Abstract
In situ hybridizations of single-copy GC-rich, gene-rich and GC-poor, gene-poor chicken DNA allowed us to localize the gene-rich and the gene-poor chromosomal regions in interphase nuclei of cold-blooded vertebrates. Our results showed that the gene-rich regions from amphibians (Rana esculenta) and reptiles (Podarcis sicula) occupy the more internal part of the nuclei, whereas the gene-poor regions occupy the periphery. This finding is similar to that previously reported in warm-blooded vertebrates, in spite of the lower GC levels of the gene-rich regions of cold-blooded vertebrates. This suggests that this similarity extends to chromatin structure, which is more open in the gene-rich regions of both mammals and birds and more compact in the gene-poor regions. In turn, this may explain why the compositional transition undergone by the genome at the emergence of homeothermy did not involve the entire ancestral genome but only a small part of it, and why it involved both coding and noncoding sequences. Indeed, the GC level increased only in that part of the genome that needed a thermodynamic stabilization, namely in the more open gene-rich chromatin of the nuclear interior, whereas the gene-poor chromatin of the periphery was stabilized by its own compact structure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Using ultracentrifugation of DNA in Cs2SO4 density gradient in the presence of sequence-specific ligands (Corneo et al. 1968), it was demonstrated that the genomes of warm-blooded vertebrates are characterized by a striking long-range compositional heterogeneity (Filipski et al. 1973; Thiery et al. 1976). Indeed, these genomes are mosaics of isochores, long (>>300 kb), compositionally fairly homogeneous regions that belong in a small number of families characterized by different GC levels (Macaya et al. 1976).
Isochores are associated with a number of functional features (for a review, see Bernardi 2004). The most important features are: (1) the GC-richest isochores (the “genome core”) have the highest gene density, the GC-poorest isochores (the “genome desert”) a very low gene density (Bernardi et al. 1985; Mouchiroud et al. 1991; Zoubak et al. 1996); (2) the GC-rich isochores replicate early in the cell cycle, the GC-poor isochores replicate late (Federico et al. 1998, 2000); and (3) isochores are related to chromosomal bands, as suggested by Cuny et al. (1981) and demonstrated by compositional mapping (Bernardi 1989). Indeed, the GC-richest and the GC-poorest isochores of the genomes of warm-blooded vertebrates are prevalently located in telomeric and nontelomeric chromosomal bands, respectively (Saccone et al. 1992, 2001; Federico et al. 2000). Moreover, these isochores are located in internal and peripheral regions of the interphase nucleus, respectively. These isochores correspond to different levels of chromatin condensation, the GC-richest isochores being remarkably more relaxed compared to the GC-poorest ones, as demonstrated by hybridization with compositional fractions of DNA or with DNA from single chromosomal bands (Saccone et al. 2002; Federico et al. 2004). This result was recently confirmed using a different approach, namely the separation of compact and open chromatin fiber structures by sucrose sedimentation (Gilbert et al. 2004) and also by other investigations (Bolzer et al. 2005). This supports a functional compartmentalization of the nucleus related to nuclear architecture (Strouboulis and Wolffe 1996). The functional compartmentalization of the cell nucleus was also demonstrated by other findings, such as the different replication timing of DNA located in the internal and peripheral part of the nucleus (Ferreira et al. 1997; Sadoni et al. 1999).
In the case of cold-blooded vertebrates, evidence is available for a bimodal distribution of genes, but GC-rich isochores have a much lower GC level than in warm-blooded vertebrates (Cacciò et al. 1994; Jabbari and Bernardi 2004; for a review, see Bernardi 2004). In the present work, we investigated the distribution of genes from cold-blooded vertebrates by hybridizing the well-characterized gene-poor and gene-rich isochores of chicken (Thiery et al. 1976; Cortadas et al. 1979; Olofsson and Bernardi 1983; Kadi et al. 1993; Smith et al. 2000; McQueen et al. 1996; Andreozzi et al. 2001; Habermann et al. 2001; Saccone et al. 2002; for review, see Bernardi 2004) on chromosomes and nuclei from the lizard Podarcis sicula and the frog Rana esculenta. The heterologous in situ hybridization allowed us to localize the gene-dense and the gene-poor regions in the genomes under study.
The choice of this strategy was due to the following considerations. Using homologous hybridization would have involved (1) the preparative fractionation of the two DNAs under study to obtain compositional fractions corresponding to the GC-poor, gene-poor isochores and to the GC-rich, gene-rich isochores and (2) the preparation of C0t 1 DNA from each genome to avoid, when fractions would be hybridized on interphase nuclei, any interference by repeated sequences. Obviously, this approach is more laborious than the one we used. An alternative possibility would have been to use as probes the GC-richest and GC-poorest fractions from a mammalian genome, like the human genome. This was not done because we have shown a perfectly parallel behavior of human and chicken fractions in similar yet homologous hybridizations, and because the phylogenetic distance between chicken and lizard is shorter than that between human and lizard.
Materials and methods
DNA probes
Chicken DNA was fractionated by preparative ultracentrifugation in Cs2SO4 density gradients in the presence of the sequence-specific ligand 3,6-bis(acetato-mercuri-methyl) dioxane, BAMD, as previously described (Cortadas et al. 1979). The GC-poorest (ρ°=1.6978) and GC-richest (ρ°=1.7135) DNA fractions (see Andreozzi et al. 2001) were labeled with biotin or digoxigenin using a nick translation kit (Roche, Germany).
Chromosomes and nuclei preparation
R. esculenta and P. sicula were collected in Sicily (Italy). Metaphase chromosomes and nuclei were prepared from bone marrow and fibroblast primary cell line cultures according to techniques previously described (Olmo et al. 1986; Yonenaga-Yassuda et al. 1988; Capriglione et al. 1991). Harvesting for cytogenetic preparations was performed following standard procedures using hypotonic treatment with KCl 0.075 M and methanol/acetic acid fixation.
In situ hybridization and detection
In situ hybridizations were performed with 200 ng of each of the two labeled probes, the GC-richest and the GC-poorest fractions, according to Andreozzi et al. (2001) with some modifications. The hybridization mixtures were 25% formamide / 2×SSC (standard saline citrate, 0.15 M NaCl; 0.015 M sodium citrate) / 10% dextran sulfate / 50 mM phosphate buffer, pH 7.0. In addition, 500× excess of unlabeled sheared chicken DNA was added to the probe. The hybridizations were performed for 72 h at 32°C and posthybridization washes were performed for three times in 2×SSC at 37°C. The low stringency was necessary considering the high degree of divergence between the probe and the target DNAs (especially in the case of the GC-rich isochores). Signal detection was done as previously described (Andreozzi et al. 2001; Saccone et al. 2002), using avidin conjugated with tetramethylrhodamine isothiocyanate and antidigoxigenin antibody conjugated with fluorescein isothiocyanate to detect biotin and digoxigenin labeled probes, respectively. Epifluorescence microscopy, with the appropriate filters and charge-coupled device camera, was used to capture hybridization signals. Fluorescent images were recorded using the MacProbe 4.2.3 and Photoshop 5.0 softwares.
Results
The chromosomes of P. sicula
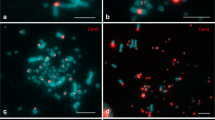
When the GC-richest and the GC-poorest isochores from the chicken genome were hybridized on the chromosomes of P. sicula, the results showed, in both cases, a nonhomogeneous distribution of the hybridization signals on the chromosomes (Fig. 1). More precisely, the chicken GC-poor DNA fractions produced no band-like hybridization signals almost anywhere, and the telomeric ends of the majority of chromosomes particularly showed no signals. In the case of the hybridization with chicken GC-richest DNA fractions, a number of internal regions, showed a general overlapping with the GC-poor hybridization signals, whereas a large number of telomeric bands only showed GC-rich hybridization signals. The nonoverlapping hybridization signals observed in most telomeric bands are also evident in the unique pair of microchromosomes, that showed signals from the GC-richest, but not from the GC-poor chicken DNA fractions. The general overlap of nontelomeric chromosomal regions could be due to the low resolution of the chromosomes under study and to the wide spreading of the hybridization signals. The lack of colocalization of the two different DNA fractions in a number of nontelomeric regions was also observed with a dual-color in situ hybridization (Fig. 2).
Hybridization of compositional DNA fractions from chicken on the P. sicula chromosomes. Metaphase chromosomes from P. sicula hybridized with a GC-poor (green signals) and b GC-rich (red signals) chicken DNA fractions. Chromosomes from lizard were stained with 4′6-diamidino-2-phenylindole 2HCl (DAPI) (blue). Panels c and d are the same metaphases shown in a and b, respectively, with the DAPI staining printed in black and white for a better visualization of the microchromosomes (indicated by the arrows)
P. sicula: nonoverlapping signal distribution on chromosomes. Dual-color in situ hybridization of P. sicula chromosomes with the chicken GC-poor (green signals) and GC-rich (red signals) DNA fractions. Some chromosomes showing a clear nonoverlapping distribution of the signals due to the above probes are shown as an example. In the chromosome shown in the bottom frames, four band-like hybridization signals are visible, with alternating GC-poor and GC-rich chicken DNA hybridizations. The white lines indicate some band-like hybridization signals with the GC-rich chicken DNA fractions (shown on the left chromosomes) and the correlation with the band-like hybridization signals with the GC-poor chicken DNA fractions (shown in the right chromosomes)
The chromosomes of R. esculenta
The GC-richest and the GC-poorest DNA fractions from chicken hybridized on very different chromosomal sites of R. esculenta (Fig. 3). Indeed, the hybridization signals of the chicken GC-richest DNA fractions were largely localized in the telomeric part of the large majority of chromosomes, whereas the hybridization due to the GC-poorest DNA was localized in nontelomeric bands. Moreover, band-like hybridization signals were generally visible, allowing us to identify chromosomes from the entire karyotype according to the chicken GC-poor and GC-rich hybridizations (Fig. 4). Finally, these hybridization-banded karyotypes are in good agreement with the early replication-banding pattern (see Fig. 5).
R. esculenta chromosomes and replication banding. Correlation between the chromosomal banding obtained with the GC-rich chicken DNA fractions (left member of each chromosome pair) and the early replicated bands (right member of each chromosome pair). The early-replicated banding is from Schempp and Schmid (1981)
The interphase nuclei of cold-blooded vertebrates
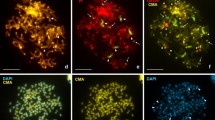
The hybridization results on interphase nuclei of both analyzed species showed (1) a more internal location of the DNA homologous with the chicken GC-richest DNA fractions and (2) a general peripheral location of DNA homologous with the chicken GC-poorest DNA fractions (Fig. 6). This result is similar to that observed in all the avian and mammalian species analyzed so far. Moreover, a high level of hybridization signals was also observed with the chicken GC-poorest isochores around the nucleolus of R. esculenta (data not shown).
The gene-dense and the gene-poor regions in vertebrate cell nuclei. Interphase nuclei from P. sicula and R. esculenta were hybridized with the chicken GC-richest (red signals) and the chicken GC-poorest (green signals) DNA fractions. Interphase nuclei from a mammal (human) and a bird (chicken) species are also shown, for a direct comparison. The human cell nucleus shows the hybridization with the homologous human isochores. a, b, c, d: nuclei from human, chicken, lizard, and frog, respectively, hybridized with the GC-rich (red signals) isochores. a″, b″, c″, d″: the same nuclei hybridized with the GC-poor (green signals) isochores. Nuclei in the middle (a′, b′, c′) show the merged hybridization signals. Nuclei were stained with DAPI (blue). The bar in the bottom-left panel is 2 μm. The results shown on the human and chicken nuclei are in agreement with the previously published compositional mapping of cell nuclei (Saccone et al. 2002)
Discussion
Considering the heterologous hybridization and the stringency conditions used, we suppose that the hybridized sequences from P. sicula and R. esculenta correspond to “unique” (largely coding) sequences, namely to sequences that are more conserved during the evolution of the genomes. In other words, we think that the identified regions (both in chromosomes and nuclei) contain the genes that increased their GC-level during the major compositional shift that occurred in the ancestors of birds and mammals (see Bernardi 2004). If this hypothesis is correct, as we believe, our results provide the first indication that the highest gene densities are in the telomeric bands of the metaphase chromosomes and in the inner part of the cell nuclei of cold-blooded vertebrates.
Indeed, the high level of correlation, shown here for the frog chromosomes, between the early replicating bands and the regions with the highest level of hybridization with the chicken GC-rich DNA fractions is a further indication of the high gene density of these regions. One of the important features related to the GC-level of the isochores is the replication timing. In mammals and birds the GC-rich and gene-rich regions are replicated at the onset of the S phase, whereas the GC-poor and gene-poor ones are late-replicating (Federico et al. 1998, 2000).
The above data suggest that the similarity between the composition of the interior and the periphery of the cell nuclei in the vertebrate species could be extended to the chromatin structure, which is more open in the gene-dense regions in both mammals and birds and more compact in the gene-poor regions (Saccone et al. 2002). In turn, this may explain why the compositional transition did not involve the entire ancestral genome but only a (small) part of it and why it involved both coding and noncoding sequences. Indeed, the GC-level increased only in that part of the genome that needed a thermodynamic stabilization, namely in the more open gene-rich chromatin of the nuclear interior, whereas the gene-poor chromatin of the periphery was stabilized by its own compact structure, as previously proposed (Saccone et al. 2002; Bernardi 2004).
References
Andreozzi L, Federico C, Motta S, Saccone S, Sazanova A, Sazanov A, Smirnov A, Galkina SA, Lukina NA, Rodionov AV, Carels N, Bernardi G (2001) Compositional mapping of chicken chromosomes and identification of the gene-richest regions. Chromosome Res 9:521–532
Bernardi G (1989) The isochore organization of the human genome. Annu Rev Genet 23:637–661
Bernardi G (2004) Structural and evolutionary genomics. Natural selection in genome evolution. Elsevier, Amsterdam
Bernardi G, Olofsson B, Filipski J, Zerial M, Salinas J, Cuny G, Meunier-Rotival M, Rodier F (1985) The mosaic genome of warm-blooded vertebrates. Science 228:953–957
Bolzer A, Kreth G, Solovei I, Koehler D, Saracoglu K, Fauth C, Müller S, Eils R, Cremer C, Speicher MR, Cremer T (2005) Three-dimensional maps of all chromosomes in human male fibroblast nuclei and prometaphase rosettes. PLoS Biol 3(5):e157, pp 826–842
Cacciò S, Perani P, Saccone S, Kadi F, Bernardi G (1994) Single-copy sequence homology among the GC-richest isochores of the genomes from warm-blooded vertebrates. J Mol Evol 39:331–339
Capriglione T, Cardone A, Odierna G, Olmo E (1991) Evolution of a centromeric satellite DNA and phylogeny of a lacertid lizards. Comp Biochem Physiol 100:641–645
Corneo G, Ginelli E, Soave C, Bernardi G (1968) Isolation and characterization of mouse and guinea pig satellite deoxyribonucleic acids. Biochemistry 7:4373–4379
Cortadas J, Olofsson B, Meunier-Rotival M, Macaya G, Bernardi G (1979) The DNA components of the chicken genome. Eur J Biochem 99:179–186
Cuny G, Soriano P, Macaya G, Bernardi G (1981) The major components of the mouse and human genomes. 1. Preparation, basic properties and compositional heterogeneity. Eur J Biochem 115:227–233
Federico C, Saccone S, Bernardi G (1998) The gene-richest bands of human chromosomes replicate at the onset of the S-phase. Cytogenet Cell Genet 80:83–88
Federico C, Andreozzi L, Saccone S, Bernardi G (2000) Gene density in the Giemsa bands of human chromosomes. Chromosome Res 8:737–746
Federico C, Saccone S, Andreozzi L, Motta S, Russo V, Carels N, Bernardi G (2004) The pig genome: compositional analysis and identification of the gene-richest regions in chromosomes and nuclei. Gene 343:245–251
Ferreira J, Paolella G, Ramos C, Lamond AI (1997) Spatial organization of large-scale chromatin domains in the nucleus: a magnified view of single chromosome territories. J Cell Biol 139:1597–1610
Filipski J, Thiery JP, Bernardi G (1973) An analysis of the bovine genome by Cs2SO4-Ag+ density gradient centrifugation. J Mol Biol 80:177–197
Gilbert N, Boyle S, Fiegler H, Woodfine K, Carter NP, Bickmore WA (2004) Chromatin architecture of the human genome: gene-rich domains are enriched in open chromatin fibers. Cell 118:555–566
Habermann FA, Cremer M, Walter J, Kreth G, von Hase J, Bauer K, Wienberg J, Cremer C, Cremer T, Solovei I (2001) Arrangements of macro- and microchromosomes in chicken cells. Chromosome Res 9:569–584
Jabbari K, Bernardi G (2004) Cytosine methylation and CpG, TpG (CpA) and TpA frequencies. Gene 333:143–149
Kadi F, Mouchiroud D, Sabeur G, Bernardi G (1993) The compositional patterns of the avian genomes and their evolutionary implications. J Mol Evol 37:544–551
Macaya G, Thiery JP, Bernardi G (1976) An approach to the organization of eukaryotic genomes at a macromolecular level. J Mol Biol 108:237–254
McQueen HA, Fantes J, Cross SH, Clark VH, Archibald AL, Bird AP (1996) CpG islands of chicken are concentrated on microchromosomes. Nat Genet 12:321–324
Mouchiroud D, D'Onofrio G, Aïssani B, Macaya G, Gautier C, Bernardi G (1991) The distribution of genes in the human genome. Gene 100:181–187
Olmo E, Odierna G, Cobron O (1986) C-band variability and phylogeny of Lacertidae. Genetica 71:63–74
Olofsson B, Bernardi G (1983) Organization of nucleotide sequences in the chicken genome. Eur J Biochem 130:241–245
Saccone S, De Sario A, Della Valle G, Bernardi G (1992) The highest gene concentrations in the human genome are in T-bands of metaphase chromosomes. Proc Natl Acad Sci U S A 89:4913–4917
Saccone S, Pavlicek A, Federico C, Paces J, Bernardi G (2001) Genes, isochores and bands in human chromosomes 21 and 22. Chromosome Res 9:533–539
Saccone S, Federico C, Bernardi G (2002) Localization of the gene-richest and the gene-poorest isochores in the interphase nuclei of mammals and birds. Gene 300:169–178
Sadoni N, Langer S, Fauth C, Bernardi G, Cremer T, Turner BM, Zink D (1999) Nuclear organization of mammalian genomes: polar chromosome territories build up functionally distinct higher order compartments. J Cell Biol 146:1211–1226
Schempp W, Schmid M (1981) Chromosome banding in Amphibia. VI. BrdU-Replication patterns in anura and demonstration of XX/XY sex chromosomes in Rana esculenta. Chromosoma 83:697–710
Smith J, Bruley CK, Paton IR, Dunn I, Jones CT, Windsor D, Morrice DR, Law AS, Masabanda J, Sazanov A, Waddington D, Fries R, Burt DW (2000) Differences in gene density on chicken macrochromosomes and microchromosomes. Anim Genet 31:96–103
Strouboulis J, Wolffe AP (1996) Functional compartmentalization of the nucleus. J Cell Sci 109:1991–2000
Thiery JP, Macaya G, Bernardi G (1976) An analysis of eukaryotic genomes by density gradient centrifugation. J Mol Biol 108:219–235
Yonenaga-Yassuda Y, Kasahara S, Chu TH, Rodrigues MT (1988) High-resolution RGB-banding pattern in the genus Tropidurus (Sauria, Iguanidae). Cytogenet Cell Genet 48:68–71
Zoubak S, Clay O, Bernardi G (1996) The gene distribution of the human genome. Gene 174:95–102
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E.A. Nigg
Rights and permissions
About this article
Cite this article
Federico, C., Scavo, C., Cantarella, C.D. et al. Gene-rich and gene-poor chromosomal regions have different locations in the interphase nuclei of cold-blooded vertebrates. Chromosoma 115, 123–128 (2006). https://doi.org/10.1007/s00412-005-0039-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-005-0039-z