Abstract
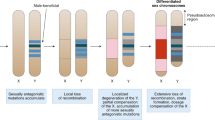
The Y and W chromosomes of mammals and birds are known to be small because most of their genetic content degenerated and were lost due to absence of recombination with the X or Z, respectively. Thus, a picture has emerged of ever-shrinking Ys and Ws that may finally even fade into disappearance. We review here the large amount of literature on sex chromosomes in vertebrate species and find by taking a closer look, particularly at the sex chromosomes of fishes, amphibians and reptiles where several groups have evolutionary younger chromosomes than those of mammals and birds, that the perception of sex chromosomes being doomed to size reduction is incomplete. Here, sex-determining mechanisms show a high turnover and new sex chromosomes appear repeatedly. In many species, Ys and Ws are larger than their X and Z counterparts. This brings up intriguing perspectives regarding the evolutionary dynamics of sex chromosomes. It can be concluded that, due to accumulation of repetitive DNA and transposons, the Y and W chromosomes can increase in size during the initial phase of their differentiation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Irrespective of the contrasting chromosomal sex-determining mechanisms, the small sizes of the mammalian Y and the chicken W chromosomes perfectly illustrate a prominent aspect of their evolutionary history. Suppression of meiotic recombination between X and Y as well as Z and W chromosomes undoubtedly is one of the first events in the evolution of sex chromosomes. This is the precondition for genetic and morphological differentiation and maintenance of sex chromosome identity (Bergero and Charlesworth 2009; Rice 1996). In making the Y and W different from the X and Z, many possibilities have been realized: inversions, accumulation of heterochromatin, deletion of heterochromatin, suppression of meiotic recombination by interference, fusions between sex chromosomes and autosomes, etc. All such mechanisms act in a very dynamic way to decrease recombination, and a general evolutionary rule seems not to exist. Because of the absence of recombination in the sex-specific region of those chromosomes, genetic alterations cannot be purged. For sex chromosomes, only the presence and functionality of the major sex-determining gene counts. Thus, although the vast majority of mutations have a negative effect on gene function, those occurring in genes linked to the sex-determining gene are not cleared during meiosis. This accumulation of deleterious changes in the Y or W is easily tolerated due to the fact that the X and Z provide corresponding intact copies, which are sufficient to fulfil the gene functions. Once a gene on the W or Y has lost its biological function, it will rapidly degenerate and its DNA sequence will disappear in the further evolution in a process described as “Muller’s ratchet”. Its space may be filled transiently by selfishly propagating transposable elements or DNA pieces inserted from autosomes, but in the long run, this process will force the Y and W to shrink to the miniature size seen in mammals and birds. The general view is that Y and W sex chromosomes will continuously decrease in gene content and in size and end up as degenerated elements which are small, gene depauperated and heterochromatic (Bachtrog 2013; Charlesworth and Charlesworth 2000; Graves 2006). Extrapolating the loss of DNA content for the mammalian Y since it originated from a chromosome similar in size to the X has led to the prediction that the human Y (and all other Ys and Ws as well) finally even will fade to total disappearance (Charlesworth and Charlesworth 2000; Graves 2006).
Taking a closer look at evolutionary younger sex chromosomes of fishes and amphibians, it clearly appears that this perception is incomplete. In these vertebrate classes, contrary to birds and mammals, sex-determining mechanisms show a high turnover and new sex chromosomes appear again and again (Ellegren 2011; Ezaz et al. 2006; Mank and Avise 2009). Also in reptiles, where some groups (Serpentes) have a conserved sex determination system, others show a high diversity (Squamata, Testudines) (Ezaz et al. 2009; Gamble 2010; Kawagoshi et al. 2014; Matsubara et al. 2013). Phylogenetically closely related species or even different populations of one and the same species may have ZW and XY sex chromosome systems. Interestingly, many of these Y and W chromosomes are considerably larger than their corresponding X and Z counterparts, sometimes being the largest chromosome of the karyotype. In the present review, cytogenetic and molecular data that exemplify this observation are compiled. It can be concluded that in the initial phase of sex chromosome evolution and sometimes for quite a long period, the Y and W sex chromosomes increase in size, and that size diminishing may be only a late attribute in the evolutionary history. As an explanation for this process, there is evidence that accumulation of repetitive DNA and transposons, often regarded as epiphenomena of genetic degeneration, plays important roles during the establishment of a sex chromosome in the first phase. Amassing such functionless sequences is a fast and effective way of making an emerging Y or W genetically different from its counterpart and preventing meiotic pairing and crossing over.
Vertebrate sex chromosome structure elucidates their evolutionary history
Currently, deep sequencing approaches in several model vertebrates are shedding light in greater depth on the evolutionary steps involved in shaping specialized Y or W sex chromosomes (Chen et al. 2014; Hughes and Rozen 2012; Hughes et al. 2010; Soh et al. 2014; Zhou et al. 2014). However, for the majority of vertebrates, the information on sex chromosomes accumulated from classic cytogenetic studies is still extremely important for determining the extent of differentiation of sex chromosomes in the heterogametic sex. The transformation from a pair of ordinary homomorphic autosomes to highly heteromorphic sex chromosomes endured a long evolution. This involved the accumulation of constitutive heterochromatin and/or the occurrence of structural changes (e.g. inversions) leading to the suppression of recombination between the homologous chromosome pairs, finally retaining a variable extent of autosomal segments in heterogametic chromosomes in different vertebrates. Remarkably, this evolutionary process can still be visualized among the karyotypes of basal vertebrates where the sex chromosomes exist either in incipient or in advanced forms. In the former case, both sex chromosomes appear homomorphic with limited meiotic isolation whereas in the latter case, they are structurally diverged by possessing small or large segments of heterochromatin in the Y or W chromosomes. In some instances, the sex chromosomes may lack any heterochromatic regions but can achieve some level of structural difference through inversions. Thus, cytological information of sex chromosomes provides a valuable help for evaluating evolutionary relationships of sex chromosomes in a wide range of vertebrates. It is a common belief that in most vertebrates, the Y and W chromosomes tend to become shorter over time compared to their counterpart (X and Z). However, exceptions to this reasoning, especially in groups with high sex chromosome turnover, have not been adequately evaluated. A detailed comparison of the cytological structure of sex chromosomes in vertebrate karyotypes may provide information whether the Y and W chromosomes always follow the general rule in evolving to small heterochromatic elements whereas the X or Z chromosomes always retain their original sizes.
Unusually structured Y chromosomes in mammals
The highly heteromorphic sex chromosomes of mammals evolved from an ordinary autosomal pair. When exactly this happened is still a matter of debate, and the estimates range from a maximum of 300 to a minimum of 166 million years ago (Bachtrog 2013; Bellott et al. 2014; Cortez et al. 2014; Lahn and Page 1999; Livernois et al. 2012). Strikingly, the X-linked genes, contrary to the genes in all autosomes, are mostly conserved in all eutherian mammals. To a large extent, the same is true for the Y chromosome, although some Y-linked genes experienced a lineage-specific loss (Bellott et al. 2014). Thus, given the remarkable conservation and long evolutionary history, the mammalian X and Y chromosomes are considered as “old” sex chromosomes.
In the vast majority of karyotypes, the so-called original-type X chromosome of most mammals constitutes approximately 5 % of the female haploid karyotype, a phenomenon consistent with the conservation of its gene content. In contrast, the Y chromosome generally is small and mostly heterochromatic, i.e. at an evolutionary endpoint largely owing to the steady decay of the ancestral Y chromosome. The process of ancestral Y chromosome degeneration may have started early on in the evolution of the Y. In basal eutherian lineages like Afrotheria (elephant) and Xenarthra (armadillos and sloths), the Y chromosomes are small and highly differentiated from the X alike the situation in the vast majority of other mammals (Table 1).
However, classical cytogenetic studies already pointed out that in several species, the X and Y can have an unusual organization (Fredga 1970). In the Chinchilla rat, Chinchilla lanigera and fin whale, Balaenoptera physalus, the X chromosome is extraordinarily large but the Y chromosome did not change and remained very small. The Y can appear as a small dot-like chromosome, being completely heterochromatic in the karyotypes of dog (Carnivora), sheep (Artiodactyla) and sable (Carnivora). On the contrary, in some rodents, like hamsters and some voles (Microtus) and also a few species of Artiodactyla, both X and Y are exceptionally large due to accumulation of heterochromatin. The Y can be from half the size of the X up to reaching the same size or can even be larger as in the tufted deer, Elaphodus cephalophus (Table 1).
The variability of the structure and/or size of the Y chromosome is most pronounced in rodent karyotypes. In the majority of mammalian species, the Y remains small and heterochromatic. But in some karyotypes, the Y becomes enlarged due to accumulation of constitutive heterochromatin, while in others, it is drastically reduced to a dot-like chromosome without exhibiting visible amounts of constitutive heterochromatin (Table 1). In the Transcaucasian mole vole, Ellobius lutescens, and two spiny rats, Tokudaia tokunoshimensis and Tokudaia osimensis, the existence of a Y chromosome in the karyotype is even hard to visualize (Honda et al. 1977; Matthey 1953) or might have been lost (for review and in depth discussion see (Just et al. 2007; Murata et al. 2012)).
The laboratory mouse is known for its relatively large Y, which comprises a full 3 % of the whole mouse genome. Sequencing of the Y chromosome revealed that it is almost entirely euchromatic and that only 2 % of the male-specific region on the Y derives from the ancestral autosome that gave rise to the mammalian sex chromosome. The majority of the sequences that make up its current size are recently acquired and massively amplified families of testis expressed genes, which are lineage specific (Soh et al. 2014).
Even more intriguing is the situation in an Argentinian rodent, Ctenomys maulinus, where the Y appears completely euchromatic and has the same size as the X (Suarez-Villota et al. 2014). Despite these features, the Y does not show extensive pairing with the X during meiosis. Thus, remarkable expansion or contraction of Y chromosome length occurred rapidly within this clade because many species of Ctenomys have heteromorphic sex chromosomes with a small heterochromatic metacentric or subtelocentric Y.
A number of other mammals, such as the Asian house shrew Suncus murinus and some hamster species, have large X and Y chromosomes with few heterochromatic regions (Table 1). However, they show extensive synapsis during meiosis, which may imply retention of a large pseudoautosomal region in some Y chromosomes. On the contrary, it may be conjectured that some autosomal segments may have been transposed to both X and Y chromosomes resulting in autosomal-like traits on the differentiated sex chromosomes. Indeed, in Zoo-FISH analysis of the chromosomes of the South African climbing mouse, Dendromus mesomelas, both large X and Y chromosomes were simultaneously labelled with mouse (Mus musculus) chromosome 15 paint (Solano et al. 2014). This indicates a scenario where both X and Y become enlarged through the acquisition of an autosomal segment, which creates a “neo-pseudoautosomal region” on the sex chromosome pair.
Y chromosome size reduction also occurs on the young neo-Y chromosomes. For example, the neo-Y chromosomes of the mongoose Herpestes sanguineus can be distinguished from X2, which is somewhat smaller with a longer p- and a shorter q-arm. In a related species, Herpestes auropunctatus, the neo-Y can hardly be detected in mitotic metaphases (Fredga 1970; Raman and Nanda 1982).
The present survey of karyotypes from many mammalian orders shows that tiny Y chromosomes are not rare (Table 1). In many instances, these tiny Ys are non-heterochromatic. The sequencing of the Y chromosome revealed that it is 99.9 % euchromatic consisting of ampliconic sequences that are expressed in the testes and distinct from what is typically referred to as repetitive sequences that constitute heterochromatin (Soh et al. 2014). Hence, though heterochromatin accumulation is considered an important step for the differentiation of sex chromosomes, it may eventually become disconnected with some differentiated Y chromosomes. They can be entirely heterochromatic with all intermediate stages up to being almost completely euchromatic (C-band negative).
In summary, mammalian Y chromosomes show quite some heterogeneity in size, ranging from tiny, dot-like Ys to unexpectedly large elements which can even reach the size of the X chromosome.
Differing extent of W chromosome degeneration in birds
As in mammals, sex determination in birds is accomplished through a stable sex chromosome system. Both Z and W chromosomes, like the mammalian XY pair, have evolved from a pair of autosomes, but from a different pair than the mammalian sex chromosomes (Marshall Graves and Shetty 2001; Nanda et al. 1999). Similar to the mammalian Y chromosome, the avian W is generally small in size, highly heterochromatic and contains palindromic sequences (Davis et al. 2010). Based on the uniform size and morphology of the Z chromosome in various avian species, Ohno (Ohno 1967) first proposed that the Z was highly conserved throughout the avian lineages, which was later confirmed by comparative FISH mapping (Nanda et al. 2008; Nishida-Umehara et al. 2007). Thus, the bird sex chromosomes like those of mammals are defined as old sex chromosomes. Though both avian Z and mammalian X chromosome derived from opposite sex-determining systems (♂ birds: homogametic ZZ, ♀ birds: heterogametic ZW; ♂ mammals: heterogametic XY, ♀ mammals: homogametic XX), both have acquired testis-specific genes and display many comparable features, which can be explained by convergent evolution (Bellott et al. 2014). Similar to the mammalian Y, the avian W evolved by losing most of its ancestral genes through recombination arrest and after continuously accumulating repetitive DNA sequences. Subsequent deletions of these highly reiterated DNA sequences apparently led to the distinctly reduced size of the W chromosome in extant avian species. Recent DNA sequence analysis in 17 bird species undoubtedly disclosed that the evolution of Z and W chromosomes, complementary to the mammalian X and Y, occurred in several strata involving inversion and recombination suppression between the two sex chromosomes (Zhou et al. 2014).
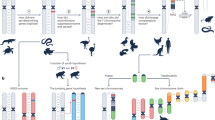
Comparative avian cytogenetics, in particular detailed comparisons of karyotypes in the two sister groups, the palaeognathous and the neognathous birds, revealed that in these vertebrates, the evolution of sex chromosomes occurred in discrete stages (Fig. 1a). In general, Z and W chromosomes of all palaeognathous birds are largely homomorphic and euchromatic. This status appears to correspond to the ancestral situation and indicates the existence of a large pseudoautosomal region in the W chromosome (Nishida-Umehara et al. 2007). However, an initial step towards the differentiation of W chromosome by heterochromatin accumulation was cytogenetically detected in the elegant crested tinamou (Tinamiformes) in which the W chromosome is partially heterochromatic and the size is already slightly diminished (Tsuda et al. 2007).
Evolutionary pathways of old and young sex chromosomes as exemplified in birds (a) and medaka (b). a Outline of the evolution of homomorphic and heteromorphic sex chromosomes in ratite (palaeognathous) and carinate (neognathous) birds. The rearrangement of ancestral sex linkage group (proto sex chromosome) through inversion and accumulation of heterochromatin may be a common mechanism underlying the evolution of “old sex chromosomes” in birds and mammals. b Unique mode of sex chromosome evolution in medaka involving duplication and insertion of a tiny chromosomal segment containing sexual regulatory locus. This mode of evolutionary strategy may contribute to rapid generation of “young” sex chromosomes. PAR pseudoautosomal region, 1 heterochromatin accumulation, 2 inversion
In the karyotypes of most neognathous birds, the W chromosome is distinctly smaller than the Z and, as a result of continuous accumulation of repetitive DNA sequences, has often acquired a completely heterochromatic structure. Since this is also the case in basal lineages as the Anseriformes and Galliformes, a rapid degeneration of the ancestral W chromosome could be deduced. However, in a comparative analysis of 200 bird species, a gradual shrinkage of the W chromosome during avian evolution was not evident and the length of the W chromosome can vary due to amplification of non-coding regions without any phylogenetic pattern observed (Rutkowska et al. 2012).
Though in many neognathous bird karyotypes, the W chromosome is generally smaller than the Z but there are some exceptions. Notably, in two species, the crimson finch, Neochmia phaeton (Passeriformes), and the paddy bird, Ardeola grayii (Pelecaniformes), the W is larger than the Z chromosome (Table 2). The W chromosome in the crimson finch is completely heterochromatic suggesting that the pseudoautosomal region is disrupted by stretches with enormous amplification of non-coding DNA. Interestingly, in the karyotypes of some hawks, eagles and kites (Accipitriformes) and owls (Strigiformes), the W chromosome is large but can be completely or only partially heterochromatic. It is noteworthy to mention that the karyotypes of most species belonging to the family Accipitridae are derived. Compared to other neognathous birds, they possess a reduced diploid chromosome number, only few microchromosomes but many bi-armed chromosomes. All these facts indicate that a process of fusion between microchromosomes and between micro- and macrochromosomes has occurred during the chromosome evolution in the Accipitridae. Hence, stretches of non-heterochromatic regions in the W chromosome of some of these species could be the result of a shift of microchromosomal DNA sequences. Such an inter-chromosomal rearrangement involving both sex chromosomes has been shown in the great reed warbler, Acrocephalus arundinaceus (Pala et al. 2012).
Further dynamics in W chromosome differentiation is evident in two parrots (the yellow-faced parrot, Salvatoria xanthops, and short-tailed parrot, Graydidascalus brachyurus) where a large part of the W chromosome is not heterochromatic although it is smaller than the Z (Caparroz and Duarte Barbanti 2004).
Similar to the mammalian Y chromosome, the avian W chromosome was subjected to extraordinary structural changes during evolution. In mammals, Y chromosome degradation seems to have been a rapid evolutionary process and in many karyotypes, the tiny Y chromosomes appear as euchromatic or poor in heterochromatin content (C-band negative). In contrast, the degradation of the ancestral W chromosome of birds was relatively slow as the W chromosomes of palaeognathous and neognathous birds demonstrate. Recent molecular analysis highlights that the pace of W chromosome degeneration also varies considerably among neognathous birds, i.e. each species has different genes retained in the non-recombining region (Zhou et al. 2014).
Large Ys and Ws among reptiles with different sex chromosome systems
Unlike mammals and birds, the sex-determining mechanisms among reptiles are extremely divergent. They range from genotypic sex determination (GSD), with either male or female heterogamety to temperature-dependent sex determination (TSD) in different lineages. Essentially, all snakes exhibit GSD with female heterogamety, whereas some lizard and turtle species are characterized by both male and female heterogamety. Some of the lineages with different modes of GSD have recently separated from their last common ancestor, as shown for the Agamidae (Ezaz et al. 2009), and given the frequent transition of sex chromosomes, it is likely that their sex chromosomes are “young” sex chromosomes. On the other hand, in Iguana lizards, the sex chromosomes appear to be evolutionary quite old (Rovatsos et al. 2014).
Historically, snakes represented important model organisms for studies on vertebrate sex chromosome evolution. The predominating hypothesis on the origin of sex chromosomes from a pair of autosomes emulated from cytogenetic studies, which demonstrated the presence of homomorphic and heteromorphic sex chromosomes in different groups of snakes (Ohno 1967). In general, Z and W chromosomes are undifferentiated and homomorphic in species of the families Boidae (Becak et al. 1964) and Pythonidae (Singh et al. 1976). An evolutionary series in which the increasing structural complexity of the sex chromosomes in the karyotypes of snakes with differing stages of W chromosome morphological differentiation was initially recognized by Becak et al. (1964). Exceptionally, the W chromosome in the Madagascar boa Acrantophis dumerili (Boidae) is structurally differentiated from the Z by a pericentric inversion that may appear as an initial structural barrier for recombination between two homomorphic sex chromosomes (Mengden and Stock 1980). Z and W in colubrid snakes were long considered to represent an intermediate stage of chromosome differentiation, but a much higher degree with complete cessation of recombination was recently shown at the molecular level (Vicoso et al. 2013). Highly degenerated and heterochromatic W chromosomes are documented in snakes belonging to the Elapidae and Viperidae.
Highly differentiated W chromosomes of snakes are usually smaller than the Z chromosomes and are enriched with repetitive DNA sequences (Jones and Singh 1985). An exception is found in two species, the Trans-Pecos rat snake Elaphe subocularis (Colubridae) and the Sind krait Bungarus walli (Elapidae), where the W chromosome is substantially larger than the Z and completely heterochromatic (Table 3). A further peculiarity is that the W chromosome can be completely differentiated from the Z but may not be entirely heterochromatic, still maintaining some level of homology with the Z as seen in the Okinawa habu Trimeresurus flavoviridis (Viperidae) (Matsubara et al. 2006).
Interestingly, the Z chromosome of snakes, like the avian Z, appears to be evolutionarily conserved (Matsubara et al. 2006). A similar situation has been found in birds, which would suggest that the ZW sex chromosomes of snakes behave similar in evolutionary terms as the ZW chromosomes of birds. However, it became clear from gene mapping studies that sex chromosomes in different reptilian groups have evolved from different ancestral autosomes. Comparative gene mapping experiments showed the existence of homoeologies between the avian Z chromosome and various autosomes of snakes, but no apparent homoeologies between the Z chromosomes of birds and snakes (Srikulnath et al. 2009).
Many lizards and some turtles have highly differentiated sex chromosomes. Contrary to the stable ZZ/ZW sex chromosomes of snakes, frequent transitions between different types of heterogamety (XY♂ → ZW♀ and vice versa) and even TSD is observed (Ezaz et al. 2009; Gamble 2010). This is conspicuously evident among lizards, especially in the family Gekkonidae, which includes species having both male and female heterogamety as well as TSD. Additionally, in lizards, the XY and ZW sex chromosomes can be homomorphic as well as heteromorphic. This diversity in sex chromosome morphology and differentiation is the result of inversions, fusions and heterochromatin accumulation (Cole et al. 1967; King 1977; Matsubara et al. 2014; Moritz 1984; Olmo et al. 1987). Furthermore, in many species, a degeneration of the Y or W chromosomes took place by deletions in their heterochromatic segments making these chromosomes distinctly smaller than the X and W chromosomes (Matsubara et al. 2013). Finally, both Z and W chromosomes show different morphological forms in different races of the purplish dtella Gehyra purpurascens, implying that the structure of sex chromosomes is subjected to rapid change (Moritz 1984).
In several instances, the Y and W chromosomes of reptiles are larger than their X and Z counterparts (Table 3). The Y chromosome is the largest chromosome in the karyotype of the patternless delma, Delma inornata. The W chromosome of the turnip-tailed gecko, Thecadactylus rapicauda, is larger than the Z, and this size difference is caused exclusively by the presence of an interstitially located heterochromatic segment in the W. A more striking situation is the extreme reduction of the size of the Z chromosome in the fat-tailed gecko, Underwoodisaurus milii (Pokorná et al. 2014), and the Malagasy giant chameleon Furcifer oustaleti, whereas the large W chromosome is completely heterochromatic (Rovatsos et al. 2015). Furthermore, in monitor lizards (Varanus spp.), the differentiated sex chromosomes have the sizes of microchromosomes. Due to accumulation of constitutive heterochromatin, the W is somewhat larger than the Z (Matsubara et al. 2014).
Although TSD is the common mode of sex determination in turtles, a few species with heteromorphic sex chromosomes are known. These sex chromosomes are found in both males and females with no common phylogenetic roots which forces the conclusion that evolutionary transitions of sex-determining modes took place in these vertebrates. In parallel, turtle sex chromosomes have also experienced considerable structural changes leading to large homomorphic XY chromosomes in the black marsh turtle Siebenrockiella crassicollis and to microchromosome-sized and differentiated Y and W sex chromosomes in the eastern long-necked turtle, Chelodina longicollis, the spiny softshell turtle, Apalone spinifera, and the Chinese soft shelled turtle, Pelodiscus sinensis (Badenhorst et al. 2013; Kawai et al. 2007). In both A. spinifera and P. sinensis, the W chromosomes appear to be larger even in these microchromosome-sized ZW chromosomes (Table 3). Moreover, the long segments in the large Y chromosomes of S. crassicollis, the Mexican giant musk turtle, Staurotypus triporcatus, and the giant musk turtle, Staurotypus salvinii, are not heterochromatic (Kawagoshi et al. 2012; Kawagoshi et al. 2014) indicating that the sex chromosomes are still in an early stage of structural differentiation. In fact, recent mapping studies exemplify the amplification of microsatellite repeat motifs is strongly correlated with the extent of differentiation and heterochromatinization of Ys and Ws in many sauropsids (Matsubara et al. 2015; Pokorná et al. 2011).
In conclusion, reptiles display a remarkable variety of different sex chromosomes and sex-determining systems. This is reflected by dynamic changes in the cytological morphology and molecular structure of the sex chromosomes. An enlargement of the W and Y chromosomes by accumulation of repetitive DNA sequences (as visible constitutive heterochromatin) has occurred in some species of snakes, lizards, turtles.
Large Y and W chromosomes in amphibians
In the vast majority of amphibians, sex is determined genotypically through either male or female heterogamety. ESD or polygenic modes of sex determination in natural populations are rare or a matter of debate (Hillis and Green 1990). Phylogenetic analyses suggested that female heterogamety was the ancestral state of sex determination and male heterogamety is thought to have subsequently and independently evolved several times in different lineages (Hillis and Green 1990). It has been proposed that the amphibian sex chromosome systems can go through transitions not only on the inter-species but also on the intra-species level. This hypothesis is based on very detailed studies in one frog species, the Japanese wrinkled frog, Rana rugosa, which exhibits ZW and XY, as well as undifferentiated sex chromosomes in different populations (Uno et al. 2008). Gene mapping data suggest that these sex chromosomes existing in different populations are largely homoeologous indicating their common origin.
Cytogenetic analyses so far revealed that most amphibian species lack morphologically differentiated sex chromosomes implying that they still have not developed supramolecular differences to an extent that this becomes evident on the cytological level. Such homomorphy, which is taken as an indication of structural collinearity and high molecular similarity, is evident in the recently diverged hylid frogs where casually recombination can occur all along the sex chromosomes (Stöck et al. 2011). In the African clawed frog Xenopus laevis, the sex-determining gene DM-W is located on the W chromosome. It is a duplicated but truncated version of the autosomal dmrt1 gene and most likely acts as a dominant negative version that abrogates the male-determining function of Dmrt1 (Yoshimoto et al. 2008). However, besides this W chromosome-linked sex-determining gene, no further evidence for molecular or cytological differentiation of the ZW sex chromosomes of X. laevis was demonstrated. In the genome of the closely related Western clawed frog, Silurana tropicalis, DM-W is not present and a different sex-determining gene appears to be in action. Again, the homomorphic sex chromosomes seen in this species do not show any sign of differentiation. Highly sensitive methods (RAD-tags and SNP analyses) failed to detect any differences on the molecular level in this species (Bewick et al. 2013), which was taken as evidence for the existence of a large pseudoautosomal region and a young age of the sex chromosomes. Recently, it was shown that in this species, even three types of sex chromosomes co-exist, which despite being indistinguishable by morphology or molecular markers act as strong genetic sex determiners either as W, Z and Y (Roco et al. 2015).
A compilation of homomorphic and heteromorphic sex chromosomes found in amphibian species was presented by Schmid et al. (Schmid et al. 2010). Although most amphibian species possess homomorphic sex chromosomes, in some anuran and urodelan species, subtle structural changes, which indicate an initial stage of morphological differentiation of sex chromosomes, are apparent. For example, in the otherwise homomorphic sex chromosomes of the edible frog, Rana esculenta, the Y chromosome can be differentiated from the X by having a late replicating region (Schempp and Schmid 1981). Likewise, in some Triturus species (Table 4), the terminal region of the Y chromosome shows heterochromatin accumulation that is not observed on the morphologically identical X chromosome (Schmid et al. 1979). Interestingly, some of the highly differentiated Y or W chromosomes detected in few amphibian species resemble those found in birds and mammals, as they are heterochromatic and are distinctly much smaller than the X and Z. For instance, the W chromosomes in the African bullfrog, Pyxicephalus adspersus, and Y chromosomes in several species of salamanders (Necturus sp.) are small and heterochromatic (Schmid 1980; Sessions 1980). Exceptional cases of XY sex chromosomes are found in the Walker’s marsupial frog, Gastrotheca walkeri (Schmid et al. 2002a; Schmid et al. 2010). In this hemiphractid frog, the Y chromosome contains less amounts of constitutive heterochromatin than the X chromosome.
The W chromosome in the common bell-ring frog, Buergeria buergeri, is found to be practically free of heterochromatin and only slightly smaller than the Z (Schmid et al. 1993). These cases provide evidence that the differentiation of amphibian sex chromosomes can proceed through very different ways and that there is not a general evolutionary rule which can be hypothesized.
As observed in many reptiles, some amphibian karyotypes show highly differentiated sex chromosomes with Y or W chromosomes, which are either slightly larger or even distinctly larger than their X or W chromosomes (Table 4). In Triturus species, both X and Y appear equal in length in conventionally stained karyotypes but after demonstration of the constitutive heterochromatin by the classical C-banding technique, the Y chromosomes turned out to be slightly larger. Several species from the genus Eleutherodactylus have W or Y chromosomes that are distinctly larger than their Z and X counterpart. Yet, it is interesting to note that the large W may be either fully or partially heterochromatic. In two closely related species, Pristimantis euphronides and Pristimantis shrevei, the W chromosome is the largest chromosome in the karyotype and is completely heterochromatic. Furthermore, in the karyotype of the South American marsupial frog, Gastrotheca riobambae, Y is the largest chromosome in the karyotype (11.5 % of the haploid genome) and is completely heterochromatic. A totally reverse pattern is seen in another anuran species, Physalaemus ephippifer, with a W larger than the Z but which is not heterochromatic. Of note, seven out of approximately 150 Eleutherodactylus species examined showed W chromosomes that were larger than the Z chromosomes indicating an explosive amplification of repetitive DNA sequences in the W chromosomes (Schmid et al. 2010).
Only 4 % of cytogenetically analysed amphibian species are reported to have structurally differentiated sex chromosomes. Remarkably, these few species exhibit a prevalence of larger Y and W chromosomes with various levels of heterochromatin accumulation. However, it should not be ignored that there are also species in which the Y and W chromosomes exhibit less amounts of constitutive heterochromatin than those present in the X and Z chromosomes.
Large Y and W chromosomes are exceedingly represented in fish lineages
Fishes exhibit the widest spectrum of sex-determining systems of all vertebrates ranging from different types of environmental sex determination to a variety of chromosomal sex-determining systems (Kobayashi et al. 2013; Volff et al. 2007).
Cytogenetic studies in the past have undoubtedly demonstrated that in the majority of gonochoristic fish species, the sex chromosomes are homomorphic and heteromorphic sex chromosomes have been encountered in roughly 10 % of species karyotyped (Devlin and Nagahama 2002). An extreme case of conserved identity between sex chromosomes is represented in the tiger pufferfish, Fugu rubripes, which has an XY sex determination system. Here the sex chromosomes differ by one non-synonymous substitution in the Amh-receptor II gene that is the sex-determining gene (Kamiya et al. 2012). No suppression of recombination around the male allele of this gene was found, which would have led to a structural differentiation of the X and Y. Remarkably, this situation is also found in two other species of Takifugu, which diverged more than 5 million years ago. Hence, over this period, suppression of recombination was not established and could not initiate the divergence of the Y chromosome from the X that is usually seen elsewhere.
In medaka, a male dominant sex determination system was described by classical genetic studies nearly 100 years ago (Aida 1921). More recent studies with molecular DNA probes revealed that, despite recombination suppression around the male-specific region on the Y (MSY), all markers were present on the X and Y with only minor sequence differences (Kondo et al. 2001). FISH probes derived from sex-linked markers identified the sex chromosome pair as the second largest in the karyotype (Matsuda et al. 2002; Nanda et al. 2002). The lack of morphological differentiation and low molecular differences were taken as evidence for a relatively young age of the medaka sex chromosomes. Indeed, it was shown that the Y chromosome of medaka is only 5 to 10 million years old (Kondo et al. 2004). Homomorphic sex chromosomes were also found in the platyfish Xiphophorus maculatus, where in different populations, Y, W and X chromosomes were identified by sex-linked phenotypes (Kallman 1984) and molecular markers (Böhne et al. 2009; Zhou et al. 2010).
All salmonid species studied so far have a XX/XY sex chromosome system (Phillips 2013). As in many other fishes, the XY chromosomes in salmonids contain a small sex-specific region and a large pseudoautosomal region and only small structural differences between X and Y were reported in some trouts (Davidson et al. 2009). FISH mapping along with genetic mapping have already indicated that the sex chromosomes belong to different linkage groups (Woram et al. 2003). In rainbow trout, the male sex-determining gene sdY was identified and found to be a truncated version of a duplicate from the interferon response factor gene irf9 (Yano et al. 2012). Evidence was provided that sdy is also the Y-linked male-determining gene in other salmonids (Yano et al. 2013). Comparative analysis of the sex-determining region shared by three salmonids (rainbow trout, Chinook salmon and Atlantic salmon) revealed a minimal region to trigger masculinization. Interestingly, this region contains all elements necessary for transposition such as transposase and RNA-directed DNA polymerase (Faber-Hammond et al. 2015). Thus, sdY may have transposed between different linkage groups and thus became linked to different chromosomes that developed into a Y (Faber-Hammond et al. 2015; Lubieniecki et al. 2015).
A striking evidence for the low degree or even absence of genetic differentiation of sex chromosomes in fishes is that fully viable and fertile sex-reversed individuals can be obtained with ease. For instance, in medaka and tilapia, XX males occur spontaneously or can be obtained when embryos are incubated at higher temperatures (Baroiller et al. 2009; Sato et al. 2005; Shinomiya et al. 2010) and a plethora of studies have been published were treatment with hormones or endocrine disruptors produces all types of sex reversals, even YY females (Devlin and Nagahama 2002; Gutierrez and Teem 2006).
Several Neotropical species display various patterns of morphologically differentiated sex chromosomes, ranging from simple sex chromosome systems to multiple sex chromosomes (de Almeida Toledo and Foresti 2001). Differentiation of sex chromosomes in these species is largely achieved by heterochromatin accumulation (Cioffi et al. 2012). Karyotypes of species belonging to the genera Characidium (Crenuchidae) and Triportheus (Characidae) are fairly conserved and present identifiable sex chromosomes. In the genus Triportheus, several species with well-differentiated W chromosomes show variable extent of heterochromatin accumulation (Artoni et al. 2001). On the other hand, in many species of the genus Characidium, the Z and W chromosomes are of similar length. The dynamic transformation of sex chromosomes can be elegantly marked in different populations of the wolf fish species complex, Hoplias malabaricus, with seven karyomorphs showing undifferentiated or differentiated XX/XY sex chromosomes and neo-Y sex chromosomes (Cioffi et al. 2011). A rare phenomenon with regard to sex chromosome differentiation is apparent in one population. Here, the Y chromosome appears smaller than the X, but its organization is unusual in the sense that the X chromosome has accumulated more heterochromatin than the Y chromosome. Similarly, in the glass knifefish, Eigenmannia virescens, which also displays an early stage of sex chromosome differentiation in some populations, heterochromatin accumulation appears to be confined to the X chromosome (de Almeida Toledo and Foresti 2001). This unusual accumulation of heterochromatin in the X chromosome is unexpected from the traditional evolutionary model of early XY/ZW differentiation in vertebrates, which predicts heterochromatization of the Y and W sex chromosome.
Strikingly, in many species with differentiated sex chromosomes, the Y or W chromosome of the heterogametic sex is larger than its X or Z counterpart (Table 5). This can range from a minor size difference that is noted only by most experienced specialists to extreme cases where the W or Y is the largest of the whole karyotype.
Some species of the genera Parodon, Apareiodon and Saccodon (Parodontidae) are known to have very large W chromosomes (Table 5) though the majority of species show simple ZZ/ZW sex chromosomes with smaller W chromosomes. In the canivete, Parodon hilarii, the W chromosome is the largest chromosome in the karyotype and highly heterochromatic (Vicente et al. 2003). In several species of Apareiodon, the W chromosomes are also larger than the Z chromosome (Table 5) but not completely heterochromatic and the euchromatic region of the W may be comparable to that on the Z (Bellafronte et al. 2012; Bellafronte et al. 2009), implying that the W chromosome in these species may be in an early stage of differentiation with a still large pseudoautosomal region. In the Neotropical genus Leporinus (Anostomidae), eight of the around 40 known species have female-specific heteromorphic sex chromosomes. In at least seven species, the W chromosome is enormously large compared to the Z owing to heterochromatin accumulation (Fig. 2a). A recent study using W chromosome-specific paints highlights the common origin of the differentiated sex chromosomes, and these W chromosome paints do not cross-react to the species with undifferentiated sex chromosomes (Parise-Maltempi et al. 2013).
Unusually large sex chromosomes of fishes. a Divergent organization of large heterogametic Y and W sex chromosomes in three different fish species. b C-banded metaphase and DAPI staining of the exceptionally large W chromosome of the Western mosquitofish. Note that except the centromeric region, the large W chromosome is not heterochromatic
In the Asian ricefish of the genus Oryzias all species have a well-defined genetic mode of sex determination but the sex chromosomes cannot be morphologically recognized except in one species, Oryzias hubbsi. This species has in contrast to the vast majority of species of Oryzias, which have a XX/XY system, acquired a ZZ/ZW system (Takehana et al. 2007). The W chromosome is larger than the Z due to an additional heterochromatic region at the terminal region, but most of part of the W chromosome appears to be euchromatic.
Stickleback fishes (Gasterosteidae) have provided important insights to sex chromosome evolution. This small teleost group shows remarkable sex chromosome diversity. Both three-spine and nine-spine sticklebacks have XX/XY sex chromosomes, but their Y chromosome organization is different. The Y chromosome in three-spine stickleback is smaller than the X and might have undergone pronounced degeneration, whereas in the nine-spine stickleback, the Y is larger than the X (Ross et al. 2009). On the other hand, the four-spine stickleback has a ZZ/ZW chromosome and the W chromosome is much larger than the Z (Ross et al. 2009).
The growth mechanism: what makes sex chromosomes grow?
Several investigations, in particular on fishes, provide some information on the question what makes sex chromosomes grow. A recent study investigated the distributions of transposable elements on the entire sex chromosomes compared to autosomes and in the genomic regions harbouring the sex determination loci of medaka, Oryzias latipes, platyfish, X. maculatus, guppy, Poecilia reticulata, tilapia, Oreochromis niloticus, the pufferfish, Takifugu rubripes, the Chinese tongue sole, Cynoglossus semilaevis, and three-spine stickleback, Gasterosteus aculeatus (Chalopin et al. 2015). In these fishes, where whole-genome sequence information is available, it is evident that transposons accumulate not only on Y or W sex determination segments but also on the corresponding regions of the X or Z. Furthermore, recent activity of some classes of transposable elements seems to be specific to these regions. This may indicate that the process of transposon accumulation on these young sex chromosomes is still ongoing. The focus on two closely related strains of medaka showed that the chromosomal region was relatively poor in transposons prior to the emergence of the new sex-determining gene. This gives support to the reasoning that young sex-determining regions sustained recent accumulations and amplifications of mobile elements. For instance, the less than 10-million-year-old Y of the three-spine stickleback has in the 3-Mb large region that contains the sex-determining gene 4.2 % of this region constituted by transposable elements, while this value is only 1 % on the corresponding region of the X (Peichel et al. 2004). The expansion of this compartment is mainly due to a massive increase in LINEs.
There are two species of fish, where extensive sequence information from heterogametic sex chromosomes is now available, namely from the medaka Y and Chinese tongue sole W. In medaka, the male sex-determining gene arose from duplication of an autosomal fragment that contains the dmrt1 gene (Nanda et al. 2002). This fragment was inserted into another chromosome, which became the proto-Y (Fig. 1b). There, the new copy of dmrt1, named dmrt1bY (or DMY), evolved to become the master regulator of male development. As predicted from the evolutionary hypothesis on sex chromosome evolution, all co-duplicated genes degenerated. Concomitantly, other duplicated gene fragments from other regions of the genomes became inserted in the male-specific region on the Y (MSY) (Kondo et al. 2006). Comparison of the MSY to the autosomal region from which it was derived also uncovered a profound expansion in the copy number of repetitive DNA, most importantly of four distinct satellite sequences and various transposable elements. This resulted in the growth of the medaka MSY over its 5 to 10 million years of existence from 45 to 250 kb. The fivefold increase in size makes the Y in principle larger than the X, but it is still too small to become apparent as a visible difference in chromosome morphology. Suppression of recombination around the MSY can be detected over a region of 3.5 Mb (Takeda 2008; Wilson et al. 2014), but this resulted only in sparse and minor sequence differences between X and Y alleles and did still not contribute to a visible change in chromosome morphology.
In the Chinese half-smooth tongue sole, a draft sequence of the W has been produced (Chen et al. 2014). Different from the medaka Y where the MSY is tiny and the pseudoautosomal region spans most of the chromosome, the tongue sole W has only 22 protein-coding genes in an approximately 650-kb region where cross-over occurs at a similar rate as on autosomes. The rest of the chromosome shows strongly reduced up to absent recombination and has the expected features of a degenerating sex chromosome. Only about one third of the original 1000 protein-coding genes are still present and appear to be intact. A 10 times higher content of pseudogenes and transposable elements than on the Z and autosomes makes the W considerably larger than its homogametic counterpart and even the largest chromosome of the whole complement. The age of the W was determined to be around 30 million years. Although two thirds of the genes were already lost over this period, this decrease in DNA content has obviously been overcompensated by accumulation of repetitive DNA and transposable elements.
From several fish species, molecular cytogenetic findings confirm such an accumulation process of repetitive sequences. In the platyfish (X. maculatus), the region around the sex-determining locus has been shown to be enriched for transposons (Chalopin et al. 2015; Volff et al. 2003). In particular, a local expansion of the XIR LTR-like transposon in the subtelomeric region of the Y occurred, which can be documented by FISH analysis (Nanda et al. 2000). Such a structure is not seen on the X. But again like in medaka, the transposon expansion on the Y is too small to result in heteromorphy of sex chromosomes in this species.
Guppies have an XY sex determination mechanism and show a very pronounced Y chromosome polymorphism, which manifests in different-sized Ys, ranging from the same size as the X to being the largest chromosome of the complement. This size polymorphism is explained by expansion of a block of constitutive heterochromatin specifically on the Y (Nanda et al. 2014). In one particular species, the Endler guppy, Poecilia wingei, the Y chromosome is enormously large arising from the heterochromatin accumulation at the distal end of the chromosome (Fig. 2a). It must be emphasized that in most cases, heterochromatin accumulation is viewed as a mechanism explaining Y chromosome degeneration, but in P. wingei, the heterochromatin accumulation hardly affected the large pseudoautosomal region. The Y chromosomes of different guppy strains show accumulation of different microsatellite repeats (Nanda et al. 1990; Nanda et al. 2014) which was taken as an indication that the size polymorphism of the heterochromatin block may be due to the expansion of different repetitive sequences.
In the redfin prochilodus, Semiprochilodus taeniatus, the W is the largest chromosome. It presents with broad areas of constitutive heterochromatin at the two terminal regions but also at pericentromeric and interstitial locations. FISH with probes for the Rex 1 transposon indicated that invasion and local expansion of this element were responsible for accumulation of heterochromatin of the W (Terencio et al. 2012). Furthermore, in the large heterochromatic W chromosomes of several Leporinus species, accumulation of Rex transposable elements is detected (Splendore de Borba et al. 2013).
In the mosquitofish, Gambusia affinis, the W chromosome is the largest chromosome of the complement. In contrast to the large W of Leporinus or the large Y chromosome of P. wingei, surprisingly it does not stain in conventional C-banding for additional heterochromatin except for the centromeric region. Also DAPI staining did not reveal specific features for the W, which may indicate that this unique large W chromosome has not undergone degeneration (Fig. 2). This does not contradict the reasoning that the large size of fish sex chromosomes is connected to TEs because in fishes, many transposons are transcribed and still active (Huang et al. 2012) and thus would not be necessarily heterochromatic.
Discussion
The Y and W chromosomes of vertebrates come with an astonishing structural plasticity ranging from tiny, dot-like and cytogenetically almost invisible ones to complete homomorphy with the X or Z, and to cases where they are gigantic and the largest chromosome of the whole complement. This variety reflects the evolutionary trajectory of sex chromosome development. From the common theories that predict that once a sex-determining gene has established itself on a proto-sex chromosome recombination is suppressed around this gene, a one-way street process of degeneration, diminution, gene loss and heterochromatization has to be expected. Those Ys and Ws that are heterochromatic and smaller than the X or Z down to the dot-like-sized ones certainly fit and are generally regarded to be of old age and to represent the endpoint of the degeneration process. This is supported by many examples from mammals and birds.
The homomorphy of many sex chromosomes is in most cases easily explained, when they are of proven young age, because then the process of decay did not go on long enough to leave its traces on the morphology and in many cases also not to a considerable extent on the level of genomic structures. Recent molecular analysis in fishes, e.g. medaka and fugu, confirm this.
An alternative hypothesis has been proposed based on studies on the homomorphic sex chromosomes of hylid frogs, where occasional cross-over between the sex chromosomes in sex-reversed individuals would lead to homogenization of both chromosomes and explain results obtained with sex chromosome specific molecular markers (Perrin 2009).
When homomorphic sex chromosomes appear in one lineage but highly degenerated ones in a parallel lineage where both are derived from a common ancestor, a reasonable explanation is a different and lineage-specific speed of the degeneration process. Such a situation is for instance found in the palaeognathous and neognathous birds.
The unexpected ones are, however, the Y and W chromosomes, which are larger than their homogametic counterparts and those which are euchromatic. To explain their existence, we propose first to categorize them as the result of secondary or primary sex chromosome growth.
A clear case of secondary growth is found in the laboratory mouse. Here it is evident from the sequencing of the whole Y that those sequences which make it larger have been added more recently to a shared common small piece with the other mammalian Ys (Soh et al. 2014). Typically, such examples of secondary growth would locate at derived positions in a phylogenetic tree, where most, and in particular the more basal species, would show the more widespread small Y type. Secondary growth would be a phenomenon to occur in groups with old sex chromosomes, but it rarely concedes the Y chromosome to exceed the size of X chromosome.
As primary growth we regard those instances, where young Ys and Ws are larger than their X or Z counterpart. We hypothesize that in the initial phase of sex chromosome evolution, the repression of recombination will favour accumulation of transposons and expansion or repetitive sequences. This is a rapid and effective mechanism to make the emerging Ys and Ws different from the other chromosomes and thus even further prevents crossovers. This process that is well documented in several young fish sex chromosomes (e.g. in medaka) will then lead to a stage where the addition of material becomes visible as a larger chromosome in the karyotype. Only in a further phase of their evolution the degeneration process and gene loss initiates the shrinking of the chromosome to the well-known small size of the mammalian and avian sex chromosomes. It should be noted that the proposed dynamics of sex chromosome morphology makes them pass a second time through a stage where both are of the same size and thus appear homomorphic.
Three different organization profiles of these large Y and W organization can be delineated Firstly, both the large Ys and Ws are nearly heterochromatic (e.g. Leporinus) that may emphasize the enormous amplification of repetitive sequences. Expansion of microsatellite arrays is known to occur in young Y chromosomes of plants (Kejnovský et al. 2013). However, this pattern of organization does not bear much importance, as it does not affect the recombination suppression. Secondly, both the large Ys and Ws are partially heterochromatic (e.g. guppy, O. hubbsi) that may signify the early stage of sex chromosome differentiation, as well as recombination within the large pseudoautosomal region. Interestingly, both highly differentiated Y and W chromosomes in old sex chromosome systems, as encountered in mammals and birds, show extreme contraction of their length, and they even loose most of the heterochromatin. This means that different levels of degeneration can affect both young and old sex chromosome. In the Chinese tongue sole, gene loss has occurred to a relatively high level in the early phase of W evolution (Chen et al. 2014), and also in plants, genetic degeneration has been documented to occur soon after suppression of recombination between the Y and X (Hough et al. 2014). Lastly, a more impressive organization is that the Y or W chromosomes are larger than their counterparts (X or Z), but they are nearly euchromatic (e.g. in G. affinis) elucidating a possible regular recombination between the sex chromosomes, unless interrupted through accumulating non-synonymous SNPs. One potential complication in this pattern of organization concerns the underlying basis of Y or W chromosome expansion without heterochromatin accumulation. It is suggested that occasionally the X or Z chromosome may undergo some level of degeneration without involving heterochromatin. Alternatively, taking the example of the large euchromatic Y in mouse (Soh et al. 2014), one may envisage expansion of amplicons in the heterochromatin free large Y or W chromosomes.
The emerging information from sequencing of still a very limited number of vertebrate sex chromosomes and the wealth of information from decades of cytogenetic investigations employing the classical staining and molecular methods of FISH and chromosome paintings have opened novel and exiting views on the amazing variety of sex chromosome structures and the dynamics of their evolution. Due to their often highly repetitive structure and genetic degeneration, sequencing of sex chromosomes still presents a major challenge and often an insurmountable problem. Nevertheless, we can hope to see in the near future sex chromosomal sequence information not only from the handful of model organisms but also from vertebrates that stand out by an unusual sex chromosome and/or genetic sex determination system.
References
Aida T (1921) On the inheritance of color in a fresh-water fish, Aplocheilus latipes Temmick and Schlegel, with special reference to sex-linked inheritance. Genetics 6:554–573
Ananias F, Modesto ADS, Mendes SC, Napoli MF (2007) Unusual primitive heteromorphic ZZ/ZW sex chromosomes in Proceratophrys boiei (Anura, Cycloramphidae, Alsodinae), with description of C-band interpopulational polymorphism. Hereditas 144:206–212
Ando K, Tagawa T, Uchida TA (1980) The C-banding pattern of 6 Japanese species of Vespertilionine bats (Mammalia, Chiroptera). Experientia 36:653–654
Andreata AA, de Almeida-Toledo LF, Oliveira C, de Almeida Toledo Filho S (1992) Chromosome studies in Hypoptopomatinae (Pisces, Siluriformes, Loricariidae) I. XX/XY Sex chromosome heteromorphism in Pseudotocinclus tietensis. Cytologia 57:369–372
Arslan A, Albayrak I, Pamukoglu N, Yorulmaz T, Toyran K (2009) C-banded karyotype and NORs of the long-eared hedgehog, Hemiechinus auritus from Turkey. Folia Zoologica 58:9–13
Artoni RF, Falcão JN, Moreira-Filho O, Bertollo LA (2001) An uncommon condition for a sex chromosome system in Characidae fish. Distribution and differentiation of the ZZ/ZW system in Triportheus. Chromosome Res 9:449–456
Azevedo NF, Svartman M, Manchester A, de Moraes-Barros N, Stanyon R, Vianna-Morgante AM (2012) Chromosome painting in three-toed sloths: a cytogenetic signature and ancestral karyotype for Xenarthra. BMC Evol Biol 12:36
Bachtrog D (2013) Y-chromosome evolution: emerging insights into processes of Y-chromosome degeneration. Nat Rev Genet 14:113–124
Badenhorst D, Stanyon R, Engstrom T, Valenzuela N (2013) A ZZ/ZW microchromosome system in the spiny softshell turtle, Apalone spinifera, reveals an intriguing sex chromosome conservation in Trionychidae. Chromosome Res 21:137–147
Baroiller JF, D’Cotta H, Saillant E (2009) Environmental effects on fish sex determination and differentiation. Sex Dev 3:118–135
Becak W, Becak ML, Nazareth HR, Ohno S (1964) Close karyological kinship between the reptilian suborder Serpentes and the class Aves. Chromosoma 15:606–617
Bed’Hom B, Coullin P, Guillier-Gencik Z, Moulin S, Bernheim A, Volobouev V (2003) Characterization of the atypical karyotype of the black-winged kite Elanus caeruleus (Falconiformes: Accipitridae) by means of classical and molecular cytogenetic techniques. Chromosome Res 11:335–343
Bellafronte E, Vicari MR, Artoni RF, Margarido VP, Moreira-Filho O (2009) Differentiated ZZ/ZW sex chromosomes in Apareiodon ibitiensis (Teleostei, Parodontidae): cytotaxonomy and biogeography. J Fish Biol 75:2313–2325
Bellafronte E, Schemberger MO, Artoni RF, Filho OM, Vicari MR (2012) Sex chromosome system ZZ/ZW in Apareiodon hasemani Eigenmann, 1916 (Characiformes, Parodontidae) and a derived chromosomal region. Genet Mol Biol 35:770–776
Bellott DW, Hughes JF, Skaletsky H, Brown LG, Pyntikova T, Cho TJ, Koutseva N, Zaghlul S, Graves T, Rock S, Kremitzki C, Fulton RS, Dugan S, Ding Y, Morton D, Khan Z, Lewis L, Buhay C, Wang Q, Watt J, Holder M, Lee S, Nazareth L, Alfoldi J, Rozen S, Muzny DM, Warren WC, Gibbs RA, Wilson RK, Page DC (2014) Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508:494–499
Bergero R, Charlesworth D (2009) The evolution of restricted recombination in sex chromosomes. Trends Ecol Evol 24:94–102
Bewick AJ, Chain FJ, Zimmerman LB, Sesay A, Gilchrist MJ, Owens ND, Seifertova E, Krylov V, Macha J, Tlapakova T, Kubickova S, Cernohorska H, Zarsky V, Evans BJ (2013) A large pseudoautosomal region on the sex chromosomes of the frog Silurana tropicalis. Genome Biol Evol 5:1087–1098
Böhne A, Schultheis C, Galiana-Arnoux D, Froschauer A, Zhou Q, Schmidt C, Selz Y, Ozouf-Costaz C, Dettai A, Segurens B, Couloux A, Bernard-Samain S, Barbe V, Chilmonczyk S, Brunet F, Darras A, Tomaszkiewicz M, Semon M, Schartl M, Volff JN (2009) Molecular analysis of the sex chromosomes of the platyfish Xiphophorus maculatus: towards the identification of a new type of master sexual regulator in vertebrates. Integr Zool 4:277–284
Borodin PM, Sablina OV, Rodionova MI (1995) Pattern of X-Y chromosome pairing in microtine rodents. Hereditas 123:17–23
Busin CS, Andrade GV, Bertoldo J, Del Grande ML, Uetanabaro M, Recco-Pimentel SM (2008) Cytogenetic analysis of four species of Pseudis (Anura, Hylidae), with the description of ZZ/ZW sex chromosomes in P. tocantins. Genetica 133:119–127
Cao X, Jiang H, Zhang X (2005) Polymorphic karyotypes and sex chromosomes in the tufted deer (Elaphodus cephalophus): cytogenetic studies and analyses of sex chromosome-linked genes. Cytogenet Genome Res 109:512–518
Caparroz R, Duarte Barbanti JM (2004) Chromosomal similarity between the scaly-headed parrot (Pionus maximiliani), the short-tailed parrot (Graydidascalus brachyurus) and the yellow-faced parrot (Salvatoria xanthops) (Psittaciformes: Aves): a cytotaxonomic analysis. Genet Mol Biol 27:522–528
Centofante L, Carlos BLA, Moreira Filho O (2002) A ZZ/ZW sex chromosome system in a new species of the genus Parodon (Pisces, Parodontidae). Caryologia 55:139–150
Chalopin D, Volff JN, Galiana-Arnoux D, Anderson J, Schartl M (2015) Transposable elements and early evolution of sex chromosomes in fish. Chromosoma Res 23:545–560
Charlesworth B, Charlesworth D (2000) The degeneration of Y chromosomes. Philos Trans R Soc Lond B Biol Sci 355:1563–1572
Chen Z-PC, Jiang X-L, Wang Y-X (1995) Karyotype and banding patterns of Fea’s tree rat (Chiromyscus chiropus). Caryologia 48:9–16
Chen S, Zhang G, Shao C, Huang Q, Liu G, Zhang P, Song W, An N, Chalopin D, Volff JN, Hong Y, Li Q, Sha Z, Zhou H, Xie M, Yu Q, Liu Y, Xiang H, Wang N, Wu K, Yang C, Zhou Q, Liao X, Yang L, Hu Q, Zhang J, Meng L, Jin L, Tian Y, Lian J, Yang J, Miao G, Liu S, Liang Z, Yan F, Li Y, Sun B, Zhang H, Zhang J, Zhu Y, Du M, Zhao Y, Schartl M, Tang Q, Wang J (2014) Whole-genome sequence of a flatfish provides insights into ZW sex chromosome evolution and adaptation to a benthic lifestyle. Nat Genet 46:253–260
Christidis L (1986a) Chromosomal evolution within the family Estrildidae (Aves). 1. The Poephilae. Genetica 71:81–97
Christidis L (1986b) Chromosomal evolution within the family Estrildidae (Aves). 2. The Lonchurae. Genetica 71:99–113
Cioffi MB, Bertollo LAC (2010) Initial steps in XY chromosome differentiation in Hoplias malabaricus and the origin of an X1X2Y sex chromosome system in this fish group. Heredity 105:554–561
Cioffi MB, Camacho JP, Bertollo LA (2011) Repetitive DNAs and differentiation of sex chromosomes in Neotropical fishes. Cytogenet Genome Res 132:188–194
Cioffi MB, Moreira-Filho O, Almeida-Toledo LF, Bertollo LA (2012) The contrasting role of heterochromatin in the differentiation of sex chromosomes: an overview from Neotropical fishes. J Fish Biol 80:2125–2139
Cole CJ, Lowe CH, Wright JW (1967) Sex chromosomes in lizards. Science 155:1028–1029
Cortez D, Marin R, Toledo-Flores D, Froidevaux L, Liechti A, Waters PD, Grutzner F, Kaessmann H (2014) Origins and functional evolution of Y chromosomes across mammals. Nature 508:488–493
da Rosa R, Bellafronte E, Moreira Filho O, Margarido VP (2006) Constitutive heterochromatin, 5S and 18S rDNA genes in Apareiodon sp. (Characiformes, Parodontidae) with a ZZ/ZW sex chromosome system. Genetica 128:159–166
Dantas SMMdM, Barros RMdS (1997) Cytogenetic study of the genus Saguinus (Callithrichidae, Primates). Braz J Genet. doi:10.1590/S0100-84551997000400014
Davidson WS, Huang TK, Fujiki K, von Schalburg KR, Koop BF (2009) The sex determining loci and sex chromosomes in the family Salmonidae. Sex Dev 3:78–87
Davis JK, Thomas PJ, Program NCS, Thomas JW (2010) A W-linked palindrome and gene conversion in New World sparrows and blackbirds. Chromosome Res 18:543–553
de Almeida Toledo LF, Foresti F (2001) Morphologically differentiated sex chromosomes in Neotropical freshwater fish. Genetica 111:91–100
Devlin RH, Nagahama Y (2002) Sex determination and sex differentiation in fish: an overview of genetic, physiological, and environmental influences. Aquaculture 208:191–364
Elder JFJ, Turner BJ, Thomerson JE, Taphorn DC (1991) Chromosomal divergence and heterogameity in two annual killifishes of the genus Pterolebias. Genome 34:674–676
Ellegren H (2011) Sex-chromosome evolution: recent progress and the influence of male and female heterogamety. Nat Rev Genet 12:157–166
Ezaz T, Stiglec R, Veyrunes F, Marshall Graves JA (2006) Relationships between vertebrate ZW and XY sex chromosome systems. Curr Biol 16:R736–743
Ezaz T, Sarre SD, O’Meally D, Graves JA, Georges A (2009) Sex chromosome evolution in lizards: independent origins and rapid transitions. Cytogenet Genome Res 127:249–260
Faber-Hammond JJ, Phillips RB, Brown KH (2015) Comparative analysis of the shared sex-determination region (SDR) among salmonid fishes. Genome Biol Evol 7:1972–1987
Fredga K (1970) Unusual sex chromosome inheritance in mammals. Philos Trans R Soc Lond B Biol Sci 259:15–36
Gamble T (2010) A review of sex determining mechanisms in geckos (Gekkota: Squamata). Sex Dev 4:88–103
Gilbert C, Goodman SM, Soarimalala V, Olson LE, O’Brien PC, Elder FF, Yang F, Ferguson-Smith MA, Robinson TJ (2007) Chromosomal evolution in tenrecs (Microgale and Oryzorictes, Tenrecidae) from the Central Highlands of Madagascar. Chromosome Res 15:1075–1091
Goldoni D, Rubini M, Fontana F (1984) Cytogenetic studies on Cervus elaphus L. Constitutive heterochromatin and nucleolus organizer regions. Caryologia 37:439–443
Gornung E, Volleth M, Capanna E, Castiglia R (2008) Comparative cytogenetics of moles (Eulipotyphla, Talpidae): chromosomal differences in Talpa romana and T. europaea. Cytogenet Genome Res 121:249–254
Graphodatsky A (1989) Appendix1. An atlas of selected karyotypes; section 1. In: Halnan CRE (ed) Cytogenetics of animals. C.A.B. International., pp 321–352
Graves JA (2006) Sex chromosome specialization and degeneration in mammals. Cell 124:901–914
Gray BA, Zori RT, McGuire PM, Bonde RK (2002) A first generation cytogenetic ideogram for the Florida manatee (Trichechus manatus latirostris) based on multiple chromosome banding techniques. Hereditas 137:215–223
Green DM (2002) Chromosome polymorphism in archey’s frog (Leiopelma archeyi) from New Zealand. Copeia 2002:204–207
Gutierrez JB, Teem JL (2006) A model describing the effect of sex-reversed YY fish in an established wild population: the use of a Trojan Y chromosome to cause extinction of an introduced exotic species. J Theor Biol 241:333–341
Hillis DM, Green DM (1990) Evolutionary changes of heterogametic sex in the phylogenetic history of amphibians. J Evol Biol 3:49–64
Honda T, Suzuk iH, Itoh M (1977) An unusual sex chromosome constitution found in the Amami spinous country-rat, Tokudaia osimensis osimensis. Jap J Genet 52:247–249
Houck ML, Kingswood SC, Kumamoto AT (2000) Comparative cytogenetics of tapirs, genus Tapirus (Perissodactyla, Tapiridae). Cytogenet Cell Genet 89:110–115
Houck ML, Kumamoto AT, Gallagher DS Jr, Benirschke K (2001) Comparative cytogenetics of the African elephant (Loxodonta africana) and Asiatic elephant (Elephas maximus). Cytogenet Cell Genet 93:249–252
Hough J, Hollister JD, Wang W, Barrett SC, Wright SI (2014) Genetic degeneration of old and young Y chromosomes in the flowering plant Rumex hastatulus. Proc Natl Acad Sci U S A 111:7713–7718
Huang CR, Burns KH, Boeke JD (2012) Active transposition in genomes. Annu Rev Genet 46:651–675
Hughes JF, Rozen S (2012) Genomics and genetics of human and primate y chromosomes. Annu Rev Genomics Hum Genet 13:83–108
Hughes JF, Skaletsky H, Pyntikova T, Graves TA, van Daalen SK, Minx PJ, Fulton RS, McGrath SD, Locke DP, Friedman C, Trask BJ, Mardis ER, Warren WC, Repping S, Rozen S, Wilson RK, Page DC (2010) Chimpanzee and human Y chromosomes are remarkably divergent in structure and gene content. Nature 463:536–539
Jones KW, Singh L (1985) Snakes and the evolution of sex-chromosomes. Trends Genet 1:55–61
Just W, Baumstark A, Suss A, Graphodatsky A, Rens W, Schafer N, Bakloushinskaya I, Hameister H, Vogel W (2007) Ellobius lutescens: sex determination and sex chromosome. Sex Dev 1:211–221
Kallman K (1984) A new look at sex determination in poeciliid fishes. In: Turner B (ed) Evolutionary genetics of fishes. Plenum, New York, pp 95–171
Kamiya T, Kai W, Tasumi S, Oka A, Matsunaga T, Mizuno N, Fujita M, Suetake H, Suzuki S, Hosoya S, Tohari S, Brenner S, Miyadai T, Venkatesh B, Suzuki Y, Kikuchi K (2012) A trans-species missense SNP in Amhr2 is associated with sex determination in the tiger pufferfish, Takifugu rubripes (fugu). PLoS Genet 8:e1002798
Kawagoshi T, Nishida C, Matsuda Y (2012) The origin and differentiation process of X and Y chromosomes of the black marsh turtle (Siebenrockiella crassicollis, Geoemydidae, Testudines). Chromosome Res 20:95–110
Kawagoshi T, Uno Y, Nishida C, Matsuda Y (2014) The Staurotypus turtles and Aves share the same origin of sex chromosomes but evolved different types of heterogametic sex determination. PLoS One 9:e105315
Kawai A, Nishida-Umehara C, Ishijima J, Tsuda Y, Ota H, Matsuda Y (2007) Different origins of bird and reptile sex chromosomes inferred from comparative mapping of chicken Z-linked genes. Cytogenet Genome Res 117:92–102
Kejnovský E, Michalovova M, Steflova P, Kejnovska I, Manzano S, Hobza R, Kubat Z, Kovarik J, Jamilena M, Vyskot B (2013) Expansion of microsatellites on evolutionary young Y chromosome. PLoS One 8:e45519
King M (1977) The evolution of sex chromosomes in lizards. In: Calaly J, Tyndale-Biscoe H (eds) Evolution and reproduction. Australian Academy of Science, Canberra, pp 55–60
King M (1990) Chromosomal and immunogenetic data: a new perspective on the origin of Australia’s reptiles. In: Olmo E (ed) Cytogenetics of amphibians and reptiles. Birkhäuser, Basel, pp 153–180
Kobayashi Y, Nagahama Y, Nakamura M (2013) Diversity and plasticity of sex determination and differentiation in fishes. Sex Dev 7:115–125
Koehler MR, Dehm D, Guttenbach M, Nanda I, Haaf T, Molina WF, Galetti PM Jr, Schmid M (1997) Cytogenetics of the genus Leporinus (Pisces, Anostomidae). 1. Karyotype analysis, heterochromatin distribution and sex chromosomes. Chromosome Res 5:12–22
Kondo M, Nagao E, Mitani H, Shima A (2001) Differences in recombination frequencies during female and male meioses of the sex chromosomes of the medaka, Oryzias latipes. Genet Res 78:23–30
Kondo M, Nanda I, Hornung U, Schmid M, Schartl M (2004) Evolutionary origin of the medaka Y chromosome. Curr Biol 14:1664–1669
Kondo M, Hornung U, Nanda I, Imai S, Sasaki T, Shimizu A, Asakawa S, Hori H, Schmid M, Shimizu N, Schartl M (2006) Genomic organization of the sex-determining and adjacent regions of the sex chromosomes of medaka. Genome Res 16:815–826
Konerat J, Bueno V, Margarido V, de Brito Portela-Castro A, Martins-Santos I (2015) Diversity sex chromosome systems in Ancistrini (Loricariidae, Hypostominae): ZZ/ZW in Ancistrus taunayi Miranda Ribeiro, 1918. Cytogen Genome Res: in press.
Lahn BT, Page DC (1999) Four evolutionary strata on the human X chromosome. Science 286:964–967
Liu JD, Yi MS, Zhao G, Zhou F, Wang DQ, Yu QX (2002) Sex chromosomes in the spiny eel (Mastacembelus aculeatus) revealed by mitotic and meiotic analysis. Cytogenet Genome Res 98:291–297
Livernois AM, Graves JA, Waters PD (2012) The origin and evolution of vertebrate sex chromosomes and dosage compensation. Heredity 108:50–58
Lubieniecki KP, Lin S, Cabana EI, Li J, Lai YY, Davidson WS (2015) Genomic instability of the sex-determining locus in Atlantic salmon (Salmo salar). G3 (Bethesda) 5:2513–2522
Mahony MJ (1991) Heteromorphic sex-chromosomes in the Australian frog Crinia bilingua (Anura, Myobatrachidae). Genome 34:334–337
Mank JE, Avise JC (2009) Evolutionary diversity and turn-over of sex determination in teleost fishes. Sex Dev 3:60–67
Marshall Graves JA, Shetty S (2001) Sex from W to Z: evolution of vertebrate sex chromosomes and sex determining genes. J Exp Zool 290:449–462
Matsubara K, Tarui H, Toriba M, Yamada K, Nishida-Umehara C, Agata K, Matsuda Y (2006) Evidence for different origin of sex chromosomes in snakes, birds, and mammals and step-wise differentiation of snake sex chromosomes. Proc Natl Acad Sci U S A 103:18190–18195
Matsubara K, Knopp T, Sarre SD, Georges A, Ezaz T (2013) Karyotypic analysis and FISH mapping of microsatellite motifs reveal highly differentiated XX/XY sex chromosomes in the pink-tailed worm-lizard (Aprasia parapulchella, Pygopodidae, Squamata). Mol Cytogenet 6:60
Matsubara K, Sarre SD, Georges A, Matsuda Y, Marshall Graves JA, Ezaz T (2014) Highly differentiated ZW sex microchromosomes in the Australian Varanus species evolved through rapid amplification of repetitive sequences. PLoS One 9:e95226
Matsubara K, O’Meally D, Azad B, Georges A, Sarre SD, Graves JA, Matsuda Y, Ezaz T (2015) Amplification of microsatellite repeat motifs is associated with the evolutionary differentiation and heterochromatinization of sex chromosomes in Sauropsida. Chromosoma: [Epub ahead of print]
Matsuda M, Nagahama Y, Shinomiya A, Sato T, Matsuda C, Kobayashi T, Morrey CE, Shibata N, Asakawa S, Shimizu N, Hori H, Hamaguchi S, Sakaizumi M (2002) DMY is a Y-specific DM-domain gene required for male development in the medaka fish. Nature 417:559–563
Matthey R (1953) La formule chromosomique et le problème de la détermination sexuelle chez Ellobius lutescens Thomas (Rodentia-Muridae-Microtinae). Arch Klaus-Stift Vererb-Forsch 28:65–73
Mengden GA, Stock AD (1980) Chromosomal evolution in Serpentes—a comparison of chromosome-G and chromosome-C banding-patterns of some colubrid and boid genera. Chromosoma 79:53–64
Mohanty MK, Bhunya SP (1990) Karyological studies in 4 species of Ardeid birds (Ardeidae, Ciconiiformes). Genetica 81:211–214
Moritz C (1984) The evolution of a highly variable sex chromosome in Gehyra purpurascens (Gekkonidae). Chromosoma 90:111–119
Murata C, Yamada F, Kawauchi N, Matsuda Y, Kuroiwa A (2012) The Y chromosome of the Okinawa spiny rat, Tokudaia muenninki, was rescued through fusion with an autosome. Chromosome Res 20:111–125
Nagamachi CY, Pieczarka JC (1988) Chromosome studies of Saguinus midas niger (Callithricidae, Primates) from Tucurui, Para, Brazil: comparison with the karyotype of Callithrix jacchus. Am J Primatology 14:277–284
Nanda I, Feichtinger W, Schmid M, Schröder JH, Zischler H, Epplen JT (1990) Simple repetitive sequences are associated with differentiation of the sex-chromosomes in the guppy fish. J Mol Evol 30:456–462
Nanda I, Shan Z, Schartl M, Burt DW, Koehler M, Nothwang H, Grutzner F, Paton IR, Windsor D, Dunn I, Engel W, Staeheli P, Mizuno S, Haaf T, Schmid M (1999) 300 million years of conserved synteny between chicken Z and human chromosome 9. Nat Genet 21:258–259
Nanda I, Volff JN, Weis S, Körting C, Froschauer A, Schmid M, Schartl M (2000) Amplification of a long terminal repeat-like element on the Y chromosome of the platyfish, Xiphophorus maculatus. Chromosoma 109:173–180
Nanda I, Kondo M, Hornung U, Asakawa S, Winkler C, Shimizu A, Shan Z, Haaf T, Shimizu N, Shima A, Schmid M, Schartl M (2002) A duplicated copy of DMRT1 in the sex-determining region of the Y chromosome of the medaka, Oryzias latipes. Proc Natl Acad Sci U S A 99:11778–11783
Nanda I, Schlegelmilch K, Haaf T, Schartl M, Schmid M (2008) Synteny conservation of the Z chromosome in 14 avian species (11 families) supports a role for Z dosage in avian sex determination. Cytogenet Genome Res 122:150–156
Nanda I, Schories S, Tripathi N, Dreyer C, Haaf T, Schmid M, Schartl M (2014) Sex chromosome polymorphism in guppies. Chromosoma 123:373–383
Nascimento J, Quindere YRSD, Recco-Pimentel SM, Lima JRF, Lourenco LB (2010) Heteromorphic Z and W sex chromosomes in Physalaemus ephippifer (Steindachner, 1864) (Anura, Leiuperidae). Genetica 138:1127–1132
Nieto LM, Kretschmer R, Ledesma MA, Garnero ADV, Gunsk iRJ (2012) Karyotype morphology suggests that the Nyctibius griseus (Gmelin, 1789) carries an ancestral ZW-chromosome pair to the order Caprimulgiformes (Aves). Comp Cytogenetics 6:379–387
Nishida-Umehara C, Tsuda Y, Ishijima J, Ando J, Fujiwara A, Matsuda Y, Griffin DK (2007) The molecular basis of chromosome orthologies and sex chromosomal differentiation in palaeognathous birds. Chromosome Res 15:721–734
Noleto RB, Amorim AP, Vicari MR, Artoni RF, Cestari MM (2009) An unusual ZZ/ZW sex chromosome system in Characidium fishes (Crenuchidae, Characiformes) with the presence of rDNA sites. J Fish Biol 75:448–453
Ohno S (1967) Sex chromosomes and sex-linked genes. Springer, Berlin
Oliveira C, Toledo LFD (2006) Evidence of an XX/XY sex chromosome system in the fish Dormitator maculatus (Teleostei, Eleotrididae). Genet Mol Biol 29:653–655
Oliveira RR, Feldberg E, Anjos MB, Zuanon J (2007) Karyotype characterization and ZZ/ZW sex chromosome heteromorphism in two species of the catfish genus Ancistrus Kner, 1854 (Siluriformes: Loricariidae) from the Amazon basin. Neotrop Ichthyol 5:301–306
Olmo E, Odierna G, Capriglione T (1987) Evolution of sex-chromosomes in lacertid lizards. Chromosoma 96:33–38
O’Meally D, Patel HR, Stiglec R, Sarre SD, Georges A, Marshall Graves JA, Ezaz T (2010) Non-homologous sex chromosomes of birds and snakes share repetitive sequences. Chromosome Res 18:787–800
Pala I, Naurin S, Stervander M, Hasselquist D, Bensch S, Hansson B (2012) Evidence of a neo-sex chromosome in birds. Heredity 108:264–272
Parise-Maltempi PP, da Silva EL, Rens W, Dearden F, O’Brien PC, Trifonov V, Ferguson-Smith MA (2013) Comparative analysis of sex chromosomes in Leporinus species (Teleostei, Characiformes) using chromosome painting. BMC Genet 14:60
Peichel CL, Ross JA, Matson CK, Dickson M, Grimwood J, Schmutz J, Myers RM, Mori S, Schluter D, Kingsley DM (2004) The master sex-determination locus in threespine sticklebacks is on a nascent Y chromosome. Curr Biol 14:1416–1424
Perrin N (2009) Sex reversal: a fountain of youth for sex chromosomes? Evolution 63:3043–3049
Phillips RB (2013) Evolution of the sex chromosomes in salmonid fishes. Cytogenet Genome Res 141:177–185
Pokorná M, Kratochvíl L, Kejnovský E (2011) Microsatellite distribution on sex chromosomes at different stages of heteromorphism and heterochromatinization in two lizard species (Squamata: Eublepharidae: Coleonyx elegans and Lacertidae: Eremias velox). BMC Genet 12:90
Pokorná M, Rens W, Rovatsos M, Kratochvíl L (2014) A ZZ/ZW sex chromosome system in the thick-tailed Gecko (Underwoodisaurus milii; Squamata: Gekkota: Carphodactylidae), a member of the ancient gecko lineage. Cytogenet Genome Res 142:190–196
Raman R, Nanda I (1982) Identification and patterns of synapsis of the autosomally translocated Y-chromosome of the Indian mongoose, Herpestes auropunctatus (Hodgson). Chromosoma 87:477–489
Raman R, Nanda I (1986) Mammalian sex chromosomes. I. Cytological changes in the chiasmatic sex chromosomes of the male musk shrew, Suncus murinus. Chromosoma 93:367–374
Rice WR (1996) Evolution of the Y sex chromosome in animals. BioScience 46:331–343
Roco AS, Olmstead AW, Degitz SJ, Amano T, Zimmerman LB, Bullejos M (2015) Coexistence of Y, W, and Z sex chromosomes in Xenopus tropicalis. Proc Natl Acad Sci U S A 112:E4752–4761
Ross JA, Urton JR, Boland J, Shapiro MD, Peichel CL (2009) Turnover of sex chromosomes in the stickleback fishes (Gasterosteidae). Plos Genetics 5:e1000391
Rossi LF, Luaces JP, Alonso FM, Merani MS (2014) Karyotype and chromosome variability in the armadillo Chaetophractus villosus in Argentina. Cytogenet Genome Res 142:101–106
Rovatsos M, Pokorna M, Altmanova M, Kratochvil L (2014) Cretaceous park of sex determination: sex chromosomes are conserved across iguanas. Biol Lett 10:20131093
Rovatsos M, Johnson Pokorna M, Altmanova M, Kratochvil L (2015) Female heterogamety in Madagascar chameleons (Squamata: Chamaeleonidae: Furcifer): differentiation of sex and neo-sex chromosomes. Sci Rep 5:13196
Rutkowska J, Lagisz M, Nakagawa S (2012) The long and the short of avian W chromosomes: no evidence for gradual W shortening. Biol Lett 8:636–638
Sato T, Endo T, Yamahira K, Hamaguchi S, Sakaizumi M (2005) Induction of female-to-male sex reversal by high temperature treatment in Medaka, Oryzias latipes. Zoolog Sci 22:985–988
Schempp W, Schmid M (1981) Chromosome-banding in Amphibia. 6. Brdu-replication patterns in Anura and demonstration of Xx-Xy Sex-chromosomes in Rana esculenta. Chromosoma 83:697–710
Schmid M (1980) Chromosome-banding in Amphibia. V. Highly differentiated ZW/ZZ sex chromosomes and exceptional genome size in Pyxicephalus adspersus (Anura, Ranidae). Chromosoma 80:69–96
Schmid M, Olert J, Klett C (1979) Chromosome banding in Amphibia. III sex chromosomes in Triturus. Chromosoma 71:29–55
Schmid M, Haaf T, Geile B, Sims S (1983) Chromosome banding in Amphibia. VIII. An unusual XY/XX-sex chromosome system in Gastrotheca riobambae (Anura, Hylidae). Chromosoma 88:69–82
Schmid M, Ohta S, Steinlein C, Guttenbach M (1993) Chromosome banding in Amphibia. XIX. Primitive ZW/ZZ sex chromosomes in Buergeria buergeri (Anura, Rhacophoridae). Cytogenet Cell Genet 62:238–246
Schmid M, Feichtinger W, Steinlein C, Nanda I, Mais C, Haaf T, Visbal Garcia R, Fernandez Badillo A (2002a) Chromosome banding in Amphibia. XXII. Atypical y chromosomes in Gastrotheca walkeri and Gastrotheca ovifera (Anura, Hylidae). Cytogenet Genome Res 96:228–238
Schmid M, Feichtinger W, Steinlein C, Rupprecht A, Haaf T, Kaiser H (2002b) Chromosome banding in Amphibia. XXIII. Giant W sex chromosomes and extremely small genomes in Eleutherodactylus euphronides and Eleutherodactylus shrevei (Anura, Leptodactylidae). Cytogenet Genome Res 97:81–94
Schmid M, Steinlein C, Bogart JP, Feichtinger W, Leon P, La Marca E, Diaz LM, Sanz A, Chen SH, Hedges SB (2010) The chromosomes of terraranan frogs. Insights into vertebrate cytogenetics. Cytogenet Genome Res 130–131:1–568
Schmid M, Steinlein C, Haaf T, Mijares-Urrutia A (2014) Nascent ZW sex chromosomes in Thecadactylus rapicauda (Reptilia, Squamata, Phyllodactylidae). Cytogenet Genome Res 143:259–267
Sessions SK (1980) Evidence for a highly differentiated sex-chromosome heteromorphism in the salamander Necturus maculosus (Rafinesque). Chromosoma 77:157–168
Shinomiya A, Otake H, Hamaguchi S, Sakaizumi M (2010) Inherited XX sex reversal originating from wild medaka populations. Heredity 105:443–448
Singh L (1972) Evolution of karyotypes in snakes. Chromosoma 38:185
Singh L, Purdom IF, Jones KW (1976) Satellite DNA and evolution of sex chromosomes. Chromosoma 59:43–62
Singh L, Purdom IF, Jones KW (1980) Sex-chromosome associated satellite DNA—evolution and conservation. Chromosoma 79:137–157
Soh YQ, Alfoldi J, Pyntikova T, Brown LG, Graves T, Minx PJ, Fulton RS, Kremitzki C, Koutseva N, Mueller JL, Rozen S, Hughes JF, Owens E, Womack JE, Murphy WJ, Cao Q, de Jong P, Warren WC, Wilson RK, Skaletsky H, Page DC (2014) Sequencing the mouse Y chromosome reveals convergent gene acquisition and amplification on both sex chromosomes. Cell 159:800–813
Solano E, Taylor PJ, Rautenbach A, Ropiquet A, Castiglia R (2014) Cryptic speciation and chromosomal repatterning in the South African climbing mice Dendromus (Rodentia, Nesomyidae). PLoS One 9:e88799
Splendore de Borba R, Lourenco da Silva E, Parise-Maltempi PP (2013) Chromosome mapping of retrotransposable elements and in Spix, 1829 species (Characiformes: Anostomidae) and its relationships among heterochromatic segments and W sex chromosome. Mob Genet Elements 3:e27460
Srikulnath K, Nishida C, Matsubara K, Uno Y, Thongpan A, Suputtitada S, Apisitwanich S, Matsuda Y (2009) Karyotypic evolution in squamate reptiles: comparative gene mapping revealed highly conserved linkage homology between the butterfly lizard (Leiolepis reevesii rubritaeniata, Agamidae, Lacertilia) and the Japanese four-striped rat snake (Elaphe quadrivirgata, Colubridae, Serpentes). Chromosome Res 17:975–986
Stöck M, Horn A, Grossen C, Lindtke D, Sermier R, Betto-Colliard C, Dufresnes C, Bonjour E, Dumas Z, Luquet E, Maddalena T, Sousa HC, Martinez-Solano I, Perrin N (2011) Ever-young sex chromosomes in European tree frogs. PLoS Biol 9:e1001062
Suarez-Villota EY, Pansonato-Alves JC, Foresti F, Gallardo MH (2014) Homomorphic sex chromosomes and the intriguing Y chromosome of Ctenomys rodent species (Rodentia, Ctenomyidae). Cytogenet Genome Res 143:232–240
Swarca AC, Fenocchio AS, Cestari MM, Bertollo LAC, Dias AL (2006) Heteromorphic sex chromosome system with an exceptionally large Y chromosome in a catfish Steindachneridion sp (Pimelodidae). Cytogenet Genome Res 112:325–328
Takeda H (2008) Draft genome of the medaka fish: a comprehensive resource for medaka developmental genetics and vertebrate evolutionary biology. Dev Growth Differ 50(Suppl 1):S157–166
Takehana Y, Naruse K, Hamaguchi S, Sakaizumi M (2007) Evolution of ZZ/ZW and XX/XY sex-determination systems in the closely related medaka species, Oryzias hubbsi and O. dancena. Chromosoma 116:463–470
Terencio ML, Schneider CH, Gross MC, Nogaroto V, de Almeida MC, Artoni RF, Vicari MR, Feldberg E (2012) Repetitive sequences associated with differentiation of W chromosome in Semaprochilodus taeniurus. Genetica 140:505–512
Terencio ML, Schneider CH, Gross MC, Vicari MR, Farias IP, Passos KB, Feldberg E (2013) Evolutionary dynamics of repetitive DNA in Semaprochilodus (Characiformes, Prochilodontidae): a fish model for sex chromosome differentiation. Sex Dev 7:325–333
Tsuda Y, Nishida-Umehara C, Ishijima J, Yamada K, Matsuda Y (2007) Comparison of the Z and W sex chromosomal architectures in elegant crested tinamou (Eudromia elegans) and ostrich (Struthio camelus) and the process of sex chromosome differentiation in palaeognathous birds. Chromosoma 116:159–173
Uno Y, Nishida C, Oshima Y, Yokoyama S, Miura I, Matsuda Y, Nakamura M (2008) Comparative chromosome mapping of sex-linked genes and identification of sex chromosomal rearrangements in the Japanese wrinkled frog (Rana rugosa, Ranidae) with ZW and XY sex chromosome systems. Chromosome Res 16:637–647
Venere PC, Souza IL, Martins C, Oliveira C (2008) Occurrence of ZZ/ZW sex chromosomes in Thoracocharax stellatus fish (Characiformes, Gasteropelecidae) from the Araguaia River, South America. Genetica 133:109–112
Vicente VE, Bertollo LA, Valentini SR, Filho OM (2003) Origin and differentiation of a sex chromosome system in Parodon hilarii (Pisces, Parodontidae). Satellite DNA, G- and C-banding. Genetica 119:115–120
Vicoso B, Emerson JJ, Zektser Y, Mahajan S, Bachtrog D (2013) Comparative sex chromosome genomics in snakes: differentiation, evolutionary strata, and lack of global dosage compensation. PLoS Biol 11:e1001643
Vistorin G, Gamperl R, Rosenkranz W (1977) Studies on sex chromosomes of four hamster species: Cricetus cricetus, Cricetulus griseus, Mesocricetus auratus, and Phodopus sungorus. Cytogenet Cell Genet 18:24–32
Volff JN, Körting C, Froschauer A, Zhou Q, Wilde B, Schultheis C, Selz Y, Sweeney K, Duschl J, Wichert K, Altschmied J, Schartl M (2003) The Xmrk oncogene can escape nonfunctionalization in a highly unstable subtelomeric region of the genome of the fish Xiphophorus. Genomics 82:470–479
Volff JN, Nanda I, Schmid M, Schartl M (2007) Governing sex determination in fish: regulatory putsches and ephemeral dictators. Sex Dev 1:85–99
Wilson CA, High SK, McCluskey BM, Amores A, Yan YL, Titus TA, Anderson JL, Batzel P, Carvan MJ 3rd, Schartl M, Postlethwait JH (2014) Wild sex in zebrafish: loss of the natural sex determinant in domesticated strains. Genetics 198:1291–1308
Woram RA, Gharbi K, Sakamoto T, Hoyheim B, Holm LE, Naish K, McGowan C, Ferguson MM, Phillips RB, Stein J, Guyomard R, Cairney M, Taggart JB, Powell R, Davidson W, Danzmann RG (2003) Comparative genome analysis of the primary sex-determining locus in salmonid fishes. Genome Res 13:272–280
Xiu-Guang M, Jin-Huan W, Wei-Ting S, Ying-Xiang W, Feng-Tang Y, Wen-Hui N (2010) Karyotypic evolution in family Hipposideridae (Chiroptera, Mammalia) revealed by comparative chromosome painting, G- and C-banding. Zool Res 31:453–460
Yamada K, Nishida-Umehara C, Matsuda Y (2004) A new family of satellite DNA sequences as a major component of centromeric heterochromatin in owls (Strigiformes). Chromosoma 112:277–287
Yano A, Guyomard R, Nicol B, Jouanno E, Quillet E, Klopp C, Cabau C, Bouchez O, Fostier A, Guiguen Y (2012) An immune-related gene evolved into the master sex-determining gene in rainbow trout, Oncorhynchus mykiss. Curr Biol 22:1423–1428
Yano A, Nicol B, Jouanno E, Quillet E, Fostier A, Guyomard R, Guiguen Y (2013) The sexually dimorphic on the Y-chromosome gene (sdY) is a conserved male-specific Y-chromosome sequence in many salmonids. Evol Appl 6:486–496
Yoshimoto S, Okada E, Umemoto H, Tamura K, Uno Y, Nishida-Umehara C, Matsuda Y, Takamatsu N, Shiba T, Ito M (2008) A W-linked DM-domain gene, DM-W, participates in primary ovary development in Xenopus laevis. Proc Natl Acad Sci U S A 105:2469–2474
Zhou Q, Braasch I, Froschauer A, Bohne A, Schultheis C, Schartl M, Volff JN (2010) A novel marker for the platyfish (Xiphophorus maculatus) W chromosome is derived from a Polinton transposon. J Genet Genomics 37:181–188
Zhou Q, Zhang J, Bachtrog D, An N, Huang Q, Jarvis ED, Gilbert MT, Zhang G (2014) Complex evolutionary trajectories of sex chromosomes across bird taxa. Science 346:1246338
Acknowledgments
Information collected on sex chromosomes of many weird species through numerous cytogenetic colleagues from many countries is highly appreciated. The list of examples for sex chromosomes discussed in this review that deviate from the classical view is certainly not complete. We apologize to all our colleagues whose work have escaped our attention and have not been mentioned appropriately. We thank Monika Niklaus-Ruiz for the help in the preparation of the manuscript. The work of the authors on sex chromosomes has been supported by the DFG through grants SCHA 408/12-1 and SCHA 408/10-1.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics and animal welfare statement
The manuscript or parts of it have not been submitted elsewhere. No data or images have been manipulated to support our conclusions. All presented data are our own; work of other authors is cited with correct references. All authors have given their consent to submit the final version. All co-authors have contributed sufficiently to the manuscript and therefore share collective responsibility and accountability for the results.
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no competing interests.
Rights and permissions
About this article
Cite this article
Schartl, M., Schmid, M. & Nanda, I. Dynamics of vertebrate sex chromosome evolution: from equal size to giants and dwarfs. Chromosoma 125, 553–571 (2016). https://doi.org/10.1007/s00412-015-0569-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-015-0569-y