Abstract
Objectives
To investigate the alterations in gene profile of placenta from pregnant women with intrahepatic cholestasis of pregnancy (ICP) and to enhance the insight of etiology and pathogenesis of ICP.
Methods
Ten pregnant women diagnosed ICP were recruited and 10 healthy pregnant women served as control. Four samples were taken from each placenta and RNA was isolated. Gene expression was analyzed with microarray and real time PCR was used to validate the differentially expressed genes.
Results
392 genes were found differentially expressed. Among these differentially expressed genes, 280 were up-regulated and 112 were down-regulated. These differentially expressed genes involved 20 categories including genes involved in transportation, cell growth, apoptosis and immune response that were putatively participated the pathogenesis of ICP.
Conclusions
293 differentially expressed genes of 20 categories were found in ICP placenta, suggesting the diversity of gene expression alteration and the complexity of etiology and pathogenesis of ICP.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intrahepatic cholestasis of pregnancy (ICP) is a pregnancy-specific complication in the second and third trimester and is characterized by pruritus, jaundice and the disturbances of liver tests, especially the elevation of bile acid [1, 2]. The maternal prognosis of ICP is usually favorable. Pruritus usually disappears in the first few days following delivery, which was accompanied by normalization of serum bile acid concentrations and other liver tests and the women with ICP generally have no hepatic sequelae [3]. However, ICP poses significant effects on fetus. It increases the risk of preterm labor [4–6], fetal distress [4, 7, 8] and even stillbirth [4, 9, 10]. The etiology and pathogenesis of ICP remain elusive and incompletely understood. It appears to be a multifactorial disease. Hormone factors, genetic predisposition and environmental factor have been thought to be involved in pathogenesis of the disease [1]. Although much work has been done to explore the etiology and pathogenesis, no specific factors, genetic or environmental, were identified.
The role of placenta in the development of ICP has been noticed because the disappearance of pruritits and the normalization of liver tests would occur after the delivery of placenta. In addition, alteration of placental gene expression was reported. Elevated expression of bax and Fas, reduced expression of bcl-2 and FasL and alteration of Bax/Bal-2 [11] and Fas/FasL [12] balance in placenta from ICP women were reported. It was also observed that the expression of tumor necrosis factor (TNF) α [13] were was significantly increased in placenta of ICP women as compared with that in normally pregnant women. Previously, we observed the presence of epidermal growth factor (EGF) receptor [14] in placental trophoblast in ICP and healthy pregnancy and found that the expression of EGFR was significantly decreased in ICP compared with healthy pregnancy. However, there was no high-throughout investigation exploring the gene expression file of ICP placenta.
Microarray technology provides a powerful tool to survey differential expression of gene products [15]. The simultaneous expression screening of thousands of transcripts facilitates the discovery of gene expression patterns that are associated with a specific condition such as that under a disease condition, and may thereby identify transcripts that may be involved in the disease development or may be altered during the disease development [16, 17].
To enhance the insight into the pathogenesis of ICP and the effect of ICP on placental gene expression, we performed a microarray analysis to identify the differentially expressed transcripts that may participate the disease development.
Materials and methods
Human subjects
This investigation was conducted after the approval of the Hospital Ethical Committee and informed consents were obtained from all the participants. All subjects were nulliparous Chinese women with singleton pregnancy. Ten pregnant women who was diagnosed ICP were recruited and ten age-matched healthy pregnant women served as control. Demographic and pregnancy characteristics the recruited subjects were presented in Table 1.
The diagnostic criteria for ICP as described previously [18–20] were (a) the presence of marked pruritus with no rash and quick disappearance after delivery, (b) biochemical evidences of intrahepatic cholestasis, including marked elevation of cholylglycine or total bile acid and mild to moderate elevation of aminotransferases in fasting serum in the second and third trimester and quick recovery after delivery, (c) absence of obstructive gallstone disease as diagnosed clinically and by ultrasonography, (d) absence of viral hepatitis as judged by negative serology to hepatitis A, B, and C, (e) absence of fever or general malaise, and (f) absence of medication that could produce the elevation of bile acid and liver enzymes.
To avoid the confusion of the diseases and multiple pregnancies on the result, the exclusion criteria were multiple pregnancies, chronic hypertension, diabetes mellitus, diseases of heart, lung, and kidney, and other underlying organ diseases. No other obstetric complications existed.
Placental tissue collection
All the placental samples both from ICP and normal pregnancy were obtained immediately after Caesarean sections. To avoid the further effect of labor on the expression profile, only placental samples that were obtained from the women who had not undergone labor were included. Four samples were taken at 0, 3, 6 and 9 o’clock of middle zone of each placenta after the deciduas and amnionic membranes were removed. We then dissected 1 × 1 × 1 cm sections of placental vili between basal and chorionic plates. After vigorous washing of the maternal blood with saline, tissues were immediately frozen in liquid nitrogen and stored at −80°C until assay.
RNA extraction and probe preparation
Total RNA was extracted from placenta tissues with Trizol (Invitrogen, Carlsbad, CA, USA) according to the manufacture’s instructions. The quality of the RNA samples was determined by electrophoresis through denaturing agarose gels and staining with ethidium bromide. The RNA was quantified and evaluated for purity by UV spectrophotometry. RNA was purified using RNeasy (Qiagen, Valencia, CA, USA). Despite the power of high through-put technologies to reveal general patterns of gene expression, variations in gene expression of among individuals must be considered. One potential approach is to make a pool of RNA for each group to reduce the in individual variation [21]. Thus RNA samples of 16 tissue samples from four women were pooled for control group or ICP group. Double stranded cDNA was synthesized using Superscript Choice system (Invitrogen Life Technologies, Carlsbad, CA) and a T7T21 oligonucleotide primer (GenSet, La Jolla, CA). Biotin-labeled RNA was synthesized by in vitro transcription using Enzo Bioarray RNA labeling kit (Enzo Diagnostics, Farmingdale, NY). Labeled cRNA was purified using the Qiagen Rneasy Mini Kit spin Columns.
Hybridization
The Affymetrix HG-U133 plus 2.0 oligonucleotide microarray (Affymetrix, Santa Clara, CA, USA) that consists of 47,000 known transcripts and expressed sequence tags (ESTs), representing approximately 38,500 human genes, was used in the current investigation. Arrays were hybridized, washed, stained, and scanned following the standard Affymetrix protocol. Hybridization was performed at 45°C for 16 h in a hybridization oven with constant rotation (60 rpm). The microarrays were then automatically washed and stained with streptavidinphycoerythrin conjugate (SAPE; Molecular Probes, Eugene, OR) in an Affymetrix Genechip Fluidics Station and fluorescence intensities were scanned with a GeneArray Scanner (Affymetrix).
Data analysis
Data analysis was performed using GCOS software (Affymetrix, Santa Clara, CA, USA). After intensity dependent normalization, the expression levels relative to the control were calculated as a ratio and the expression profile was compared between ICP and control groups. A gene with ratio more than +2.0 or less than −2.0 was considered up- or down-regulated. Each differently regulated gene was classified according to its gene ontology (GO), in which genes were organized into hiearchial categories based on the biological process, molecular function and cellular process. Genesifter software (VizX labs,Seattle,USA) was used to perform the GO analysis.
Real-time PCR
To validate the microarray data, real-time PCR was used to detect the expression of randomly selected genes and glyceraldehydes-3-phosphate dehydrogenase (GAPDH) was used as the internal control to normalize mRNA level. Total RNA was isolated as described above in 40 placental samples obtained from 10 women including those enrolled in microarray. Reverse transcription was performed in a reaction system containing buffer, oligo(dT), dNTP mixture and reverse transcriptase. PCR was conducted in duplicate using the SYBR Green I assay on ABI PRISM 7900 (Perkin-Elmer, Foster city, CA, USA). The cycling conditions were as following: 2 min at 95°C and then 30 s at 95°C, 30 s at 56°C repeats 40. The sequences of primers were listed in Table 2. The expression of detected genes was calculated as previously described. Statistical significance was determined by Student’s t test and P values of <0.05 were regarded as significant.
Results
Clinical characteristics of subjects
Demographic and pregnancy characteristics of each group were presented in Table 1. Maternal ages were similar in ICP patients and controls (26.9 ± 4.38 vs. 27.4 ± 2.91). As expected, ICP women delivered at earlier gestational age than control (35.9 ± 1.91 vs. 39 ± 0.47) and neonatal birth weight were lower in ICP group than in control (3,485 ± 348 vs. 2,775 ± 614, P = 0.005). ICP women had significantly increased level of CG compared with control (307.34 ± 178.60 vs. 4,669.97 ± 1,892, P < 0.001).
Differential expression of genes in placenta of ICP
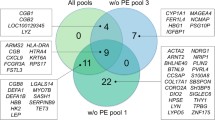
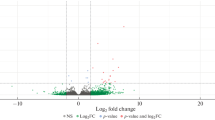
By comparison with the control, the expression of 392 genes was found significantly altered: 280 genes were up- and 112 down-regulated in ICP group. The names and gene IDs of differentially expressed genes were listed in the supporting material.
By performing GO analysis, the differentially expressed genes were categorized into 20 categories according to their biological process. As listed in Table 3, cell process, metabolic process, biological regulation, regulation of biological process, multicellular organismal process, developmental process, response to stimulus, biological adhesion, immune system process, growth, reproductive process and reproduction were among the these categories.
Because placental transportation abnormality, growth dysfunction and apoptosis, as well as immune maladaptation and disordered lipid metabolism were proposed to be important pathophysiology of ICP and were possibly involved in the pathogenesis of disease, differentially expressed genes of these categories were outlined in Table 4.
To verify the results of microarray, the mRNA levels of IL-1RL1, PLSCR-1 and PAEP were analyzed in 40 samples of 10 ICP patients and compared with appropriate controls. We found that the expression of IL-1RL1 and PLSCR-1 were significantly elevated while the expression of PAEP decreased in ICP group compared with that of control P < 0.05 for all, Fig. 1). The change trend was consistent with that found in microassay analysis, which validated the results of microarray to certain degree.
Discussion
By conducting the current investigation, we revealed for the first time the profile of differentially expressed genes in placenta of ICP women. It was found that 280 genes were up-regulated and 122 down-regulated in ICP placenta and that these gene were categorized into 20 categories, indicating that alterations in placenta of ICP patients were multifaceted. Our study provided the evidences that confirmed the complexity of the pathogenesis of ICP and role of placenta in the disease development. To the best of our knowledge, this is the first report of a genome-wide analysis of gene expression in ICP placenta. However, previous studies have documented that abnormal immunity and cell growth, lipid metabolism, placental apoptosis and probably altered material transportation are involved in the disease pathogenesis.
Abnormal immunity has been described in ICP and considered an important pathophysiology of ICP. Previously, we reported that serum neopterin, a marker of monocyte/macrophage activity, and soluble interleukin-2 receptor (sIL-2R), a product of activated T lymphocyte and a quantitative marker of T lymphocyte activity, were significantly increased in ICP women relative to normal pregnancy, which suggested the abnormal activation of both monocyte/macrophage and T lymphocyte and the abnormality of innate and adaptive immunity [22]. Peng and Liu [23] found that the production of Th1-type cytokine by peripheral lymphocyte was increased and that of Th2-type cytokine decreased and that the balance of Th1/Th2 shifted toward Th1 in ICP women. Ling et al. [24] observed increased NK cells and NKT cells, decreased T cells and over-secretion of IFN-γ in the decidual parietalis of ICP women, suggesting the involvement of cell-mediated immunity imbalance in the pathophysiology of ICP. We also found the suppressed mixed lymphocyte reaction between the mother and fetus in ICP setting, implying abnormality of maternal-fetal immunity recognition [20]. These findings point to the importance of altered immunity in the pathophysiology of ICP.
In the current investigation, at least six genes were found differentially expressed in ICP placenta relative to normal placenta. Adenosine deaminase (ADA), “class II, major histocompatibility complex, transactivator” (CIIA), chemokine (C-X-C motif) ligand1 (CXCL1), chemokine (C-X-C motif) ligand 4 (CXCL4) and chemokine (C-X-C motif) ligand 7 (CXCL7) were up-regulated and Interleukin 1 receptor-like 1 (IL1RL1) were down-regulated. Among these genes, the change in the expression of and IL1RL1 was confirmed by real-time PCR.
Adenosine deaminase (ADA) is an enzyme of purine metabolism that catalyzes the deamination of adenosine to inosine and deoxyadenosine to deoxyinosine. ADA is a marker of T-cell activation and considered important for lymphocyte differentiation and growth [25, 26]. IL1RL1, also known as ST2, is a member of the IL-1R family and is expressed preferentially on Th2 effector cells but not on Th1 cells [27–29]. The interaction of IL1RL1 (ST2) and soluble fusion protein ST2L resulted in the suppression of Th2 cell differentiation, suggesting a functional role for ST2L and ST2 in the development of Th2 cells [30]. The up-regulation of ADA and down-regulation of IL1RL1 pointed to abnormality of cell-mediated immunity and the imbalance of Th1/Th2 at the maternal–fetal interface in ICP. Our finding of a group of differentially expressed genes of immunity indicates complex alteration of immunity at maternal–fetal interface.
Abnormal lipid metabolism and altered lipid profile are another important pathophysiology of ICP. It was documented that serum levels of low-density lipoprotein cholesterol (LDL-Ch), apolipoprotein B-100 and total cholesterol were significantly increased while that of high-density lipoprotein cholesterol (HDL-Ch) decreased in ICP compared with normal pregnancy [31]. However, the underlying mechanisms have not been investigated. In the current study, six genes involved in lipid metabolism process were found differentially expressed in the placenta of ICP: LPL (lipoprotein lipase), PLA2G4C (phospholipase A2, group IVC), ELOVL4 (elongation of very long chain fatty acids -like 4), APOD (apolipoprotein D), LIPF (lipase, gastric) and DHCR7 (7-dehydrocholesterol reductase).
DHCR7 is rate-limiting enzyme of cholesterol synthesis. The up-regulation of DHCR7 indicates the increase of cholesterol synthesis. APOD was identified an HDL component and was in association with LCAT, apoA-I and CETP (cholesteryl ester transfer protein), where it could be part of a complex responsible for the transport of cholesterol from peripheral tissues to the liver for its further catabolism [32]. The down-regulation of APOD reduces the transportation and metabolism of cholesterol in peripheral tissues and may lead to the increase of cholesterol in circulation. Our findings of differentially expressed genes involved in lipid metabolism provided evidence that placenta may participate the disordered lipid metabolism in ICP and indicated that the alteration in lipid metabolism is multifaceted.
Previous studies revealed that the expression of vascular endothelial growth factor (VEGF) [33] and epidermal growth factor receptor (EGFR) [14] in placenta was significantly lower in ICP than in normal pregnancy, indicating the altered expression of VEGF and EGFR may be involved in ICP. Microarray analysis did not detect the alteration of VEGF and EGFR expression but detected the up-regulation of insulin-like growth factor binding protein 1 (IGFBP1) and insulin-like growth factor 2 (IGF2) and the down-regulation of nephroblastoma overexpressed gene (NOV).
The insulin-like growth factor (IGF)—IGF-binding protein (IGFBP) system is crucial for placental development and fetal growth. Fetal growth was retarded in Igf 1 -/- or Igf 2 -/- mouse [34, 35]. By binding IGF, which prevents from being degraded or prevents from interacting with it receptor, IGFBP affects IGF activity. NOV, also known as CCN3, is a member of CCN family of angiogenic regulators, that consists of six members (CCN1-6) and that are extracellular matrix proteins involved in the regulation of cellular processes like adhesion, migration, proliferation and differentiation[36, 37]. In early onset pre-eclamptic placenta, the expression of NOV at both mRNA and protein level was significantly decreased compared with normal pregnancy [38].
Chen et al. [39] reported that the apoptotic cell increased and that pro-apoptotic gene bax was up-regulated while anti-apoptotic gene bcl-2 was down-regulated in ICP placenta, resulting in the increase in bax/bcl-2 ratio. Perez [40] found that maternal obstructive cholestasis during pregnancy (OCP) causes apoptosis in rat placenta by demonstrating the increase in the ratio of Bax/Bcl-2 mRNA. In the current study, we found that deoxyribonuclease I-like 3 (DNASE1L3), tumor protein p73-like (TP73L), ring finger protein 36 (RNF36) and KIAA0367 were differentially expressed. These genes are all pro-apoptotic and were found up-regulated in ICP placenta. All these findings indicate that apoptosis of placental trophoblast is an important pathology of ICP, although the cause-effect relationship is not known yet.
The alteration in transportation carrier of bile acid was certainly the focus of investigation in this field. Huang et al. [41] observed the expression of farnesoid X receptor (FXR) and bile salt excretory pump (BSEP) in ICP placenta with immunohistochemistry and found that the expression of FXR was increased and that of BSEP was decreased relative to normal pregnancy. In this study, we did not find any differentially expressed gene of transportation carrier of bile acid, however, we did find some differentially expressed genes involved in material transportation. These differentially expressed genes included transient receptor potential cation channel, subfamily V, member 5 (TRPV5), Solute carrier family 6, member 17 (SLC16A17), ATPase, Class II, type 9A (ATP9A) and others. Although the involvement of these genes in ICP remains unknown, it is reasonable to speculate that the abnormalities of placental material transportation carrier might be important pathophysiology of ICP.
Mutations or polymorphisms of genes expressing hepatobiliary transport proteins, at least including ABCB4 (MDR3), FIC1 (ATP8B1) and ABCB11 (BSEP), may contribute to the development and/or severity of ICP [42, 43]. These genes were expressed in human placentas probably at a low level [44, 45]. The low expression of these genes, limited sensitivity of microarray to identify differentiation of lowly expressed genes and relatively small sample size might be among the causes for lack of differential expression of these genes.
Except for above-mentioned genes, genes of categories at least including signal transduction, response to stimulus, reproduction, multi-organism process, biological adhesion, developmental process were differentially expressed, indicating the diversity of gene expression in placenta of ICP and the complexity of etiology and pathogenesis of ICP. Although the current investigation was primary, we believe that our experiment provided some clue for the further investigation of ICP etiology and pathogenesis.
In summary, 293 differentially expressed genes of 20 categories were found in ICP placenta, which suggests the diversity of gene expression alteration and the complexity of etiology and pathogenesis of ICP.
References
Lammert F, Marschall HU, Glantz A, Matern S (2000) Intrahepatic cholestasis of pregnancy: molecular pathogenesis, diagnosis and management. J Hepatol 33(6):1012–1021
Beuers U, Pusl T (2006) Intrahepatic cholestasis of pregnancy—a heterogeneous group of pregnancy-related disorders? Hepatology 43(4):647–649
Riely CA, Bacq Y (2004) Intrahepatic cholestasis of pregnancy. Clin liver dis 8(1):167–176
Fisk NM, Storey GN (1988) Fetal outcome in obstetric cholestasis. Br J Obstet Gynaecol 95(11):1137–1143
Bacq Y, Sapey T, Bréchot MC, Pierre F, Fignon A, Dubois F (1997) Intrahepatic cholestasis of pregnancy: a French prospective study. Hepatology 26(2):358–364
Rioseco AJ, Ivankovic MB, Manzur A et al (1994) Intrahepatic cholestasis of pregnancy: a retrospective case-control study of perinatal outcome. Am J Obstet Gynecol 170(3):890–895
Laatikainen T, Ikonen E (1975) Fetal prognosis in obstetric hepatosis. Ann Chir Gynaecol Fenn 64(3):155–164
Shaw D, Frohlich J, Wittmann BA, Willms M (1982) A prospective study of 18 patients with cholestasis of pregnancy. Am J Obstet Gynecol 142(6 Pt 1):621–625
Alsulyman OM, Ouzounian JG, Ames-Castro M, Goodwin TM (1996) Intrahepatic cholestasis of pregnancy: perinatal outcome associated with expectant management. Am J Obstet Gynecol 175(4 Pt 1):957–960
Williamson C, Hems LM, Goulis DG et al (2004) Clinical outcome in a series of cases of obstetric cholestasis identified via a patient support group. BJOG 111(7):676–681
Zhao YJ, Yue YF, Liu XQ, Li SH, Liu ZY, Wang Y (2004) Study on relationship between serum cholylglycine and placental apoptosis in patients with intrahepatic cholestasis of pregnancy (in Chinese). Zhonghua Fu Chan Ke Za Zhi 39(7):446–448
Wang DM, La XL, Ding L (2003) Study on apoptosis and expression of Fas, FasL in placenta of intrahepatic cholestasis of pregnancy (in Chinese). Chin J Perinat Med 6(4):199–201
Zhang SJ (2007) Immunohistochemical study on the expression of TNF-α in placenta in intrahepatic cholestasis of pregnancy (in Chinese). Acta Acad Med Xuzhou 27(7):451–452
Dong MY, He J, Wang ZP (2003) Placental expression of epidermal growth factor receptor in intrahepatic cholestasis of pregnancy. Chin J Obstet Gynecol 38(2):106–107
Watson A, Mazumder A, Stewart M, Balasubramanian S (1998) Technology for microarray analysis of gene expression. Curr Opin Biotechnol 9:609–614
Smith L, Greenfield A (2003) DNA microarrays and development. Hum Mol Genet 12(1):R1–R8
Schena M, Shalon D, Davis RW, Brown PO (1995) Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 270(20):467–470
Dong M, Shi Y (1996) Evaluation of cholic glycine in serum and hemorrheological parameters in women with intrahepatic cholestasis of pregnancy (in Chinese). Curr Adv Obstet Gynecol 5:251–253
Dong M, Wang Z, He J, Shi Y, Yang P, Pan Y (1999) Increased anticardiolipin antibody in intrahepatic cholestasis of pregnancy (in Chinese). Chin J Obstet Gynecol 34:491
Dong M, Xie X, Wang Z, He J, Zhou J (2002) Impaired mixed lymphocyte reaction in intrahepatic cholestasis of pregnancy (in Chinese). Gynecol Obstet Invest 54:191–195
Cristofalo VJ (2000) A DNA chip off aging block. Nat Med 6(5):507
Wang ZP, Dong MY, Chu HN, He J (2004) Increased serum levels of neopterin and soluble interleukin-2 receptor in intrahepatic cholestasis of pregnancy. Acta Obstet Gynecol Scand 83(11):1067–1070
Peng B, Liu SY (2002) Study of relationship between T helper cell type1 and type2 cytokines and intrahepatic cholestasis of pregnancy (in Chinese). Zhonghua Fu Chan Ke Za Zhi 37(9):516–518
Ling B, Yao FQ, Zhou Y, Chen ZZ, Shen GD, Zhu YY (2007) Cell-mediated immunity imbalance in patients with intrahepatic cholestasis of pregnancy. Cell Mol Immunol 4(1):71–75
Koehler LH, Benz EJ (1962) Serum adenosine deaminase: methodology and clinical applications. Clin Chem 8:133–140
Ungerer JP, Oosthuizen HM, Bissbort SH, Vermaak WJ (1992) Serum adenosine deaminase: isoenzymes and diagnostic application. Clin Chem 38(7):1322–1326
Trajkovic V, Sweet MJ, Xu D (2004) T1/ST2—an IL-1 receptor-like modulator of immune responses. Cytokine Growth Factor Rev 15(2–3):87–95
Xu D, Chan WL, Leung BP et al (1998) Selective expression of a stable cell surface molecule on type 2 but not type 1 helper T cells. J Exp Med 187(5):787–794
Löhning M, Stroehmann A, Coyle AJ et al (1998) T1/ST2 is preferentially expressed on murine Th2 cells, independent of interleukin 4, interleukin 5, and interleukin 10, and important for Th2 effector function. Proc Natl Acad Sci USA 95(12):6930–6935
Coyle AJ, Lloyd C, Tian J et al (1999) Crucial role of the interleukin 1 receptor family member T1/ST2 in T helper cell type 2-mediated lung mucosal immune responses. J Exp Med 190(7):895–902
Dann AT, Kenyon AP, Wierzbicki AS, Seed PT, Shennan AH, Tribe RM (2006) Plasma lipid profiles of women with intrahepatic cholestasis of pregnancy. Obstet Gynecol 107(1):106–114
Spreyer P, Schaal H, Kuhn G et al (1990) Regeneration-associated high level expression of apolipoprotein D mRNA in endoneurial fibroblasts of peripheral nerve. EMBO J 9(8):2479–2484
Xia SY, Chen ZQ, Li L (2003) Change of vascular endothelial growth factor (VEGF) and its significance in placenta for patients with intrahepatic cholestasis in pregnancy (in Chinese). J Chin Phys 5(8):1068–1070
Liu JP, Baker J, Perkins AS, Robertson EJ, Efstratiadis A (1993) Mice carrying null mutations of the genes encoding insulin-like growth factor I (Igf-1) and type 1 IGF receptor (Igf1r). Cell 75(1):59–72
DeChiara TM, Efstratiadis A, Robertson EJ (1990) A growth-deficiency phenotype in heterozygous mice carrying an insulin-like growth factor II gene disrupted by targeting. Nature 345(6270):78–80
Lin CG, Leu SJ, Chen N et al (2003) CCN3 (NOV) is a novel angiogenic regulator of the CCN protein family. J Biol Chem 278(26):24200–24208
Ellis PD, Chen Q, Barker PJ, Metcalfe JC, Kemp PR (2000) Nov gene encodes adhesion factor for vascular smooth muscle cells and is dynamically regulated in response to vascular injury. Arterioscler Thromb Vasc Biol 20(8):1912–1919
Gellhaus A, Schmidt M, Dunk C, Lye SJ, Kimmig R, Winterhager E (2006) Decreased expression of the angiogenic regulators CYR61 (CCN1) and NOV (CCN3) in human placenta is associated with pre-eclampsia. Mol Hum Reprod 12(6):389–399
Chen M, Sun LZ, Ding HJ, Wu WP (2007) The relationship between intrahepatic cholestasis and placenta cells apoptosis in pregnancy (in Chinese). Mod Med J 35(6):428–431
Perez MJ, Macias RI, Marin JJ (2006) Maternal cholestasis induces placental oxidative stress and apoptosis. Protective effect of ursodeoxycholic acid. Placenta 27(1):34–41
Huang JR, Liu J, Chang SF (2006) Mechanism of farnesoid X receptor and bile salt excretory pump on bile acid transport in placenta of intrahepatic cholestasis of pregnany (in Chinese). Chongqing Med J 174:9–1950
Floreani A, Carderi I, Paternoster D et al (2008) Hepatobiliary phospholipid transporter ABCB4, MDR3 gene variants in a large cohort of Italian women with intrahepatic cholestasis of pregnancy. Dig Liver Dis 40(5):366–370
Arrese M, Macias RI, Briz O, Perez MJ, Marin JJ (2008) Molecular pathogenesis of intrahepatic cholestasis of pregnancy. Expert Rev Mol Med 28:10:e9
Serrano MA, Macias RIR, Briz O et al (2007) Expression in human trophoblast and choriocarcinoma cell lines, BeWo, Jeg-3 and JAr of genes involved in the hepatobiliary-like excretory function of the placenta. Placenta 28:107–117
Patel P, Weerasekera N, Hitchins M, Boyd CAR, Johnston DG, Williamson C (2003) Semi quantitative expression analysis of MDR3, FIC1, BSEP, OATP-A, OATP-C, OATP-D, OATP-E and NTCP gene transcripts in 1st and 3rd trimester human placenta. Placenta 24:39–44
Acknowledgment
This study was financially supported by ZJNSF (No Y205297).
Author information
Authors and Affiliations
Corresponding author
Additional information
J. Wei and H. Wang were contributed equally.
Rights and permissions
About this article
Cite this article
Wei, J., Wang, H., Yang, X. et al. Altered gene profile of placenta from women with intrahepatic cholestasis of pregnancy. Arch Gynecol Obstet 281, 801–810 (2010). https://doi.org/10.1007/s00404-009-1156-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00404-009-1156-3