Abstract
In Drosophila melanogaster, the two chromosomal proteins HP1 and HP2 colocalize on heterochromatic and euchromatic sites in polytene chromosomes. Mutations in the HP2 gene act as dominant suppressors of position effect variegation, demonstrating a role for HP2 in the formation or maintenance of heterochromatin. In this paper, we investigated whether a putative homolog of the D. melanogaster HP2 is involved in the facultative heterochromatinization process in mealybugs. Using an antibody raised against the Drosophila HP2, we identified in the mealybug Planococcus citri a cross-reactive epitope, which we refer to as HP2-like. We investigated the HP2-like pattern during the male embryo development where the entire paternal haploid chromosome set becomes heterochromatic. The HP2 antibody heavily decorates the chromocenters, where it localizes with HP1, and marks the chromatin before it acquires the full cytological characteristics of the male-specific heterochromatin. In euchromatic chromosomes, HP2-like is mainly concentrated at telomeric sites. The interplay between HP2-like and HP1-like was studied by dsRNA interference experiments. Extinguishing HP1-like expression by RNAi does not prevent the association of HP2-like with facultative heterochromatin, implying that HP2-like binds to chromatin in a HP1-independent manner. Our results confirm and extend the structural and functional conservation of proteins involved in heterochromatin assembly.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The chromatin is not a static structure, rather it is a dynamic complex able to change from a decondensed and active state (euchromatin) to a packaged and inactive one (heterochromatin) and vice versa.
Position-effect variegation (PEV) and facultative heterochromatinization demonstrate the functional consequences of dynamic changes in chromatin. Juxtaposition of euchromatic genes to heterochromatic domains, either by chromosome rearrangements or transposition events, generally results in silencing of the gene in a subset of the cells in which it is normally expressed, giving rise to a variegating phenotype (PEV; reviewed in Wakimoto 1998). Facultative heterochromatinization is the developmentally regulated and tissue-specific cis-spreading of a heterochromatic state onto an euchromatic region, leading to the inactivation of all the genes it harbors. Two prominent examples are the inactivation of one of the two X chromosomes in female mammals (reviewed in Lyon 1992) and the heterochromatinization of the entire, paternally derived haploid chromosome set in males of lecanoid coccids, or mealybugs (Brown and Nur 1964; Nur 1990). In these species, it is possible to follow the earliest events of heterochromatin formation during male embryo development (Bongiorni et al. 2001). Furthermore, in some adult male tissues, male-specific heterochromatin undergoes reversal (Nur 1967), thus allowing to analyze the very rare process of deheterochromatinization. By contrast, in female embryos, both the paternal and the maternal complements remain euchromatic throughout ontogeny (Brown and Nur 1964).
Both facultative heterochromatinization and PEV provide paradigmatic examples of epigenetic regulation. The heterochromatin assembly depends on the dosage of its individual constituents. The best-studied example of a heterochromatic protein is the D. melanogaster heterochromatin protein 1 (HP1; James and Elgin 1986). Loss-of-function mutations of the Hp1 gene (named Su(var)205), are associated with suppression of PEV, whereas over expression of HP1 results in PEV enhancement, indicating that HP1 is involved in the spreading of heterochromatin (Eissenberg et al. 1992). HP1 is highly conserved in organisms as distant as fission yeast and mammals where it acts in chromatin-mediated gene repression (Singh et al. 1991; Eissenberg and Elgin 2000; Singh and Georgatos 2002). In addition to this role, HP1 has also a telomere-capping function that prevents telomere fusions (Fanti et al. 1998) and seems to be required for transcripts elongation (Piacentini et al. 2003; Vakoc et al. 2005). HP1 is a nonhistone chromosomal protein with two highly conserved domains. The amino-terminal “chromodomain” has the capacity to bind tri-methylated lysine 9 of histone H3 (Bannister et al. 2001; Lachner et al. 2001). The carboxy-terminal “chromoshadow” domain is involved in mediating protein–protein interactions (Eissenberg and Elgin 2000). This latter domain interacts with a Drosophila heterochromatin protein 2 (HP2) in a yeast two-hybrid assay (Shaffer et al. 2002). HP2 coprecipitates with HP1 in D. melanogaster embryo extracts, colocalizes with HP1 on both heterochromatic and euchromatic sites on polytene chromosomes of four Drosophila species, and is recruited to ectopic HP1 sites in vivo (Shaffer et al. 2002; Stephens et al. 2005). Mutations in the HP2 encoding gene (Su(var)2-HP2) act as dominant suppressors of PEV, implying that HP2 is involved in the formation and spreading of heterochromatin (Shaffer et al. 2002, 2006). Very recently, it has been shown that HP2 is also involved in the structural organization of chromosomes (Shaffer et al. 2006).
We recently identified a HP1-like protein in the mealybug Planococcus citri, which is preferentially associated with the male-specific heterochromatin. In early blastoderm embryos, the HP1-like appearance precedes the onset of heterochromatinization that advances as a wave from one pole toward the other one (Bongiorni et al. 2001; Bongiorni and Prantera 2003). Moreover, we found that in mealybugs, facultative heterochromatinization is accompanied by specific histone tail posttranscriptional modifications (Cowell et al. 2002; Kourmouli et al. 2004).
To further dissect the facultative heterochromatinization process, in the present paper, we used the antibody raised against the D. melanogaster HP2 protein as a probe to test its possible contribution to the chromatin remodeling events leading to the heterochromatinization of the paternal chromosome set in P. citri.
Through immunoblot and immunocytological experiments, we provide evidence for the presence of a HP2 cross-reactive epitope in P. citri, which is preferentially associated with the male-specific facultative heterochromatin.
To study the interaction between HP1-like and the presumptive HP2-like protein, we compared the behavior of the two proteins during both the formation of facultative heterochromatin and its reversion to euchromatin that occurs in some adult male tissues. Moreover, the interaction between the two chromosomal proteins has been functionally investigated in Hp1-like mutant phenocopies produced by dsRNAi.
Materials and methods
Mealybug cultures
The mealybugs P. citri (Homoptera, Coccoidea) were raised in our laboratory on sprouting potatoes at 26°C inside glass food containers covered with gauze. The potatoes were kept in the dark to sprout for 1 month before use.
Cytological preparations
Chromosome spreads from embryos were obtained as previously described (Bongiorni et al. 1999, 2001). Chromosome spreads from adult male tissues were obtained fixing male third instar larvae in Bradley–Carnoy. Fixed males were then dissected in a drop of 45% glacial acetic acid on siliconized coverslips and then squashed on microscope slides. Slides were frozen in liquid nitrogen, and the coverslips were popped off with a razor blade (Bongiorni et al. 2004).
Immunofluorescence microscopy
Immediately after preparation, slides were rehydrated in a 1× phosphate buffer solution. The primary antibodies used were the anti-HP1 mouse monoclonal C1A9 antibody (isolated by James and Elgin (1986) and kindly provided by B. Wakimoto; 1:50 dilution), and the anti-HP2 rabbit polyclonal antibody (isolated by Shaffer et al. (2002) and kindly provided by Sarah C. R. Elgin), 1:50 dilution, which is specific for both HP2-L and HP2-S. The immunostaining was performed according to Bongiorni et al. (2001). The secondary antibodies used were Alexa 488-conjugated goat anti-mouse antibody (Molecular Probes, Eugene, OR, 1:100 dilution) and Alexa 594-conjugated goat anti-rabbit antibody (Molecular Probes, 1:200 dilution). Slides were counterstained with 0.2 μg/ml 4′,6-diamidino-2-phenylindole (DAPI; Boehringer, Mannheim) in 2× salt–sodium citrate for 5 min and mounted in an antifade medium (DABCO, Sigma). Negative controls were obtained by incubating slides with the secondary antibodies only. Immunofluorescent preparations were observed and documented as previously described (Bongiorni et al. 1999, 2001), using filter combinations suitable for the different fluorochromes.
Hp1-like dsRNA production
The pchet2 (Hp1-like) cDNA (Bongiorni et al. 2007) cloned into pGEM-T vector (Promega) between the T7 and SP6 promoter sequences was used as a template for in vitro transcription using the RiboMAX large-scale RNA production system T7 (Promega). In this system, dsRNA production requires a T7 RNA polymerase promoter sequence at both 5′ ends of pchet2 cDNA antiparallel strands. Moreover, the transcription efficiency increases if the two T7 promoter sequences are 5′ flanked from a short DNA sequence. Consequently, first of all, we stepwise replaced the 3′ SP6 promoter with a T7 promoter by two polymerase chain reactions (PCRs), at the same time adding two short sequences at both ends of the amplicon. For the first PCR, we used the following primers (underlined: T7 sequence; italic: vector sequence):
-
Forward: 5′-GACGGCCAGTGAATTG TAATACGA-3′
-
Reverse 1: 5′-CACTATAGGG TACTCAAGC-3′
The product of this first PCR was amplified again using the following primers (underlined: T7 sequence; italic: vector sequence):
-
Forward: 5′-GACGGCCAGTGAATTG TAATACGA-3′
-
Reverse 2: 5′-CCAAGTAATACGACTCACT ATAGGG-3′
The pchet2 cDNA, together with the 5′ and 3′ T7 flanking promoters, was then amplified using the forward primer and the reverse 2 primer. The PCR product was purified and directly used for in vitro RNA transcription (Ribomax system; Promega). The DNA template was removed after RNA synthesis adding RQ1 RNase-free DNase (1 U/μg). The RNA mixture that contained both strands of RNA was then denaturated at 65°C and allowed to anneal slowly by cooling to room temperature (Somma et al. 2002). For quality control, an aliquot of each dsRNA was analyzed on standard nondenaturing agarose gel to confirm the size and integrity of the dsRNA.
dsRNA treatment of mealybug embryos (RNAi)
The dsRNAs were precipitated with ammonium acetate and isopropanol and then dissolved in a soaking buffer (Maeda et al. 2001). Embryos were soaked in 500 μl of a 20- to 40-μg/ml dsRNAi solution, and incubated at 26°C for 4 h. Embryos were dissected and immunostained, and the phenotypes were observed under a fluorescence microscope (as described in Bongiorni et al. 2001). The negative controls involved soaking the embryos in the buffer alone, without dsRNA. The exposure time to the soaking solution was limited to a maximum of 4 h because embryos do not survive for longer times after releasing from the mother, even in a mock-interfering solution.
Western blot analysis
Whole protein extracts were prepared from 25–30 fertilized adult females of P. citri according to Bongiorni et al. 2001. To extract HP2 from D. melanogaster nuclei, the pellet was soaked in 25 mM N-2-hydroxyethylpiperazine-N′-2-ethanesulfonic acid pH 8, 400 mM NaCl, 1 mM ethylenediamine tetraacetic acid, 10% glycerol, and 2% TritonX100 for half an hour on ice. Proteins were loaded onto 8% sodium dodecyl sulfate (SDS)-polyacrylamide gel and run at 120 V by using the Mini-Protean II system (BioRad, Richmond, CA) until the 50-kDa marker is near the bottom of the gel. The protein mass ladder was the See Blue from Invitrogen. Proteins were electrically transferred to a nitrocellulose membrane Hybond enzymatic chemiluminescence (ECL; Amersham Biosciences) at 120 V on ice for 1 h and a half, by using a semidry electroblotting apparatus (BioRad) with standard buffers. After transfer, the filter was blocked at room temperature for an hour with 5% nonfat dried milk in TBST (0.05% Tween 20 + 1× Tris-buffered saline: 10 mM Tris–HCl, pH 8.0, 150 mM NaCl). The filter was incubated with chicken anti-HP2 antibody (Stephens et al. 2005; kindly provided by Sarah C. R. Elgin), 1:1,000 in TBST, overnight at 4°C. The filter was then washed in TBST three times for 10 min each and then incubated 1 h with a 1:2,000 dilution in TBST of goat anti-chicken IgG-horseradish peroxidase-conjugated secondary antibody (Santa Cruz Biotechnology, Santa Cruz, CA). After three additional washes of 10 min each in TBST, the filter was incubated with the ECL-plus reagent (Amersham Biosciences). Chemioluminescent bands were detected by exposing the filter to an X-ray film (Kodak, Rochester, NY) for times ranging from 1 to 5 min. Negative controls were performed by incubating protein samples with secondary antibody only.
Results
Drosophila anti-HP2 antibody identifies a cross-reactive antigen in mealybug nuclei
The presence of a presumptive HP2-like protein in cytological preparations of the mealybug P. citri (2n = 10) was investigated by immunoblotting and immunofluorescence analysis. In our analysis, we used the polyclonal antibodies raised against the D. melanogaster HP2 and characterized by Shaffer et al. (2002) and Stephens et al. (2005). These authors showed that Drosophila presents two HP2 isoforms, with predicted molecular weights of 356 and 176 kDa, this latter migrating at an apparent molecular weight around 250 kDa (Stephens et al. 2005). In mealybugs, the anti-HP2 antibodies recognize cross-reactive epitopes in both protein extracts and cytological preparations (Fig. 1). Figure 1a shows a Western blot of P. citri protein extracts probed with the chicken anti-HP2 antibody. In mealybug extracts, the antibody recognizes two protein species, one migrating as the larger Drosophila isoform and one migrating with an apparent molecular weight around 200 kDa. Isolation and sequencing of the P. citri Hp2 gene is required as a definitive proof to assess that the bands detected by Western analysis correspond to the P. citri ortholog of the D. melanogaster Hp2. However, as described below, cytological studies show that the anti-HP2 antibody recognizes a chromosomal protein whose behavior is consistent with that expected of an HP2-like molecule. Hence, we refer to the P. citri protein recognized by the antibody as HP2-like. Figure 1b–d illustrates the DAPI staining of male and female interphase nuclei, which can be easily distinguished for the presence, in male cells, of a conspicuous, brightly fluorescent chromocenter. In male nuclei, the rabbit anti-HP2 antibody localizes almost exclusively with the male-specific heterochromatin (Fig. 1b,c), whereas in female tissues, the HP2 antibody signal is scattered throughout the chromatin (Fig. 1d). In male nuclei, the coincidence of the HP2 signal with the chromocenter is particularly apparent where the chromocenter is splitted into two masses (arrows in Fig. 1c). In male mitotic cells, the cytological differentiation between paternal and maternal sets is still apparent at early (Fig. 2a) and middle metaphase (Fig. 2b), whereas at late metaphase, the euchromatic and the heterochromatic chromosome sets reach the same degree of condensation and are no longer distinguishable on a morphological basis only (Fig. 2c). The association of the HP2 antibody signal with the male-specific heterochromatin persists throughout the mitotic phases. At early (Fig. 2a) and middle metaphase (Fig. 2b), the fluorescent HP2 antibody signal is predominantly localized over the five paternally derived heterochromatic chromosomes (recognizable by the intense DAPI staining). At late metaphase, the HP2 antibody signal is still preferentially associated to five chromosomes, presumably the paternal heterochromatic ones (the leftmost five chromosomes in Fig. 2c). The five maternal euchromatic chromosomes show preferential HP2 antibody staining at telomeric regions (arrows) throughout metaphase stages, except for one chromosome that is labeled only at an interstitial site (arrowheads).
D. melanogaster anti-HP2 antibody recognizes a conserved epitope in mealybug nuclei. a Western blot of protein extracts from D. melanogaster and P. citri probed with the D. melanogaster anti-HP2 antibody as primary antibody. As expected, in Drosophila two bands are evidenced (Stephens et al. 2005). In addition, in mealybug lane, two isoforms are evidenced: one with a mobility very similar to the D. melanogaster HP2-L and the other with an apparent MW around 200 kDa. M indicates the molecular weight marker. b–d Immunostaining of P. citri cytological preparations with the D. melanogaster anti-HP2 antibody as primary antibody. Left: DAPI staining. Middle: anti-HP2 antibody immunostaining. Right: merged image, with DAPI fluorescence in red and HP2 immunolabeling in green. In b and c, male interphase nuclei are shown: note the prominent colocalization of the HP2 antibody signal with the DAPI-bright chromocenters. The localization of the HP2 antibody with the chromocenter is particularly apparent in the upper nucleus in c where the chromocenter is splitted in two (arrows). In d, female interphase nuclei are shown, with the HP2 immunofluorescence displaying a scattered distribution over the chromatin. Scale bar: 10 μm
HP2 antibody pattern in mitotic male cells. Left: DAPI staining. Middle: anti-HP2 immunostaining. Right: merged image, with DAPI fluorescence in red and HP2 immunostaining in green. a–c Chromosomes from male nuclei in early (a), middle (b), and late (c) metaphase. The HP2 antibody signal is preferentially concentrated over heterochromatic chromosomes (identified in a and b because of their higher DAPI intensity), whereas the five euchromatic chromosomes show the presence of HP2 labeling at interstitial (arrowheads) and telomeric (arrows) sites. Note that, in late metaphase (c), where the differentiation between euchromatic and heterochromatic chromosomes by DAPI staining is no longer apparent, five chromosomes show a heavier HP2 labeling than the other five ones. Scale bar: 10 μm
In female mitotic cells, where all the chromosomes have an euchromatic appearance, the HP2 antibody signal is localizes preferentially at telomeres (Fig. 3a,b). At late metaphase, the signal often surrounds as a “cloudy” cap one of the two telomeres of nearly all the chromosomes (Fig. 3b).
HP2 antibody pattern in mitotic female cells. Left: DAPI staining. Middle: anti-HP2 immunostaining. Right: merged image, with DAPI fluorescence in red and HP2 immunostaining in green. The ten euchromatic chromosomes of the female complement are shown at early (a) and late (b) metaphase. HP2 antibody signals are telomeric and interstitial in a and mostly telomeric in b, where the signal appears as a sort of “cloud” that covers one of the telomeres of most chromosomes. Scale bar: 10 μm
From immunoblotting and immunolocalization studies, we infer that a HP2 cross-reactive protein there exists in P. citri, which preferentially targets the male-specific facultative heterochromatin and telomeres.
HP2 antibody signal localizes with HP1 and its appearance precedes the onset of heterochromatinization
In a preceding paper, we showed that a HP1-like protein is present in mealybugs and plays a role in facultative heterochromatin formation (Bongiorni et al. 2001). To investigate a possible relationship between HP1-like and the presumptive HP2-like, we set up coimmunolocalization experiments in which embryo cytological preparations were simultaneously labeled with antibodies against Drosophila HP1 and HP2. In male interphase nuclei, HP1 and HP2 antibodies colocalize with each other and with the paternally derived heterochromatic set (Fig. 4a). However, the HP2 antibody signal consistently appears more extensive than that of the HP1-like.
Comparison of HP1-like and HP2 antibody patterns in mealybug cells. a Male interphase cells. From left to right: DAPI staining, anti-HP1 immunostaining, anti-HP2 immunostaining, and merged image with DAPI in blue, HP1 immunolabeling in green, and HP2 immunolabeling in red. HP1 and HP2 antibody signals show a remarkable coincidence with each other and with the chromocenter (see merged image). Note that the HP2-like signal is more diffused over the euchromatin than that of the HP1-like signal. b Female prometaphase: DAPI staining. In c, the anti-HP1- (green) and anti-HP2-immunolabeling (red) of the same prometaphase is shown. Note in the insert, a magnified view of a detail of the chromosomes showing the nonperfect colocalization of the two proteins. Scale bars: 10 μm
In female chromosomes, HP1-like and HP2 antibody signals are concentrated, although not fully coincident, over telomeric or subtelomeric regions (Fig. 4b, see insert). Furthermore, in this case, the HP2 antibody signals seem more diffused than those of the HP1-like.
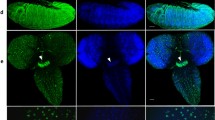
The timing of HP2-like appearance in relation to heterochromatinization onset was analyzed by immunostaining of P. citri embryos at different developmental stages. In a previous work, we showed that in male embryos, the heterochromatinization process occurs like a wave proceeding from one pole of the embryo toward the other one, between the seventh and the eighth nuclear divisions. We also showed that the appearance of the HP1-like protein precedes the appearance of a cytologically differentiated chromocenter (Bongiorni et al. 2001). At this embryo stage, it is thus possible to observe in a single embryo, some nuclei with a fully developed chromocenter (Fig. 5c, DAPI staining) and others with no evidence of a chromocenter (Fig. 5a,b, DAPI staining) or with chromocenters in formation that appear as slightly differentiated chromatin regions (arrows in Fig. 5a,b, DAPI staining). In all the nuclei that have undergone heterochromatinization, not only the HP1- but also the HP2-immunostaining signal is already present and colocalizes with the chromocenters (Fig. 5c). However, in nuclei with no cytologically apparent or only slightly differentiated heterochromatin, the HP1 antibody signal forms in all cases conspicuous spots (Fig. 5a,b, HP1) over still undifferentiated chromatin regions or over the chromocenter rudiments (Fig. 5a,b, merged), whereas the HP2-immunostaining shows two different patterns. In some nuclei, the HP2 antibody signals exhibit a more dispersed pattern than that of the HP1-like (Fig. 5a, HP2 and merged), whereas in other nuclei, also the HP2 antibody signals appear concentrated in a single conspicuous spot that localizes with the HP1-like (Fig. 5b, HP2 and merged).
HP1-like and HP2 antibody patterns in male embryos at the onset of facultative heterochromatinization (128–256 nuclei stage). Left: Male embryo at onset of heterochromatin formation that occurs like a wave starting from one pole of the embryo (sector c) toward the other one (sector a). In a–c, from the left to the right: DAPI staining, anti-HP1 immunostaining, anti-HP2 immunostaining, and merged image with DAPI in blue, and anti-HP1 and anti-HP2 fluorescence in green and red, respectively. a, b Two embryo sectors where most of the nuclei do not exhibit a fully mature chromocenter (arrows). Both HP1 and HP2 antibody signals are already present: HP1-like forms a single conspicuous spot in a and b, whereas HP2 antibody signal is dispersed in a and aggregated in a single spot, coincident with that of the HP1-like signal in b. The nuclei in c exhibits a mature chromocenter heavily stained by both anti-HP1 and anti-HP2 antibodies. Scale bar: 10 μm
During heterochromatin reversion, the HP2 antibody signal becomes delocalized
In P. citri adult males, the nuclei of some tissues, such as the skeletal muscles, the intestinal tract, and the Malpighian tubules, lack the heterochromatic set as a result of the reversal of heterochromatinization (Nur 1967). In these tissues, it is thus possible to follow the unique process of deheterochromatinization.
We immunostained gut tissue cytological preparations from third instar males with the antibody anti-HP2. In the hindgut, we can observe patches of tissue in which some cells have undergone reversal of heterochromatinization and some others have not (Epstein et al. 1992). Figure 6 shows patches of the same hindgut tissue displaying three categories of nuclei: nuclei where the reversal of heterochromatinization was completed (Fig. 6a, DAPI staining, arrowheads), nuclei that still retain a distinct chromocenter (Fig. 6a, DAPI staining, arrows), and nuclei where the reversal of heterochromatinization is in progress (Fig. 6b, DAPI staining). In these latter nuclei, the facultative heterochromatin lost its high degree of compaction, and consequently, a DAPI-bright chromocenter is no longer visible, although chromocenter remnants are somewhat still recognizable (Fig. 6b, DAPI staining, arrows). Both HP1 and HP2 antibody signals are localized with the male specific heterochromatin in nuclei that have not yet reverted (Fig. 6a, arrows), although they are diffused over the chromatin of nuclei that already completed the reversal of heterochromatinization (Fig. 6a, arrowheads). However, in nuclei where the reversal of heterochromatinization is still in progress, a different behavior of the two proteins can be observed (Fig. 6b). HP1-like signals retain a weak concentration over the chromocenter remnants, although they are mainly scattered throughout the whole chromatin (Fig. 6b, HP1), whereas HP2 signals do not show any preferential localization (Fig. 6b, HP2).
HP1-like and HP2 antibody patterns in adult male tissues where the reversal of heterochromatinization occurs. From left to right: DAPI staining, anti-HP1 immunostaining, anti-HP2 immunostaining, and merged image, with DAPI in blue, anti-HP1 in green, and anti-HP2 in red. a A mosaic tissue with some nuclei that have undergone reversal of heterochromatinization (arrowheads; note the absence of chromocenter) and some that have not (arrows indicate the presence of the chromocenter). Both HP1-like and HP2 antibody signals are dispersed throughout the chromatin of nuclei that already underwent reversion. b Some nuclei where reversal of heterochromatinization is in progress. Note the loss of DAPI-brightness of chromocenter remnants (indicated by arrows). HP1 immunofluorescence is still mainly associated to the chromocenter remnants, whereas HP2 immunofluorescence is scattered throughout the chromatin. Scale bar: 10 μm
dsHp1-like RNAi does not alter HP2 antibody heterochromatic pattern
In Drosophila, HP2 was isolated as HP1 interacting protein in a yeast two-hybrid assay and in coprecipitation experiments of embryo extracts (Shaffer et al. 2002). The P. citri gene encoding for HP1-like is named pchet2 (Epstein et al. 1992; Bongiorni et al. in preparation). To investigate a possible interaction between HP1-like and the presumptive HP2-like in mealybugs, we analyzed the HP2-like presence and pattern in cytological preparations from embryos treated for RNAi with the dsRNA of pchet2. The RNAi experiments show that the HP1-like shortage does not influence the presence and pattern of HP2 antibody signals, which are still abundant over slightly damaged chromocenters, whereas the HP1-like immunostaining is, as expected, abolished (Fig. 7). In parallel experiments, the HP1-like depletion in protein extracts from pchet2 RNAi-treated embryos was demonstrated by Western blot (data not shown).
HP2 antibody pattern in HP1-like knockout embryos. A sector of a male embryo depleted of HP1 by means of Hp1 dsRNA interference. From left to right: DAPI staining, anti-HP1 immunostaining, anti-HP2 immunostaining, and merged image with DAPI in blue, HP1 immunolabeling in green, and HP2 immunolabeling in red. Note the absence of HP1 antibody signal whereas the HP2 antibody signal is still present and exhibits a wild-type pattern. Scale bar: 10 μm
Discussion
The coccid chromosome system offers a powerful tool for gaining insights into the structure of facultative heterochromatin and the epigenetic mechanisms of its developmentally regulated formation. In previous studies, we showed that a HP1-like protein is involved in the regulation of facultative heterochromatinization in coccids (Bongiorni et al. 2001, 2007; Bongiorni and Prantera 2003). In this work, we focused our study to ascertain the presence and the potential role in facultative heterochromatinization in mealybugs of an ortholog of the heterochromatin protein HP2 (Shaffer et al. 2002, 2006; Stephens et al. 2005).
In D. melanogaster, HP2 shows two isoforms (HP2-L and HP2-S, 365 and 176 kDa, respectively), resulting from alternative splicing events (Shaffer et al. 2002; Stephens et al. 2005). The chicken antibody raised against HP2 recognizes both isoforms; the smaller one migrates on SDS-PAGE with an apparent molecular weight of 250 kDa (Stephens et al. 2005). In mealybugs, the HP2 antibodies recognize a cross-reactive nuclear epitope, as shown for both nuclear protein extracts and cytological preparations. The Western blot, probed with the chicken antibody, reveals two protein bands. The larger band migrates as the Drosophila HP2-L, whereas the smaller band migrates with an apparent molecular weight around 200 kDa. In immunocytological experiments, performed with the rabbit antibody, the anti-HP2 signals show a close association with the male-specific heterochromatin. As a whole, these results are consistent with the presence of a HP2-like heterochromatin protein in mealybugs. Hp2 homologs have been identified in D. willistoni, D. virilis, and D. pseudoobscura (Stephens et al. 2005) but not in the sequenced genomes of more distant species, like the honey bee and mosquito. A more extensive molecular analysis is required to determine if the protein identified in coccids by the HP2 antibody is actually an ortholog of the Drosophila HP2. Hence, our designation of the coccid cross reactive epitope as HP2-like must be considered tentative in the absence of sequence verification.
In mealybugs, the enrichment of the HP2-like protein at telomeres of euchromatic homologs in males and, more prominently, at most of the telomeres of female chromosomes, would suggest some telomere functions for HP2 in mealybugs. In D. melanogaster polytene chromosomes, HP2 localizes with telomeres in a HP1-dependent manner (Shaffer et al. 2006). However, differently from HP1 mutants (Fanti et al. 1998; Cenci et al. 2003), D. melanogaster HP2 mutants do not present any evidence of an impaired telomere-capping function in mitotic chromosomes (Shaffer et al. 2006).
In this study, the double immunostaining with the antibodies raised against the D. melanogaster HP1 and HP2 shows the strict colocalization of the two proteins over the chromocenters in male mealybug embryo cells. In the euchromatic regions, however, HP2-like signal is reliably more diffused than that of the HP1-like. These observations are consistent with those reported for Drosophila mitotic chromosomes, where the two heterochromatin proteins colocalize over heterochromatic regions, whereas HP2 only is enriched in euchromatin (Shaffer et al. 2006).
The relationships between HP1-like and HP2-like have been further investigated during male embryo development at the onset of the heterochromatinization process and during deheterochromatinization in adult male tissues. In male mealybugs, the HP1-like protein concentrates in large spots over specific chromatin areas before they acquire the full cytological features of heterochromatin, being thus a candidate agent of facultative heterochromatin assembly (Bongiorni et al. 2001, 2007). In this work, we showed that the epitope recognized by the HP2 antibody accumulates in large blocks in male nuclei, before a mature chromocenter is formed. Whether the HP2-like binding to chromatin is a prerequisite for facultative heterochromatin assembly is an interesting possibility that must be tested by inactivation of Hp2-like. However, HP2 antibody signal aggregates over distinct chromatin areas, which identify the future chromocenters, after they have already been bound by HP1-like. This suggests that the recruitment of HP2-like to the potential heterochromatic domains depends on HP1-like. Nevertheless, in adult tissues where the heterochromatinization reversal occurs, the HP2-like epitope is lost by the chromocenter remnants before HP1-like, which thus seems to be not sufficient per se to anchor HP2-like to chromatin. Moreover, the strict colocalization of HP2-like with the chromocenters is not abolished in Hp1-like knockout embryos. These results are in perfect agreement with those in D. melanogaster, where the pattern of HP2 on mitotic chromosomes is not affected in Hp1 homozygous mutants (Shaffer et al. 2006).
Our results represent the first indication of the involvement of a HP2-like molecule in facultative heterochromatinization. They also suggest that the structural and functional conservation of proteins involved in heterochromatin assembly might extend to HP2.
References
Bannister AJ, Zegerman P, Partridge JF, Miska EA, Thomas JO, Allshire RC, Kouzarides T (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410:120–124
Bongiorni S, Prantera G (2003) Imprinted facultative heterochromatization in mealybugs. Genetica 117:271–279
Bongiorni S, Cintio O, Prantera G (1999) The relationship between DNA methylation and chromosome imprinting in the coccid Planococcus citri. Genetics 151:1471–1478
Bongiorni S, Mazzuoli M, Masci S, Prantera G (2001) Facultative heterochromatization in parahaploid male mealybugs: involvement of a heterochromatin-associated protein. Development 128:3809–3817
Bongiorni S, Fiorenzo P, Pippoletti D, Prantera G (2004) Inverted meiosis and meiotic drive in mealybugs. Chromosoma 112:331–341
Bongiorni S, Pasqualini B, Taranta M, Singh PB, Prantera G (2007) Epigenetic regulation of facultative heterochromatinisation in the mealybug. Planococcus citri via the Me(3)K9H3/HP1/Me(3)K20H4 pathway. J Cell Sci (in press)
Brown SW, Nur U (1964) Heterochromatic chromosomes in the coccids. Science 145:130–136
Cenci G, Siriaco G, Raffa GD, Kellum R, Gatti M (2003) The Drosophila HOAP protein is required for telomere capping. Nat Cell Biol 5:82–84
Cowell IG, Aucott R, Mahadevaiah SK, Burgoyne PS, Huskisson N, Bongiorni S, Prantera G, Fanti L, Pimpinelli S, Wu R, Gilbert DM, Shi W, Fundele R, Morrison H, Jeppesen P, Singh PB (2002) Heterochromatin, HP1 and methylation at lysine 9 of histone H3 in animals. Chromosoma 111:22–36
Eissenberg JC, Elgin SC (2000) The HP1 protein family: getting a grip on chromatin. Curr Opin Genet Dev 10:204–210
Eissenberg JC, Morris GD, Reuter G, Hartnett T (1992) The heterochromatin-associated protein HP-1 is an essential protein in Drosophila with dosage-dependent effects on position-effect variegation. Genetics 13:345–352
Epstein H, James TC, Singh PB (1992) Cloning and expression of Drosophila HP1 homologs from a mealybug, Planococcus citri. J Cell Sci 101:463–474
Fanti L, Giovinazzo G, Berloco M, Pimpinelli S (1998) The heterochromatin protein 1 prevents telomere fusions in Drosophila. Mol Cell 2:527–538
James TC, Elgin SC (1986) Identification of nonhistone chromosomal protein associated with heterochromatin in Drosophila and its gene. Mol Cell Biol 6:3862–3872
Kourmouli N, Jeppesen P, Mahadevhaiah S, Burgoyne P, Wu R, Gilbert DM, Bongiorni S, Prantera G, Fanti L, Pimpinelli S, Shi W, Fundele R, Singh PB (2004) Heterochromatin and tri-methylated lysine 20 of histone H4 in animals. J Cell Sci 117:2491–2501
Lachner M, O’Carroll D, Rea S, Mechtler K, Jenuwein T (2001) Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410:116–120
Lyon MF (1992) Some milestones in the history of X-chromosome inactivation. Annu Rev Genet 26:16–28
Maeda I, Kohara Y, Yamamoto M, Sugimoto A (2001) Large-scale analysis of gene function in Caenorhabditis elegans by high-throughput RNAi. Curr Biol 11:171–176
Nur U (1967) Reversal of heterochromatization and the activity of the paternal chromosome set in the male mealy bug. Genetics 56: 375–389
Nur U (1990) Heterochromatization and euchromatization of whole genomes in scale insects (Coccoidea: Homoptera). Development [Suppl]:29–34
Piacentini L, Fanti L, Berloco M, Perrini B, Pimpinelli S (2003) Heterochromatin protein 1 (HP1) is associated with induced gene expression in Drosophila euchromatin. J Cell Biol 161:707–714
Shaffer CD, Stephens GE, Thompson BA, Funches L, Bernat JA, Craig, CA, Elgin SC (2002) Heterochromatin protein 2 (HP2), a partner of HP1 in Drosophila heterochromatin. Proc Natl Acad Sci USA 99:14332–14337
Shaffer CD, Cenci G, Thompson B, Stephens GE, Slawson EE, Adu-Wusu K, Gatti M, Elgin SCR (2006) The large isoform of Drosophila melanogaster heterochromatin protein 2 plays a critical role in gene silencing and chromosome structure. Genetics 174:1189–1204
Singh PB, Georgatos SD (2002) HP1: facts, open questions, and speculation. J Struct Biol 140:10–16
Singh PB, Miller JR, Pearce J, Kothary R, Burton RD, Paro R, James TC, Gaunt SJ (1991) A sequence motif found in a Drosophila heterochromatin protein is conserved in animals and plants. Nucleic Acids Res 19:789–794
Somma MP, Fasulo B, Cenci G, Cundari E, Gatti M (2002) Molecular dissection of cytokinesis by RNA interference in Drosophila cultured cells. Mol Biol Cell 13:2448–2460
Stephens GE, Slawson EE, Craig CA, Elgin SC (2005) Interaction of heterochromatin protein 2 with HP1 defines a novel HP1-binding domain. Biochemistry 44:13394–13403
Vakoc CR, Mandat SA, Olenchock BA, Blobel GA (2005) Histone H3 lysine 9 methylation and HP1 gamma are associated with transcription elongation through mammalian chromatin. Mol Cell 19:381–391
Wakimoto BT (1998) Beyond the nucleosome: Epigenetic aspects of position effect variegation in Drosophila. Cell 93:321–324
Acknowledgments
We thank Sarah Elgin for supplying us with the HP2 antibody and for an insightful discussion. We greatly appreciated Gena Stephens for helpful suggestions. We are grateful to Barbara Wakimoto for the critical reading of the manuscript and the C1A9 antibody. This work was supported by a grant from the University of Tuscia to GP.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by S. Pimpinelli
Silvia Volpi and Silvia Bongiorni contributed equally to the work.
Rights and permissions
About this article
Cite this article
Volpi, S., Bongiorni, S. & Prantera, G. HP2-like protein: a new piece of the facultative heterochromatin puzzle. Chromosoma 116, 249–258 (2007). https://doi.org/10.1007/s00412-007-0095-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-007-0095-7