Abstract
Genes and proteins form complex dynamical systems or gene regulatory networks (GRN) that can reach several steady states (attractors). These may be associated with distinct cell types. In plants, the ABC combinatorial model establishes the necessary gene combinations for floral organ cell specification. We have developed dynamic gene regulatory network (GRN) models to understand how the combinatorial selection of gene activity is established during floral organ primordia specification as a result of the concerted action of ABC and non-ABC genes. Our analyses have shown that the floral organ specification GRN reaches six attractors with gene configurations observed in primordial cell types during early stages of flower development and four that correspond to regions of the inflorescence meristem. This suggests that it is the overall GRN dynamics rather than precise signals that underlie the ABC model. Furthermore, our analyses suggest that the steady states of the GRN are robust to random alterations of the logical functions that define the gene interactions. Here we have updated the GRN model and have systematically altered the outputs of all the logical functions and addressed in which cases the original attractors are recovered. We then reduced the original three-state GRN to a two-state (Boolean) GRN and performed the same systematic perturbation analysis. Interestingly, the Boolean GRN reaches the same number and type of attractors as reached by the three-state GRN, and it responds to perturbations in a qualitatively identical manner as the original GRN. These results suggest that a Boolean model is sufficient to capture the dynamical features of the floral network and provide additional support for the robustness of the floral GRN. These findings further support that the GRN model provides a dynamical explanation for the ABC model and that the floral GRN robustness could be behind the widespread conservation of the floral plan among eudicotyledoneous plants. Other aspects of evolution of flower organ arrangement and ABC gene expression patterns are discussed in the context of the approach proposed here.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
Gene interactions are prevalent and key during development, and thus development may bias, restrict, or mediate the variation that evolutionary forces may act on. Therefore development should be explicitly considered in evolutionary models that aim at understanding the origin and evolutionary dynamics of phenotypical traits (Lewontin 1974; Salazar-Ciudad and others 2000).
Complex networks of interacting components at different levels of biological organization are being uncovered at the molecular level. Genes and proteins form complex dynamical systems or gene regulatory networks (GRN) that can reach several steady states (attractors; Figure 1). These have been associated with distinct cell types (Kauffman 1969), and this proposition has been recently supported experimentally (Acar and others 2005; Huang and others 2005). Our aim is to contribute to a general framework for understanding the interplay between the structure, function, and evolution of GRN and their role in pattern formation that is central for understanding development.
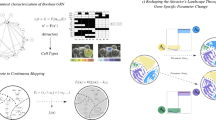
Floral structure of Arabidopsis thaliana, ABC model and GRN. (A) Basin of attraction of the petal-2 point attractor. (B) Pattern of expression of the genes, where 0 means “off” and 1 means “on.” (C) The same pattern depicted on the network; the dark circles indicate that the corresponding node is “on” and the rest are “off.” (D) The pattern of petal-2 corresponds to the overlapping activity of A + B genes. (E) The structure emerging from this pattern of activity: petals are depicted in red in the longitudinal and radial floral diagrams.
The approach we have followed has been to construct and validate qualitative models of specific biological processes that have been functionally characterized. The long-term aim is to infer novel and general regulatory principles of cellular networks by analyzing the structural and dynamical features of the specific gene regulatory networks studied and modeled. We think that this approach will complement the more theoretical ones that aim at uncovering general principles for network assemblage and dynamics using mathematical and simulation models or intend to infer global GRN architectures relying on functional genomics data. All approaches should provide feedback and complement one another.
Structural and dynamical analyses of well-characterized subnetworks grounded on solid experimental data are also useful to integrate experimental data and to find holes and inconsistencies in such data. Gene regulatory network models grounded on experimental data are also important to validate inferences of network connectivity from functional genomic data. Indeed, the network that we proposed for cell fate determination during floral organ specification (Mendoza and Alvarez-Buylla 1998) has been used by several researchers to validate methods of GRN architecture inference (Perkins and others 2004; Aracena and Demongeot 2004).
We have focused on plant systems to pursue our aims. In plants, multicellularity evolved independently from animals. A comparative approach of multicellular development in its broadest sense should therefore consider plant as well as animal systems (Meyerowitz 2002). Moreover, plants are relatively simpler than animals in their cellular organization, and therefore it will be more feasible to propose models that scale from network behavior to cell and morphogenetic patterns. Within plants, comparative analyses of networks controlling cell-fate determination or pattern formation may help uncover subnetworks that have been fairly fixed during evolution, modules that are co-opted for several cell-fate determination functions within or among plant species, and critical network motifs or subnetworks underlying important functions and evolutionary innovations.
The ABC’s of Flower Development
Flowers are the most complex and showy multicellular structures of angiosperms, and they have attracted the attention of many researchers. Upon induction to flowering, the shoot apical meristem that only produces vegetative structures from its flanks turns into an inflorescence meristem that also produces flowers. Adult floral morphology originates from floral meristems that arise from the flanks of the inflorescence meristem in a spiral arrangement. One day after flower induction a flower buttress arises, and 1.25 days afterwards the flower primordium forms in the flanks of each floral meristem. The sepal primordia arise first in the periphery of the floral meristem at stage three (3 days) of flower development; then the petal and stamen primordia appear in the second and third whorls, respectively, almost simultaneously. Last, the carpel primordium is formed from the remaining central portion of the floral meristem (Bowman 1994).
Thus, a few days after flower induction, the floral meristem is partitioned into four regions, each composed of primordial sepal, petal, stamen, and carpel cells that are sequentially differentiated from the outer whorl to the center of the meristem. Floral organ primordia continue to grow and differentiate into the distinct cell types that constitute each of the mature floral organs. The models that we have developed refer to the GRN that underlie primordial cell fate determination and hence underlie the combinatorial gene activities that characterize the four regions into which the flower meristem is partitioned during the early stages of flower development.
A wealth of data on the molecular genetic mechanisms underlying different aspects of flower development have accumulated during the last 15 years (Coen and Meyerowitz 1991). The genes controlling cell fate determination during floral organ specification have yielded the ABC model of flower development that was derived from genetic analyses of floral organ homeotic mutants in two plant study systems: Antirrhinum majus L. and Arabidopsis thaliana (L.) Heynh. (Coen and Meyerowitz 1991). These species have the stereotypical floral arrangement of almost all eudicotyledoneous species (see Vergara-Silva and others 2003; Ambrose and others 2006 for an exception), with sepals in the outermost whorl, then petals, stamens, and carpels in the flower center. The ABC model states that the identities of the floral organ types are established by combinations of genes grouped in three main classes, A, B, and C. A genes alone determine sepal identity; A plus B, petal identity; B plus C, stamen identity; and C alone, carpel identity (Figure 1). However, the ABC model does not provide an explanation for how such combinatorial selection of gene activity is established during floral organ primordia specification, and how the spatiotemporal pattern of ABC and non-ABC gene expression is established. Furthermore, the conserved pattern of floral organ determination and also the overall conservation of the ABC gene patterns of expression among eudicotyledoneous species suggest a robust mechanism underlying such combinatorial selection of gene activities. The ABC model by itself does not provide an explanation for such robustness either. We have been developing qualitative GRN models for tackling these questions
Discrete Gene Regulatory Network Models: The Formalism
Regulatory network models are abstractions of the multiple regulatory interactions among genes during cell-fate determination and pattern formation. Each gene undergoes a network of mRNA transcription, translation, protein modifications, and protein transport until its final functional destiny. Most data available for plant development is at the transcriptional regulatory level, although recent findings suggest that miRNA regulation is important during plant development (Chen 2004).
Gene regulatory network models may incorporate continuous or discrete functions for the gene-activation kinetics. Because the temporal scales of the molecular processes of the pathway that goes from DNA transcription to protein function are relatively short in comparison to those of the processes of pattern formation, it is reasonable to focus only on the qualitative regulatory interactions. Continuous implementations can incorporate more detail and can yield quantitative predictions. However, experimental data that provide estimates of such models are lacking. Furthermore, formal analyses of equivalent continuous and discrete models have analytically shown that both yield equivalent results (Thomas and others 1995). Furthermore, in systems of many components with many non-linearities, the behavior of the system depends mostly on the qualitative aspects of the GRN topology rather than on the kinetic details for each interaction and component. Therefore, we postulate models that assume discrete functions for the kinetics of gene activation. In the simplest case, Boolean functions that may only have two values: 0 (“off”) or 1 (“on”) are assumed. The activity of each gene depends on updating rules that are formalized as logical functions grounded on experimental data. Boolean models have also been successfully used in animal systems. For example, Albert and Othmer (2003) have shown that a Boolean GRN recovers the same patterns for the Drosophila melanogaster segment polarity genes as those recovered by a continuous model (Von Dassow and others 2000) analysis that had already been shown to be robust to parameter perturbations.
Discrete models are further justified because recent experimental evidence suggests that gene expression is digital and stochastic at the individual cell level rather than continuous. In cell aggregates, gene expression may appear to change continuously as an inductive signal increases. However, recent experimental data at the individual cell level suggests that it is the probability of a particular template to be active within a certain time window, rather than the rate of transcription from this template, that seems to be subject to regulation. Hence, genes within individual cells have a distinct probability of responding to a given concentration of stimulus of transcription, and the gene is either “on” or “off” in a certain time window (Hume 2000; Ozbudak and others 2002; Elowitz and others 2002; Blake and others 2003; Paulsson 2004; Walters and others 1995; Fiering and others 2000; Ho and others 1996). Rossi and collaborators (2000) have proposed a mechanism that is necessary and sufficient to account for the binary response in inducible gene expression that proposes that for each site in a promoter region there are transcription factors with opposing functions (for example, activation and repression) for the same target promoter motif.
Since their proposal (Kauffman 1969), random Boolean networks (RBN) have successfully described in a qualitative way several important aspects of gene regulation and cell differentiation processes (Kauffman 1993, 1995). Discrete GRN grounded on experimental data have been inspired in the RBN, but in experimentally grounded GRN genes are not connected by chance as in a RBN. Recent data are suggesting that the connectivity of real GRN obeys a power law (Aldana and Cluzel 2003). The type of discrete GRN models that we are developing consist of a set of discrete variables, ε1, ε2,...,ε N , each acquiring a discrete value corresponding to the two states of gene expression. The state of each gene is regulated by a set of K other genes, ε n1, ε n2,...,ε nK , according to logical rules that are grounded on experimental data (see example in Espinosa-Soto and others 2004). The network dynamics are then given by:
where f n is a logical rule associated with the nth element of the network. These rules are derived from experimental data.
A particular GRN has a large number of possible gene activity configurations or states, and each one always leads to another state or to itself, in which case that state is a point attractor (steady state). Eventually all states, the Ω space, will reach an attractor. A system may have more than one attractor. The set of all states that lead to a specific attractor is the basin of that attractor. The basins of attraction and attractors of a GRN depend on the number of elements, the number of possible states of each element, the topology of the network, and on the logical rules of each gene (Figure 2). Consequently, the dynamics of the system are deterministic, a condition in which the fates of all states are known.
Basins of attraction and point attractors of the four floral primordial cell types (sepal, petal, stamen, and carpel) around the flower diagram for A. thaliana. (A) Boolean network model. (B) Three-state network model. The basins of attraction end in the point attractors that correspond to gene expression profiles that characterize sepal (lower right), petal (upper left), stamen (lower left), and carpel (upper right) primordial cells in both A and B. Note that the state space is much larger in the second case.
Gene Regulatory Network Models for Arabidopsis thaliana (L.) Heynh. Cell Type Determination during Organ Type Specification
We are interested in understanding the GRN structure and dynamics underlying cell-fate determination during floral organ specification. We have mainly focused on Arabidopsis thaliana, for which most data are available. On a first approach, Mendoza and Alvarez-Buylla (1998) proposed a GRN dynamic model with 11 nodes that included ABC and non-ABC genes that attained six attractors that corresponded to the genetic activities proposed in the ABC model. However, in this first model there was circularity in the interaction weights estimates, and some attractors did not correspond to observed gene activity combinations. Mendoza and collaborators (1999) then analyzed the dynamic behavior of the same model in terms of positive and negative feedback circuits. Interestingly, they recovered the same attractors as the ones found in their first approach, and they were able to identify the functional positive feedback loops underlying such attractors. However, once again interaction parameters were not grounded on experimental data. Nonetheless, this latter approach shows that positive regulatory circuits play a central role in cell differentiation and that modularity that may be described in terms of functional loops is a central characteristic of GRN (Thieffry and Sánchez 2004). It is also useful to detect possible missing nodes and/or connections that could form additional functional positive loops with nodes within the GRN or in other GRN that could explain observed gene configurations or cell behaviors (Thomas 1991).
Recently, in Espinosa-Soto and collaborators (2004), we integrated all available data on ABC and non-ABC genes in a GRN model of 15 nodes that significantly improves previous studies on GRN modeling for flower organ specification in our laboratory. The GRN was modeled as a three-state discrete model to capture all the experimental data at hand, and the problem of circularity was avoided by explicitly establishing the logical rules based on the experimental data available. The updating was assumed synchronous, making it possible to avoid subjective interpretations concerning the temporal order with which genes are actualized. Of the total number of initial combinations (139,968), the network attained only 10 states, which coincide with the gene-expression profiles documented experimentally in cells of A. thaliana inflorescence meristems and floral organ primordia, and it also reproduces mutants’ gene-expression patterns.
These data suggest that the gene regulatory model incorporates the key components of a developmental module or subnetwork that underlies the combinatorial gene activities predicted in the ABC model. Thus it provides a dynamical explanation for the ABC model of floral organ determination, showing that precise signaling pathways are not required to restrain organ primordia cell types during A. thaliana flower development. Our model suggests that these are rather determined by overall gene network topology and dynamics. This was confirmed by robustness analyses of random perturbations of gene-interaction parameters (Espinosa-Soto and others 2004).
One value of the approach that we pursue is that it provides an opportunity to integrate all previously published experimental evidence on particular genetic interactions, and this might aid in detecting contradictions and holes in the experimental evidence, as well as guiding functional genomic approaches. Although our GRN studies are not historical accounts of all work that has been done, but rather a formal qualitative model of the behavior of the gene network that regulates floral organ cell differentiation, the work should nonetheless incorporate the critical evidence for each interaction. Since we published the Espinosa-Soto and others (2004) article we have detected a few instances in which some key research papers that first documented a particular interaction were not cited, and we update these cases here. We have also detected a few minor errors in the article of Espinosa-Soto and others (2004). We list the additions and corrections in Appendix 1 and show that these do not affect the model or the study results in any way.
Updated Floral Network Structure and Dynamics: Predictions Met and New Interactions
We present here an updated GRN for cell-fate determination during floral organ specification (Figure 3). Two recent experimental findings confirmed two predictions made by our analyses (Espinosa-Soto and others 2004). A publication that came out during the review process of our work showed that the SEP1, 2, 3 genes are indeed activated by LFY (Schmid and others 2003). We had predicted that a floral meristem identity gene should activate the SEP, or an inflorescence identity gene should repress them. We had postulated that TFL1 repressed them. Hence, we have now deleted this interaction and instead postulated that LFY activates SEP (Figure 3; detailed tables of logical rules are available on request). Second, we had predicted that AG should have a positive direct or indirect feedback loop, and this has been confirmed in a recent article (Gómez-Mena and others 2005).
Updated floral network of A. thaliana. Nodes correspond to genes or proteins, and edges correspond to regulatory interactions (arrows are positive and blunt-end are negative). See also Espinosa-Soto and others (2004).
New experimental evidence suggests that the activity of AP2 is postranscriptionally repressed by a specific miRNA (Chen 2004), whose abundance depends on the activity of the DICER-like protein HUA ENHANCER 1 (HEN1, Park and others 2002). However, even though HEN1 is expressed throughout the plant (Chen and others 2002), miRNA-dependent AP2 repression seems to occur only in the inner floral whorls (Chen 2004). This suggests that a cofactor, expressed in the inner floral whorls, is required to give specificity to HEN1-dependent repression of AP2. One possibility is that this cofactor is downstream of AG activity, supporting the mutual repression by A and C genes, proposed in the original ABC model (Coen and Meyerowitz 1991). We tested this possibility in our model. Because HEN1 is expressed throughout the plant, we can consider it to be permanently “on,” and hence it is only required to add, in the AP2 logical rules, that the inactivity of AG is needed for AP2 function. The only change we observed in this simulation was that in the carpel and stamen attractors AP2 is “off.”
Because both systems—the one that considers AP2 repression by AG and the one that does not—are so similar, we decided to work with the latter to avoid what may be unrealistic assumptions.
It has also been found that FT expression favors SEP3 and FUL activity (Teper-Bamnolker and Samach 2005). However, these interactions could be mediated by genes such as LFY and TFL1, and because this explanation is more parsimonious, we decided to avoid these additional interactions.
Interestingly, the updated GRN behavior is almost identical to that of the previously published (Espinosa-Soto and others 2004) network. We obtained the same number of point attractors as in the previous network. All the basins of attraction are of the same size and contain the same set of initial conditions as in the previous GRN, except for the sepal and the petal-1 basins, in which eight states of the petal-1 basin now belong to the sepal basin.
Reduction of the Three-state to a Boolean GRN
The original network had three states for some genes: inactive (0), medium active (1), and active (2). We reduced this three-state GRN to a Boolean network. We performed this analysis as an additional robustness analysis and also to test if it is useful to incorporate intermediate levels of gene activity (see Appendix 2 for details in GRN reduction). Although it is obvious that the basins of attraction are larger and more complicated in the three-state than in the two-state GRN, interestingly, the same attractors are reached in both cases (Figure 2). We then addressed if both GRN respond in the same way to systematic perturbations of the logical rules.
We mutated each of the outputs of all logical rules and followed the dynamics, counting the number of the original point attractors that were conserved after the perturbation (Table 1). Although the Ω space is quite different for both GRN, it is interesting to note that the qualitative behavior of the two networks is almost identical with respect to the number of point attractors recovered, and both GRN are quite robust. As expected, genes with more outputs are the most stable ones, whereas the genes with fewer outputs are the most sensitive ones. Although it makes functional sense that attractors are buffered against perturbations to most connected genes, these could also be older and hence have had more time to accumulate connections. Alternatively, robustness of the GRN to perturbations in these gene logical functions may have been selected for. As discussed in Espinosa-Soto and others (2004), it seems that the new attractors are not very different from the original ones.
Themes and Variations in Eudicotyledoneous Flower Evolution
Our previous studies had suggested that the GRN that underlies cell-fate determination during floral organ specification constitutes a robust developmental module or subnetwork that may underlie the widely conserved basic floral plan of eudicotyledoneous flowering plants. Furthermore, comparative analyses of flower patterning and floral organ determination gene network components and interactions suggest that this subnetwork has persisted over long periods irrespective of the evolution of the rest and largely uncovered molecular components that interact with this subnetwork (Ferrario and others 2004). These other components should underlie other, relatively more variable aspects of flower morphology (symmetry, organ number, organ color, etc). If this hypothesis is true, then the floral GRN should be conserved among most eudicotyledoneous species, and this could be tested experimentally.
The GRN model may also be a basis for explorations of evolutionary differences in network architecture between A. thaliana and other species. Petunia hybrida Vilm. is another experimental plant system in which the overall network of cell fate determination during flower organ development seems to be conserved with respect to A. thaliana (Ferrario and others 2004). Nevertheless, P. hybrida B-gene mutant phenotypes are different from those described in A. thaliana because these genes are duplicated in the former species. Interestingly, the A. thaliana GRN with duplicated B genes recovered the observed expression profiles observed in P. hybrida floral organ primordial cells under wild type and also when the node corresponding to the P. hybrida AP3 ortholog was simulated as mutated (Espinosa-Soto and others 2004). This result further validates the model and suggests that the GRN underlying floral organ cell-fate determination is robust also to gene duplications.
Models that couple the floral GRN dynamics in explicit spatial domains will be useful in exploring possible mechanisms underlying the divergent floral patterns and the role of ABC genes among basal angiosperms. Preliminary analyses of a single-cell stochastic version of the GRN model considered here suggest that the temporal sequence with which floral organ primordia fate is determined may result, at least in part, from the dynamics of the organ specification regulatory network studied in the face of stochastic noise (Alvarez-Buylla, Chaos, Cortés, Espinosa-Soto, and Padilla-Longoria, unpublished work).
An extension of this model will be useful to postulate hypotheses underlying the variations of floral organs and ABC gene expression patterns observed among angiosperms. Probably the most striking floral organ arrangement is that observed in the endemic heterotrophic species of the monocotyledons, Lacandonia schismatica E. Martínez & Ramos (Vergara-Silva and others 2003; Ambrose and others 2006), which has a homeotic inversion of stamens and carpels with the latter in the third rather than the fourth central whorl. Spatiotemporal models that explicitly consider the type of GRN model proposed here will be needed to address which mutations may underlie this unique floral phenotype.
Although 75% of angiosperms are eudicotyledoneous, ABC gene-expression patterns and floral organ identities and arrangements are more variable among the basal flowering plants. Recently, Kim and collaborators (2005) reported that (1) the expression pattern of AG (a C class gene) of basal angiosperms is quite similar to that observed in core eudicotyledoneous species, (2) AP3 and PI (B class genes) homologs seem to have a wider spatial expression pattern in basal angiosperms than that observed in eudicotyledoneous species, and (3) the expression pattern of AP1 (an A class gene) differs greatly from that observed in eudicotyledoneous plants. Coincidentally, the floral organs that are specified by the A and B genes also show the most divergent patterns with respect to those characteristic of the core eudicotyledoneous species. For example, in the basal angiosperm Nuphar advena (Aiton) W. T. Aiton, the outer perianth organs are green and sepal-like whereas the inner perianth organs are more like petals because they are brightly colored. However, another basal angiosperm, Asimina longifolia Kral., has a perianth that is well differentiated into sepals and petals (Kim and others 2005).
This leads to the question of whether GRN models represent useful frameworks for unravelig the molecular evolution of flower patterns. Interestingly, the basins of attraction of the reproductive organs that seem to be the most stable among angiosperms are also the largest and, in consequence, the most stable in our simulations. This could have a historical explanation, and it could imply that the genes and/or interactions involved in the development of stamens and carpels evolved first, whereas those underlying sepal and petal specification arose later during evolution. The variation observed among basal angiosperms can reveal the unstable footsteps of the assemblage of the floral organ specification GRN that occurred during the evolution of the early lineages represented by extant basal angiosperms. The use of formal GRN models may help explore these hypotheses.
Our formal and dynamic GRN approaches suggest that the emergence of structures could have been the result of relatively slight modifications in GRN. New floral organs may have arisen as additional genes were added to a primitive network that originally yielded only reproductive structures. Data from gymnosperms (Sundstrom and Engstrom 2002) does indeed suggest that part of the angiosperm floral organ GRN could have been present before the origin of flowering plants. Adding a gene to the GRN may multiply the attractors and enlarge the potential adaptative landscape. The origin of sepals and petals could have been the result of such a process, and this could have expanded the morphospace on which natural selection could act, yielding new evolutionary possibilities. If our rationale holds, perturbation analyses of an explicitly spatiotemporal model of coupled GRN could reveal which are the most stable steady states of the network and hence the probable primitive core within it that has remained unchanged during long evolutionary times. Such an approach not only allows for much more flexibility but also seems more realistic in evolutionary terms than postulating that the emergence of a new structure has relied on addition of genes from A to Z endlessly. It also allows for more specific predictions that could be first tested in silico and then in vivo by pursuing comparative analyses of key genes within the GRN in species selected according to their phylogenetic position in the tree of angiosperm evolution.
Cell identity attainment can be conceptualized as the product of a series of developmental decisions in which alternative gene activity patterns are acquired. The ABC model succeeded as a mechanistic proposal of floral organ specification because it showed that floral organ cell identity could be interpreted as the outcome of two developmental decisions: A or C gene activity, and B gene activity or inactivity (hereafter “ABC decisions”). Despite new discoveries on genes involved in floral morphogenesis and their interactions, the logic behind the ABC model remains unaltered. We now know that, in wild-type plants, the ABC decisions depend on an earlier decision: the floral or non-floral meristem decision (F-NF decision), in which genes such as LFY and TFL1 are involved. A consequence of the F-NF decision is the upregulation of the SEP genes, because LFY promotes their expression (Schmid and others 2003). In later events, on the other hand, ovule identity genes, such as SEEDSTICK (Pinyopich and others 2002) seem to require C class gene activity as a necessary condition, because C mutants lack ovules. Hence, a hierarchical series of decisions in which some can only be taken if there is a certain outcome in a previous one seems to account for floral cell identity attainment. Therefore, as data accumulate, it is becoming clear that the developmental decisions during flower development considered in the ABC model hold, despite the fact that additional earlier and later developmental decisions have been discovered.
On the other hand, the inclusion of factors that provide positional information and that bias the ABC decisions, such as UFO (Parcy and others 1998) and WUS (Lenhard and others 2001; Lohman and others 2001), suggest that these genes may be key inputs for determining the specific time and site where the combinations of gene activities considered in the ABC model are established. Explicitly spatial models that consider this will be useful to understand the mechanisms underlying the observed spatiotemporal patterns of ABC gene expression. In this context, the ABC model promises to continue to be a valid and useful core framework for understanding floral organ pattern, development, and evolution
References
Acar M, Becskei A, Van Oudenaarden A. 2005. Enhancement of cellular memory by reducing stochastic transitions. Nature 435:228–232
Aracena J, Demongeot J. 2004. Mathematical methods for inferring regulatory networks interactions: application to genetic regulation. Acta Biotheor 52:391–400
Albert R, Othmer HG. 2003. The topology of the regulatory interactions predicts the expression pattern of the segment polarity genes in Drosophila melanogaster. J Theor Biol 223:1–18
Aldana M, Cluzel P. 2003. A natural class of robust networks. Proc Natl Acad Sci USA 100:8710–8714
Ambrose BA, Espinosa-Matías S, Vázquez-Santana S, Vergara-Silva F, Martínez E, et al. 2006. Comparative developmental series of the mexican triurids support a euanthial interpretation for the unusual reproductive axes of Lacandonia schismatica (Pandanales, Liliopsida). Am J Bot 93:15–35
Blake W, Kaern M, Cantor C, Collins J. 2003. Noise in eukaryotic gene expression. Nature 422:633–637
Bowman JL, Smyth DR, Meyerowitz EM. 1989. Genes directing flower development in Arabidopsis. Plant Cell 1:37–52
Bowman J, Smyth D, Meyerowitz E. 1991. Genetic interactions among floral homeotic genes of Arabidopsis. Development 112:1–20
Bowman J 1994. Arabidopsis–an Atlas of Morphology and Development. Springer-Verlag, Berlin Heidelberg New York, NY, USA, p 480
Bowman JL, Alvarez J, Weigel D, Meyerowitz EM, Smyth D. 1993. Control of flower development in Arabidopsis thaliana by APETALA1 and interacting genes. Development 119:721–743
Chen X, Liu J, Cheng Y, Jia D. 2002. HEN1 functions pleiotropically in Arabidopsis development and acts in C function in the flower. Development 129:1085–1094
Chen X. 2004. A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303:2022–2025
Coen ES, Meyerowitz E. 1991. The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Deyholos MK, Sieburth LE. 2000. Separable whorl-specific expression and negative regulation by enhancer elements within the AGAMOUS second intron. Plant Cell 12:1799–1810
Elowitz M, Levine A, Siggia E, Swain P. 2002. Stochastic gene expression in a single cell. Science 297:1183–1186
Espinosa-Soto C, Padilla-Longoria P, Alvarez-Buylla E. 2004. A gene regulatory network model for cell-fate differentiation during Arabidopsis thaliana flower development that is robust and recovers experimental gene expression profiles. Plant Cell 16:2923–2939
Ferrario S, Immink RG, Angenent GC. 2004. Conservation and diversity in flower land. Curr Opin Plant Biol 7:84–91
Fiering S, Whitelaw E, Martin D. 2000. To be or not to be active: the stochastic nature of enhancer action. BioEssays 22:381–387
Gómez-Mena C, de Folter S, Costa MMR, Angenent GC, Sablowski R. 2005. Transcriptional program controlled by the floral homeotic gene AGAMOUS during early organogenesis. Development 132:429–438
Hill TA, Day CD, Zondlo SC, Thackeray AG, Irish VF. 1998. Discrete spatial and temporal cis-acting elements regulate transcription of the Arabidopsis floral homeotic gene APETALA3. Development 125:1711–1721
Ho S, Biggar S, Spencer D, Schreiber S, Crabtree G. 1996. Dimeric ligands define a role for transcriptional activation domains in reinitiation. Nature 382:822–826
Honma T, Goto K. 2000. The Arabidopsis floral homeotic gene PISTILLATA is regulated by discrete cis-elements responsive to induction and maintenance signals. Development 127:2021–2030
Huang S, Eichler G, Bar-Yam Y, Ingber DE. 2005. Cell fates as high-dimensional attractor states of a complex gene regulatory network. Physical Rev Lett 94:128701–128704
Hume D. 2000. Probability in transcriptional regulation and its implications for leukocyte differentiation and inducible gene expression. Blood 96:2323–2328
Kauffman SA. 1969. Metabolic stability and epigenesis in randomly constructed genetic nets. J Theor Biol 22:437–467
Kauffman SA. 1993. The Origins of Order: Self-organization and Selection in Evolution. Oxford University Press, Oxford, England, UK
Kauffman SA. 1995. At Home in the Universe. New York, NY, USA, Oxford University Press
Kim S, Koh J, Yoo M, Kong H, Hu Y, et al. 2005. Expression of floral MADS-box genes in basal angiosperms: implications for the evolution of floral regulators. Plant J 43:722–744
Lenhard M, Bonhert A, Jurgens G, Laux T. 2001. Termination of stem cell maintenance in Arabidopsis floral meristems by interactions between WUSCHEL and AGAMOUS. Cell 105:805–808
Lewontin R. 1974. The Genetic Basis of Evolutionary Change. Columbia University Press, New York, NY, USA, p 352
Lohmann JU, Hong RL, Hobe M, Busch MA, Parcy F, et al. 2001. A molecular link between stem cell regulation and floral patterning in Arabidopsis. Cell 105:793–803
Mayer KFX, Schoof H, Haecker A, Lenhard M, Jürgens G, et al. 1998. Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 95:805–815
Mendoza L, Alvarez-Buylla ER. 1998. Dynamics of the genetic regulatory network of Arabidopsis thaliana flower morphogenesis. J Theor Biol 193:307–319
Mendoza L, Thieffry D, Alvarez-Buylla ER. 1999. Genetic control of flower morphogenesis in Arabidopsis thaliana: a logical analysis. Bioinformatics 15:593–606
Meyerowitz E. 2002. Plants compared to animals: the broadest comparative study of development. Science 295:1482–1485
Ozbudak EM, Thattai M, Kurtser I, Grossman AD, Van Oudenaarden A. 2002. Regulation of noise in the expression of a single gene. Nat Genet 31:69–73
Parcy F, Nilsson O, Busch MA, Lee I, Weigel D. 1998. A genetic framework for floral patterning. Nature 395:561–566
Paulsson J. 2004. Summing up the noise in gene networks. Nature 427:415–418
Park W, Li J, Song R, Messing J, Chen X. 2002. CARPEL FACTORY, a Dicer homolog, and HEN1, a novel protein, act in microRNA metabolism in Arabidopsis thaliana. Curr Biol 12:1484–1495
Perkins T, Hallet M, Glass L. 2004. Inferring models of gene expression dynamics. J Theor Biol 230:289–299
Piñeiro M, Coupland G. 1998. The control of flowering time and floral identity in Arabidopsis. Plant Physiol 117:1–8
Pinyopich A, Ditta GS, Savidge B, Liljegren SJ, Baumann E, et al. 2002. Assessing the redundancy of MADS-box genes during carpel and ovule development. Nature 424:85–88
Rossi FMV, Kringstein AM, Spicher A, Guicherit OM, Blau HM. 2000. Rheostat mechanism of transcriptional regulation converted to ON/OFF switch. Mol Cell 6:723–728
Salazar-Ciudad I, García-Fernández J, Solé RV. 2000. Gene networks capable of pattern formation: from induction to reaction-diffusion. J Theor Biol 205:587–603
Schmid M, Uhlenhaut NH, Godard F, Denmar M, Bressan R, et al. 2003. Dissection of floral induction pathways using global expression analysis. Development 130:6001–6012
Sundstrom J, Engstrom P. 2002. Conifer reproductive development involves B-type MADS-box genes with distinct and different activities in male organ primordial. Plant J 31:161–169
Teper-Bamnolker P, Samach A. 2005. The flowering integrator FT regulates SEPALLATA3 and FRUITFULL accumulation in Arabidopsis leaves. Plant Cell 17:2661–2675
Thieffry D, Sánchez L. 2004. Qualitative analysis of gene networks: toward the delineation of cross-regulatory modules, pp. 222–243 In: Schlosser G, Wagner G, editors. Modularity in Development and Evolution. Chicago, IL, USA, The University of Chicago Press, 600 p
Thomas R. 1991. Regulatory networks seen as asynchronous automata: a logical description. J Theor Biol 153:1–23
Thomas R, Thieffry D, Kaufman M. 1995. Dynamical behaviour of biological regulatory networks—I. Biological role of feedback loops and practical use of the concept of the loop-characteristic state. Bull Math Biol 57:247–276
Vergara-Silva F, Espinosa-Matías S, Ambrose BA, Vázquez-Santana S, Martínez-Mena A, Márquezand et al. 2003. Inside-out flowers characteristic of Lacandonia schismatica evolved at least before its divergence from a closely related taxon, Triuris brevistylis. Int J Plant Sci 164:345–357
von Dassow G, Meir E, Munro EM, Odell GM. 2000. The segment polarity network is a robust developmental module. Nature 406:188–193
Wagner D, Sablowski RWM, Meyerowitz EM. 1999. Transcriptional activation of APETALA1 by LEAFY. Science 285:582–584
Walters MC, Fiering S, Eidemiller J, Magis W, Groudine M, et al. 1995. Enhancers increase the probability but not the level of gene expression. Proc Natl Acad Sci USA 92:7125–7129
Weigel D, Meyerowitz EM. 1993. Activation of floral homeotic genes in Arabidopsis. Science 261:1723–1726
Acknowledgments
We acknowledge R. Vicencio and A. Navarrete for support in various tasks while preparing this manuscript. Financial support was from PAPIIT (Programa de Apoyo a Proyectos de Investigación e Innovación Tecnológica), UNAM (Universidad Nacional Autónoma de México; IN 230002, IN212995 and IN221406), and CONACYT (Consejo Nacional de Ciencia y Tecnología; 41848-Q and 0435/B-1) grants to E.A.B; E.A.B. also was supported by a SFI International fellowship while completing this article, and M.A. received CONACyT grant P47836-F. A.C. and C.E.S. received PhD fellowships from Consejo Nacional de Ciencia y Tecnología and Universidad Nacional Autónoma de México.
Author information
Authors and Affiliations
Corresponding author
Additional information
Álvaro Chaos, Max Aldana and Elena Alvarez-Buylla contributed equally to this work.
Appendices
Appendix I. Corrections of GRN in Espinosa-Soto, Padilla-Longoria and Alvarez-Buylla 2004
We list some errors detected in Espinosa-Soto and others (2004). The truth table for FT on page 2925 has two outcomes, 0 or 1, which defines FT as a binary gene; nevertheless, on the same page in the truth table of AP1, FT has three values which is a contradiction. A closer look at the later truth table reveals that for the first rule of AP1 the state of FT has no effect on AP1 because it can be any state. For the second and the third rules, the state of FT is the same, which is active. In addition, for the other rules (3, 4 and 5) the state of FT can be non-active or medium active. Therefore, to avoid that contradiction between the truth table of FT and its states defining the truth table of AP1 we made some changes. The first rule remains the same. In the second and the third rules of AP1, we substituted the value of 2 by 1, which means it is active. As the other rules of AP1 do not make a distinction between the state of no activity and the state with medium activity, we changed them by 0 which means no activity. This contradiction does not affect the results obtained in Espinosa-Soto and others (2004).
The truth table of WUS (page 2926) has three inputs, which are WUS, AG, and SEP. The gene network architecture on the same page depicts that the inputs of WUS are just SEP and AG, so it needs a connection between WUS and itself to be coherent with the truth table. The printed truth table of PI in Espinosa-Soto and others (2004) had a few mistakes and here we present the correct truth table of PI (Table A1). In the two latter cases the correct truth tables shown here were used in the simulations supporting the results of Espinosa-Soto and others (2004).
In the logical rules for LFY the evidence for the fact that in ap1 and ap1cauliflower (cal) double mutants, LFY expression is reduced, Bowman and others (1993) besides Piñeiro and Coupland (1998) should be cited. In the logical rules for AP1 the evidence for the direct activation of AP1 by LFY that binds to the AP1 promoter was first documented by Parcy and collaborators (1998) and then by Wagner and collaborators (1999). On APETALA3 logical rules, the evidence for the fact that in lfy ap1 double mutants, AP3 mRNA is not detected, but in ap1 mutants, AP3 is expressed as in wild-type plants Weigel and Meyerowitz (1993) as well as Hill and collaborators (1998) should be cited. On PI logical rules, the evidence that the level of PI mRNA and its domain of expression are severely reduced in lfy-6 null mutants is found in Weigel and Meyerowitz (1993) as well as in Honma and Goto (2000). On AGAMOUS logical rules, the evidence that in strong ap2 mutant alleles, AG RNA accumulates in the four floral whorls, and in ap2-2 and ap2-9 alleles, 1st whorl organs are converted into carpels or carpelloid sepals, and 2nd whorl organs are absent was first presented in Bowman and collaborators (1989, 1991) and later by Deyholos and Sieburth (2000). Also, support for the fact that AG, PI, AP3, WUS, and others are transcriptions factors is also found in Parcy and collaborators (1998) and Lohmann and collaborators (2001) as well as in Coen and Meyerowitz (1991) and Mayer and collaborators (1998).
Appendix II. Reduction of the Three-state to a Boolean GRN
To perform this reduction we had to decide to which state, active or inactive, the medium active state corresponded. To translate each rule for all truth tables we proceeded as follows: if the state was 2 it was changed to 1, if it was 0 it remained in that state. For some rules a combination of two states, gave the same output. Those cases were combinations of 0 and 1, and 1 and 2. In such cases, the first one was translated to 0 because there was no distinction between inactivation and intermediate activation, and in the second one it was translated to 1 because the rule gave the same output when the input was either 1 or 2. If the state was just 1 we assigned an uncertainty indicator because we could not assign a state of activity (1) or inactivity (0). For all possible inputs of each gene (for example, for a gene with three input genes there are 23 different inputs), we assigned an output according to the translated rules (that is, the rule that corresponds to that specific combination of inputs). Because of the reduction of the states it is possible that a rule governs an input that is already governed by other rules. In these cases there are two possibilities. Overlapping rules can be either redundant, if all rules give the same outputs, or contradictory, if some of all the overlapping rules yield different outputs. If the outputs of all the rules for a particular combination of inputs are unassigned, we call it a total ambiguity. If the output of one rule has a defined state, but one rule governing the same combination of inputs has an undefined state, we took the output of the defined rule and we call this case a partial ambiguity.
The total number of combinations of inputs of all genes from the original network with three states was 5500, whereas the Boolean network has only 772 combinations of inputs for all genes. For example, FUL was regulated by two genes with three entries for each in the three-state case, that yielded 9 different combinations of inputs. In contrast, in the Boolean translation, FUL is still regulated by the same two genes, but now these two have only two states each, yielding four combinations.
After assigning all states of all genes as mentioned, we found 96 total ambiguities and only 1 contradiction. We tested several combinations for these 97 problematic cases to try to recover the same behavior of the network with 3 states, starting with all of them with the state of inactivity (0), continuing with different combinations of 0 and 1, and finally assigning all of them a state of activity (1). When we assigned to all the problematic states the value of activity, we obtained exactly the same dynamics as for the original network: 10 point attractors with the same pattern of gene activity.
This can be understood in two ways. The observation that some products of certain genes differ in concentrations was the cause of the abstraction of the 3 states. If a particular threshold to activate a gene is surpassed, then the gene will enter in a state of activation. As we do not know the activation threshold of the genes, it could happen that regardless there is a marked difference in the concentration of the products of the genes; in both cases, medium activation and activation, the concentration surpasses the specific gene activation threshold. Consequently, we will obtain the same dynamics with the three-state network as that from the two-state network. On the other side, it is possible that the observed difference in gene product quantities is important and meaningful in the context of the system, but the robustness of the architecture network is sufficient to tackle this alteration of the parameters in gene product concentrations and to perform a normal behavior. This approach is a robustness analysis because the network is robust to this magnitude of change.
Rights and permissions
About this article
Cite this article
Chaos, Á., Aldana, M., Espinosa-Soto, C. et al. From Genes to Flower Patterns and Evolution: Dynamic Models of Gene Regulatory Networks. J Plant Growth Regul 25, 278–289 (2006). https://doi.org/10.1007/s00344-006-0068-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-006-0068-8