Abstract
KNL1 is an evolutionarily conserved kinetochore-associated protein essential for accurate chromosome segregation in eukaryotic cells. This large scaffold protein, predicted to be almost entirely unstructured, is involved in diverse mitotic processes including kinetochore assembly, chromosome congression, and mitotic checkpoint signaling. How this kinetochore “hub” coordinates protein–protein interactions spatially and temporally during mitosis to orchestrate these processes is an area of active investigation. Here we summarize the current understanding of KNL1 and discuss possible mechanisms by which this protein actively contributes to multiple aspects of mitotic progression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
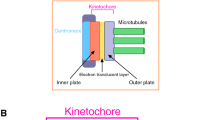
During mitosis, replicated chromosomes attach to spindle microtubules (MTs) in order to congress to the spindle equator and correctly segregate to opposite spindle poles during anaphase. Attachment between replicated chromosomes and MTs is mediated by the kinetochore, a complex network of proteins assembled on centromeric DNA of each mitotic sister chromatid. The kinetochore also orchestrates a sophisticated signaling network called the spindle assembly checkpoint (SAC), a fail-safe mechanism that prevents mitotic exit until all chromosomes are attached to spindle MTs. Kinetochores are comprised of over 100 proteins, many of which are evolutionarily conserved and exhibit high interdependency and functionally diverse roles (Hori and Fukagawa 2012; Przewloka and Glover 2009; Santaguida and Musacchio 2009). Despite this complexity, we are beginning to understand the molecular basis for kinetochore assembly (Hori and Fukagawa 2012; Santaguida and Musacchio 2009), chromosome congression (Kops et al. 2010; McIntosh et al. 2012; Walczak and Heald 2008), and SAC signaling (Jia et al. 2013; Lara-Gonzalez et al. 2012; Musacchio 2012). The scaffold protein KNL1, a relatively recently described kinetochore protein, plays an important role in these three critical mitotic processes; however, its specific contributions to each are just beginning to be appreciated. Here we provide an overview of the current state of understanding of KNL1, a protein that is emerging as a key integrator of multiple kinetochore activities.
KNL1 history
The gene encoding the human protein KNL1 (also known as D40, AF15q14, CASC5 for Cancer Susceptibility Candidate 5, and Blinkin for Bub1 linking kinetochore protein) was originally identified as an interacting partner for the transcription factor GCF (GC factor) in a yeast two-hybrid assay (Wei et al. 1999). Later, a small fragment of the KNL1 gene was described as a fusion with the oncogene MLL (mixed lineage leukemia) in acute leukemia (Hayette et al. 2000). Early studies showed that human KNL1 mRNA is expressed in thymus, testis, and bone marrow in adult tissues, in a variety of human cancer cell lines and in primary tumors, and ubiquitously expressed in fetal tissues (Hayette et al. 2000; Takimoto et al. 2002). An RNAi-based functional genomic screen in Caenorhabditis elegans identified C02F5.1 (later named KNL1) as a gene encoding a previously undescribed protein required for the fidelity of mitotic cell division (Gönczy et al. 2000). Soon after, a detailed functional study in C. elegans demonstrated that RNAi-mediated depletion of C02F5.1 resulted in failure of kinetochores to recruit multiple proteins, premature spindle pole separation, and chromosome segregation failure (Desai et al. 2003). Based on this phenotype, which was previously observed upon depletion of the inner kinetochore proteins CENP-C and CENP-A (Oegema et al. 2001), C02F5.1 was categorized as “kinetochore null” and renamed KNL-1 (Desai et al. 2003). In parallel to its discovery as a kinetochore protein in C. elegans, KNL1/Spc105 was also characterized as a kinetochore component in budding yeast (Nekrasov et al. 2003).
In human cells, KNL1 was first recognized as a kinetochore protein in 2004, and its conserved role in kinetochore assembly was later demonstrated (Cheeseman et al. 2004, 2008; Kiyomitsu et al. 2007, 2011). Since its discovery, KNL1 has been recognized as a critical platform for kinetochore protein recruitment and as a linker between centromeric DNA and the plus ends of spindle MTs not only in human cells but also in yeast, worms, chicken, and flies (Cheeseman et al. 2008; Kerres et al. 2004; Nekrasov et al. 2003; Przewloka et al. 2007; Schittenhelm et al. 2009).
KNL1: the unknown structure
Despite being evolutionarily conserved, KNL1 displays low overall sequence similarity between species, and according to folding prediction algorithms, it is mostly intrinsically disordered (FoldIndex, Phyre). Human KNL1 for example, with over 2,300 amino acids, has only a single region predicted to fold into an ordered structure. This region, which resides at the far C-terminus of the protein, is composed of a globular domain containing tandem RWD motifs (A. Musacchio, personal communication) and a coiled coil domain (Petrovic et al. 2010). The RWD motifs mediate direct interaction with the Mis12 complex, while the coiled coil domain mediates interaction with Zwint-1. Both the Mis12 complex and Zwint-1 are important for proper kinetochore assembly (Fig. 1). Within the remaining “unstructured” region of the protein, several motifs have been identified. The RVSF (RVXF) and SILK ([SG]ILK) motifs, both of which reside at the far N-terminus of KNL1, mediate binding of the phosphatase PP1 (Liu et al. 2010). Two KI motifs, KI1 and KI2 (KI[DN]XXXF[LI]XXLK), also localized in the N-terminus of KNL1, interact with the essential SAC proteins Bub1 and BubR1, respectively (Bolanos-Garcia et al. 2011; Kiyomitsu et al. 2011; Krenn et al. 2012). More recently, MELT motifs (M[ED][ILVM][ST]) in the N-terminal and central regions of KNL1 have been identified and demonstrated as mediators for recruitment of Bub1 and Bub3, another essential SAC component (London et al. 2012; Primorac et al. 2013; Shepperd et al. 2012; Yamagishi et al. 2012) (Fig. 1). A search for KNL1 orthologs across eukaryotes recently identified KNL1 in three out of the six eukaryotic supergroups (Vleugel et al. 2012). Although KNL1 primary sequence similarity across species is poor and the predicted protein size is quite variable, the C-terminal region, SILK, RVSF, and MELT motifs are well conserved. The number of MELT motifs across species, however, is widely variable (Vleugel et al. 2012) (Fig. 2).
Domain architecture of human KNL1. Key regions involved in KNL1 function are indicated schematically (top), and the corresponding amino acid sequences for each region in human KNL1 are listed (bottom). Aurora B kinase phosphorylates the N-terminus of KNL1 to inhibit association with PP1 (Liu et al. 2010). Mps1 phosphorylates the KNL1 MELT repeats to promote association with Bub1 and Bub3 (London et al. 2012; Primorac et al. 2013; Shepperd et al. 2012; Yamagishi et al. 2012). Amino acid residues 1–68 indicate the MT binding domain (Welburn et al. 2010), and underlined is a basic patch of residues (RRRH), whose analogous residues in C. elegans have been implicated in direct MT interaction (Espeut et al. 2012)
Evolutionary conservation of KNL1 domain architecture. Key KNL1 functional regions are shown for several organisms. The respective number of amino acids in KNL1 for the various organisms is shown (right). Two values in this column indicate multiple isoforms (H. sapiens). The green region in D. melanogaster indicates the presence of repetitive motifs that differ from the MELT consensus motif observed in other organisms (Schittenhelm et al. 2009)
Full-length C. elegans KNL1 and fragments of yeast and human KNL1 have been successfully reconstituted in vitro (Bolanos-Garcia et al. 2011; Cheeseman et al. 2006; Espeut et al. 2012; Krenn et al. 2012; Shepperd et al. 2012; Yamagishi et al. 2012). However, there is a dearth of structural information for KNL1 as only small fragments of the human N-terminal region containing the KI motifs (Bolanos-Garcia et al. 2011; Krenn et al. 2012) and small fragments of yeast KNL1 (Spc105) containing MELT motifs have been crystallized (Primorac et al. 2013). In these atomic structures, only a handful of residues (~15) showed significant electron density, presumably due to the disordered nature of KNL1. The residues identified with confidence in the structures of both KNL1 KI motifs fold into short alpha-helices when bound to Bub1 or BubR1 (Bolanos-Garcia et al. 2011; Krenn et al. 2012). Circular dichroism experiments indicate that these regions of KNL1 by themselves lack secondary structure (Bolanos-Garcia et al. 2011); thus, it appears that KNL1 transitions from a disordered configuration to an ordered one upon binding to other proteins.
The importance of intrinsically disordered proteins in a diverse number of cellular processes is only beginning to be recognized (Dyson and Wright 2005; Tompa 2012). Hypotheses regarding the molecular and evolutionary advantage of this class of proteins have emerged over the last decade. For instance, it is proposed that largely disordered regions provide greater surface area for interaction with multiple binding partners and more local conformational flexibility, allowing fine control of their binding affinity (Dyson and Wright 2005). Intrinsic lack of structure has also been implicated in force sensing and stretching during mechano-transduction (Pan et al. 2012). Furthermore, disordered proteins may be less sensitive to environmental perturbations, allowing them to provide stability to complex regulatory networks (Gunasekaran et al. 2003; Huang and Liu 2010). How KNL1 folds and binds to its kinetochore partners in the cell, how it stabilizes multiple kinetochore protein networks, and what signaling events result from these interactions are compelling questions that remain to be addressed.
KNL1 and the KMN network
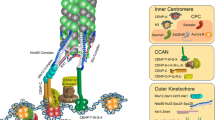
KNL1 is a component of the “KMN network,” a complex of proteins that makes up the primary MT binding interface at the kinetochore. The KMN network is comprised of the KNL1 protein, the Mis12 complex (Mis12, Dsn1, Nsl1, and Nnf1), and the Ndc80 complex (Hec1, Nuf2, Spc24, and Spc25) (DeLuca and Musacchio 2012; Tooley and Stukenberg 2011; Varma and Salmon 2012). Zwint1, a protein involved in checkpoint signaling, is sometimes considered a member of the KMN network since it binds directly to KNL1 and co-purifies with the rest of the KMN network (Lin et al. 2006; Pagliuca et al. 2009; Wang et al. 2004). The high binding affinity measured in vitro between several KMN components (K d~nM range) (Petrovic et al. 2010), their apparent cooperative binding to MTs (Cheeseman et al. 2006), and their substantial co-dependency for kinetochore recruitment in vivo (Varma and Salmon 2012) (Fig. 3) suggest that these proteins function together during mitosis. The most recognized function of the KMN network at kinetochores is linkage of the inner kinetochore (chromatin proximal region) to plus ends of spindle MTs. Association with the inner kinetochore region requires binding between the Mis12 complex and the constitutive centromere associated network (CCAN), a large group of centromere-associated proteins responsible for establishing and maintaining centromere integrity (McAinsh and Meraldi 2011). (The CCAN is composed of 16 proteins: CENP-B, C, -H, -I, -K, -L, -M, -N, -O, -P, -R, -S, -T, -U, -W, and -X.) The KMN–MT interaction, on the other hand, is primarily mediated by Hec1, a component of the Ndc80 complex (Alushin et al. 2010; Cheeseman et al. 2006; Ciferri et al. 2008; DeLuca et al. 2006; Powers et al. 2009). KNL1 also directly binds MTs in vitro; however, a significant role for this MT binding activity in the formation of end-on, force-generating kinetochore–MT attachments has not been demonstrated (discussed in the following section). Although the KMN network is considered a functional unit (Cheeseman et al. 2006), it is important to note that depletion of individual KMN components results in different phenotypes. For instance, Mis12 and KNL1 depletion from human cells leads to a partial chromosome alignment phenotype (Kline et al. 2006; Cheeseman et al. 2008; Pagliuca et al. 2009), while depletion of any of the Ndc80 complex subunits leads to severe kinetochore–MT attachment defects and completely unaligned chromosomes (Tooley and Stukenberg 2011). To what extent the components of the KMN network rely on each other to carry out their functions, independent of their role in network connectivity, is an interesting question for further study.
Requirements for KNL1 kinetochore localization
KNL1 localizes to kinetochores from prophase to early telophase in human cells and has also been weakly observed near centromere regions in interphase nuclei (Cheeseman et al. 2008; Kiyomitsu et al. 2007). The KNL1 kinetochore localization domain resides in its C-terminus, the region that mediates direct binding to Zwint-1 and the Mis12 complex (Kiyomitsu et al. 2007, 2011; Petrovic et al. 2010) (Fig. 1). Specifically, amino acids 2106–2316 of human KNL1 directly bind to Nsl1 (a component of the four-protein Mis12 complex) (Petrovic et al. 2010). Although this region of KNL1 (2106–2316) does not directly interact with Zwint-1, a longer KNL1 fragment containing C-terminal amino acids 1904–2316 mediates Zwint-1 binding in vitro (Petrovic et al. 2010). A GFP-tagged KNL1 fragment of amino acids 1833–2316 localizes to kinetochores in human cells (Kiyomitsu et al. 2007, 2011) as does a KNL1 fragment containing the minimal Mis12 binding region (2106–2316) (Caldas et al. 2013), indicating that KNL1 localizes to kinetochores via binding the Mis12 complex. In support of this, depletion of the Mis12 complex component Dsn1 leads to a significant reduction in KNL1 kinetochore localization in human cells, chicken cells, and C. elegans (Cheeseman et al. 2004, 2008). It was recently shown that Zwint-1 depletion also results in a considerable reduction of kinetochore-localized KNL1 (~60 % reduction) (Varma et al. 2013). Therefore, it is possible that both Zwint-1 and the Mis12 complex contribute to kinetochore recruitment of KNL1 through direct binding with the KNL1 C-terminal region. It remains to be tested, however, if Mis12 localization requires Zwint-1, which would alternatively explain the dependency of KNL1 kinetochore recruitment on Zwint-1. Importantly, KNL1 depletion results in significant kinetochore delocalization of Zwint-1 and Mis12 complex components (Cheeseman et al. 2008; Varma et al. 2013), demonstrating a high level of co-dependency between these proteins.
Despite the close association between KMN network components and KNL1's dependency on the Mis12 complex for kinetochore localization, depletion of Ndc80 complex components in human cells does not prevent KNL1 kinetochore association (Cheeseman et al. 2008; Miller et al. 2008; Pagliuca et al. 2009). Additionally, a high level of co-dependency between the Mis12 complex and KNL1 kinetochore localization seems to be conserved through evolution, but co-dependency between the Ndc80 complex and KNL1 does not follow the same pattern. Consistent with this, biochemical analysis revealed that while the Ndc80 complex and KNL1 both directly interact with the Mis12 complex, KNL1 and the Ndc80 complex do not bind to each other (Cheeseman et al. 2006; Petrovic et al. 2010). It is important to note, however, that the Ndc80 subunit Hec1 is required for the recruitment of Zwint-1 to kinetochores (Lin et al. 2006), suggesting that the Ndc80 complex may indirectly contribute to KNL1 kinetochore association. Thus, while the Mis12 complex is an absolute requirement for KNL1 kinetochore localization, the contribution of the Ndc80 complex is likely less significant.
Role of KNL1 in kinetochore assembly
In C. elegans, depletion of KNL1 by RNAi results in loss of kinetochore-localized Hec1 (CeNdc80), Nuf2 (HIM10), Spc25 (KBP3), Bub1, CENP-F (HCP-1), and CLASP2 and reduced kinetochore localization of Dsn1 (KNL3) and Mis12 (Desai et al. 2003; Cheeseman et al. 2004) (Fig. 3). Of those tested, total protein levels were maintained in KNL1-depleted embryos, indicating that KNL1 is required for their recruitment and/or maintenance at kinetochores but not for their stability (Desai et al. 2003; Cheeseman et al. 2004). In addition, Zwilch (ZWL-1, a component of the RZZ complex which additionally contains ZW10 and Rod), Spindly (SPDL-1), and PP1 kinetochore localization is also dependent on KNL1 in C. elegans (Espeut et al. 2012; Gassmann et al. 2010).
Depletion of KNL1 from human cells results in a similar loss of kinetochore proteins as observed in C. elegans, with some variations (Fig. 3). Bub1, Zwint-1, and RZZ components are lost from kinetochores in KNL1-depleted cells (Cheeseman et al. 2008; Kiyomitsu et al. 2007; Varma et al. 2013). BubR1, Cenp-E, Cenp-F, Mad1, and Mad2 kinetochore localization relies on Bub1 (Johnson et al. 2004). Therefore, logic dictates that depletion of KNL1 correspondingly results in kinetochore delocalization of Bub1 downstream targets. Not surprisingly, BubR1, Cenp-F, and Mad2 kinetochore localization is decreased in KNL1-depleted cells (Cheeseman et al. 2008; Kiyomitsu et al. 2007; Pagliuca et al. 2009), and we observe a reduction of kinetochore-associated Cenp-E upon KNL1 depletion from human cells (G.C. and J.D., unpublished data). Presumably, although untested, kinetochore recruitment of the dynein–dynactin complex, which depends on the RZZ complex, is also perturbed after KNL1 depletion.
Kinetochore levels of the Mis12 complex component Dsn1 are reduced after KNL1 depletion from human cells (Cheeseman et al. 2008). The same study reported that depletion of KNL1 from DT40 chicken cells resulted in a 45 % reduction of Hec1 at kinetochores, but in human cells Hec1 localization was not affected (Cheeseman et al. 2008). In contrast, a recent study found that Hec1 levels decreased by ~40 % after KNL1 depletion from HeLa cells (Caldas et al. 2013). Despite the disparity, it is evident that the Ndc80 complex retains some ability to localize to kinetochores in KNL1-depleted human cells. This is not surprising as it has been demonstrated that the CCAN component CENP-T also contributes to kinetochore recruitment of the Ndc80 complex in human cells (Gascoigne et al. 2011). Such KNL1-independent Ndc80 complex kinetochore localization does not follow a pattern through evolution since, as in C. elegans, Hec1 relies on KNL1 for localization in Drosophila (Desai et al. 2003; Schittenhelm et al. 2009), but this dependency is not observed in fission yeast (Kerres et al. 2007).
Its predicted high level of structural disorder and its ability to bind multiple proteins suggest that KNL1 is a kinetochore “hub” or “scaffold” protein (Haynes et al. 2006). The functionality of scaffold proteins with unrecognized catalytic activity such as KNL1 is usually attributed to their most basic role as structural organizers of components involved in specific signaling pathways. Co-localizing and concentrating proteins to a defined region may enhance the specificity of a signal, providing a significant level of regulation. Thus, it is generally assumed that disturbance of such scaffolds results in disruption of molecular processes in an indirect manner. However, there is growing evidence that scaffold proteins may play a more complex role in signal transduction (Pan et al. 2012; Shaw and Filbert 2009). The characteristic flexibility in domain architecture of scaffold proteins may not only allow interaction with multiple proteins but may also directly affect their activities. For example, it has been suggested that binding to scaffold proteins could result in allosteric changes in signaling proteins that either enhance or inhibit their activity or, alternatively, could prevent the degradation of active proteins (Ferreon et al. 2013; Laine et al. 2008). Whether KNL1 acts as a catalytic scaffold to coordinate multiple activities at the kinetochore and functions beyond a platform for kinetochore assembly is an exciting possibility that has not yet been investigated.
Role of KNL1 in chromosome congression
In all organisms studied to date, depletion of KNL1 results in chromosome segregation defects. In C. elegans embryos, KNL1 depletion precludes metaphase chromosome congression and results in premature spindle pole separation, ultimately leading to random distribution of chromosomes into daughter cells (Cheeseman et al. 2004; Desai et al. 2003). In both budding and fission yeast, KNL1 temperature-sensitive mutant strains exhibit chromosome bi-orientation defects and spindle abnormalities (Kerres et al. 2007; Pagliuca et al. 2009). Drosophila cells lacking KNL1 are able to form metaphase plates but exhibit lagging chromosomes during anaphase (Schittenhelm et al. 2009). Although complete chromosome misalignment was observed in one study using HeLa cells (Kiyomitsu et al. 2007), later studies reported a partial alignment phenotype after KNL1 depletion (Caldas et al. 2013; Cheeseman et al. 2008; Pagliuca et al. 2009). Thus, with the exception of C. elegans, the consensus phenotype resulting from KNL1 depletion is a partial alignment phenotype, in which a population of chromosomes is able to congress to the spindle equator but a significant number of uncongressed chromosomes remain stranded near the spindle poles. The molecular mechanisms that facilitate complete alignment of chromosomes to the spindle equator are not well understood (Kops et al. 2010; McIntosh et al. 2012), however, perturbation of attachments between kinetochores and MTs impairs directed chromosome movement and metaphase plate formation (Santaguida and Musacchio 2009; Tooley and Stukenberg 2011). We discuss three hypotheses to explain how loss of KNL1 might perturb kinetochore-MT attachments, resulting in congression defects.
-
1.
Insubstantial kinetochore–MT attachments—Recent studies have shown that stable, end-on kinetochore–MT attachments are not explicitly required to congress chromosomes to the spindle equator and that either transient end-on attachments or lateral attachments can suffice (Cai et al. 2009; Magidson et al. 2011). However, there may be an additional requirement for increased kinetochore–MT attachment strength to congress chromosomes positioned substantially far away from the spindle equator upon nuclear envelope breakdown. As such, it is possible that KNL1-depleted cells harbor weak kinetochore–MT attachments, sufficient to congress chromosomes located between the two spindle poles, relatively near the spindle equator, but insufficient to congress polar chromosomes, resulting in a partial alignment phenotype. Kinetochores with reduced MT binding ability in KNL1-depleted cells could arise in multiple ways: First, KNL1 directly binds MTs through its N-terminus (Cheeseman et al. 2006; Espeut et al. 2012; Pagliuca et al. 2009; Welburn et al. 2010); therefore, loss of KNL1 MT binding activity after KNL1 depletion could directly reduce KMN MT binding affinity, preventing the formation of robust kinetochore–MT connections. Interestingly, in C. elegans, disruption of the KNL1 MT binding domain does not significantly affect the formation of kinetochore–MT attachments, chromosome congression, or chromosome segregation but instead impairs spindle checkpoint silencing (discussed in the following section) (Espeut et al. 2012). Based on this evidence, it is unlikely that defective chromosome congression observed in human cells results from lack of KNL1 MT binding activity, but this remains to be formally tested. A second explanation for weakened kinetochore–MT attachments is the reduction in Ndc80 complex proteins at kinetochores after KNL1 depletion. This is not likely the case since previous studies have demonstrated that human cells with >30 % of wild-type Hec1 levels are able to properly congress chromosomes to the spindle equator (DeLuca et al. 2002) and Ndc80 components are only moderately reduced after KNL1 depletion (Caldas et al. 2013; Cheeseman et al. 2008). A third possibility is that KNL1 depletion decreases the ability of the Ndc80 complex to generate load-bearing kinetochore–MT attachments by impairing its activity or conformation. In support of this, in vitro experiments have demonstrated that KNL1 works synergistically with the Mis12 and Ndc80 complexes to bind MTs, presumably by enhancing the MT binding affinity of the Ndc80 complex (Cheeseman et al. 2006).
-
2.
Defects in MT motor-driven congression—Kinetochore recruitment of the plus-end directed MT motor Cenp-E, implicated in chromosome congression, depends on KNL1 (Fig. 3). Cenp-E promotes kinetochore-mediated capture of MTs and drives congression of mono-oriented chromosomes to the spindle equator, likely on MT “tracks” present in nearby kinetochore fibers (Kapoor et al. 2006; Kim et al. 2008). Reminiscent of KNL1 depletion, depletion of Cenp-E results in a “cruciform” phenotype, in which chromosome congression is largely achieved, with a small population of chromosomes remaining clustered near the spindle poles (McEwen et al. 2001; Schaar et al. 1997; Wood et al. 1997). Therefore, it is possible that in KNL1 depleted cells, loss of kinetochore-localized MT motor proteins such as Cenp-E contributes to inefficient movement of distally positioned chromosomes.
-
3.
Failure to correct erroneous kinetochore–MT attachments—Increased MT turnover during early mitosis prevents the premature stabilization of kinetochore–MTs (Kabeche and Compton 2013). This activity depends on Aurora B kinase, the enzymatic component of the chromosomal passenger complex, which phosphorylates kinetochore substrates to increase kinetochore–MT turnover (Carmena et al. 2012). Defective Aurora B activity results in the accumulation of erroneous kinetochore–MT connections, including syntelic attachments, in which both sister kinetochores remain bound to MTs emanating from the same spindle pole (Cimini et al. 2006; Ditchfield et al. 2003; Hauf et al. 2003). KNL1 is required for kinetochore localization of proteins that both increase (Bub1) and counteract (PP1, PP2A via BubR1) Aurora B activity (Fig. 4). Therefore, depletion of KNL1 likely affects Aurora B kinase-mediated kinetochore–MT attachment regulation. In support of this, KNL1 is required for Bub1-mediated histone H2A phosphorylation (Caldas et al. 2013), which promotes centromere and kinetochore recruitment of Aurora B (Caldas et al. 2013; Yamagishi et al. 2010; Kawashima et al. 2010) and Aurora B activity. Accordingly, KNL1 depletion results in decreased levels of centromere and kinetochore-localized active Aurora B and decreased phosphorylation of kinetochore substrates required for kinetochore–MT regulation (Fig. 4) (Caldas et al. 2013). Thus, chromosome misalignment after KNL1 depletion may result, in part, from reduced Aurora B kinase activity. However, the prevalence of erroneous kinetochore–MT attachments in KNL1-depleted cells has not been directly tested.
Fig. 4 Proposed model for the function of KNL1 in kinetochore–MT attachment regulation and the SAC. In early mitosis, KNL1 promotes the kinetochore recruitment of SAC proteins Bub1, BubR1, Bub3, Mad1, and Mad2, which leads to generation and amplification of the “wait anaphase” signal that inhibits APC/CCdc20 activation. Association of Bub1 with KNL1 promotes Bub1 activity, leading to centromere and kinetochore Aurora B kinase recruitment and activation. Aurora B phosphorylates outer kinetochore substrates, which prevents premature kinetochore–MT stabilization. In late mitosis, KNL1-mediated recruitment of phosphatases antagonizing Aurora B promotes kinetochore–MT attachment stability. PP1 dephosphorylates KNL1, resulting in Bub1 dissociation and subsequent loss of SAC proteins. PP1 binding to KNL1 and KNL1 MT binding additionally promotes SAC silencing. With attached kinetochores no longer amplifying the “wait anaphase” signal, disassembled SAC inhibitory complexes lead to APC/CCdc20 activation, cyclin B and securin degradation, and mitotic exit. The cartoon shows the Mis12 complex and CENP-T simultaneously binding to the Ndc80 complex for simplicity. However, it has been demonstrated that these two kinetochore components compete for binding to the Ndc80 complex (Bock et al. 2012; Nishino et al. 2013; Schleiffer et al. 2012)
Proper congression of chromosomes to the spindle equator, which is key for the accurate distribution of replicated chromosomes, involves the coordination of multiple mechanisms that are not completely understood. KNL1 may potentially be involved in a number of these processes, including establishment of end-on kinetochore–MT attachments, KMN-independent chromosome movement, and regulation of kinetochore–MT attachment strength. Counter-intuitively, depletion of KNL1, a key component of the KMN network with direct MT binding activity, does not completely prevent chromosome alignment to the spindle equator. Determining the mechanisms that remain intact in KNL1-depleted cells and those that are disrupted will be important for the greater understanding of the complex process of chromosome congression.
Role of KNL1 in SAC activation
KNL1 serves as a binding platform, both directly and indirectly, for proteins involved in the SAC, the safety mechanism that prevents mitotic exit until all kinetochores are correctly attached to spindle MTs. The SAC delays anaphase by promoting and sustaining inhibition of the anaphase promoting complex/cyclosome (APC/C), an E3 ubiquitin ligase that, upon activation by Cdc20, targets securin and cyclin B for degradation, which ultimately drives mitotic exit (Jia et al. 2013; Lara-Gonzalez et al. 2012; Musacchio and Salmon 2007). The core components of the SAC are the evolutionarily conserved proteins Bub1, BubR1/Mad3, Bub3, Mad1, Mad2, and Mps1. All of these SAC components, except for Mps1, have been shown to rely on KNL1 for kinetochore localization (Fig. 4). Bub1, Bub3, and BubR1 directly interact with KNL1 (Bolanos-Garcia et al. 2011; Krenn et al. 2012; Primorac et al. 2013; Shepperd et al. 2012; Taylor et al. 1998; Yamagishi et al. 2012). Mad1 and Mad2 kinetochore localization depends on Bub1 (Klebig et al. 2009; Rischitor et al. 2007; Sharp-Baker and Chen 2001) and the RZZ complex (Karess 2005); therefore, it is expected that KNL1 depletion compromises Mad1 and Mad2 kinetochore localization (Pagliuca et al. 2009).
KNL1 was first found to associate with Bub1 and BubR1 through yeast two-hybrid experiments (Kiyomitsu et al. 2007, 2011). Later experiments demonstrated that the two closely related but distinct KI motifs in KNL1, KI1 and KI2, directly bind the TPR (tetratricopeptide repeat) domains of Bub1 and BubR1, respectively (Bolanos-Garcia et al. 2011; Kiyomitsu et al. 2011; Krenn et al. 2012). Interestingly, the KI/TPR interactions are dispensable for both Bub1 and BubR1 kinetochore targeting in human cells (Krenn et al. 2012). However, that is not the case in fission yeast, where a ΔTPR Bub1 mutant does not efficiently target to kinetochores (Vanoosthuyse et al. 2004). It is possible that in human cells, the KI/TPR interaction enhances, but is not essential for SAC protein recruitment and/or function. Recent studies in yeast and human cells established that Mps1-mediated phosphorylation of the KNL1 MELT repeats is required for Bub1/Bub3 kinetochore targeting (London et al. 2012; Primorac et al. 2013; Shepperd et al. 2012; Yamagishi et al. 2012). Although Bub1 and Bub3 require each other for KNL1-mediated kinetochore localization, the inter-dependency of BubR1 with Bub1 and/or Bub3 for kinetochore localization and function remains unresolved. In addition, whether all MELT repeats, specific repeats, or a threshold number of repeats are critical for recruiting Bub1 and Bub3 and if MELT phosphorylation also mediates direct BubR1 recruitment are outstanding, unanswered questions.
Although we are closer to understanding how the checkpoint proteins interact with KNL1, how these interactions impact their function in SAC signaling is a fundamental issue that remains to be addressed. Bub1 is required for kinetochore localization of BubR1, Bub3, Mad1, and Mad2 (Lara-Gonzalez et al. 2012), and accordingly, Bub1's role in SAC signaling has been primarily attributed to its scaffolding function. Consistent with this, Bub1 localizes to kinetochores in late prophase and exhibits a relatively low turnover rate at kinetochores compared to other checkpoint proteins (Howell et al. 2004; Shah et al. 2004). A logical assumption is that Bub1 must itself localize to kinetochores to carry out its scaffolding role. In support of this, in fission yeast, a ΔTPR Bub1 mutant deficient in kinetochore localization and unable to recruit Bub3 and BubR1 exhibited a defective SAC (Vanoosthuyse et al. 2004). In human cells, however, a Bub1 mutant lacking the TPR domain was able to partially mediate a SAC response (Klebig et al. 2009), presumably due to its ability to interact with KNL1 in a MELT-mediated manner. It will be important to determine if a KNL1 mutant completely disrupted for MELT-mediated Bub1 kinetochore association supports SAC signaling in human cells.
KNL1 also directly binds the pseudokinase BubR1 (Bolanos-Garcia et al. 2011; Krenn et al. 2012), whose function in the SAC has been more clearly defined (Jia et al. 2013). BubR1 serves as a pseudosubstrate inhibitor of APC/CCdc20, and it requires Bub3 for kinetochore association. In one study, BubR1-depleted HeLa cells expressing a GLEBS-domain BubR1 mutant (defective for Bub3 binding and kinetochore localization) failed to mount a SAC response in the presence of unattached kinetochores (Elowe et al. 2010). However, subsequent studies in human cancer cell lines, primary human cells, and mouse embryonic fibroblasts demonstrated that BubR1 GLEBS domain mutants were able, in fact, to support SAC activity (Malureanu et al. 2009; Ding et al. 2013). Based on the latter evidence, we favor a model in which BubR1 kinetochore localization is not a requirement for SAC function.
KNL1 also plays a role in recruiting the essential SAC proteins Mad1 and Mad2 to kinetochores, likely through indirect pathways. Mad1, the kinetochore receptor for Mad2, requires both Bub1 and the RZZ complex for kinetochore localization (Kops and Shah 2012). Direct kinetochore-binding receptors for Mad1, however, have not been identified. Unattached kinetochores accumulate complexes of Mad1–Mad2 that catalyze the conversion of inactive, cytoplasmic Mad2 (“open” Mad2) to a form of Mad2 that binds Cdc20 (“closed” Mad2) and inhibits APC/C activity (Musacchio 2012) (Fig. 4). Cdc20-bound Mad2, at the kinetochore or in the cytoplasm, also potentially mediates Mad2 catalytic conversion, resulting in further inhibition of APC/CCdc20. Mad2-based amplification of the APC/CCdc20 inhibitor is thought to be critical for delaying mitotic exit in the presence of unattached kinetochores (Chen et al. 1996; De Antoni et al. 2005; Kulukian et al. 2009; Luo et al. 2004; Nezi et al. 2006). Although it is clear that Mad1 and Mad2 are indispensable for SAC function, whether their kinetochore localization is explicitly required is not yet resolved (Musacchio and Salmon 2007; Przewloka and Glover 2009). In a recent study, human cells expressing a version of Mad1 that exhibited constitutive kinetochore localization accumulated in mitosis (approximately fivefold increase in mitotic index). Introduction of mutations in Mad1 that abolished the interaction between kinetochore-localized Mad1 and Mad2 led to accumulation of mitotic cells, albeit to a lesser extent (~2.5-fold increase in mitotic index), suggesting that (Maldonado and Kapoor 2011), suggesting that SAC activation is not entirely dependent on Mad1–Mad2 association at kinetochores.
Since it is not clear to what extent most SAC proteins require kinetochore localization to function, it is not surprising that the role of KNL1 in SAC activation is also not clear. Kiyomitsu et al. (2007) demonstrated that depletion of KNL1 from HeLa cells resulted in a failure of cells to arrest after incubation with microtubule poisons. Paradoxically, although we observe a less robust SAC after KNL1 depletion from human cells, we do not observe SAC abrogation despite a penetrant RNAi-mediated depletion (GC and JD, unpublished data). Examining the SAC response after KNL1 perturbation in other organisms reveals mixed results. Drosophila cells depleted of KNL1 exhibit a mitotic delay in response to unaligned chromosomes and accumulate in mitosis upon addition of the MT poison colchicine (Schittenhelm et al. 2009). Similarly, populations of chicken DT40 cells depleted of KNL1 delay in mitosis in response to unaligned chromosomes, indicative of a functioning SAC (Cheeseman et al. 2008). However, in budding yeast, cells harboring a KNL1 temperature-sensitive mutant of Spc105 do not delay mitosis in the presence of mono-oriented chromosomes and fail to arrest upon incubation with MT poisons (Pagliuca et al. 2009). Additionally, KNL1-depleted C. elegans embryos, which exhibit complete chromosome congression failure, exit mitosis with similar timing as control embryos, suggesting defective SAC activity (Desai et al. 2003). These differences could be explained by the extent of dependency of SAC proteins on KNL1 for kinetochore localization in different organisms..
An interesting question arising from these observations is how, without a platform for stable kinetochore binding, might SAC proteins sustain checkpoint activity in KNL1-depleted cells? Based on the fact that APC/C inhibitory complexes are found in the cytoplasm, even before mitosis (Sudakin et al. 2001), and that essential components of these complexes including BubR1 appear to function in a manner that is independent of kinetochore localization (Ding et al. 2013; Malureanu et al. 2009), it is possible that a less potent SAC is able to form in the cytoplasm of KNL1-depleted cells. Additionally, increasing evidence points to a role for KNL1 in SAC silencing (discussed in the following section) (Espeut et al. 2012; Meadows et al. 2011; Rosenberg et al. 2011). Thus, a reasonable explanation for the mitotic delay observed in KNL1-depleted cells and for the SAC-mediated arrest upon treatment with MT drugs is that these cells are unable to reliably silence a weakened SAC. Clearly, resolving the mechanism by which the SAC responds to unattached kinetochores to inhibit mitotic exit requires an understanding of the interactions between SAC proteins and the kinetochore platform. Identifying kinetochore receptors for SAC proteins and the specific domains that mediate kinetochore localization will allow the design of experiments to test to what extent SAC activation relies on kinetochores.
Role of KNL1 in SAC silencing
The RVSF and SILK motifs in the KNL1 N-terminus mediate direct binding to PP1 (Fig. 1), a phosphatase that counteracts Aurora B kinase activity (Lesage et al. 2011). Aurora B kinase itself regulates the interaction between KNL1 and PP1 as binding is disrupted by Aurora B-mediated phosphorylation of the RVSF motif (Liu et al. 2010). In human cells, PP1 binding to KNL1, together with KNL1-BubR1-mediated PP2A kinetochore recruitment, is proposed as a mechanism by which outer kinetochore substrates are dephosphorylated during late mitosis, allowing stabilization of kinetochore–MT attachments (Foley et al. 2011; Liu et al. 2010; Suijkerbuijk et al. 2012). Interestingly, in both budding and fission yeast and in C. elegans, KNL1-mediated PP1 binding contributes to checkpoint silencing (Espeut et al. 2012; Meadows et al. 2011; Pinsky et al. 2009; Rosenberg et al. 2011; Vanoosthuyse and Hardwick 2009). In budding yeast, it was recently demonstrated that a KNL1 mutant unable to bind PP1 (RVSF→RASA) was not viable (Rosenberg et al. 2011). This mutant became viable upon Mad2 depletion, indicating that lethality resulted from persistent SAC activation and suggesting that the persistent SAC response was due to impaired PP1 binding to KNL1 (Rosenberg et al. 2011). In the same study, a non-phosphorylatable RVSF mutant (RVAF), which in principle constitutively recruits PP1 to kinetochores, did not prematurely silence the SAC, suggesting that PP1 recruitment alone is not sufficient for this process. In agreement with this, the MT binding region of KNL1 is necessary for SAC silencing in C. elegans (Fig. 1) (Espeut et al. 2012). Embryos containing mutations in the MT binding domain of KNL1 that disrupted MT interaction in vitro were able to congress chromosomes to the metaphase plate without obvious kinetochore–MT attachment defects but exhibited delayed anaphase onset (Espeut et al. 2012). Furthermore, KNL1 MT binding mutants only experienced an extended mitotic arrest when MTs were present, further suggesting that KNL1 MT binding activity contributes to SAC silencing (Espeut et al. 2012). It remains to be tested if PP1 binding and/or MT binding activities are crucial for SAC silencing in human cells. Specifically, it will be informative to test if a KNL1 mutant unable to recruit SAC proteins, but with intact MT- and PP1-binding activities, reverses the mitotic delay observed upon KNL1 depletion. Experimental manipulation of KNL1, which acts as both a platform for SAC-activating and SAC-silencing proteins, will help elucidate how checkpoint activation and amplification are coupled with checkpoint silencing and how MT attachment modulates this complex kinetochore machinery.
Closing remarks and future directions
The success of mitosis relies on the (1) formation of kinetochore–MT attachments able to facilitate both correct congression and segregation of all replicated chromosomes and (2) production of a temporally controlled, SAC-generated APC/CCdc20 inhibitor that prevents mitotic exit until all chromosomes are properly attached to spindle MTs. Despite considerable progress in the understanding of key proteins and molecular mechanisms involved in these two processes, there is no unified view of how they are linked. KNL1 is necessary for the establishment of a reliable kinetochore–MT interface that attaches dynamic spindle MT plus-ends to mitotic chromosomes. KNL1 also ensures that SAC proteins required to inhibit APC/CCdc20 accumulate at unattached kinetochores to amplify the inhibitory signal and delay mitotic exit when appropriate. Moreover, KNL1 is required to quench APC/CCdc20 inhibition and silence the SAC to promote mitotic exit when the cell is ready to divide. As a kinetochore hub, playing a central role in the organization of kinetochore proteins and their interactivity, it is not surprising that KNL1 localization to kinetochores is critical for these processes. Regardless, the possibility that this large and “unstructured” protein plays a more direct role in coupling these fundamental mitotic processes cannot be dismissed and demands further investigation. A major challenge for the future is to determine how KNL1 influences the activities of individual signaling molecules and how, when accumulated to the same kinetochore platform, these molecules exert their functions.
References
Alushin GM, Ramey VH, Pasqualato S, Ball DA, Grigorieff N, Musacchio A, Nogales E (2010) The Ndc80 kinetochore complex forms oligomeric arrays along microtubules. Nature 467(7317):805–810. doi:10.1038/nature09423
Bock LJ, Pagliuca C, Kobayashi N, Grove RA, Oku Y, Shrestha K, Alfieri C, Golfieri C, Oldani A, Dal Maschio M, Bermejo R, Hazbun TR, Tanaka TU (2012) De Wulf P Cnn1 inhibits the interactions between the KMN complexes of the yeast kinetochore. Nat Cell Biol 14(6):614–24. doi:10.1038/ncb2495
Bolanos-Garcia VM, Lischetti T, Matak-Vinković D, Cota E, Simpson PJ, Chirgadze DY, Spring DR, Robinson CV, Nilsson J, Blundell TL (2011) Structure of a Blinkin–BUBR1 complex reveals an interaction crucial for kinetochore–mitotic checkpoint regulation via an unanticipated binding site. Structure 19(11):1691–1700. doi:10.1016/j.str.2011.09.017
Cai S, O'Connell CB, Khodjakov A, Walczak CE (2009) Chromosome congression in the absence of kinetochore fibres. Nat Cell Biol 11(7):832–838. doi:10.1038/ncb1890
Caldas, GV, DeLuca, KF, DeLuca, JG (2013) KNL1 facilitates phosphorylation of outer kinetochore proteins by promoting aurora b kinase activity. J Cell Biol, in press
Carmena M, Wheelock M, Funabiki H, Earnshaw WC (2012) The chromosomal passenger complex (CPC): from easy rider to the godfather of mitosis. Nat Rev Mol Cell Biol 13(12):789–803. doi:10.1038/nrm3474
Cheeseman IM, Niessen S, Anderson S, Hyndman F, Yates JR 3rd, Oegema K, Desai A (2004) A conserved protein network controls assembly of the outer kinetochore and its ability to sustain tension. Genes Dev 18(18):2255–2268
Cheeseman IM, Chappie JS, DAWilson-Kubalek EM (2006) The conserved KMN network constitutes the core microtubule binding site of the kinetochore. Cell 127:983–997
Cheeseman IM, Hori T, Fukagawa T, Desai A (2008) KNL1 and the CENP-H/I/K complex coordinately direct kinetochore assembly in vertebrates. Mol Biol Cell 19(2):587–594
Chen RH, Waters JC, Salmon ED, Murray AW (1996) Association of spindle assembly checkpoint component XMAD2 with unattached kinetochores. Science 274(5285):242–246
Ciferri C, Pasqualato S, Screpanti E, Varetti G, Santaguida S, Dos Reis G, Maiolica A, Polka J, De Luca JG, De Wulf P, Salek M, Rappsilber J, Moores CA, Salmon ED, Musacchio A (2008) Implications for kinetochore–microtubule attachment from the structure of an engineered Ndc80 complex. Cell 133(3):427–439. doi:10.1016/j.cell.2008.03.020
Cimini D, Wan X, Hirel CB, Salmon ED (2006) Aurora kinase promotes turnover of kinetochore microtubules to reduce chromosome segregation errors. Curr Biol 16(17):1711–1718
De Antoni A, Pearson CG, Cimini D, Canman JC, Sala V, Nezi L, Mapelli M, Sironi L, Faretta M, Salmon ED, Musacchio A (2005) The Mad1/Mad2 complex as a template for Mad2 activation in the spindle assembly checkpoint. Curr Biol 15(3):214–225
DeLuca JG, Musacchio A (2012) Structural organization of the kinetochore–microtubule interface. Curr Opin Cell Biol 24(1):48–56. doi:10.1016/j.ceb.2011.11.003
DeLuca JG, Moree B, Hickey JM, Kilmartin JV, Salmon ED (2002) hNuf2 inhibition blocks stable kinetochore–microtubule attachment and induces mitotic cell death in HeLa cells. J Cell Biol 159(4):549–555
DeLuca J, Gall W, Ciferri C, Cimini D, Musacchio A, Salmon E (2006) Kinetochore microtubule dynamics and attachment stability are regulated by Hec1. Cell 127(5):969–982
Desai A, Rybina S, Müller-Reichert T, Shevchenko A, Shevchenko A, Hyman A, Oegema K (2003) KNL-1 directs assembly of the microtubule-binding interface of the kinetochore in C. elegans. Genes Dev 17(19):2421–2435
Ding Y, Hubert CG, Herman J, Corrin P, Toledo CM, Skutt-Kakaria K, Vazquez J, Basom R, Zhang B, Risler JK, Pollard SM, Nam DH, Delrow JJ, Zhu J, Lee J, DeLuca J, Olson JM, Paddison PJ (2013) Cancer-specific requirement for BUB1B/BUBR1 in human brain tumor isolates and genetically transformed cells. Cancer Discov 3(2):198–211. doi:10.1158/2159-8290.CD-12-0353
Ditchfield C, Johnson VL, Tighe A, Ellston R, Haworth C, Johnson T, Mortlock A, Keen N, Taylor SS (2003) Aurora B couples chromosome alignment with anaphase by targeting BubR1, Mad2, and Cenp-E to kinetochores. J Cell Biol 161(2):267–280
Dyson HJ, Wright PE (2005) Intrinsically unstructured proteins and their functions. Nat Rev Mol Cell Biol 6(3):197–208
Elowe S, Dulla K, Uldschmid A, Li X, Dou Z, Nigg EA (2010) Uncoupling of the spindle-checkpoint and chromosome-congression functions of BubR1. J Cell Sci 123(Pt 1):84–94. doi:10.1242/jcs.056507
Espeut J, Cheerambathur DK, Krenning L, Oegema K, Desai A (2012) Microtubule binding by KNL-1 contributes to spindle checkpoint silencing at the kinetochore. J Cell Biol. doi:10.1083/jcb.201111107
Ferreon AC, Ferreon JC, Wright PE, Deniz AA (2013) Modulation of allostery by protein intrinsic disorder. Nature 498(7454):390–394. doi:10.1038/nature12294
Foley EA, Maldonado M, Kapoor TM (2011) Formation of stable attachments between kinetochores and microtubules depends on the B56-PP2A phosphatase. Nat Cell Biol 13(10):1265–1271. doi:10.1038/ncb2327
Gascoigne KE, Takeuchi K, Suzuki A, Hori T, Fukagawa T, Cheeseman IM (2011) Induced ectopic kinetochore assembly bypasses the requirement for CENP-A nucleosomes. Cell 145(3):410–422. doi:10.1016/j.cell.2011.03.031
Gassmann R, Holland AJ, Varma D, Wan X, Civril F, Cleveland DW, Oegema K, Salmon ED, Desai A (2010) Removal of Spindly from microtubule-attached kinetochores controls spindle checkpoint silencing in human cells. Genes Dev 24(9):957–971. doi:10.1101/gad.1886810
Gönczy P, Echeverri C, Oegema K, Coulson A, Jones SJ, Copley RR, Duperon J, Oegema J, Brehm M, Cassin E, Hannak E, Kirkham M, Pichler S, Flohrs K, Goessen A, Leidel S, Alleaume AM, Martin C, Ozlü N, Bork P, Hyman AA (2000) Functional genomic analysis of cell division in C. elegans using RNAi of genes on chromosome III. Nature 408(6810):331–336
Gunasekaran K, Tsai CJ, Kumar S, Zanuy D, Nussinov R (2003) Extended disordered proteins: targeting function with less scaffold. Trends Biochem Sci 28(2):81–85
Hauf S, Cole RW, LaTerra S, Zimmer C, Schnapp G, Walter R, Heckel A, van Meel J, Rieder CL, Peters JM (2003) The small molecule Hesperadin reveals a role for Aurora B in correcting kinetochore microtubule attachment and in maintaining the spindle assembly checkpoint. J Cell Biol 161(2):281–294
Hayette S, Tigaud I, Vanier A, Martel S, Corbo L, Charrin C, Beillard E, Deleage G, Magaud JP, Rimokh R (2000) AF15q14, a novel partner gene fused to the MLL gene in an acute myeloid leukaemia with a t(11;15)(q23;q14). Oncogene 19(38):4446–4450
Haynes C, Oldfield CJ, Ji F, Klitgord N, Cusick ME, Radivojac P, Uversky VN, Vidal M, Iakoucheva LM (2006) Intrinsic disorder is a common feature of hub proteins from four eukaryotic interactomes. PLoS Comput Biol 2(8):e100. doi:10.1371/journal.pcbi.0020100
Hori T, Fukagawa T (2012) Establishment of the vertebrate kinetochores. Chromosome Res 20(5):547–561. doi:10.1007/s10577-012-9289-9
Howell BJ, Moree B, Farrar EM, Stewart S, Fang G, Salmon ED (2004) Spindle checkpoint protein dynamics at kinetochores in living cells. Curr Biol 14(11):953–964
Huang Y, Liu Z (2010) Smoothing molecular interactions: the “kinetic buffer” effect of intrinsically disordered proteins. Proteins 78(16):3251–3259. doi:10.1002/prot.22820
Jia L, Kim S, Yu H (2013) Tracking spindle checkpoint signals from kinetochores to APC/C. Trends Biochem Sci 38(6):302–311
Johnson VL, Scott MIF, Holt SV, Hussein D, Taylor SS (2004) Bub1 is required for kinetochore localization of BubR1, Cenp-E, Cenp-F and Mad2, and chromosome congression. J Cell Sci 117(Pt8):1577–1589. doi:10.1242/jcs.01006
Kabeche L, Compton DA (2013) Cyclin A regulates kinetochore microtubules to promote faithful chromosome segregation. Nature 502(7469):110–113. doi:10.1038/nature12507
Kapoor TM, Lampson MA, Hergert P, Cameron L, Cimini D, Salmon ED, McEwen BF, Khodjakov A (2006) Chromosomes can congress to the metaphase plate before biorientation. Science 311(5759):388–391
Karess R (2005) Rod-Zw10-Zwilch: a key player in the spindle checkpoint. Trends Cell Biol 15(7):386–392
Kawashima SA, Yamagishi Y, Honda T, Ishiguro K, Watanabe Y (2010) Phosphorylation of H2A by Bub1 prevents chromosomal instability through localizing Shugoshin. Science 327(5962):172–177. doi:10.1126/science.1180189
Kerres A, Vietmeier-Decker C, Ortiz J, Karig I, Beuter C, Hegemann J, Lechner J, Fleig U (2004) The fission yeast kinetochore component Spc7 associates with the EB1 family member Mal3 and is required for kinetochore–spindle association. Mol Biol Cell 15(12):5255–5267
Kerres A, Jakopec V, Fleig U (2007) The conserved Spc7 protein is required for spindle integrity and links kinetochore complexes in fission yeast. Mol Biol Cell 18(7):2441–2454
Kim Y, Heuser JE, Waterman CM, Cleveland DW (2008) CENP-E combines a slow, processive motor and a flexible coiled coil to produce an essential motile kinetochore tether. J Cell Biol 181(3):411–419. doi:10.1083/jcb.200802189
Kiyomitsu T, Obuse C, Yanagida M (2007) Human Blinkin/AF15q14 is required for chromosome alignment and the mitotic checkpoint through direct interaction with Bub1 and BubR1. Dev Cell 13(5):663–676. doi:10.1016/j.devcel.2007.09.005
Kiyomitsu T, Murakami H, Yanagida M (2011) Protein interaction domain mapping of human kinetochore protein Blinkin reveals a consensus motif for binding of spindle assembly checkpoint proteins Bub1 and BubR1. Mol Cell Biol 31(5):998–1011. doi:10.1128/MCB.00815-10
Klebig C, Korinth D, Meraldi P (2009) Bub1 regulates chromosome segregation in a kinetochore-independent manner. J Cell Biol 185(5):841–858. doi:10.1083/jcb.200902128
Kline SL, Cheeseman IM, Hori T, Fukagawa T, Desai A (2006) The human Mis12 complex is required for kinetochore assembly and proper chromosome segregation. J Cell Biol 173(1):9–17
Kops GJ, Shah JV (2012) Connecting up and clearing out: how kinetochore attachment silences the spindle assembly checkpoint. Chromosoma 121(5):509–525. doi:10.1007/s00412-012-0378-5
Kops GJ, Saurin AT, Meraldi P (2010) Finding the middle ground: how kinetochores power chromosome congression. Cell Mol Life Sci 67(13):2145–2161. doi:10.1007/s00018-010-0321-y
Krenn V, Wehenkel A, Li X, Santaguida S, Musacchio A (2012) Structural analysis reveals features of the spindle checkpoint kinase Bub1–kinetochore subunit Knl1 interaction. J Cell Biol 196(4):451–467. doi:10.1083/jcb.201110013
Kulukian A, Han J, Cleveland D (2009) Unattached kinetochores catalyze production of an anaphase inhibitor that requires a Mad2 template to prime Cdc20 for BubR1 binding. Dev Cell 16(1):105–117. doi:10.1016/j.devcel.2008.11.005
Laine O, Streaker ED, Nabavi M, Fenselau CC, Beckett D (2008) Allosteric signaling in the biotin repressor occurs via local folding coupled to global dampening of protein dynamics. J Mol Biol 381(1):89–101. doi:10.1016/j.jmb.2008.05.018
Lara-Gonzalez P, Westhorpe FG, Taylor SS (2012) The spindle assembly checkpoint. Curr Biol 22(22):R966–R980. doi:10.1016/j.cub.2012.10.006
Lesage B, Qian J, Bollen M (2011) Spindle checkpoint silencing: PP1 tips the balance. Curr Biol 21(21):R898–R903. doi:10.1016/j.cub.2011.08.063
Lin YT, Chen Y, Wu G, Lee WH (2006) Hec1 sequentially recruits Zwint-1 and ZW10 to kinetochores for faithful chromosome segregation and spindle checkpoint control. Oncogene 25(52):6901–6914
Liu D, Vleugel M, Backer CB, Hori T, Fukagawa T, Cheeseman IM, Lampson MA (2010) Regulated targeting of protein phosphatase 1 to the outer kinetochore by KNL1 opposes Aurora B kinase. J Cell Biol 188(6):809–820. doi:10.1083/jcb.201001006
London N, Ceto S, Ranish JA, Biggins S (2012) Phosphoregulation of Spc105 by Mps1 and PP1 regulates Bub1 localization to kinetochores. Curr Biol CB 22(10):900–906. doi:10.1016/j.cub.2012.03.052
Luo X, Tang Z, Xia G, Wassmann K, Matsumoto T, Rizo J, Yu H (2004) The Mad2 spindle checkpoint protein has two distinct natively folded states. Nat Struct Mol Biol 11(4):338–345
Magidson V, O'Connell CB, Lončarek J, Paul R, Mogilner A, Khodjakov A (2011) The spatial arrangement of chromosomes during prometaphase facilitates spindle assembly. Cell 146(4):555–567. doi:10.1016/j.cell.2011.07.012
Maldonado M, Kapoor TM (2011) Constitutive Mad1 targeting to kinetochores uncouples checkpoint signalling from chromosome biorientation. Nat Cell Biol. doi:10.1038/ncb2223
Malureanu LA, Jeganathan KB, Hamada M, Wasilewski L, Davenport J, van Deursen JM (2009) BubR1 N terminus acts as a soluble inhibitor of cyclin B degradation by APC/C(Cdc20) in interphase. Dev Cell 16(1):118–131. doi:10.1016/j.devcel.2008.11.004
McAinsh AD, Meraldi P (2011) The CCAN complex: linking centromere specification to control of kinetochore–microtubule dynamics. Semin Cell Dev Biol 22(9):946–952. doi:10.1016/j.semcdb.2011.09.016
McEwen BF, Chan GK, Zubrowski B, Savoian MS, Sauer MT, Yen TJ (2001) CENP-E is essential for reliable bioriented spindle attachment, but chromosome alignment can be achieved via redundant mechanisms in mammalian cells. Mol Biol Cell 12(9):2776–2789
McIntosh JR, Molodtsov MI, Ataullakhanov FI (2012) Biophysics of mitosis. Q Rev Biophys 45(2):147–207. doi:10.1017/S0033583512000017
Meadows JC, Shepperd LA, Vanoosthuyse V, Lancaster TC, Sochaj AM, Buttrick GJ, Hardwick KG, Millar JBA (2011) Spindle checkpoint silencing requires association of PP1 to both Spc7 and kinesin-8 motors. Dev Cell 20(6):739–750. doi:10.1016/j.devcel.2011.05.008
Miller SA, Johnson ML, Stukenberg PT (2008) Kinetochore attachments require an interaction between unstructured tails on microtubules and Ndc80 (Hec1). Curr Biol 18(22):1785–1791. doi:10.1016/j.cub.2008.11.007
Musacchio A (2012) Spindle assembly checkpoint: the third decade. Philos Trans R Soc Lond B Biol Sci 366(1584):3595–3604. doi:10.1098/rstb.2011.0072
Musacchio A, Salmon ED (2007) The spindle-assembly checkpoint in space and time. Nat Rev Mol Cell Biol 8(5):379–393
Nekrasov VS, Smith MA, Peak-Chew S, Kilmartin JV (2003) Interactions between centromere complexes in Saccharomyces cerevisiae. Mol Biol Cell 14(12):4931–4946
Nezi L, Rancati G, De Antoni A, Pasqualato S, Piatti S, Musacchio A (2006) Accumulation of Mad2–Cdc20 complex during spindle checkpoint activation requires binding of open and closed conformers of Mad2 in Saccharomyces cerevisiae. J Cell Biol 174(1):39–51
Nishino T, Rago F, Hori T, Tomii K, Cheeseman IM, Fukagawa T (2013) CENP-T provides a structural platform for outer kinetochore assembly. EMBO J 32(3):424–436. doi:10.1038/emboj.2012.348
Oegema K, Desai A, Rybina S, Kirkham M, Hyman AA (2001) Functional analysis of kinetochore assembly in Caenorhabditis elegans. J Cell Biol 53(6):1209–1226
Pagliuca C, Draviam VM, Marco E, Sorger PK, De Wulf P (2009) Roles for the conserved Spc105p/Kre28p complex in kinetochore–microtubule binding and the spindle assembly checkpoint. PLoS ONE 4(10):e7640. doi:10.1371/journal.pone.0007640
Pan CQ, Sudol M, Sheetz M, Low BC (2012) Modularity and functional plasticity of scaffold proteins as p(l)acemakers in cell signaling. Cell Signal 24(11):2143–2165. doi:10.1016/j.cellsig.2012.06.002
Petrovic A, Pasqualato S, Dube P, Krenn V, Santaguida S, Cittaro D, Monzani S, Massimiliano L, Keller J, Tarricone A, Maiolica A, Stark H, Musacchio A (2010) The MIS12 complex is a protein interaction hub for outer kinetochore assembly. J Cell Biol 190(5):835–852
Pinsky BA, Nelson CR, Biggins S (2009) Protein phosphatase 1 regulates exit from the spindle checkpoint in budding yeast. Curr Biol 19(14):1182–1187. doi:10.1016/j.cub.2009.06.043
Powers AF, Franck AD, Gestaut DR, Cooper J, Gracyzk B, Wei RR, Wordeman L, Davis TN, Asbury CL (2009) The Ndc80 kinetochore complex forms load-bearing attachments to dynamic microtubule tips via biased diffusion. Cell 136(5):865–875. doi:10.1016/j.cell.2008.12.045
Primorac I, Weir JR, Chiroli E, Gross F, Hoffmann I, van Gerwen S, Ciliberto A, Musacchio A (2013) Bub3 reads phosphorylated MELT repeats to promote spindle assembly checkpoint signaling. Elife 2:e01030. doi:10.7554/eLife.01030
Przewloka MR, Glover DM (2009) The kinetochore and the centromere: a working long distance relationship. Annu Rev Genet 43:439–465. doi:10.1146/annurev-genet-102108-134310
Przewloka MR, Zhang W, Costa P, Archambault V, D'Avino PP, Lilley KS, Laue ED, McAinsh AD, Glover DM (2007) Molecular analysis of core kinetochore composition and assembly in Drosophila melanogaster. PLoS ONE 2(5):e478. doi:10.1371/journal.pone.0000478
Rischitor PE, May KM, Hardwick KG (2007) Bub1 is a fission yeast kinetochore scaffold protein, and is sufficient to recruit other spindle checkpoint proteins to ectopic sites on chromosomes. PLoS One 2(12):e1342
Rosenberg JS, Cross FR, Funabiki H (2011) KNL1/Spc105 recruits PP1 to silence the spindle assembly checkpoint. Curr Biol CB 21(11):942–947. doi:10.1016/j.cub.2011.04.011
Santaguida S, Musacchio A (2009) The life and miracles of kinetochores. EMBO J 28(17):2511–2531. doi:10.1038/emboj.2009.173
Schaar BT, Chan GK, Maddox P, Salmon ED, Yen TJ (1997) CENP-E function at kinetochores is essential for chromosome alignment. J Cell Biol 139(6):1373–1382
Schittenhelm RB, Chaleckis R, Lehner CF (2009) Intrakinetochore localization and essential functional domains of Drosophila Spc105. EMBO J 28(16):2374–2386. doi:10.1038/emboj.2009.188
Schleiffer A, Maier M, Litos G, Lampert F, Hornung P, Mechtler K, Westermann S (2012) CENP-T proteins are conserved centromere receptors of the Ndc80 complex. Nat Cell Biol 14(6):604–613. doi:10.1038/ncb2493
Shah JV, Botvinick E, Bonday Z, Furnari F, Berns M, Cleveland DW (2004) Dynamics of centromere and kinetochore proteins; implications for checkpoint signaling and silencing. Curr Biol 14(11):942–952
Sharp-Baker H, Chen RH (2001) Spindle checkpoint protein Bub1 is required for kinetochore localization of Mad1, Mad2, Bub3, and CENP-E, independently of its kinase activity. J Cell Biol 153(6):1239–1250
Shaw AS, Filbert EL (2009) Scaffold proteins and immune-cell signalling. Nat Rev Immunol 9(1):47–56. doi:10.1038/nri2473
Shepperd LA, Meadows JC, Sochaj AM, Lancaster TC, Zou J, Buttrick GJ, Rappsilber J, Hardwick KG, Millar JBA (2012) Phosphodependent recruitment of Bub1 and Bub3 to Spc7/KNL1 by Mph1 kinase maintains the spindle checkpoint. Curr Biol CB 22(10):891–899. doi:10.1016/j.cub.2012.03.051
Sudakin V, Chan GK, Yen TJ (2001) Checkpoint inhibition of the APC/C in HeLa cells is mediated by a complex of BUBR1, BUB3, CDC20, and MAD2. J Cell Biol 154(5):925–936
Suijkerbuijk SJ, Vleugel M, Teixeira A, Kops GJ (2012) Integration of kinase and phosphatase activities by BUBR1 ensures formation of stable kinetochore–microtubule attachments. Dev Cell 23(4):745–755. doi:10.1016/j.devcel.2012.09.005
Takimoto M, Wei G, Dosaka-Akita H, Mao P, Kondo S, Sakuragi N, Chiba I, Miura T, Itoh N, Sasao T, Koya RC, Tsukamoto T, Fujimoto S, Katoh H, Kuzumaki N (2002) Frequent expression of new cancer/testis gene D40/AF15q14 in lung cancers of smokers. Br J Cancer 86(11):1757–1762
Taylor SS, Ha E, McKeon F (1998) The human homologue of Bub3 is required for kinetochore localization of Bub1 and a Mad3/Bub1-related protein kinase. J Cell Biol 142(1):1–11
Tompa P (2012) Intrinsically disordered proteins: a 10-year recap. Trends Biochem Sci 37(12):509–516. doi:10.1016/j.tibs.2012.08.004
Tooley J, Stukenberg PT (2011) The Ndc80 complex: integrating the kinetochore's many movements. Chromosome Res 19(3):377–391. doi:10.1007/s10577-010-9180-5
Vanoosthuyse V, Hardwick KG (2009) A novel protein phosphatase 1-dependent spindle checkpoint silencing mechanism. Curr Biol 19(14):1176–1181. doi:10.1016/j.cub.2009.05.060
Vanoosthuyse V, Valsdottir R, Javerzat JP, Hardwick KG (2004) Kinetochore targeting of fission yeast Mad and Bub proteins is essential for spindle checkpoint function but not for all chromosome segregation roles of Bub1p. Mol Cell Biol 24(22):9786–9801
Varma D, Salmon ED (2012) The KMN protein network—chief conductors of the kinetochore orchestra. J Cell Sci 125(Pt 24):5927–5936. doi:10.1242/jcs.093724
Varma D, Wan X, Cheerambathur D, Gassmann R, Suzuki A, Lawrimore J, Desai A, Salmon ED (2013) Spindle assembly checkpoint proteins are positioned close to core microtubule attachment sites at kinetochores. J Cell Biol 202(5):735–746. doi:10.1083/jcb.201304197
Vleugel M, Hoogendoorn E, Snel B, Kops GJ (2012) Evolution and function of the mitotic checkpoint. Dev Cell 23(2):239–250. doi:10.1016/j.devcel.2012.06.013
Walczak CE, Heald R (2008) Mechanisms of mitotic spindle assembly and function. Int Rev Cytol 265:111–158. doi:10.1016/S0074-7696(07)65003-7
Wang H, Hu X, Ding X, Dou Z, Yang Z, Shaw AW, Teng M, Cleveland DW, Goldberg ML, Niu L, Yao X (2004) Human Zwint-1 specifies localization of Zeste White 10 to kinetochores and is essential for mitotic checkpoint signaling. J Biol Chem 279(52):54590–54598
Wei G, Takimoto M, Yoshida I, Mao PZ, Koya RC, Miura T, Kuzumaki N (1999) Chromosomal assignment of a novel human gene D40. Nucleic Acids Symp Ser (42):71–2
Welburn JPI, Vleugel M, Liu D, Yates JR, Lampson MA, Fukagawa T, Cheeseman IM (2010) Aurora B phosphorylates spatially distinct targets to differentially regulate the kinetochore–microtubule interface. Mol Cell 38(3):383–392. doi:10.1016/j.molcel.2010.02.034
Wood KW, Sakowicz R, Goldstein LS, Cleveland DW (1997) CENP-E is a plus end-directed kinetochore motor required for metaphase chromosome alignment. Cell 91(3):357–366
Yamagishi Y, Honda T, Tanno Y, Watanabe Y (2010) Two histone marks establish the inner centromere and chromosome bi-orientation. Science 330(6001):239–243
Yamagishi Y, Yang CH, Tanno Y, Watanabe Y (2012) MPS1/Mph1 phosphorylates the kinetochore protein KNL1/Spc7 to recruit SAC components. Nat Cell Biol 14(7):746–752. doi:10.1038/ncb2515
Acknowledgments
The authors thank Daniela Cimini, O'Neil Wiggan, Keith DeLuca, and Eric Tauchman for critical comments on the manuscript. JGD is supported by National Institutes of Health (NIH) grant R01GM088371.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Caldas, G.V., DeLuca, J.G. KNL1: bringing order to the kinetochore. Chromosoma 123, 169–181 (2014). https://doi.org/10.1007/s00412-013-0446-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-013-0446-5