Abstract
Mice homozygous for the dreher (dr) mutation are characterized by pigmentation and skeletal abnormalities and striking behavioral phenotypes, including ataxia, vestibular deficits, and hyperactivity. The ataxia is associated with a cerebellar malformation that is remarkably similar to human Dandy-Walker malformation. Previously, positional cloning identified mutations in LIM homeobox transcription factor 1 alpha gene (Lmx1a) in three dr alleles. Two of these alleles, however, are extinct and unavailable for further analysis. In this article we report a new spontaneous dr allele and describe the Lmx1a mutations in this and six additional dr alleles. Strikingly, deletion null, missense, and frameshift mutations in these alleles all cause similar cerebellar malformations, suggesting that all dr mutations analyzed to date are null alleles.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mice homozygous for mutations at the dreher (dr) locus are characterized by a complex phenotype that includes circling behavior, balance abnormalities, hyperactivity, deafness, sterility, and pigmentation and tail abnormalities (Lyon 1961; Sweet and Wahlsten 1983; Washburn and Eicher 1986). The ataxia demonstrated by dr mutants is caused by a distinct phenotype of cerebellar hypoplasia and abnormal foliation preferentially affecting the cerebellar vermis. Interestingly, cerebellar abnormalities in dr/dr mice are very similar to those observed in patients with Dandy-Walker malformation (reviewed in Parisi and Dobyns 2003), suggesting that dreher mice represent a model for this common human birth defect. Other internal abnormalities in homozygous dr mice include neocortical, hippocampal, and spinal cord defects as well as skeletal defects and hypoplasia of Mullerian duct derivatives (Manzanares et al. 2000; Sekiguchi et al. 1992, 1994; Washburn and Eicher 1986), making dreher mice an excellent model to study neural, skeletal, and reproductive development.
Ten spontaneous dr mutant alleles have been described to date. The first dr allele was discovered in four abnormal mice captured in a factory in Detmold, Germany (Falconer and Sierth-Roth 1951). Subsequently, nine additional dr alleles have been identified (reviewed in Bergstrom et al. 1999). It is unknown if all dr alleles engender the same abnormal phenotype or if some phenotypic abnormalities are unique to specific dr alleles.
Using a positional cloning strategy, Lmx1a was identified as the dr gene (Millonig et al. 2000). The protein product of Lmx1a has two LIM domains and one homeodomain and belongs to a family of LIM-homeodomain transcription factors (Hobert and Westphal 2000; Hunter and Rhodes 2005). To date, Lmx1a mutations have been described in three dr alleles: dr sst, dr 3J, and dr J. Both dr sst and dr 3J alleles inactivate the Lmx1a gene by deletions that remove the 5′ portion of the gene, including the ATG translation initiation codon (Millonig et al. 2000). Both dr sst and dr 3J are extinct and not available for further analysis. Currently, dr J is the only molecularly defined allele available and mutant mice with this allele have been used extensively in embryologic and neurologic studies (Chizhikov and Millen 2004a; Costa et al. 2001; Manzanares et al. 2000; Millen et al. 2004; Millonig et al. 2000; Sekiguchi et al. 1992, 1994). These studies have led to several important conclusions regarding the role of this gene in dorsal central nervous system (CNS) development (reviewed in Chizhikov and Millen 2004b, 2005). Because the dr J mutation changes a conserved cysteine that is critical for zinc-finger integrity within the LIM1 domain of LMX1A, it is presumed that dr J is a null allele, although others have questioned this interpretation (Manzanares and Krumlauf 2000). Identification of a true null Lmx1a allele therefore is clearly important for further understanding the in vivo role of Lmx1a.
In this study we report a new dr allele (dr Kjmi) and determine the nature of the Lmx1a mutation in this and six other previously identified, yet molecularly uncharacterized, dr alleles (dr, dr 2J, dr 4J, dr 6J, dr 7J, and dr 8J). Through comparison of the cerebellar phenotypes caused by the dr Kjmi , dr 7J, and dr 8J alleles with that of the well-studied dr J allele, we conclude that despite different Lmx1a mutations in each allele, all dr homozygous mutant mice have comparable cerebellar phenotypes and all dr alleles most likely represent null alleles of Lmx1a. The Lmx1a mutations described in this study provide valuable information about specific residues and domains required for LMX1A function in vivo and will be useful for exploring this locus as a candidate gene for human Dandy-Walker malformation.
Materials and methods
Animals
The origin and genetic background of mice carrying the dr alleles are summarized in Table 1. dr J/+ mice (B6C3Fe a/a-Lmx1a dr-J) were obtained from The Jackson Laboratory (Bar Harbor, ME) and maintained by sib-pair mating. For six other alleles (dr, dr 2J, dr 4J, dr 6J, dr 7J, and dr 8J ) only genomic DNA and/or brain tissue was obtained from homozygous mutants and normal littermate controls. The dr kjmi mutation arose spontaneously on a C57BL/6J-129Sv mixed background within our own animal facility at the University of Chicago. Allelism of the new mutation was determined by mating heterozygous carriers to heterozygous dr J/+ mice. All animal studies were conducted according to IACUC guidelines.
DNA extraction, PCR amplification, and DNA sequencing
Genomic DNA was prepared from tail biopsies by standard protocols with proteinase K digestion/phenol extraction (Sambrook and Russell 2001). PCR was performed according to standard protocols. Primers and conditions used in the current study are available upon request. Bidirectional sequencing of Sephadex G-50 (Amersham Biosciences, Buckinghamshire, UK) purified PCR products was performed with the ABI BigDye Terminator Sequencing Kit (Applied Biosystems, Foster City, CA) and an automated capillary array sequencer (ABI 3100, Applied Biosystems).
The sequencing was performed on independent PCR reactions.
Cerebellar histology
To investigate the cerebellar mutant phenotype of dr mutant mice, cardiac perfusion was performed under anesthesia on dr J /dr J (n = 10), dr kjmi /dr kjmi (n = 7), dr 7J /dr 7J (n = 3), and dr 8J /dr 8J (n = 3) mice together with several of their normal littermates at 4–5 weeks of age. Cerebella were immediately removed and immediately fixed in Bouin’s solution [71.4% picric acid solution (1.2% wt/vol)/23.8% formalin/4.8% glacial acetic acid] for 24 h and washed for 2 days in 70% ethanol. The tissues were dehydrated in ethanol, embedded in paraffin, sagittally serially sectioned at 10 μM, and stained with hematoxylin and eosin. Alternatively, some were fixed in 4% PFA in PBS for 12 h and transferred to sucrose buffer (30% sucrose in PBS) for cryoprotection. Before sectioning, the cerebella were embedded in sucrose-gelatin (30% sucrose, 10% gelatin in distilled water]. The gelatin blocks were fixed in a sucrose-formalin solution (30% sucrose, 10% formalin in PBS). Brains were sagittally serially sectioned at 50 μM and stained with cresyl violet.
Results
Mutations
Table 1 describes the seven dr alleles investigated in the current study. Six of these (dr, dr 2J, dr 4J, dr 6J, dr 7J, and dr 8J) were previously established as dr alleles (Bergstrom et al. 1999 and references therein). The seventh allele was recently identified in our mouse colony at the University of Chicago, spontaneously occurring on a C57BL6/J-129Sv/J mixed background. The new mutation was inherited as an autosomal recessive locus with complete penetrance. Affected animals survive until adulthood but are hyperactive, uncoordinated, and ataxic. All have patches of white hair on the ventrum, a short or blunted tail, and a small cerebellum with abnormal foliation (Fig. 4 and data not shown). Because their phenotype was strikingly similar to the dr phenotype, we mated known heterozygotes to dr J/+ mice. Of the 27 offspring analyzed, 7 were affected, indicating allelism. We have designated our new dr allele as dr Kjmi.
Molecular characterization
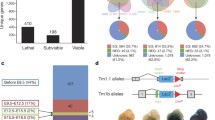
We first examined the structural integrity of the Lmx1a gene by amplifying genomic DNA from homozygous mutant mice with primer pairs specific to each of eight exons encoding full-length LMX1A (eF and eR, Fig. 1A, B). We successfully amplified all Lmx1a exons from seven control mice and from mice homozygous for the dr, dr Kjmi, and dr 2J alleles (Fig. 1C and data not shown). Most exons also were amplified from four other dr alleles, except exon 6 in dr 4J, exons 4-8 in dr 6J, exons 1 and 2 in dr 7J, and exons 4 and 5 in dr 8J (Fig. 1C), suggesting that these exons were deleted in the corresponding alleles. Loss of all exons was confirmed by additional independent primer pairs (eF′ and eR′, Fig. 1B). Also, in every case of a suspected deletion, we failed to amplify adjacent intronic DNA located both upstream (using primers uF and uR, Fig. 1B) and downstream (using primers dF and dR, Fig. 1B) of the deleted exons (data not shown), further supporting our conclusion of complete loss of the corresponding exons.
Lmx1a mutations in dreher alleles. (A) Schematic diagram of the mouse Lmx1a gene. Exons (1-8) encoding the complete Lmx1a protein are represented by solid boxes. Introns are illustrated by thin lines. The ATG translation initiation and the TGA termination codons are indicated. The LIM1 (exon 2) and LIM2 (exon 3) regions are yellow. The homeodomain (exons 4, 5, and 6) is red. (B) Location of four primer pairs used to assess deletions of each of Lmx1a exon in each of the dr alleles. Two primer pairs (eF and eR, and eF′ and eR′) are located within an exon. uF and uR are located within the intronic sequence immediately upstream of each exon. dF and dR are located within the intronic sequence immediately downstream of each exon. (C) PCR amplification of DNA from homozygous dr mice with primers eF and eR specific to each of the eight Lmx1a exons. dr alleles are indicated at the top. Exon numbers are indicated at the left. Wt-positive control: DNA from a wild-type mouse [although strain-specific control was used in every case, only one of them (B6/129Sv) is shown]. Neg: negative control (without DNA). (D–F) Chromatograms showing molecular alterations in dr (D), dr Kjmi (E), and dr 2J (F) alleles. Wild-type and mutant nucleotide and amino acid sequence are shown above each chromatogram. Base pair substitutions found in dr and dr Kjmi alleles are indicated by asterisks. Single base pair deletion found in dr 2J allele is indicated by an arrow. (G) Schematic representation of molecular alterations found in dr, dr Kjmi, dr 2J, dr 4J, dr 6J, dr 7JI, and dr 8J alleles. Exons (1-8) encoding the complete Lmx1a protein are represented by solid boxes. Introns are illustrated by thin lines. The ATG translation initiation and the TGA termination codons are indicated. The LIM1 (exon 2) and LIM2 (exon 3) domains are yellow. The homeodomain (exons 4, 5, and 6) is red. Deletions are indicated by dotted lines. Point mutations are indicated by asterisks.
The intact exons and flanking intronic regions were sequenced using genomic DNA from dr, dr 2J, dr 4J, dr 6J, dr 7J, dr 8J, and dr Kjmi homozygous mutant mice and several of their normal littermates. To ensure that the observed nucleotide changes were segregating with the phenotype and did not simply represent polymorphisms in the parental genetic background, we also sequenced DNA from the parental strains for each allele except the original dr allele because the original background of this allele is unknown. Single base pair substitutions were found in the dr and dr Kjmi alleles (Fig. 1D,E). A single base pair deletion was also detected in the dr 2J allele (Fig. 1E). No mutations were detected in the remaining intact exons of the dr 4J, dr 6J, dr 7J, and dr 8J alleles (Table 2).
Based on the Lmx1a mutations identified in this study, we can make several predictions regarding their effects on the LMX1A protein. Two mutations (dr 7J and dr 6J) most likely prevent effective LMX1A protein production. In the dr 7J allele, deletion of 5′ portion of the gene removes the ATG translation initiation codon and the Kozak sequence (as well as the sequence corresponding to the LIM1 domain). The dr 6J deletion removes the 3′ portion of the gene, including the sequence corresponding to the homeodomain and C-terminal portion of the protein in addition to the TGA transcriptional terminator codon and the polyadenylation site (Fig. 1G). In the dr 2J allele, deletion of one base pair occurs immediately after the ATG translation initiation codon. This allele is predicted to produce a very short protein consisting of 17 amino acids followed by 11 aberrant amino acids and a premature stop codon (Figs. 1F and 2). dr 4J and dr 8J have internal deletions of the Lmx1a gene, and if the normal splice sites are used for the intact exons, they are predicted to produce truncated LMX1A proteins (Fig. 2). Direct analysis of the Lmx1a transcripts and LMX1A proteins produced by dr 4J and dr 8J alleles is impossible because no tissue is available for these two mouse lines. The remaining two alleles (dr and dr Kjmi) encode missense mutations. The dr mutation results in an amino acid substitution that alters a conserved Asp (D) 44 to Val (V) in the LIM1 domain (Figs. 1D, 2, and 3A). The dr Kjmi mutation changes a conserved Arg (R) 127 to Pro (P) in the LIM2 domain (Figs. 1E, 2, and 3B). A summary of the entire dr allelic series is provided in Table 2.
Schematic representation of predicted LMX1A proteins produced by five dr alleles compared with normal mouse LMX1A protein. Amino acid sequence derived from each exon (1-8) is represented by boxes. The ATG translation initiation and the TGA termination codons are indicated. The LIM1 (exon 2) and LIM2 (exon3) regions are yellow. The homeodomain region (exons 4, 5, and 6) is red. Aberrant amino acid sequences are represented by hatched boxes. Missense mutations are indicated by asterisks. The structure of LMX1A proteins produced by dr 4J and dr 8J alleles is predicted based on the assumption that normal splice sites are used for the intact Lmx1a exons. Since we cannot experimentally test if normal splice sites are used in these alleles, hypothetical dr 4J and dr 8J LMX1A proteins are shown in brackets.
Multiple sequence alignment of LIM1 (A) and LIM2 (B) domains of representative LIM-homeodomain proteins. Only the N-terminal part of LIM1 and medial part of LIM2 domains are shown. Conserved residues are red letters. Nonconserved residues are black letters. Residues mutated in dr and dr Kjmi alleles are blue and are indicated by arrows.
Cerebellar abnormalities
One of the most striking features of Lmx1a mutants is abnormal cerebellar development resulting in a severely malformed adult cerebellum that contributes to the characteristic ataxic phenotype. The developing and adult cerebellar phenotypes are particularly well characterized in homozygous mutant dr J animals (Millonig et al. 2000), which was, until recently, the only publicly available dr allele. We therefore examined cerebellar foliation and cytoarchitecture in three additional dreher alleles (dr Kjmi, dr 7J and dr 8J) and compared them with cerebellar abnormalities found in homozygous dr J mice. Although different mutations in Lmx1a were found for each dr allele, homozygous mutant mice for each allele did not have significantly different cerebellar phenotypes. In each case, cerebellar foliation was consistently disrupted in homozygous mutants, with reduced cerebellar size and absent posterior vermis. The lateral cerebellar hemispheres were only moderately dysmorphic (Fig. 4). Despite disrupted folial patterning, histologic staining shows that the lamina of the adult cerebellum, including the molecular, Purkinje cell, and internal granule cell layers (Chizhikov and Millen 2003; Goldowitz and Hamre 1998), were present in homozygous mutants for all four alleles.
Comparison of cerebellar morphology of dr J, dr Kjmi, dr 7J, and dr 8J homozygous mutant mice. Dorsal views of dissected cerebella from adult wild-type (wt) (A), dr J (C), dr Kjmi (E), dr 7J (G), and dr 8J (I) mice. (B, D, F, H, J) Hematoxylin and eosin (H&E) or cresyl violet-stained sagittal sections of the vermis of cerebella shown in A, C, E, G, I.
Discussion
Series of allelic mutations in the mouse are valuable genetic tools for revealing the full spectrum of gene function in vivo and can mimic the range of human mutations and polymorphisms. Both truncations and more subtle amino acid changes are informative, revealing structural requirements for biological function. In this report we describe the molecular alterations in seven spontaneous dr alleles, including six previously identified dr alleles (dr, dr 2J,dr 4J, dr 6J, dr 7J, and dr 8J ) and one new allele (dr Kjmi). Together with the three previously characterized dr alleles, dr J , dr sst, and dr 3J (Millonig et al. 2000), these dr alleles represent an allelic series of ten defined mutations of the mouse Lmx1a gene. One additional reported dr allele, dr 5J, is extinct and no DNA is available for analysis.
Based on sequence analysis, five of ten molecularly defined dr alleles to date (dr sst, dr 3J , dr 7J, dr 2J, and dr 6J) most likely represent Lmx1a null alleles because they harbor deletions of the ATG translation initiation codon (dr sst, dr 3J, and dr 7J), the transcription termination TGA codon, and polyadenylation stop signal (dr 6J) or cause a frameshift with very early termination of the protein (dr 2J). In contrast, three other dr alleles (dr, dr J, and dr Kjmi) are predicted to generate an altered protein. They provide important information on critical residues and domains that are required for LMX1A function in vivo. Only one published report has examined the structural requirement for LMX1A functional activity to date (German et al. 1992). This study determined that by interacting with other promoter-binding proteins, LMX1A can activate transcription from the insulin promoter in vitro. Deletion of one or both of the LIM domains removed LMX1A activity, arguing that both LIM domains are critical for LMX1A-dependent transcriptional activation in this experimental system. The physiologic relevance of LMX1A binding to the insulin promoter, however, is unclear. Missense mutations in the LIM1 (dr and dr J) and the LIM2 (dr Kjmi) domains, however, suggest that each LIM domain is critical for LMX1A activity in vivo. Alternatively, point mutations observed in dr, dr J, and dr Kjmi alleles may lead to instability of the protein in vivo. Additional experiments are required to understand the exact role of these residues for LMX1A function.
To directly compare the phenotypic consequences of the different Lmx1a mutations, we compared the adult cerebellar phenotypes of available dr Kjmi, dr 7J, and dr 8J homozygous mutant mice to the cerebellar phenotype of homozygous mutant mice with the most studied allele, dr J. In every allele, the cerebellum was significantly reduced in size with comparable dysmorphic foliation. The posterior vermis was absent while the cerebellar hemispheres were less affected. Although the cerebellar phenotype was somewhat variable between multiple animals with the same genotype (data not shown), there were no significant differences in cerebellar phenotype across all genotypes and the variation between alleles was well within the range of variation that we have observed for dr J. Furthermore, there were no significant differences between the cerebellar phenotypes observed in this study and the published cerebellar phenotypes of mutant mice with the dr sst allele (Wahlsten et al. 1983) or the original dr allele (Falconer and Sierth-Roth 1951). Both dr sst and dr 7J are likely null alleles based on the sizes and locations of the genomic deletions. A Lmx1a null mutation was recently described in the rat. These mutant animals also have a strikingly similar adult cerebellar phenotype to dr mice (Kuwamura et al. 2005). Our finding that the cerebellar phenotypes of Lmx1a null mutants are comparable to the cerebellar phenotypes of mice carrying missense alleles dr J and dr Kjmi is extremely significant and argues that these missense mutations completely abrogate Lmx1a in vivo function. Supporting this conclusion is our previous finding that the Lmx1a protein with the dr J missense change, unlike wild-type Lmx1a, is incapable of inducing ectopic roof plate in chick spinal cord overexpression assays (Chizhikov and Millen 2004a).
In the mouse, many spontaneous mutations are represented by only one allele. The 11 mouse Lmx1a mutations have independently and spontaneously arisen during the last 60 years (Table 2). Two hypotheses may explain the frequent appearance of Lmx1a mutations. The Lmx1a locus may be extremely accessible to mutation representing a mutation “hot spot” in the vertebrate genome. Indeed, Lmx1a is a large gene, with exons encoded across 157 kb of genomic DNA (http://www.genome.ucsc.edu). A potentially more attractive hypothesis, however, is that the Lmx1a locus has a normal mutation rate, but the remarkable and very obvious behavioral and pigmentation phenotypes of homozygous mutants allow for easy detection.
The uniqueness of the dr phenotype is noteworthy. All 11 mutants in mice and one rat display characteristic neurologic phenotypes and associated pigmentation and skeletal phenotypes. All mutants with this phenotype are allelic to the dr locus and no other spontaneous or induced mutant mice have a similar phenotype. This is surprising because one can imagine that mutations affecting upstream or downstream components of the Lmx1a regulatory network may produce “dreher-like” phenotypes. We have previously shown that in the CNS Lmx1a expression is activated by peptides of the bone morphogenetic protein (BMP) family. LMX1A, in turn, also activates expression of multiple BMPs as well as members of the WNT family (Chizhikov and Millen 2004a, b; Millonig et al. 2000). Both the BMP and WNT families consist of multiple members that have redundant activities. Inactivation of single members of these families frequently has no obvious phenotype in mice, suggesting that LMX1A acts at the intersection of multiple signaling pathways, each of which is regulated by multiple redundant molecules.
The uniqueness of the dr phenotype also has implications for human neurogenetics, especially Dandy-Walker malformation, the most common malformation of the human cerebellum (Parisi and Dobyns 2003). Although heterozygous deletion of two linked genes, ZIC1 and ZIC4, has recently been reported as a cause of Dandy-Walker malformation, less than 5% of human Dandy-Walker malformation cases can be attributed to loss of these genes (Grinberg et al. 2004). The fact that the dr cerebellar phenotype is so remarkably similar to the cerebellar phenotype of Dandy-Walker patients leads to the prediction that Lmx1a alterations may also play a role in the pathogenesis of this important yet poorly understood human birth defect.
References
Bergstrom DE, Gagnon LH, Eicher EM (1999) Genetic and physical mapping of the dreher locus on mouse chromosome 1. Genomics 59:291-299
Chizhikov V, Millen KJ (2003) Development and malformations of the cerebellum in mice. Mol Genet Metab 80:54-65
Chizhikov VV, Millen KJ (2004a) Control of roof plate formation by Lmx1a in the developing spinal cord. Development 131:2693-7205
Chizhikov VV, Millen KJ (2004b) Mechanisms of roof plate formation in the vertebrate CNS. Nat Rev Neurosci 5:808-812
Chizhikov VV, Millen KJ (2005) Roof plate-dependent patterning of the vertebrate dorsal central nervous system. Dev Biol 277:287-295
Costa C, Harding B, Copp AJ (2001) Neuronal migration defects in the Dreher (Lmx1a) mutant mouse: role of disorders of the glial limiting membrane. Cereb Cortex 11:498-505
Falconer DS, Sierth-Roth U (1951) Dreher, a new gene of the waltzer-shaker group in the house mouse. Z Indukt Abstamm Vererbungsl 84:71-73
Hobert O, Westphal H (2000) Functions of LIM-homeobox genes. Trends Genet 16:75-83
Hunter CS, Rhodes SJ (2005) LIM-homeodomain genes in mammalian development and human disease. Mol Biol Rep 32:67-77
German MS, Wang J, Chadwick RB, Rutter WJ (1992) Synergistic activation of the insulin gene by a LIM-homeodomain protein and a basic helix-loop-helix protein: building a functional insulin minienhancer complex. Genes Dev 6:2165-2176
Goldowitz D, Hamre K (1998) The cells and molecules that make a cerebellum. Trends Neurosci 21:375-382
Grinberg I, Northrup H, Ardinger H, Prasad C, Dobyns WB, et al. (2004) Heterozygous deletion of the linked genes ZIC1 and ZIC4 is involved in Dandy-Walker malformation. Nat Genet 36:1053-1055
Kuwamura M, Muraguchi T, Matsui T, Ueno M, Takenaka S, et al. (2005) Mutation at the Lmx1a locus provokes aberrant brain development in the rat. Brain Res Dev Brain Res 155:99–106
Lyon MF (1961) Linkage relations and some pleiotropic effects of the dreher mutant of the house mouse. Genet Res 2:92-95
Manzanares M, Krumlauf R (2000) Developmental biology. Raising the roof. Nature 403:720-721
Manzanares M, Trainor PA, Ariza-McNaughton L, Nonchev S, Krumlauf R (2000) Dorsal patterning defects in the hindbrain, roof plate and skeleton in the dreher (dr(J)) mouse mutant. Mech Dev 94:147-156
Millonig JH, Millen KJ, Hatten ME (2000) The mouse Dreher gene Lmx1a controls formation of the roof plate in the vertebrate CNS. Nature 403:764-769
Millen KJ, Millonig JH, Hatten ME (2004) Roof plate and dorsal spinal cord dl1 interneuron development in the dreher mutant mouse. Dev Biol 270:382-392
Parisi MA, Dobyns WB (2003) Human malformations of the midbrain and hindbrain: review and proposed classification scheme. Mol Genet Metab 80:36-53
Sambrook J, Russel DW (2001) In Molecular cloning: a laboratory manual, 2nd edn. (Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press)
Sekiguchi M, Shimai K, Guo H, Nowakowski RS (1992) Cytoarchitectonic abnormalities in hippocampal formation and cerebellum of dreher mutant mouse. Brain Res Dev Brain Res 67:105-112
Sekiguchi M, Abe H, Shimai K, Huang G, Inoue T, et al. (1994) Disruption of neuronal migration in the neocortex of the dreher mutant mouse. Brain Res Dev Brain Res 77:37-43
Sweet HO, Wahlsten D (1983) Shaker short-tail (sst). Mouse News Lett 69:26
Wahlsten D, Lyons JP, Zagaja W (1983) Shaker short-tail, a spontaneous neurological mutant in the mouse. J Hered 74:421-425
Washburn L, Eicher EM (1986) A mutation at the dreher locus (dr2J). Mouse News Lett 75:28-29
Acknowledgments
The authors thank Eva M. Eicher for providing dr, dr 2J,dr 4, dr 6J, dr 7J, and dr 8J genomic DNA and for her valuable comments on the manuscript. They also thank Kanayo Tatsumi for her expert histologic assistance, and Jennifer Schmidt, Marissa Blank, Anne Lindgren, and Rumen Slavkov for helpful discussions and insights. This work was supported by NIH grants RR01183 to Eva M. Eicher and R01 NS044262 to KJM.
Author information
Authors and Affiliations
Corresponding author
Additional information
Victor Chizhikov and Ekaterina Steshina contributed equally to this work.
Rights and permissions
About this article
Cite this article
Chizhikov, V., Steshina, E., Roberts, R. et al. Molecular definition of an allelic series of mutations disrupting the mouse Lmx1a (dreher) gene. Mamm Genome 17, 1025–1032 (2006). https://doi.org/10.1007/s00335-006-0033-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00335-006-0033-7