-A deposition and appropriate targeting, but are involved in exerting CENP-A function in building the centromere (Fachinetti et al. 2013; Foltz et al. 2009; Logsdon et al. 2015). Likewise, while the CATD is important for targeting CENP-A to the centromere, the two amino acid “bulge” Arg-80 and Gly-81 in the structure contributes to CCAN recruitment (Sekulic et al. 2010; Tachiwana et al. 2011). Overall, differences in structural rigidity of the CENP-A nucleosome may also distinguish CENP-A from general chromatin and help serve its unique function (Black and Cleveland 2011; Black et al. 2010; Maddox et al. 2012; Stellfox et al. 2013).
Posttranslational modifications of CENP-A and histone H3. A comparison of domains and posttranslational modifications of CENP-A and histone H3, which include methylation, phosphorylation, acetylation, and ubiquitylation. * In original articles, the initiating M cleavage was not taken into consideration and the position of these residues is shifted by + 1. The correct position of these residues is one unit less than depicted the diagram
Posttranslational modifications of canonical histone H3 regulate its function and affect a wide range of cellular processes, including cell differentiation, chromatin condensation, gene expression, and DNA replication and repair (Fig. 1) (Xu et al. 2014). The canonical histone H3 amino terminus is rich in lysine and arginine (especially at the N–terminus), which are frequently modified by acetylation and methylation. On the other hand the lysine content of CENP-A is extremely low compared with H3; therefore, the available sites of modification are fewer. And, although the N–terminus of CENP-A is also rich in arginine, these residues do not appear to be frequently modified, if at all. Thus, the degree to which CENP-A is posttranslationally modified is significantly less than H3. This suggests that in the case of lysine modification, including activating and repressing marks such as H3K4 and H3K27 methylation, CENP-A has evolved to be refractory to these types of control mechanisms (Bannister and Kouzarides 2011; Vakoc et al. 2006). Moreover, where H3 undergoes at least 17 different types of modifications (Xu et al. 2014), only four types of modifications, methylation, acetylation, phosphorylation, and ubiquitylation, have been identified for CENP-A (Fig. 1). Nevertheless, recent discoveries demonstrate the significance of CENP-A PTMs in centromere biology (Table 1). In the subsequent sections of this review, we will describe how these PTMs contribute to the function of CENP-A.
Role of posttranslational modifications of CENP-A in mitosis and CCAN recruitment
Early work from Sullivan and colleagues showed that CENP-A is phosphorylated at Ser-7 (Zeitlin et al. 2001a). Canonical histone H3 is phosphorylated at Ser-10 by the aurora kinases, which is a hallmark of mitosis (Sawicka and Seiser 2012). It was hypothesized that CENP-A Ser-7 could also be subjected to phosphorylation by aurora kinases based on the similarly with histone H3 Ser-10. Despite their similarities, the temporal pattern of H3 and CENP-A phosphorylations appear to be slightly different during G2/M phase of the cell cycle. H3 phosphorylation begins in early G2 and persists throughout mitosis; whereas, CENP-A Ser-7 phosphorylation initiates in prophase, peaks during prometaphase and disappears in anaphase (Fig. 2a) (Zeitlin et al. 2001a). Both mitotic kinases Aurora-A and Aurora-B have been reported to phosphorylate CENP-A at Ser-7. While Aurora-A initiates CENP-A phosphorylation in prophase, Aurora-B is required for the maintenance of phosphorylation (Fig. 2a) (Kunitoku et al. 2003; Zeitlin et al. 2001b). Subsequent mass spectrometry-based studies failed to identify Ser-7 phosphorylation (Bailey et al. 2013) perhaps because Ser-7 phosphorylation is highly labile and was not significantly maintained in mass spectrometry-based experiments. Alternatively, only a small proportion of the CENP-A nucleosomes may be phosphorylated at Ser-7 during mitosis. Mass spectrometry analysis of yeast Cse4 revealed the phosphorylation of Ser-22, Ser-33, Ser-40, and Ser-105 (Boeckmann et al. 2013; Hoffmann et al. 2017) (Fig. 4). The phosphorylations at these four sites is mediated by Ipl1, the yeast homolog of Aurora-B. Such N–terminus phosphorylation of Cse4 by Ipl1/Aurora-B is reminiscent of human CENP-A Ser-7 phosphorylation and it has been shown that phosphorylation of Cse4 facilitates the destabilization of defective kinetochore to ensure the proper chromosome segregation (Boeckmann et al. 2013). Thus these data suggest the functional conservation of Aurora-B phosphorylation of CENP-A.
N-terminal modifications of CENP-A facilitate CCAN recruitment. a NRMT1 trimethylates CENP-A at Gly-1 in interphase; the proportion of which is increased during mitosis. Aurora-A initiates the Ser-7 phosphorylation in prophase, the mark is then maintained by Aurora B. During anaphase, Ser-7 phosphorylation is reduced. b CENP-A trimethylation facilitates the recruitment of CCAN components CENP-T and CENP-I. Ser-7 phosphorylation mediates the recruitment of CENP-C through 14-3-3 proteins. The CENP-A C-terminal tail also directly recruits CENP-C independent of Aurora phosphorylation
Studies performed using a human CENP-A phospho-defective mutant indicate the importance of Ser-7 phosphorylation in mitosis, but present contradictory results. Zeitlin et al. demonstrated that the prevention of Ser-7 phosphorylation does not affect kinetochore function, but leads to increased midbody persistence and size, and thus delays the completion of cytokinesis (Zeitlin et al. 2001b). On the other hand, Kunitoku et al. subsequently showed that non-phosphorylatable CENP-A results in misalignment of chromosomes and impaired kinetochore attachment to microtubules (Kunitoku et al. 2003). Although these studies differ in their conclusions regarding the specific function of Ser-7 phosphorylation, both the studies show Ser-7 phosphorylation to be important for Aurora-B localization at centromeres. In this way, CENP-A Ser-7 phosphorylation and Aurora-B appear to function bi-directionally, where Aurora-B maintains the Ser-7 phosphorylation initiated by Aurora-A and phospho-CENP-A in turn recruits Aurora-B to the inner centromere. Mutational analysis further substantiates the role of CENP-A Ser-7 phosphorylation in chromosome segregation and cytokinesis, as CENP-A phospho-defective mutants fail to rescue the mitotic defects caused by the loss of endogenous CENP-A (Goutte-Gattat et al. 2013).
Amino-terminal trimethylation of CENP-A, which was identified through high-resolution mass spectrometry, has emerged as a regulator of CENP-A function and is required for precise chromosome segregation during mitosis (Bailey et al. 2013; Sathyan et al. 2017). Similar to histone H3, the initiating methionine is constitutively removed from CENP-A and the enzyme NRMT1 trimethylates the alpha amino group of the exposed glycine residue on CENP-A (Fig. 2a). Notably, this modification is not observed on canonical H3. Although a subset of CENP-A is methylated in randomly cycling cells, almost the entire pool of nucleosomal CENP-A is trimethylated by the time cells enter mitosis (Fig. 2a) (Bailey et al. 2013). Preventing CENP-A methylation in conjunction with loss of the p53 tumor suppressor leads to multipolar spindles. The CCAN protein CENP-T is reduced in cells expressing CENP-A that cannot be methylated on its amino terminus and reduced levels of CENP-T in these cells cause spindle multipolarity. Chromosome missegregation errors also occur when CENP-A cannot be methylated, but these errors are independent of p53 status. CENP-A methylation mutants leads to uncontrolled cell proliferation, and when expressed in p53 −/− null cells, results in early onset of tumors in nude mice, suggesting that misregulation of CENP-A methylation may have implications in cancer (Sathyan et al. 2017).
As a master component of the centromere, CENP-A recruits the CCAN, which is required for correct attachment of microtubules to the kinetochores and proper chromosome segregation during mitosis (Carroll et al. 2010; Carroll et al. 2009; Cheeseman and Desai 2008; Fachinetti et al. 2013; Foltz et al. 2006; Guse et al. 2011; Hori et al. 2008; Izuta et al. 2006; Kato et al. 2013; Okada et al. 2006). CENP-A phosphorylation, acetylation, and trimethylation appear to affect the recruitment of CCAN-associated proteins to the centromere. Previously, Ser-7 phosphorylation was found to be involved in loading of CENP-C onto centromeres through phospho-binding protein 14-3-3, which acts as an intermolecular bridge between phospho-CENP-A and CENP-C (Goutte-Gattat et al. 2013) (Fig. 2). More recently, both N– and C–termini of CENP-A have been shown to be independently involved in CENP-C recruitment to the centromere. The carboxyl tail facilitates direct recruitment of CENP-C, whereas the N–terminus recruits CENP-C via an interaction with CENP-B (Fachinetti et al. 2013; Fachinetti et al. 2015; Guse et al. 2011). Thus, the CENP-A Ser-7 phosphorylation may affect the loading of only a subset of CENP-C. Acetylation of CENP-A at Lys-124 has been suggested to impede the accessibility of CENP-C; however, it is not clear if this selectively affects N- or C-terminally associated CENP-C (Bui et al. 2017). Gly-1 trimethylation, on the other hand, regulates the localization of other components of CCAN, namely CENP-T and CENP-I without affecting CENP-C (Fig. 2b) (Sathyan et al. 2017).
CENP-A posttranslational modifications and deposition at centromeres
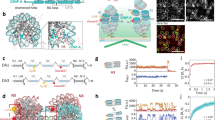
CENP-A nucleosome deposition occurs in G1 phase and is directed by the chaperone HJURP (Bernad et al. 2011; Dunleavy et al. 2009; Foltz et al. 2009; Jansen et al. 2007; Shuaib et al. 2010). HJURP recognizes the CENP-A CATD (CENP-A targeting domain) through its N-terminal SCM3 homology domain and facilitates the loading of CENP-A-containing nucleosomes into DNA (Barnhart et al. 2011; Bassett et al. 2012; Dunleavy et al. 2009; Foltz et al. 2009; Jansen et al. 2007; Sanchez-Pulido et al. 2009; Shuaib et al. 2010). CENP-A deposition is regulated by the phosphorylation of its deposition machinery by two kinases, PLK1 and CDK1/2 (McKinley and Cheeseman 2014; Silva et al. 2012; Stankovic et al. 2017). While PLK1 is a positive regulator of CENP-A deposition, CDK1/2 is inhibitory. Both PLK1 and CDK are the regulators of CENP-A assembly factor Mis18 complex. The Mis18 complex, that includes Mis18BP1, Mis18α and Mis18β, is localized to centromeres during late telophase and early G1 and is required for HJURP recruitment and new CENP-A deposition (Barnhart et al. 2011; Fujita et al. 2007; Hayashi et al. 2004; Nardi et al. 2016; Wang et al. 2014). PLK1 phosphorylates the Mis18 complex and promotes its centromere localization (McKinley and Cheeseman 2014). In contrast, CDK activity regulates multiple components of the deposition machinery to inhibit CENP-A deposition until mitosis is completed. Mis18BP1 is a substrate for CDK1 phosphorylation, and modification of the protein disrupts the association of the protein with the Mis18α/β hexamer (Pan et al. 2017; Silva et al. 2012; Spiller et al. 2017). CDK1 also phosphorylates HJURP to inhibit CENP-A deposition (Muller et al. 2014; Stankovic et al. 2017). Thus the negative regulation by CDK1 ensures CENP-A deposition does not occur in G2, but is restricted to early G1, immediately following satisfaction of the mitotic checkpoint in human cells.
While the CENP-A CATD is sufficient to bind HJURP and directs CENP-A deposition to centromeres, the posttranslational modifications of two evolutionarily conserved residues, Ser-68 and Lys-124, that lie outside the CATD, have been demonstrated to affect CENP-A localization at the centromeres, although the functional aspects of these modifications remain a point of dispute and are discussed below (Hu et al. 2011; Niikura et al. 2015; Wang et al. 2017; Yu et al. 2015). More recently, another modification, Ser-18 phosphorylation that also exists outside CATD has been shown to negatively regulate CENP-A deposition (Takada et al. 2017). Modifications on histone H4 present in the prenucleosomal complex are also involved in CENP-A deposition. Histone H4 Lys-5 and Lys-12 acetylations occur in the pre-deposition complex, are removed after CENP-A nucleosome formation, and appear to be critical for deposition (Fukagawa 2017; Shang et al. 2016).
The crystal structure of the HJURP-CENP-A-H4 complex indicates that HJURP binds the CENP-A-H4 dimer, and this binding may be influenced by Ser-68 of CENP-A (Hu et al. 2011). In vivo, Ser-68 phosphorylation is governed by the opposing actions of the kinase CDK1 and phosphatase PP1α (Yu et al. 2015). Consistent with the relative activities of CDK1/PP1α, CENP-A Ser-68 phosphorylation occurs during G2/M phase and as the cell cycle proceeds through mitosis and G1, Ser-68 is dephosphorylated (Fig. 3).
CENP-A posttranslational modifications during cell cycle. A diagram depicting modifications on CENP-A primarily involved in interaction with HJURP and centromere deposition. K124 undergoes multiple types of mutually exclusive modifications that include ubiquitylation, acetylation, and methylation. Phosphorylation of the CENP-A amino terminus at Ser-18 and Ser-68 are cell cycle regulated and may contribute to restricting the timing of HJURP binding
Canonical histone H3 contains glutamine at the position comparable to CENP-A Ser-68, which suggests that phosphorylation of Ser-68 confers regulatory function unique to CENP-A. Substitution of Ser-68 with the bulky glutamine or phospho-mimetic glutamic acid in CENP-A abolishes HJURP interaction and prevents its localization to centromeres, whereas the S68A mutant shows a robust binding with HJURP (Logsdon et al. 2015; Yu et al. 2015). The CATD in the context of the rest of histone H3 (H3CATD) is sufficient to bind HJURP and direct its deposition to centromeres (Black et al. 2004; Black et al. 2007; Hu et al. 2011). CENP-A equivalent substitutions in this mutant (H3CATD-Q68S) significantly enhances its ability to interact with HJURP (Logsdon et al. 2015; Yu et al. 2015). Nevertheless, the H3 mutant containing Ser-68 without the CATD remains unable to bind to HJURP, clearly reinforcing the idea that CATD is the primary determinant for CENP-A deposition to centromeres (Logsdon et al. 2015; Yu et al. 2015).
Similar to Ser-68, Ser-18 is also not essential for CENP-A deposition, but Ser-18 phosphorylation by Cyclin E/CDK2 complex appears to negatively regulate centromeric localization of CENP-A (Takada et al. 2017). Given that Cyclin E levels remain low during mitosis to G1 transition, a hypophosphorylated state of Ser-18 may facilitate the specific timing of CENP-A deposition at centromeres in G1 phase.
While phosphorylation of Ser-68 generates a potential steric hindrance and impairs CENP-A interaction with HJURP, this interaction is facilitated by Lys-124 ubiquitylation (Fig. 3) (Niikura et al. 2015). The SGT1-HSP90 complex facilitates the recognition of CENP-A by COPS8 and Lys-124 undergoes ubiquitylation by the CUL4A-RBX1-COPS8 complex (Niikura et al. 2016; Niikura et al. 2015; Niikura et al. 2017b) (Fig. 3). Fusion of ubiquitin to the C–terminus of CENP-A–K124R mutant restores the CENP-A interaction with HJURP and localization at centromere (Niikura et al. 2015).
In contrast to the dynamic nature of Ser-68 phosphorylation, so far, no deubiquitylating enzyme has been identified for Lys-124. However, Lys-124 has been proposed to possess different modifications at different times during cell cycle. Ubiquitylation, which takes place at mitotic exit and entry into G1 phase switches to acetylation at G1/S phase, which is further exchanged for monomethylation during S phase (Bui et al. 2012; Bui et al. 2017; Niikura et al. 2017a; Niikura et al. 2015). The interplay among E3 ligase, acetyl, and methyl transferases (and their opposing counterparts, if there are any for CENP-A) that determines the modification status of Lys-124 has not been mechanistically defined; however, these studies intimate that acetylation and monomethylation may check Lys-124 ubiquitylation until M/G1 phase when CENP-A deposition occurs (Fig. 3).
Despite the biochemical evidence described above, it is unclear whether modifications of Ser-68 and Lys-124 are essential for CENP-A function. In order to address this question, Fachinetti and colleagues attempted to rescue CENP-A knockout RPE (retinal pigment epithelium) cells by expressing CENP-A modification mutants by viral transduction. In these experiments, individual Ser-68 phospho-mimetic mutants, or unmodifiable Lys-124 mutant, rescued cell viability resulting from loss of endogenous CENP-A (Fachinetti et al. 2017). Potential caveats of these experiments, including the degree to which mutant CENP-A overexpression may compensate for loss of CENP-A modifications have been proposed to support the importance of the Lys–124 and Ser-68 modifications at the cellular level (Niikura et al. 2017a; Wang et al. 2017). Indeed, Fachinetti and colleagues observed that LacI-fused CENP-A–S68Q mutant reduced HJURP recruitment at LacO array, which is in agreement with a negative role for Ser-68 phosphorylation in HJURP binding as reported by Guohong and colleagues. In contrast, these same experiments showed that the CENP-A–K124R mutant was as efficient as wild-type CENP-A in recruiting HJRUP to the LacO site (Fachinetti et al. 2017), casting additional doubt on whether K124 affects CENP-A deposition through the proposed model. The contradiction between the apparent biochemical impact of these mutations and the fact that they are dispensable for cell viability may suggest that while Ser-68 and Lys-124 are not absolutely required to regulate CENP-A recognition by HJURP, these modifications are likely to play a subtle modulatory role in CENP-A deposition than originally described. Further studies are warranted to resolve the remaining discrepancies.
Diversity of CENP-A modifications across species
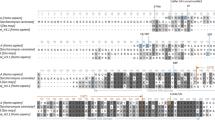
CENP-A shows a much higher degree of evolutionary divergence than its histone H3.1 counterpart, and these changes may be driven by co-evolution with the underlying centromeric DNA (Malik 2009; Rosin and Mellone 2017). The CENP-A N–terminus is the most variable region of the protein (Fig. 4). While the N-terminal sequence of CENP-A shows some conservation within the vertebrate lineage, more distantly related species are highly divergent (Fig. 4). Given the high degree of divergence, it is not surprising that the Ser-7 and Ser-16/18 modifications are only partially conserved in vertebrates, and mostly lost in other eukaryotes including flies, budding yeast, and nematodes. The histone-fold domains of CENP-A homologs are well conserved, as are the Ser-68 and Lys-124 modification sites within these domains. Caenorhabditis elegans, which does not contain an HJURP homolog, also does not show conservation at these sites. Interestingly, amino acid changes observed in zebrafish, flies, and fission yeast retain the charge at amino acid 124, but contain an arginine that would not undergo ubiquitylation or acetylation, suggesting that amino acid charge may be critical at this site (Fig. 4).
Conservation of human CENP-A posttranslational modifications across species. Schematic representation of CENP-A proteins in different organisms. Percent identity with respect to human CENP-A was computed through pairwise alignment using EMBOSS Needle program. CENP-A protein sequences in these organisms are also aligned through Clustal Omega and zoomed to focus on posttranslationally modified sites
Consistent with the divergence of the CENP-A N–terminus, several modifications have been identified that are unique to individual species. The Drosophila counterpart of human CENP-A (a.k.a. dCENP-ACID) has been shown to be acetylated at Lys-105, located within the N–terminus, exclusively in cytosolic prenucleosomal fraction (Boltengagen et al. 2016). In addition, dCENP-ACID also undergoes phosphorylations at Ser-20, Ser-75, and Ser-77 (Boltengagen et al. 2016). The phosphorylation of dCENP-ACID Ser-75/77 is suggestive of human CENP-A Ser-16/18 phosphorylation; however, the functional consequences of these phosphorylation events appear to be different in Drosophila and human. While prenucleosomal cytosolic dCENP-ACID exhibits unphosphorylated, mono- or di-phosphorylated (Ser-20 and Ser-75) forms, the nucleoplasmic dCENP-ACID is enriched in an additional phosphorylation mark at Ser-77 (Boltengagen et al. 2016). Both acetylation and phosphorylation of Drosophila dCENP-ACID occur on the N–terminus, and based on their differential patterns in prenucleosomal and nucleosomal fractions, have been suggested to determine the localization of dCENP-ACID (Boltengagen et al. 2016).
dCENP-ACID has also been reported to be ubiquitylated, although the site of ubiquitylation is undetermined, and the effect of ubiquitylation is unique from human CENP-A. Ubiquitylation of dCENP-ACID is distinguished by the fact that the E3 ligase CUL3/RDX1 complex directly interacts with the functional homolog of HJURP in flies, called CAL1, which serves as an adaptor for the enzymatic reaction (Bade et al. 2014; Chen et al. 2014; Erhardt et al. 2008). CAL1 itself does not undergo ubiquitylation; nonetheless, both CAL1 and dCENP-ACID are stabilized by the CUL3/RDX complex, and loss of RDX leads to fragmented chromosomes and perturbs centromere maintenance.
The yeast homolog of human CENP-A, Cse4 is also posttranslationally modified. In addition to the functionally conserved Ipl1/Aurora-B sites found in the amino terminus of Cse4 (discussed earlier in the manuscript), the protein is methylated at Arg-37 and acetylated at Lys-49 (Boeckmann et al. 2013; Samel et al. 2012) (Fig. 4). While the function of Lys-49 acetylation is currently unknown, Arg-37 methylation is required for the recruitment of kinetochore components at centromeres, and its prevention leads to impaired chromosome segregation and growth defects in Saccharomyces cerevisiae (Samel et al. 2012).
CENP-A retention and its consistent levels at centromeres are crucial for genomic integrity. On the other hand, CENP-A overexpression is associated with its mislocalization outside the centromere to euchromatic regions in tumorigenesis (Allshire and Karpen 2008; Amato et al. 2009; Athwal et al. 2015; Filipescu et al. 2017; Hu et al. 2010; Lacoste et al. 2014; Li et al. 2011; Shrestha et al. 2017; Tomonaga et al. 2003; Wu et al. 2012; Zhang et al. 2016). Therefore, in normal conditions, there must exist a mechanism to eliminate CENP-A from non-centromeric loci. Ubiquitylation has been reported to prevent the ectopic localization of Cse4 by triggering its degradation, and the E3 ubiquitin ligases Psh1, Rcy1, Slx5, and Ubr1 have been shown to ubiquitylate Cse4 (Fig. 5) (Au et al. 2013; Cheng et al. 2017; Cheng et al. 2016; Collins et al. 2004; Hewawasam et al. 2010; Ohkuni et al. 2016; Ranjitkar et al. 2010). While Psh1 ubiquitylates Lys-4, Lys-131, Lys-155, Lys-163 and Lys-172 (Hewawasam et al. 2010), Slx5-mediated proteolysis of Cse4 is directed by sumoylation at Lys-65 (Ohkuni et al. 2018) (Fig. 4). The SUMO E3 ligases Siz1 and Siz2 target Cse4 for sumoylation, which further undergoes ubiquitylation by the ubiquitin ligase Slx5 (Fig. 5) (Ohkuni et al. 2016).
These E3 ubiquitin ligases function independently (Cheng et al. 2017). Despite Psh1 localization to centromeres, centromeric Cse4 is precluded from ubiquitylation through its interaction with the Scm3 chaperone (Hewawasam et al. 2010). Psh1 activity towards Cse4 is facilitated by casein kinase 2-mediated phosphorylation and its association with the chromatin modifying FACT (facilitates chromatin transcription/transactions) complex (Deyter and Biggins 2014; Hewawasam et al. 2014), providing additional levels of regulation.
Although human CENP-A is subjected to proteasomal degradation, the identity of the E3 ubiquitin ligases that act on it remains unknown (Earnshaw 2015; Lomonte et al. 2001). While Psh1 is not evolutionarily conserved, Rcy1, Slx5, and Ubr1 have human homologs, and loss of the human STUBL (SUMO-targeted ubiquitin ligase) ortholog RNF4 leads to chromosome segregation defects, suggesting the possibility that human CENP-A is regulated similar to Cse4 (van de Pasch et al. 2013). Moreover, the finding that Psh1 targets Cse4 through the functionally conserved CATD domain (Ranjitkar et al. 2010) supports the possibility for the existence of proteolytic regulation of human CENP-A akin to that which acts on Cse4. Taken together, the yeast studies have provided a tantalizing idea that changes in CENP-A stability may contribute to CENP-A overexpression and mislocalization in cancers, and it will be interesting to investigate whether this is the result of deregulation in the ubiquitin machinery targeting CENP-A.
Perspective
Intrinsic or induced differences in the structure of the CENP-A nucleosome also contribute to distinguishing CENP-A chromatin from general chromatin (Falk et al. 2015; Sekulic et al. 2010). CENP-A posttranslational modifications that include Lys-124 acetylation and Ser-16/18 phosphorylation have the potential to influence the intrinsic properties of the CENP-A nucleosome or the CENP-A-containing chromatin (Bailey et al. 2013; Bui et al. 2017). CENP-A has been proposed to form a salt-bridged secondary structure through intra- and intermolecular association between phosphorylated Ser-16 and –18 and nearby arginine residues, both of which are highly conserved (Bailey et al. 2013). In vitro, Ser-16/18 salt bridging alters the organization of CENP-A-containing nucleosome arrays. In vivo, mutations that block CENP-A phosphorylation at these sites lead to chromosome missegregation, suggesting that organization of centromeric chromatin through the CENP-A amino-terminal tail may be important for centromere function. Lys-124 is located close to the pseudo-dyad DNA axis of the centromeric nucleosome and acetylation of this residue occurs during G1/S phase of the cell cycle (Bui et al. 2017). Based on the similarity with canonical H3 K122 acetylation and molecular dynamic simulations, it has been proposed that acetylation of Lys-124 tightens the CENP-A nucleosome, potentially contributing to its unique function. However, clear experimental data is not yet available to substantiate the structural changes induced by Lys-124 modification.
CENP-A PTMs regulate CENP-A deposition at centromeres, protein stability, and CCAN recruitment, and thus, these PTMs play an important role in faithful segregation of chromosomes during cell division. Growing evidence suggests the potential alteration of CENP-A PTMs associated with cancer progression. The observation that a CENP-A methylation defective mutant together with loss of p53, promotes tumor development in nude mice and the correlation of Ser-18 hyperphophorylation with cyclin E-driven tumors, indicate a potential role of CENP-A modifications in cancer (Sathyan et al. 2017; Takada et al. 2017). Large-scale proteomic studies have revealed several additional modifications; the biological relevance of which is yet to be discovered (please see http://www.phosphositeplus.com). Clearly, our current knowledge of CENP-A PTMs is just the tip of the iceberg and offers enormous possibility of future research.
References
Allshire RC, Karpen GH (2008) Epigenetic regulation of centromeric chromatin: old dogs, new tricks? Nat Rev Genet 9:923–937
Amato A, Schillaci T, Lentini L, Di Leonardo A (2009) CENPA overexpression promotes genome instability in pRb-depleted human cells. Mol Cancer 8:119
Athwal RK, Walkiewicz MP, Baek S, Fu S, Bui M, Camps J, Ried T, Sung MH, Dalal Y (2015) CENP-A nucleosomes localize to transcription factor hotspots and subtelomeric sites in human cancer cells. Epigenetics Chromatin 8:2
Au WC, Dawson AR, Rawson DW, Taylor SB, Baker RE, Basrai MA (2013) A novel role of the N terminus of budding yeast histone H3 variant Cse4 in ubiquitin-mediated proteolysis. Genetics 194:513–518
Bade D, Pauleau AL, Wendler A, Erhardt S (2014) The E3 ligase CUL3/RDX controls centromere maintenance by ubiquitylating and stabilizing CENP-A in a CAL1-dependent manner. Dev Cell 28:508–519
Bailey AO, Panchenko T, Sathyan KM, Petkowski JJ, Pai PJ, Bai DL, Russell DH, Macara IG, Shabanowitz J, Hunt DF et al (2013) Posttranslational modification of CENP-A influences the conformation of centromeric chromatin. Proc Natl Acad Sci U S A 110:11,827–11,832
Bannister AJ, Kouzarides T (2011) Regulation of chromatin by histone modifications. Cell Res 21:381–395
Barnhart MC, Kuich PH, Stellfox ME, Ward JA, Bassett EA, Black BE, Foltz DR (2011) HJURP is a CENP-A chromatin assembly factor sufficient to form a functional de novo kinetochore. J Cell Biol 194:229–243
Bassett EA, DeNizio J, Barnhart-Dailey MC, Panchenko T, Sekulic N, Rogers DJ, Foltz DR, Black BE (2012) HJURP uses distinct CENP-A surfaces to recognize and to stabilize CENP-A/histone H4 for centromere assembly. Dev Cell 22:749–762
Bergmann JH, Rodriguez MG, Martins NM, Kimura H, Kelly DA, Masumoto H, Larionov V, Jansen LE, Earnshaw WC (2011) Epigenetic engineering shows H3K4me2 is required for HJURP targeting and CENP-A assembly on a synthetic human kinetochore. EMBO J 30:328–340
Bernad R, Sanchez P, Rivera T, Rodriguez-Corsino M, Boyarchuk E, Vassias I, Ray-Gallet D, Arnaoutov A, Dasso M, Almouzni G et al (2011) Xenopus HJURP and condensin II are required for CENP-A assembly. J Cell Biol 192:569–582
Black BE, Cleveland DW (2011) Epigenetic centromere propagation and the nature of CENP-a nucleosomes. Cell 144:471–479
Black BE, Foltz DR, Chakravarthy S, Luger K, Woods VL Jr, Cleveland DW (2004) Structural determinants for generating centromeric chromatin. Nature 430:578–582
Black BE, Jansen LE, Maddox PS, Foltz DR, Desai AB, Shah JV, Cleveland DW (2007) Centromere identity maintained by nucleosomes assembled with histone H3 containing the CENP-A targeting domain. Mol Cell 25:309–322
Black BE, Jansen LE, Foltz DR, Cleveland DW (2010) Centromere identity, function, and epigenetic propagation across cell divisions. Cold Spring Harb Symp Quant Biol 75:403–418
Blower MD, Sullivan BA, Karpen GH (2002) Conserved organization of centromeric chromatin in flies and humans. Dev Cell 2:319–330
Boeckmann L, Takahashi Y, Au WC, Mishra PK, Choy JS, Dawson AR, Szeto MY, Waybright TJ, Heger C, McAndrew C et al (2013) Phosphorylation of centromeric histone H3 variant regulates chromosome segregation in Saccharomyces cerevisiae. Mol Biol Cell 24:2034–2044
Boltengagen M, Huang A, Boltengagen A, Trixl L, Lindner H, Kremser L, Offterdinger M, Lusser A (2016) A novel role for the histone acetyltransferase Hat1 in the CENP-A/CID assembly pathway in Drosophila melanogaster. Nucleic Acids Res 44:2145–2159
Bui M, Dimitriadis EK, Hoischen C, An E, Quenet D, Giebe S, Nita-Lazar A, Diekmann S, Dalal Y (2012) Cell-cycle-dependent structural transitions in the human CENP-A nucleosome in vivo. Cell 150:317–326
Bui M, Pitman M, Nuccio A, Roque S, Donlin-Asp PG, Nita-Lazar A, Papoian GA, Dalal Y (2017) Internal modifications in the CENP-A nucleosome modulate centromeric dynamics. Epigenetics Chromatin 10:17
Carroll CW, Silva MC, Godek KM, Jansen LE, Straight AF (2009) Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nat Cell Biol 11:896–902
Carroll CW, Milks KJ, Straight AF (2010) Dual recognition of CENP-A nucleosomes is required for centromere assembly. J Cell Biol 189:1143–1155
Cheeseman IM, Desai A (2008) Molecular architecture of the kinetochore-microtubule interface. Nat Rev. Mol Cell Biol 9:33–46
Chen CC, Dechassa ML, Bettini E, Ledoux MB, Belisario C, Heun P, Luger K, Mellone BG (2014) CAL1 is the Drosophila CENP-A assembly factor. J Cell Biol 204:313–329
Cheng H, Bao X, Rao H (2016) The F-box protein Rcy1 is involved in the degradation of histone H3 variant Cse4 and genome maintenance. J Biol Chem 291:10,372–10,377
Cheng H, Bao X, Gan X, Luo S, Rao H (2017) Multiple E3s promote the degradation of histone H3 variant Cse4. Sci Rep 7:8565
Choo KH (2000) Centromerization. Trends Cell Biol 10:182–188
Cleveland DW, Mao Y, Sullivan KF (2003) Centromeres and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell 112:407–421
Collins KA, Furuyama S, Biggins S (2004) Proteolysis contributes to the exclusive centromere localization of the yeast Cse4/CENP-A histone H3 variant. Curr Biol 14:1968–1972
Deyter GM, Biggins S (2014) The FACT complex interacts with the E3 ubiquitin ligase Psh1 to prevent ectopic localization of CENP-A. Genes Dev 28:1815–1826
Drinnenberg IA, de Young D, Henikoff S, Malik HS (2014) Recurrent loss of CenH3 is associated with independent transitions to holocentricity in insects. Elife 3.
Dunleavy EM, Roche D, Tagami H, Lacoste N, Ray-Gallet D, Nakamura Y, Daigo Y, Nakatani Y, Almouzni-Pettinotti G (2009) HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell 137:485–497
Earnshaw WC (2015) Discovering centromere proteins: from cold white hands to the A, B, C of CENPs. Nat Rev. Mol Cell Biol 16:443–449
Erhardt S, Mellone BG, Betts CM, Zhang W, Karpen GH, Straight AF (2008) Genome-wide analysis reveals a cell cycle-dependent mechanism controlling centromere propagation. J Cell Biol 183:805–818
Fachinetti D, Folco HD, Nechemia-Arbely Y, Valente LP, Nguyen K, Wong AJ, Zhu Q, Holland AJ, Desai A, Jansen LE et al (2013) A two-step mechanism for epigenetic specification of centromere identity and function. Nat Cell Biol 15:1056–1066
Fachinetti D, Han JS, McMahon MA, Ly P, Abdullah A, Wong AJ, Cleveland DW (2015) DNA sequence-specific binding of CENP-B enhances the fidelity of human centromere function. Dev Cell 33:314–327
Fachinetti D, Logsdon GA, Abdullah A, Selzer EB, Cleveland DW, Black BE (2017) CENP-A Modifications on Ser68 and Lys124 are dispensable for establishment, maintenance, and long-term function of human centromeres. Dev Cell 40:104–113
Falk SJ, Guo LY, Sekulic N, Smoak EM, Mani T, Logsdon GA, Gupta K, Jansen LE, Van Duyne GD, Vinogradov SA et al (2015) Chromosomes. CENP-C reshapes and stabilizes CENP-A nucleosomes at the centromere. Science 348:699–703
Filipescu D, Naughtin M, Podsypanina K, Lejour V, Wilson L, Gurard-Levin ZA, Orsi GA, Simeonova I, Toufektchan E, Attardi LD et al (2017) Essential role for centromeric factors following p53 loss and oncogenic transformation. Genes Dev 31:463–480
Foltz DR, Jansen LE, Black BE, Bailey AO, Yates JR 3rd, Cleveland DW (2006) The human CENP-A centromeric nucleosome-associated complex. Nat Cell Biol 8:458–469
Foltz DR, Jansen LE, Bailey AO, Yates JR 3rd, Bassett EA, Wood S, Black BE, Cleveland DW (2009) Centromere-specific assembly of CENP-a nucleosomes is mediated by HJURP. Cell 137:472–484
Fujita Y, Hayashi T, Kiyomitsu T, Toyoda Y, Kokubu A, Obuse C, Yanagida M (2007) Priming of centromere for CENP-A recruitment by human hMis18alpha, hMis18beta, and M18BP1. Dev Cell 12:17–30
Fukagawa T (2017) Critical histone post-translational modifications for centromere function and propagation. Cell Cycle 16:1259–1265
Fukagawa T, Earnshaw WC (2014) The centromere: chromatin foundation for the kinetochore machinery. Dev Cell 30:496–508
Garcia Del Arco A, Erhardt S (2017) Post-translational modifications of centromeric chromatin. Prog Mol Subcell Biol 56:213–231
Giam M, Rancati G (2015) Aneuploidy and chromosomal instability in cancer: a jackpot to chaos. Cell Div 10:3
Goutte-Gattat D, Shuaib M, Ouararhni K, Gautier T, Skoufias DA, Hamiche A, Dimitrov S (2013) Phosphorylation of the CENP-A amino-terminus in mitotic centromeric chromatin is required for kinetochore function. Proc Natl Acad Sci U S A 110:8579–8584
Guse A, Carroll CW, Moree B, Fuller CJ, Straight AF (2011) In vitro centromere and kinetochore assembly on defined chromatin templates. Nature 477:354–358
Hayashi T, Fujita Y, Iwasaki O, Adachi Y, Takahashi K, Yanagida M (2004) Mis16 and Mis18 are required for CENP-A loading and histone deacetylation at centromeres. Cell 118:715–729
Henikoff S, Dalal Y (2005) Centromeric chromatin: what makes it unique? Curr Opin Genet Dev 15:177–184
Henikoff S, Furuyama T (2010) Epigenetic inheritance of centromeres. Cold Spring Harb Symp Quant Biol 75:51–60
Hewawasam G, Shivaraju M, Mattingly M, Venkatesh S, Martin-Brown S, Florens L, Workman JL, Gerton JL (2010) Psh1 is an E3 ubiquitin ligase that targets the centromeric histone variant Cse4. Mol Cell 40:444–454
Hewawasam GS, Mattingly M, Venkatesh S, Zhang Y, Florens L, Workman JL, Gerton JL (2014) Phosphorylation by casein kinase 2 facilitates Psh1 protein-assisted degradation of Cse4 protein. J Biol Chem 289:29,297–29,309
Hoffmann G, Samel-Pommerencke A, Weber J, Cuomo A, Bonaldi T, Ehrenhofer-Murray AE (2017) A role for CENP-A/Cse4 phosphorylation on serine 33 in deposition at the centromere. FEMS Yeast Res.
Holland AJ, Cleveland DW (2009) Boveri revisited: chromosomal instability, aneuploidy and tumorigenesis. Nat Rev. Mol Cell Biol 10:478–487
Hori T, Amano M, Suzuki A, Backer CB, Welburn JP, Dong Y, McEwen BF, Shang WH, Suzuki E, Okawa K et al (2008) CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. Cell 135:1039–1052
Hu Z, Huang G, Sadanandam A, Gu S, Lenburg ME, Pai M, Bayani N, Blakely EA, Gray JW, Mao JH (2010) The expression level of HJURP has an independent prognostic impact and predicts the sensitivity to radiotherapy in breast cancer. Breast Cancer Res 12:R18
Hu H, Liu Y, Wang M, Fang J, Huang H, Yang N, Li Y, Wang J, Yao X, Shi Y et al (2011) Structure of a CENP-A-histone H4 heterodimer in complex with chaperone HJURP. Genes Dev 25:901–906
Izuta H, Ikeno M, Suzuki N, Tomonaga T, Nozaki N, Obuse C, Kisu Y, Goshima N, Nomura F, Nomura N et al (2006) Comprehensive analysis of the ICEN (interphase centromere complex) components enriched in the CENP-A chromatin of human cells. Genes Cells 11:673–684
Jansen LE, Black BE, Foltz DR, Cleveland DW (2007) Propagation of centromeric chromatin requires exit from mitosis. J Cell Biol 176:795–805
Kato H, Jiang J, Zhou BR, Rozendaal M, Feng H, Ghirlando R, Xiao TS, Straight AF, Bai Y (2013) A conserved mechanism for centromeric nucleosome recognition by centromere protein CENP-C. Science 340:1110–1113
Kunitoku N, Sasayama T, Marumoto T, Zhang D, Honda S, Kobayashi O, Hatakeyama K, Ushio Y, Saya H, Hirota T (2003) CENP-A phosphorylation by Aurora-A in prophase is required for enrichment of Aurora-B at inner centromeres and for kinetochore function. Dev Cell 5:853–864
Lacoste N, Woolfe A, Tachiwana H, Garea AV, Barth T, Cantaloube S, Kurumizaka H, Imhof A, Almouzni G (2014) Mislocalization of the centromeric histone variant CenH3/CENP-A in human cells depends on the chaperone DAXX. Mol Cell 53:631–644
Li Y, Zhu Z, Zhang S, Yu D, Yu H, Liu L, Cao X, Wang L, Gao H, Zhu M (2011) ShRNA-targeted centromere protein A inhibits hepatocellular carcinoma growth. PLoS One 6:e17794
Lo AW, Craig JM, Saffery R, Kalitsis P, Irvine DV, Earle E, Magliano DJ, Choo KH (2001) A 330 kb CENP-A binding domain and altered replication timing at a human neocentromere. EMBO J 20:2087–2096
Logsdon GA, Barrey EJ, Bassett EA, DeNizio JE, Guo LY, Panchenko T, Dawicki-McKenna JM, Heun P, Black BE (2015) Both tails and the centromere targeting domain of CENP-A are required for centromere establishment. J Cell Biol 208:521–531
Lomonte P, Sullivan KF, Everett RD (2001) Degradation of nucleosome-associated centromeric histone H3-like protein CENP-A induced by herpes simplex virus type 1 protein ICP0. J Biol Chem 276:5829–5835
Maddox PS, Corbett KD, Desai A (2012) Structure, assembly and reading of centromeric chromatin. Curr Opin Genet Dev 22:139–147
Malik HS (2009) The centromere-drive hypothesis: a simple basis for centromere complexity. Prog Mol Subcell Biol 48:33–52
McKinley KL, Cheeseman IM (2014) Polo-like kinase 1 licenses CENP-A deposition at centromeres. Cell 158:397–411
Melters DP, Paliulis LV, Korf IF, Chan SW (2012) Holocentric chromosomes: convergent evolution, meiotic adaptations, and genomic analysis. Chromosome Res 20:579–593
Mendiburo MJ, Padeken J, Fulop S, Schepers A, Heun P (2011) Drosophila CENH3 is sufficient for centromere formation. Science 334:686–690
Muller S, Montes de Oca R, Lacoste N, Dingli F, Loew D, Almouzni G (2014) Phosphorylation and DNA binding of HJURP determine its centromeric recruitment and function in CenH3(CENP-A) loading. Cell Rep 8:190–203
Nardi IK, Zasadzinska E, Stellfox ME, Knippler CM, Foltz DR (2016) Licensing of centromeric chromatin assembly through the Mis18alpha-Mis18beta heterotetramer. Mol Cell 61:774–787
Neumann P, Pavlikova Z, Koblizkova A, Fukova I, Jedlickova V, Novak P, Macas J (2015) Centromeres off the hook: massive changes in centromere size and structure following duplication of CenH3 gene in Fabeae species. Mol Biol Evol 32:1862–1879
Niikura Y, Kitagawa R, Ogi H, Abdulle R, Pagala V, Kitagawa K (2015) CENP-A K124 ubiquitylation is required for CENP-A deposition at the centromere. Dev Cell 32:589–603
Niikura Y, Kitagawa R, Kitagawa K (2016) CENP-A Ubiquitylation is inherited through dimerization between cell divisions. Cell Rep 15:61–76
Niikura Y, Kitagawa R, Kitagawa K (2017a) CENP-A Ubiquitylation is required for CENP-A deposition at the centromere. Dev Cell 40:7–8
Niikura Y, Kitagawa R, Ogi H, Kitagawa K (2017b) SGT1-HSP90 complex is required for CENP-A deposition at centromeres. Cell Cycle, 1–12
Ohkuni K, Takahashi Y, Fulp A, Lawrimore J, Au WC, Pasupala N, Levy-Myers R, Warren J, Strunnikov A, Baker RE, et al. (2016) SUMO-Targeted Ubiquitin Ligase (STUbL) Slx5 regulates proteolysis of centromeric histone H3 variant Cse4 and prevents its mislocalization to euchromatin. Mol Biol Cell
Ohkuni K, Levy-Myers R, Warren J, Au WC, Takahashi Y, Baker RE, Basrai MA (2018) N-terminal sumoylation of centromeric histone H3 variant Cse4 regulates its proteolysis to prevent mislocalization to non-centromeric chromatin. G3 (Bethesda)
Okada M, Cheeseman IM, Hori T, Okawa K, McLeod IX, Yates JR 3rd, Desai A, Fukagawa T (2006) The CENP-H-I complex is required for the efficient incorporation of newly synthesized CENP-A into centromeres. Nat Cell Biol 8:446–457
Pan D, Klare K, Petrovic A, Take A, Walstein K, Singh P, Rondelet A, Bird AW, Musacchio A (2017) CDK-regulated dimerization of M18BP1 on a Mis18 hexamer is necessary for CENP-A loading. Elife 6
Perpelescu M, Fukagawa T (2011) The ABCs of CENPs. Chromosoma 120:425–446
Pfau SJ, Amon A (2012) Chromosomal instability and aneuploidy in cancer: from yeast to man. EMBO Rep 13:515–527
Ranjitkar P, Press MO, Yi X, Baker R, MacCoss MJ, Biggins S (2010) An E3 ubiquitin ligase prevents ectopic localization of the centromeric histone H3 variant via the centromere targeting domain. Mol Cell 40:455–464
Rosin LF, Mellone BG (2017) Centromeres drive a hard bargain. Trends Genet 33:101–117
Saffery R, Irvine DV, Griffiths B, Kalitsis P, Wordeman L, Choo KH (2000) Human centromeres and neocentromeres show identical distribution patterns of >20 functionally important kinetochore-associated proteins. Hum Mol Genet 9:175–185
Samel A, Cuomo A, Bonaldi T, Ehrenhofer-Murray AE (2012) Methylation of CenH3 arginine 37 regulates kinetochore integrity and chromosome segregation. Proc Natl Acad Sci U S A 109:9029–9034
Sanchez-Pulido L, Pidoux AL, Ponting CP, Allshire RC (2009) Common ancestry of the CENP-A chaperones Scm3 and HJURP. Cell 137:1173–1174
Sathyan KM, Fachinetti D, Foltz DR (2017) Alpha-amino trimethylation of CENP-A by NRMT is required for full recruitment of the centromere. Nat Commun 8:14,678
Sawicka A, Seiser C (2012) Histone H3 phosphorylation—a versatile chromatin modification for different occasions. Biochimie 94:2193–2201
Sekulic N, Bassett EA, Rogers DJ, Black BE (2010) The structure of (CENP-A-H4)(2) reveals physical features that mark centromeres. Nature 467:347–351
Sen S (2000) Aneuploidy and cancer. Curr Opin Oncol 12:82–88
Shang WH, Hori T, Westhorpe FG, Godek KM, Toyoda A, Misu S, Monma N, Ikeo K, Carroll CW, Takami Y et al (2016) Acetylation of histone H4 lysine 5 and 12 is required for CENP-A deposition into centromeres. Nat Commun 7:13,465
Shrestha RL, Ahn GS, Staples MI, Sathyan KM, Karpova TS, Foltz DR, Basrai MA (2017) Mislocalization of centromeric histone H3 variant CENP-A contributes to chromosomal instability (CIN) in human cells. Oncotarget 8:46,781–46,800
Shuaib M, Ouararhni K, Dimitrov S, Hamiche A (2010) HJURP binds CENP-A via a highly conserved N-terminal domain and mediates its deposition at centromeres. Proc Natl Acad Sci U S A 107:1349–1354
Silva MC, Bodor DL, Stellfox ME, Martins NM, Hochegger H, Foltz DR, Jansen LE (2012) Cdk activity couples epigenetic centromere inheritance to cell cycle progression. Dev Cell 22:52–63
Spiller F, Medina-Pritchard B, Abad MA, Wear MA, Molina O, Earnshaw WC, Jeyaprakash AA (2017) Molecular basis for Cdk1-regulated timing of Mis18 complex assembly and CENP-A deposition. EMBO reports 18:894–905
Stankovic A, Guo LY, Mata JF, Bodor DL, Cao XJ, Bailey AO, Shabanowitz J, Hunt DF, Garcia BA, Black BE et al (2017) A dual inhibitory mechanism sufficient to maintain cell-cycle-restricted CENP-A assembly. Molecular Cell 65:231–246
Stellfox ME, Bailey AO, Foltz DR (2013) Putting CENP-A in its place. Cell Mol Life Sci 70:387–406
Sullivan KF, Hechenberger M, Masri K (1994) Human CENP-A contains a histone H3 related histone fold domain that is required for targeting to the centromere. J Cell Biol 127:581–592
Tachiwana H, Kagawa W, Shiga T, Osakabe A, Miya Y, Saito K, Hayashi-Takanaka Y, Oda T, Sato M, Park SY et al (2011) Crystal structure of the human centromeric nucleosome containing CENP-A. Nature 476:232–235
Tachiwana H, Kagawa W, Kurumizaka H (2012) Comparison between the CENP-A and histone H3 structures in nucleosomes. Nucleus 3:6–11
Takada M, Zhang W, Suzuki A, Kuroda TS, Yu Z, Inuzuka H, Gao D, Wan L, Zhuang M, Hu L, et al. (2017) FBW7 loss promotes chromosomal instability and tumorigenesis via cyclin E1/CDK2-mediated phosphorylation of CENP-A. Cancer Res
Tomonaga T, Matsushita K, Yamaguchi S, Oohashi T, Shimada H, Ochiai T, Yoda K, Nomura F (2003) Overexpression and mistargeting of centromere protein-A in human primary colorectal cancer. Cancer Res 63:3511–3516
Vakoc CR, Sachdeva MM, Wang H, Blobel GA (2006) Profile of histone lysine methylation across transcribed mammalian chromatin. Mol Cell Biol 26:9185–9195
van de Pasch LA, Miles AJ, Nijenhuis W, Brabers NA, van Leenen D, Lijnzaad P, Brown MK, Ouellet J, Barral Y, Kops GJ et al (2013) Centromere binding and a conserved role in chromosome stability for SUMO-dependent ubiquitin ligases. PLoS One 8:e65628
Wang J, Liu X, Dou Z, Chen L, Jiang H, Fu C, Fu G, Liu D, Zhang J, Zhu T et al (2014) Mitotic regulator Mis18beta interacts with and specifies the centromeric assembly of molecular chaperone holliday junction recognition protein (HJURP). J Biol Chem 289:8326–8336
Wang K, Yu Z, Liu Y, Li G (2017) Ser68 phosphorylation ensures accurate cell-cycle-dependent CENP-A deposition at centromeres. Dev Cell 40:5–6
Warburton PE (2004) Chromosomal dynamics of human neocentromere formation. Chromosome Res 12:617–626
Westhorpe FG, Straight AF (2014) The centromere: epigenetic control of chromosome segregation during mitosis. Cold Spring Harb Perspect Biol 7:a015818
Wu Q, Qian YM, Zhao XL, Wang SM, Feng XJ, Chen XF, Zhang SH (2012) Expression and prognostic significance of centromere protein A in human lung adenocarcinoma. Lung Cancer 77:407–414
Xu YM, Du JY, Lau AT (2014) Posttranslational modifications of human histone H3: an update. Proteomics 14:2047–2060
Yu Z, Zhou X, Wang W, Deng W, Fang J, Hu H, Wang Z, Li S, Cui L, Shen J et al (2015) Dynamic phosphorylation of CENP-A at Ser68 orchestrates its cell-cycle-dependent deposition at centromeres. Dev Cell 32:68–81
Zeitlin SG, Barber CM, Allis CD, Sullivan KF (2001a) Differential regulation of CENP-A and histone H3 phosphorylation in G2/M. J Cell Sci 114:653–661
Zeitlin SG, Shelby RD, Sullivan KF (2001b) CENP-A is phosphorylated by Aurora B kinase and plays an unexpected role in completion of cytokinesis. J Cell Biol 155:1147–1157
Zhang W, Mao JH, Zhu W, Jain AK, Liu K, Brown JB, Karpen GH (2016) Centromere and kinetochore gene misexpression predicts cancer patient survival and response to radiotherapy and chemotherapy. Nat Commun 7:12,619
Acknowledgments
We thank Foltz lab members Ann Hogan and Ewelina Zasadzinska for their helpful discussion and inputs in the preparation of the manuscript.
Funding
D.R.F. was supported by NIH R01GM111907 and by a Zell Scholar award from the Robert H. Lurie Comprehensive Cancer Center.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Srivastava, S., Foltz, D.R. Posttranslational modifications of CENP-A: marks of distinction. Chromosoma 127, 279–290 (2018). https://doi.org/10.1007/s00412-018-0665-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-018-0665-x