Abstract
A recent study reported a negative association between a putatively functional dopamine (DA) polygenic score, indexing higher levels of DA signaling, and depressive symptoms. We attempted to replicate this association using data from the Duke Neurogenetics Study. Our replication attempt was made in a subsample of 520 non-Hispanic Caucasian volunteers (277 women, mean age 19.78 ± 1.24 years). The DA polygenic score was based on the following five loci: rs27072 (SLC6A3/DAT1), rs4532 (DRD1), rs1800497 (DRD2/ANKK1), rs6280 (DRD3), and rs4680 (COMT). Because the discovery sample in the original study consisted mostly of Asian participants, we also conducted a post hoc analysis in a smaller subsample of Asian volunteers (N = 316, 179 women, mean age 19.61 ± 1.32 years). In the primary sample of non-Hispanic Caucasians, a linear regression analysis controlling for sex, age, socioeconomic status (SES), body mass index, genetic ancestry, and both early and recent life stress, revealed that higher DA polygenic scores were associated with higher self-reported symptoms of depression. This was in contrast to the original association of higher DA polygenic scores and lower depressive symptoms. However, the direction of the association in our Asian subsample was consistent with this original finding. Our results also suggested that compared to the Asian subsample, the non-Hispanic Caucasian subsample was characterized by higher SES, lower early and recent life stress, and lower depressive symptoms. These differences may have contributed to the observed divergence in associations. Collectively, the current findings add to evidence that specific genetic associations may differ between populations and further encourage explicit modeling of race/ethnicity in examining the polygenic nature of depressive symptoms and depression.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Multiple lines of evidence implicate dopamine (DA) signaling in the pathophysiology of depression [1, 2]. Accordingly, two recent genome-wide association studies (GWAS) have implicated common sequence variation in DA-related genes in depression [3, 4]. Consistent with this literature, Pearson-Fuhrhop et al. [5] reported a significant association between depressive symptoms and a putatively functional DA polygenic score consisting of common polymorphisms across five different DA-related genes. Specifically, they found that higher DA polygenic scores were significantly associated with lower depressive symptoms. As higher DA polygenic scores putatively index higher in vivo DA signaling, the direction of their reported association is consistent with some earlier research linking relatively low DA with depressive symptoms, such as anhedonia and amotivation [6, 7].
The current study represents our attempt to replicate the association between higher DA polygenic scores and lower depressive symptoms in the Duke Neurogenetics Study (DNS). To avoid population stratification biases and achieve maximum statistical power, we focused our analyses on the largest ethnic subsample in the DNS, non-Hispanic Caucasians (N = 520). However, as the original association reported by Pearson-Fuhrhop et al. [5] emerged from a discovery sample (N = 273) with a preponderance of Asian participants (49%), we also present findings from an exploratory post hoc analysis in a small subsample of 316 students of Asian descent.
Methods and materials
Participants
Study participants included a subset of individuals having completed the DNS, which was designed to identify biomarkers of risk for psychopathology amongst 18- to 22-year-old full-time university students. Participants were recruited through posted flyers on the Duke University campus and through a Duke University listserv. All procedures were approved by the Institutional Review Board of the Duke University Medical Center, and participants provided informed consent before study initiation. All participants were free of the following study exclusions: (1) medical diagnoses of cancer, stroke, diabetes requiring insulin treatment, chronic kidney or liver disease, or lifetime history of psychotic symptoms; (2) use of psychotropic, glucocorticoid, or hypolipidemic medication; and (3) conditions affecting cerebral blood flow and metabolism (e.g., hypertension).
Our primary sample consisted of 520 self-reported non-Hispanic Caucasian participants (277 women, mean age 19.78 ± 1.24 years) from the DNS for whom there were complete data on genotypes, depressive symptoms, and all covariates. Exploratory post hoc analyses were performed on a smaller Asian subsample (N = 316, 179 women, mean age 19.61 ± 1.32 years) from the DNS, also with complete data on study variables.
Current and lifetime DSM-IV axis I or select axis II disorders (antisocial personality disorder and borderline personality disorder) were assessed with the electronic Mini International Neuropsychiatric Interview [8] and Structured Clinical Interview for the DSM-IV axis II subtests [9]. Of the 520 non-Hispanic Caucasians included in our analyses, 117 individuals had at least one DSM-IV diagnosis. Of the 316 Asian participants, 42 individuals had at least one DSM-IV diagnosis. Importantly, neither current nor lifetime diagnosis was an exclusion criterion, as the DNS sought to establish broad variability in multiple behavioral phenotypes related to psychopathology. However, no participants, regardless of diagnosis, were taking any psychoactive medication during or at least 14 days prior to their participation.
Socioeconomic status (SES)
We controlled for possible SES effects using the “social ladder” instrument [10], which asks participants to rank themselves relative to other people in the United States (or their origin country) on a scale from 0 to 10, with people who are best off in terms of money, education, and respected jobs, at the top (10) and people who are worst off at the bottom (0).
Genetic ancestry
To further minimize the risk of population stratification, an analysis of identity-by-state of whole-genome single nucleotide polymorphisms (SNPs) was performed in PLINK [11]. Multidimensional scaling was done separately for the two subgroups included in the current analyses (i.e., non-Hispanic Caucasians and Asians), and the first two components for each group were used in each respective analysis as covariates.
Depressive symptoms
The 20-item Center for Epidemiologic Studies Depression Scale (CES-D) was used to asses depressive symptoms in the past week [12]. All items were summed to create a total depressive symptoms score.
Life stress
We controlled for the effects of early life stress using the Childhood Trauma Questionnaire [CTQ; 13], and of stress during the year prior to study participation using the Life Events Scale for Students [LESS; 14]. All items from the CTQ were summed to create a total score of early life stress. For the LESS, we summed the 38 negatively valenced items for a total score of recent stress [as described in 15, 16].
Body mass index (BMI)
BMI was calculated based on the height and weight of the participants (pounds/inches2 × 703). We included BMI as a covariate because it is a known risk factor for depression [17].
Genotyping
DNA was isolated from saliva using Oragene DNA self-collection kits (DNA Genotek) customized for 23andMe (www.23andme.com). Genotyping was done through 23andMe which uses the sevices/labs of the National Genetics Institute (NGI), a CLIA-certified clinical laboratory and subsidiary of Laboratory Corporation of America. One of the two different Illumina arrays with custom content was used to provide genome-wide SNP data, the HumanOmniExpress (N = 524) or HumanOmniExpress-24 [N = 276; 18–20]. We performed a K-means analysis based on the first two multidimensional scaling components of the Asian participants, dividing them into two subgroups. Hardy–Weinberg equilibrium (HWE) tests were performed in PLINK [11] on the created Asian groups and on the non-Hispanic Caucasians separately. All SNPs were in HWE (p > 0.05).
Dopamine polygenic scores
As per the strategy of Pearson-Fuhrhop et al. [5], we derived a DA polygenic score for each participant based on the following five loci: rs27072 (SLC6A3/DAT1), rs4532 (DRD1), rs1800497 (DRD2/ANKK1), rs6280 (DRD3), and rs4680 (COMT). Scoring details are provided in Table 1. Of note, we used a putatively functional SNP in SLC6A3/DAT1 instead of the variable number tandem repeat (VNTR) used in the original study, because VNTRs were not genotyped in our sample.
Statistical analyses
Descriptive statistics and univariate analyses of variance were conducted using SPSS version 24. Mplus version 7 [21] was used to conduct a linear regression analysis with participants’ sex (coded as 0 = males, 1 = females), age, two genetic ancestry principal components, SES, BMI, and both early and recent life stress, as covariates. Maximum likelihood estimation with bias-corrected bootstrapping (set to 10,000) was used in all models to generate non-symmetric 95% confidence intervals (CIs).
Results
Allele frequencies and descriptive statistics for the non-Hispanic Caucasian and Asian subsamples are presented in Tables 1 and 2, respectively. The allele frequencies are comparable to those observed in the 1000 genomes sample for Europeans and East Asians.
DA polygenic scores and depressive symptoms
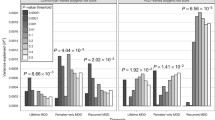
Of the covariates, only early life stress (b = 0.24, SE = 0.046, p < 0.001; CI 0.15–0.33) and recent life stress (b = 0.67, SE = 0.15, p < 0.001; CI 0.39–0.96), significantly predicted depressive symptoms; so, that higher stress levels were associated with higher depressive symptoms. The DA polygenic score significantly and positively predicted depressive symptoms (b = 0.43, SE = 0.20, p = 0.03; CI 0.05–0.82; Fig. 1a). The R2 of the model was 0.166. Comparing the R2 of the model with the DA polygenic score to the R2 of a model without it indicated that the variance explained by the polygenic score was small (R2 = 0.007).
Dopamine polygenic scores are differentially associated with depressive symptoms in a non-Hispanic Caucasians and b Asians. To better depict the linear regression described in the text, the standardized residuals of depressive symptoms, as assessed by the CES-D, are shown (controlling for sex, age, two multidimensional scaling ancestry components, SES, BMI, and early and recent life stress)
Exploratory post hoc analyses in the Asian subsample
Since Pearson-Fuhrhop et al. [5] used a discovery sample with a majority of Asian participants, we also examined the association in a smaller Asian subsample as a post hoc analysis. Descriptive statistics for the Asian participants are presented in Table 2. Comparisons of the non-Hispanic Caucasian and Asian participants on the main study variables pointed to many differences between the groups. Non-Hispanic Caucasians were more likely to have a higher SES [F(1, 834) = 15.47, p < 0.001], less likely to experience early life stress [F(1, 834) = 44.65, p < 0.001], more likely to have a higher BMI [F(1, 834) = 6.21, p = 0.013], less likely to experience depressive symptoms [F(1, 834) = 9.92, p = 0.002], somewhat more likely to experience recent stress [F(1, 834) = 3.74, p = 0.054], and somewhat more likely to be older [F(1, 834) = 3.74, p = 0.054]. Lastly, the non-Hispanic Caucasian participants had higher DA polygenic scores than the Asian participants [F(1, 834) = 67.93, p < 0.001], suggesting differences in allele frequencies between populations.
In the Asian subsample, similar to the primary non-Hispanic Caucasian subsample, early and recent life stress were significant predictors of depressive symptoms (early life stress: b = 0.33, SE = 0.072, p < 0.001, CI 0.20–0.48; recent life stress: b = 0.54, SE = 0.17, p < 0.001, CI 0.21–0.90). However, in contrast to the non-Hispanic Caucasian subsample, SES also significantly predicted depressive symptoms (b = − 0.80, SE = 0.29, p = 0.006, CI − 1.39 to − 0.24), so that individuals from lower SES experienced higher levels of depressive symptoms. Lastly, in the Asian subsample, the association between the DA polygenic scores and depressive symptoms was significant and negative (b = − 0.68, SE = 0.34, p = 0.047; CI − 1.38 to − 0.021), so that higher DA polygenic scores predicted lower levels of depressive symptoms (Fig. 1b). The R2 of the model was 0.214. Comparing the R2 of the model with the DA polygenic score to the R2 of a model without it indicated that the variance explained by the polygenic score was small (R2 = 0.012).
The effect of each SNP is presented in Table 3 for both the non-Hispanic Caucasian and Asian subsamples. In the non-Hispanic Caucasian subsample, the SNP in the SLC6A3/DAT1 gene (rs27072) was significant (b = 1.62, SE = 0.63, p = 0.01; CI 0.50–3.001), explaining 0.015 of the variance. In the Asian subsample, the SNP in the DRD1 gene (rs4532) was significant (b = − 1.83, SE = 0.80, p = 0.022; CI − 3.49 to − 0.36), explaining 0.013 of the variance.
Discussion
In the current study, we attempted to replicate earlier work by Pearson-Fuhrhop et al. [5], reporting an association between higher DA polygenic scores and lower levels of depressive symptoms. While we did find a significant association between DA polygenic scores and depressive symptoms in our primary sample of non-Hispanic Caucasians, it was in the opposite direction to the one reported in the original study: higher DA polygenic scores were associated with higher depressive symptoms. However, the pos hoc analysis in our smaller subsample of Asian participants revealed an association consistent with that of the original report: higher DA polygenic scores were associated with lower depressive symptoms. When each SNP was examined separately, there were specific loci that showed an independent significant effect on depressive symptoms, but these differed between our subsamples as well as between our study and the original study, indicating that individual SNP effects are less reliable, as replication studies of single candidate gene associations have previously shown [e.g., 22]. Lastly, the variance explained by the DA polygenic scores in both of our models (non-Hispanic Caucasians R2 = 0.007, Asians R2 = 0.012) was smaller than what was reported in the original discovery sample (R2 = 0.085). Overestimating the true effect in discovery samples and finding smaller effect sizes in replication samples is a common observation usually explained as a regression to the mean [23].
The potential moderation of an association between DA polygenic scores and depressive symptoms is not inconsistent with the original report of Pearson-Fuhrhop et al. [5]. In that report, there was a successful replication, albeit with a DA polygenic score that only included three of the original variants, in a sample of 1267 adult Caucasians having a clinical diagnosis of non-psychotic MDD. However, a second replication attempt with a DA polygenic score comprising four of the original variants, in a sample of healthy 381 young adult Caucasians, was not successful. Thus, the link between DA polygenic scores and depressive symptoms may be moderated by clinical diagnosis and age. Indeed, the presence of a clinically significant disease has been shown to moderate the effect of DA-related genes [24]. Additionally, there is an age-related decline in DA signaling [25], which could contribute to differences in the effects of dopaminergic genes on behavior across development.
Our findings, although exploratory, further suggest that race/ethnicity may also moderate the link between DA polygenic scores and depressive symptoms. This observation is in line with previous findings demonstrating population differences in the effects of DA-related polymorphisms. For example, in a meta-analysis of 30 studies [26], the putatively functional 7-repeat allele of the dopamine receptor D4 exon III variable nucleotide tandem repeat was positively linked to attention deficit hyperactivity disorder (ADHD) in European Caucasian and South American samples, but negatively linked to ADHD in Middle-Eastern samples. Similarly, the Met allele of the catechol-o-methyltransferase rs4680 has been associated with larger hippocampal volume in Caucasians [27], but smaller hippocampal volume in Han Chinese [28]. Theoretically, such differences can reflect variation in allele frequencies, linkage disequilibrium (LD) patterns, and/or non-genetic (e.g., cultural) effects. However, allele frequencies may be less likely to explain the current observed differences, because the effect in our non-Hispanic Caucasian subsample was not stronger, as may be expected, but reversed. It is possible that some of the examined SNPs only tag causal polymorphisms and consequently LD differences may affect the observed findings. Additionally, our two groups differed in stress levels and socioeconomic background, which suggest cultural and experience-related influences that could potentially moderate the genetic effects (e.g., through epigenetic modifications that affect gene expression [29] and/or through modifications of brain structure and function [30]). Indeed, previous studies have demonstrated that culture can act as a moderator of genetic influences [31, 32]. Nevertheless, as with any association study, replication across different age groups and populations, and in clinical and healthy samples is necessary.
Our study has several limitations. First, we were not able to directly assess the functionality of the DA polygenic score and establish links with in vivo DA signaling. Although previous studies have demonstrated the functionality of each individual SNP, research that validates the functionality of the polygenic score is needed. Second, the analysis in the Asian subsample was done post hoc and should be treated with caution. Third, we did not have a large sample of Asians nor were we able to examine other racial/ethnic groups. Fourth, the current polygenic score is limited to five SNPs and future studies may choose to extend it by implementing findings from open resources such as the Genotype-Tissue Expression project [33]. Notably, however, here we were specifically interested in attempting to replicate the finding of Pearson-Fuhrhop et al. [5]. Lastly, our findings should be viewed as tentative until further replication is reported.
In studies seeking mechanistic insight, polygenic scores similar to the one used in the current study may be preferable to either single candidate genes, which often fail to replicate [e.g., 22], or to GWAS-based scores, which may explain more phenotypic variance, but are often limited in providing insight into the molecular mechanisms underlying the phenotype of interest [34]. If replicated, our results suggest that efforts to discover the biological mechanisms underlying depression and identify treatment targets should consider race/ethnicity as a moderator. Previous studies have indeed shown that racial/ethnic differences can affect treatment response to various diseases [35,36,37,38].
References
Grace AA (2016) Dysregulation of the dopamine system in the pathophysiology of schizophrenia and depression. Nat Rev Neurosci 17:524
Belujon P, Grace AA (2017) Dopamine system dysregulation in major depressive disorders. Int J Neuropsychopharmacol 20:1036–1046
Howard DM, Adams MJ, Shirali M et al (2018) Genome-wide association study of depression phenotypes in UK biobank identifies variants in excitatory synaptic pathways. Nat Commun 9:1470
Wray NR, Ripke S, Mattheisen M et al (2018) Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat Genet 50:668–681
Pearson-Fuhrhop KM, Dunn EC, Mortero S et al (2014) Dopamine genetic risk score predicts depressive symptoms in healthy adults and adults with depression. PLoS One 9:e93772
Wise R (2004) Dopamine, learning and motivation. Nat Rev Neurosci 5:483–494
Wise R (2008) Dopamine and reward: the anhedonia hypothesis 30 years on. Neurotox Res 14:169–183
Lecrubier Y, Sheehan DV, Weiller E et al (1997) The mini international neuropsychiatric interview (mini). A short diagnostic structured interview: reliability and validity according to the CIDI. Eur Psychiatry 12:224–231
First MB, Gibbon M, Spitzer RL et al (1997) Structured clinical interview for DSM-IV axis II personality disorders, (SCID-II). American Psychiatric Press, Washington, DC
Adler NE, Epel ES, Castellazzo G et al (2000) Relationship of subjective and objective social status with psychological and physiological functioning: preliminary data in healthy, white women. Health Psychol 19:586
Purcell S, Neale B, Todd-Brown K et al (2007) Plink: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81:559–575
Radloff LS (1977) The CES-D scale: a self-report depression scale for research in the general population. Appl Psychol Meas 1:385–401
Bernstein DP, Stein JA, Newcomb MD et al (2003) Development and validation of a brief screening version of the childhood trauma questionnaire. Child Abuse Negl 27:169–190
Clements K, Turpin G (1996) The life events scale for students: validation for use with british samples. Pers Individ Differ 20:747–751
Avinun R, Nevo A, Knodt AR et al (2017) Reward-related ventral striatum activity buffers against the experience of depressive symptoms associated with sleep disturbances. J Neurosci 37:9724–9729
Nikolova YS, Bogdan R, Brigidi BD et al (2012) Ventral striatum reactivity to reward and recent life stress interact to predict positive affect. Biol Psychiatry 72:157–163
Johnston E, Johnson S, McLeod P et al (2004) The relation of body mass index to depressive symptoms. Can J Public Health 95:179–183
Do CB, Tung JY, Dorfman E et al (2011) Web-based genome-wide association study identifies two novel loci and a substantial genetic component for parkinson’s disease. PLoS Genet 7:e1002141
Eriksson N, Macpherson JM, Tung JY et al (2010) Web-based, participant-driven studies yield novel genetic associations for common traits. PLoS Genet 6:e1000993
Tung JY, Do CB, Hinds DA et al (2011) Efficient replication of over 180 genetic associations with self-reported medical data. PLoS One 6:e23473
Muthén LK, Muthén BO (2007) Mplus user’s guide. Muthén & Muthén, Los Angeles
Avinun R, Nevo A, Knodt AR et al (2018) Replication in imaging genetics: the case of threat-related amygdala reactivity. Biol Psychiatry 84:148–159
van Aert RC, van Assen MA (2018) Examining reproducibility in psychology: a hybrid method for combining a statistically significant original study and a replication. Behav Res Methods 50:1515–1539
Savitz J, Hodgkinson CA, Martin-Soelch C et al (2013) DRD2/ANKK1 Taq1A polymorphism (rs1800497) has opposing effects on d2/3 receptor binding in healthy controls and patients with major depressive disorder. Int J Neuropsychopharmacol 16:2095–2101
Karrer TM, Josef AK, Mata R et al (2017) Reduced dopamine receptors and transporters but not synthesis capacity in normal aging adults: a meta-analysis. Neurobiol Aging 57:36–46
Nikolaidis A, Gray JR (2009) ADHD and the DRD4 exon III 7-repeat polymorphism: an international meta-analysis. Soc Cogn Affect Neurosci 5:188–193
Cerasa A, Gioia MC, Labate A et al (2008) Impact of catechol-o-methyltransferase Val108/158 met genotype on hippocampal and prefrontal gray matter volume. Neuroreport 19:405–408
Wang Y, Li J, Chen C et al (2013) Comt rs4680 met is not always the ‘smart allele’: Val allele is associated with better working memory and larger hippocampal volume in healthy chinese. Genes Brain Behav 12:323–329
Klengel T, Binder EB (2015) Epigenetics of stress-related psychiatric disorders and gene × environment interactions. Neuron 86:1343–1357
Belujon P, Grace AA (2015) Regulation of dopamine system responsivity and its adaptive and pathological response to stress. Proc R Soc B Biol Sci 282:20142516
Kim HS, Sherman DK, Mojaverian T et al (2011) Gene–culture interaction: oxytocin receptor polymorphism (OXTR) and emotion regulation. Soc Psychol Pers Sci 2:665–672
Avinun R, Davidov M, Mankuta D et al (2018) Predicting the use of corporal punishment: child aggression, parent religiosity, and the BDNF gene. Aggress Behav 44:165–175
GTEx Consortium (2013) The genotype-tissue expression (GTEX) project. Nat Genet 45:580
Boyle EA, Li YI, Pritchard JK (2017) An expanded view of complex traits: from polygenic to omnigenic. Cell 169:1177–1186
Ray LA, Bujarski S, Chin PF et al (2011) Pharmacogenetics of naltrexone in Asian Americans: a randomized placebo-controlled laboratory study. Neuropsychopharmacology 37:445
Kato M, Zanardi R, Rossini D et al (2009) 5-HT2A gene variants influence specific and different aspects of antidepressant response in japanese and italian mood disorder patients. Psychiatry Res 167:97–105
Porcelli S, Fabbri C, Serretti A (2012) Meta-analysis of serotonin transporter gene promoter polymorphism (5-HTTLPR) association with antidepressant efficacy. Eur Neuropsychopharmacol 22:239–258
Shen H, He MM, Liu H et al (2007) Comparative metabolic capabilities and inhibitory profiles of CYP2D6. 1, CYP2D6. 10, and CYP2D6. 17. Drug Metab Dispos 35:1292–1300
Huang W, Ma JZ, Payne TJ et al (2008) Significant association of DRD1 with nicotine dependence. Hum Genet 123:133–140
Ota VK, Spindola LN, Gadelha A et al (2012) Drd1 rs4532 polymorphism: a potential pharmacogenomic marker for treatment response to antipsychotic drugs. Schizophr Res 142:206–208
Ferrari M, Comi C, Marino F et al (2016) Polymorphisms of dopamine receptor genes and risk of visual hallucinations in Parkinson’s patients. Eur J Clin Pharmacol 72:1335–1341
Thompson J, Thomas N, Singleton A et al (1997) D2 dopamine receptor gene (DRD2) taq1 a polymorphism: reduced dopamine d2 receptor binding in the human striatum associated with the a1 allele. Pharmacogenetics 7:479–484
Noble E, Blum K, Ritchie T et al (1991) Allelic association of the d2 dopamine receptor gene with receptor-binding characteristics in alcoholism. Arch Gen Psychiatry 48:648–654
Stice E, Spoor S, Bohon C et al (2008) Relation between obesity and blunted striatal response to food is moderated by TaqIA A1 allele. Science 322:449–452
Jeanneteau F, Funalot B, Jankovic J et al (2006) A functional variant of the dopamine d3 receptor is associated with risk and age-at-onset of essential tremor. Proc Natl Acad Sci USA 103:10753–10758
Savitz J, Hodgkinson CA, Martin-Soelch C et al (2013) The functional DRD3 ser9gly polymorphism (rs6280) is pleiotropic, affecting reward as well as movement. PLoS One 8:e54108
Lachman HM, Papolos DF, Saito T et al (1996) Human catechol-o-methyltransferase pharmacogenetics: description of a functional polymorphism and its potential application to neuropsychiatric disorders. Pharmacogenetics 6:243–250
Dreher J-C, Kohn P, Kolachana B et al (2009) Variation in dopamine genes influences responsivity of the human reward system. Proc Natl Acad Sci USA 106:617–622
Pinsonneault JK, Han DD, Burdick KE et al (2011) Dopamine transporter gene variant affecting expression in human brain is associated with bipolar disorder. Neuropsychopharmacology 36:1644–1655
Acknowledgements
The authors would like to thank the Duke Neurogenetics Study participants and the staff of the Laboratory of NeuroGenetics, especially Annchen R. Knodt. The Duke Neurogenetics Study was supported by Duke University as well as US-National Institutes of Health Grants R01DA033369 and R01DA031579. RA and ARH received further support from US-National Institutes of Health Grant R01AG049789.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing financial or other interests.
Rights and permissions
About this article
Cite this article
Avinun, R., Nevo, A., Radtke, S.R. et al. Divergence of an association between depressive symptoms and a dopamine polygenic score in Caucasians and Asians. Eur Arch Psychiatry Clin Neurosci 270, 229–235 (2020). https://doi.org/10.1007/s00406-019-01040-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00406-019-01040-x