Abstract
Objectives
Radiomics of soft tissue sarcomas (STS) is assumed to correlate with histologic and molecular tumor features, but radiogenomics analyses are lacking. Our aim was to identify if distinct patterns of natural evolution of STS obtained from consecutive pre-treatment MRIs are associated with differential gene expression (DGE) profiling in a pathway analysis.
Methods
All patients with newly diagnosed STS treated in a curative intent in our sarcoma reference center between 2008 and 2019 and with two available pre-treatment contrast-enhanced MRIs were included in this retrospective study. Radiomics features (RFs) were extracted from fat-sat contrast-enhanced T1-weighted imaging. Log ratio and relative change in RFs were calculated and used to determine grouping of samples based on a consensus hierarchical clustering. DGE and oncogenesis pathway analysis were performed in the delta-radiomics groups identified in order to detect associations between delta-radiomics patterns and transcriptomics features of STS. Secondarily, the prognostic value of the delta-radiomics groups was investigated.
Results
Sixty-three patients were included (median age: 63 years, interquartile range: 52.5–70). The consensus clustering identified 3 reliable delta-radiomics patient groups (A, B, and C). On imaging, group B patients were characterized by increase in tumor heterogeneity, necrotic signal, infiltrative margins, peritumoral edema, and peritumoral enhancement before the treatment start (p value range: 0.0019–0.0244), and, molecularly, by downregulation of natural killer cell–mediated cytotoxicity genes and upregulation of Hedgehog and Hippo signaling pathways. Group A patients were characterized by morphological stability of pre-treatment MRI traits and no local relapse (log-rank p = 0.0277).
Conclusions
This study highlights radiomics and transcriptomics convergence in STS. Proliferation and immune response inhibition were hyper-activated in the STS that were the most evolving on consecutive imaging.
Key Points
• Three consensual and stable delta-radiomics clusters were identified and captured the natural patterns of morphological evolution of STS on pre-treatment MRIs.
• These 3 patterns were explainable and correlated with different well-known semantic radiological features with an ascending gradient of pejorative characteristics from the A group to C group to B group.
• Gene expression profiling stressed distinct patterns of up/downregulated oncogenetic pathways in STS from B group in keeping with its most aggressive radiological evolution.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Soft tissue sarcomas (STS) are rare malignant mesenchymal tumors characterized by their clinical, radiological, and molecular heterogeneity, which can lead to diagnostic uncertainties [1]. Consequently, the time elapsed from initial symptoms to final histological diagnosis can reach months during which the tumor can evolve naturally and grow without constraints [2].
MRI is the imaging of reference for the local staging of STS, treatment response assessment, and local relapse detection [3]. MRI enables to evaluate classical prognostic features such as size and depth, which are included in nomograms, as well as the surgical operability that guides the therapeutic strategy [3,4,5]. Moreover, baseline MRI features (i.e., peritumoral enhancement, necrotic signal, or intra-tumoral heterogeneity on conventional sequences) have been correlated with histologic grade and survivals [6, 7]. However, these “semantic” features are subjective [8] and do not account for the natural changes (i.e., in the absence of any medical intervention) observed when multiple MRIs are performed during the diagnostic interval.
Yet, significant improvements in the quantification of the tumor radiological phenotype (or “radiophenotype”) have been achieved through radiomics [9, 10]. Basically, radiomics correspond to the extensive quantification of the radiophenotype thanks to mathematical operators applied on any imaging modality. A key hypothesis is that the heterogeneity captured by radiomics radiophenotyping reflects the tumor histologic and molecular characteristics. The resulting radiomics features (RFs) are incorporated in supervised machine learning pipelines to predict the patients’ outcomes or to identify associations with relevant histologic and molecular status, and unsupervised algorithms (i.e., algorithms that analyze and cluster unlabeled dataset according to the similarity of their RFs) in order to identify hidden patterns and subgroups of similar observations when there is no a priori hypothesis [9].
So far, radiomics approaches have been successfully applied to STS to predict the histologic grade [11,12,13,14], metastatic relapse and cancer-related death [12, 15,16,17], and treatment response [18,19,20]. Moreover, the interest of evaluating the changes in the STS radiophenotype during treatments through “delta-radiomics” approaches (i.e., quantitative variation in RFs between two evaluations) has been demonstrated during neoadjuvant chemotherapy and radiotherapy [18, 19].
However, as opposed to other neoplasms (such as glioma, urothelial carcinoma, lung adenocarcinoma, clear cell carcinoma, or ovarian cancer), correlations between radiomics, delta-radiomics, and transcriptomics features (i.e., corresponding to the expression levels and profiles of multiple ribonucleic acids [RNA] transcripts) in sarcoma are missing.
Considering the rapid evolution and diagnostic delays seen in STS patients, multidisciplinary boards in sarcoma reference centers sometimes require to update the imaging in order to refine the local staging. Comparing these pre-treatment MRIs acquired before any medical intervention highlights distinct patterns of “natural” evolution of STS, from global morphological stability to appearance of strong intra-tumoral heterogeneity and infiltrative margins, which suggests different biological pathways involved in tumorigenesis. Our group has recently investigated the clinical interest of assessing natural radiological and radiomics changes in STS before starting initial treatments [21]. Surprisingly, the natural changes in RFs were not associated with histological type or simple histologic markers of proliferation like Ki-67 or mitotic count, which prompted us to hypothesize that delta-radiomics would rather reflect other oncogenetic pathways and could complement standard radiological and histologic analyses.
Consequently, our aim was to establish a reliable classification of the radiophenotype of STS based on the natural changes in RFs before treatment and using unsupervised algorithms, and to investigate whether those delta-radiomics radiophenotypes could be biologically explained through gene expression profiling.
Materials and methods
Study design
This single-center retrospective study was IRB-approved. Patients were retrieved from our institutional database between 2008 and 2019. The inclusion criteria were age ≥ 16 years, newly diagnosed histopathologically proven STS of extremities, with two available contrast-enhanced (CE) MRIs before the treatment beginning (named MRI0 and MRI1), and treated in a curative intent including a surgical resection in our sarcoma reference center. The exclusion criteria were lack of fat-sat (FS) turbo spin echo (TSE) CE-T1-weighted imaging (WI) in at least one of the two pre-treatment MRIs, non-diagnostic image quality due to MRI artifacts, delay between pre-treatment MRIs below 3 weeks, and atypical lipomatous tumors. Figure 1 shows the study flowchart. Sixty out of the 63 finally included patients were previously analyzed in another preliminary study [21], which focused on the morphological changes using standard radiological analyses, depiction of delta-radiomics calculation, and associations with basic histologic features.
The data collection consisted of age at diagnosis, sex, World Health Organization Performance Status (WHO-PS), tumor location and depth, pre-treatment histological type, and grade (according to the “Fédération Nationale des Centres de Lutte Contre le Cancer” [FNCLCC] grading system) [22]. The initial management was also recorded as follows: surgical margins (R0 versus R1-R2), radiotherapy, and chemotherapy.
Radiomics pipeline
All MRI examinations were acquired on 1.5-T magnets from 9 radiological centers (with most MRI1 in our sarcoma reference center) and included T1-WI, T2-WI, and TSE FS-CE-T1-WI. The MR systems were Optima Jem MR450W, Discovery MR450, and SIGNA Explorer (General Electric Healthcare); Achieva and Ingenia (Philips Healthcare); and Magnetom Aera and Amira (Siemens Healthineers).
The radiomics analysis was only achieved on the FS-CE-T1-WI, as this sequence had the most homogeneous acquisition scheme at the two time points over the entire cohort, and was the most informative regarding the tumor radiological architecture. The ranges of acquisition parameters were echo/repetition times: 8–20/350 – 1000 ms for the T1-WI and CE-T1-WI, and 40–120/2000 – 9000 ms for the T2-WI, respectively; in plane resolution: 0.75 × 0.75 − 1.4 × 1.4 mm, and thickness: 1–7 mm. Contrast agents were gadoterate meglumine (Dotarem®, Guerbet), gadobutrol (Gadovist®, Bayer), and gadobenate dimeglunine (Multihance®, Bracco).
Since FS-CE-T1-WI were obtained on different MRI systems and given the lack of standardized units for conventional MRI, a post-processing pipeline was achieved to homogenize the MRI dataset by using the ITK library and, then, to extract 108 3D RFs from the 3D volume of each STS on MRI0 and MRI1 by using the Olea Sphere software (v3.0-SP16, Olea Medical) [23,24,25]. The full radiomics pipeline and quality control, and RF definitions are detailed in Supplementary Data S1 and S2.
Conventional radiological analysis
The morphological changes occurring between MRI0 and MRI1 were also evaluated using a standard radiological approach. Three radiologists (2 seniors and 1 fellow) reviewed in consensus the tumor longest diameter (LD) and the 6 following items characterizing the tumor changes in signal, compartments, and surrounding tissues: change in intra-tumoral heterogeneity on FS-CE-T1-WI (categorized as homogeneous or similar heterogeneity versus increase); change in necrotic signal (defined as high, fluid-like signal on T2-WI and no enhancement on FS-CE-T1-WI, and categorized as no necrosis or similar amount versus increase); change in hemorrhagic signal (defined as high signal on T1-WI and no enhancement on FS-CE-T1-WI, and categorized as no hemorrhage or similar amount versus increase); change in MRI growth pattern on FS-CE-T1-WI (categorized as perfectly well-delineated tumor versus < 50% of tumor circumference with ill-defined margins versus ≥ 50% of tumor circumference with ill-defined margins) [26]; change in peritumoral edema on T2-WI (categorized as absent or similar versus increase) [6, 7]; and change in peritumoral enhancement on FS-CE-T1-WI (categorized as absent or similar versus increase) [6, 7]. Visual examples are shown in Supplementary Data S3.
RNA sequencing
All patients with frozen material before treatment and contemporary to MRI1 were included in this analysis. After whole-RNA sequencing (Integragen), the produced RNA sequences were quality controlled and aligned to the transcriptome. Gene expression was then estimated by the counts of high-quality sequences aligned per gene. Finally gene expression counts were normalized using the Voom method (Supplementary Data S4) [27].
Statistical analysis
Statistical analyses were performed with R (v4.1.0). All tests were two-tailed. A p value < 0.05 was deemed significant.
Methods to calculate delta-radiomics
Five mathematical methods have been proposed to calculate the change in RFs during the MRI0-MRI1 interval and were applied to the cohort, namely absolute change (AC), time-weighted absolute change (TWAC), relative change (RT), time-weighted relative change (TWRC), and log ratio (LR) (Supplementary Data S5) [21]. We then assessed how the delta-radiomics distributions could be affected by the scale and shape bias potentially associated to each method in order to filter the delta RFs on which the unsupervised algorithms would correctly be applied. Briefly, AC was excluded because it could scale-bias the selection of significant delta RFs by promoting high-scale features and penalizing low-scale ones. Regarding TWAC and TWRC, a bias could result from time delay variability across patients and from how time could potentially non-linearly affect each RF (i.e., plateau effect, cycle effect). RC gives equal odds to both small- and large-scale features and it respects the symmetric and Gaussian nature of the delta-radiomics distribution. Similarly, LR is not affected by scale bias and confers a bell-like shape to the distribution. Consequently, RC and LR delta RFs were used in the subsequent analysis while AC, TWAC, and TWRC delta-RFs were removed.
Consensus clustering
We performed two consensus clustering independently, one with RF delta-radiomics calculated with LR and one with RC. Each clustering was resampled 10,000 times by leave-one-out of 40% of the samples [28]. Both clustering provided 3 groups. To find a final agreement between the two grouping schemes, we created the confusion matrix between the labeling of each sample generated by the LR and RC clustering analyses. We defined the agreement groups as those that maximized the counts of samples placed in the diagonal of the matrix (Supplementary Data S6).
Explaining the delta-radiomics groups with the radiological features
Associations between the delta-radiomics groups and the radiological features, LD on MRI0, the relative change in LD, and the delays between the two pre-treatment MRIs were assessed using the Kruskal-Wallis test. Associations with the main histological types were assessed with the chi-2 test. Linear trends in the proportion of the radiological features across the delta-radiomics groups were investigated by using the chi-square test for trend in proportion.
Associations with the transcriptomics features
Differential gene expression (DGE) and geneset enrichment analyses between the final delta-radiomics groups are detailed in Supplementary Data S7. Exploratory DGE was performed by t-test calculation per gene. To discriminate significant up/downregulated genes, the fold change was set to 1.5. The p value cut-off of 0.05 was not adjusted as this study was exploratory on a small cohort of heterogeneous histotypes. However, to confirm that the results were anyway statistically significant, we applied an alternative method named significance analysis of microarrays (SAM), which has been designed to investigate significance in the setting of small size cohorts for -omics data. The principles and application are detailed in Supplementary Data S8 [29].
We then assessed geneset enrichment in biological pathways based on Broad Institute’s Molecular Signature database and the CIBERSORT LM22 immuno-genesets [30, 31].
Survival analysis
Metastatic and local relapse-free survivals (MFS and LFS, respectively) were defined as the time elapsed between surgical resection and histologically proven metastatic and local relapses, respectively, or death or last patient contact. Overall survival (OS) was defined as the time elapsed from surgery to death due to disease or last patient contact. The Kaplan-Meier curves for MFS, LFS, and OS depending on the delta-radiomics clusters were drawn and the differences in survivals were tested with the log-rank test. Univariate Cox regressions were performed to estimate the hazard ratio (HR) with 95% confidence interval (CI) of the delta-radiomics groups.
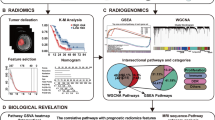
The overall study workflow is presented on Fig. 2.
Results
Study population
The study population finally included 63 patients with complete delta-radiomics data (median age: 63 years, range: 21–95, interquartile range [IQR]: 52.5–70, 24/63 [38.1%] women) (Table 1 and Fig. 1). The most frequent histological type was undifferentiated pleomorphic sarcoma (UPS, 29/63 [46%]).
The median delay between the two pre-treatment MRIs was 74 days (range: 24–939, IQR: 55–105). The median LDs were 98 mm (range: 12–199, IQR: 67–121) on MRI0 and 120 mm (range: 22–270, IQR: 89–153.5) in MRI1, which corresponded to a median increase of 21.8% (range: 0–158.3, IQR: 9.3–44) over the inter-MRIs delay.
Defining and explaining the delta-radiomics groups
We independently performed iterative unsupervised hierarchical clustering on both relative change and on log ratio delta-radiomics as these two methods were theoretically the less prone to distribution and scaling bias (Fig. 3a). To check the consensus given by the two clusterings (Fig. 3b), an agreement table was created, which finally defined 3 consensus groups called A, B, and C (n = 16, 15, and 21, respectively), for a total of 52 patients (Fig. 3c). The 11 patients who did not achieve consensus were excluded. We did not find associations between the delta-radiomics groups and LD on MRI0, the relative change in LD, and the MRI0-MRI1 delay (p = 0.5912, 0.4503, 0.4256, and 0.4114, respectively), as well as associations with the main histological types (as categorized as UPS, myxoid/round cells liposarcomas, and others, p = 0.3594) and the histologic grade assessed on whole surgical specimen (categorized as grade I-II versus grade III, p = 0.2273).
Definition of the radiomics groups. a Assessment of the distribution of the delta-radiomics features (RF) computed according of the five proposed methods. b Hierarchical clustering based on relative change and log ratio delta RFs identified 3 groups with good reproducibility via 10,000 times cross-validation repeated on the patient samples. Relative change groups: A, B, and C. Log ratio groups: A′, B′, C′. c Agreement table between the relative change and log ratio clustering schemes
Confronting the delta-radiomics groups and the conventional radiological analysis stressed the largest proportion of morphological changes in the B group, namely increases in tumor heterogeneity (10/15, 66.7%), necrotic signal (14/15, 93.3%), hemorrhagic signal (10/15, 66.7%), infiltrative margins (9/15, 60%), peritumoral edema (8/15, 53.3%), and peritumoral enhancement (9/15, 60%) (Fig. 4, Supplementary Data S9). Significant ascending linear trends were found from A group, to C group, to B group for the following proportions: increase in tumor heterogeneity (p = 0.0199), necrotic signal (p = 0.0244), infiltrative margins (p = 0.0032), peritumoral edema (p = 0.0019), and peritumoral enhancement (p = 0.0166).
Associations between the radiomics three groups and the changes in the radiological features of interest during the diagnostic interval. (a) Color-coded matrix indicating the proportion of patients in each radiomics groups (column) with the radiological features of interest (row). §: The p value indicated in the last column corresponds to the test assessing significant trend from group A to group B to group C. *: p < 0.05; **: p < 0.005. Abbreviations: n.s., not significant; CE-T1-WI, contrast-enhanced T1-weighted imaging. Prototypal changes observed on contrast-enhanced MRI of soft tissue sarcomas from group A (b), global stability of the tumor and its surrounding tissues), group C (c), increase in necrotic signal and heterogeneity with a global stability of the surrounding tissues), and group B (d), increase in necrotic signal, tumor heterogeneity, peritumoral enhancement, and worse definition of the tumor margin, i.e., increase in infiltrative growth pattern)
Correlating delta-radiomics and gene expression
Twenty-one patients had available tumor tissue for RNA sequencing of patients of group A (5/21 [23.8%]), group B (8/21 [38.1%]), and group C (8/21 [38.1%]). To identify the transcriptomics features associated to tumors with aggressive outcome against the others, we performed a supervised analysis by comparing the gene expression patterns of the tumors from group B versus the pool of tumors of group A plus C. This revealed 566 differentially expressed genes and 105 enriched pathways (Fig. 5a, Supplementary Data S10). The alternative statistical method, SAM, allowed us to confirm that those genes (except for one, MAGEA3) were statistically significant, with a q value ≤ 2.16% (Supplementary Data S11). Table 2 shows the most significantly enriched genesets. Overall, B group tumors were characterized by a downregulation of natural killer (NK) cell–mediated cytotoxicity genes and an upregulation of genes involved in the glycoproteins, sphingolipids, and peptide hormone biosynthesis, the Hedgehog pathway (cell differentiation during embryogenesis), the Hippo signaling pathway (regulation of cell proliferation and apoptosis), the FAS pathway (regulation of cell death), Ephrin B reverse signaling (cell adhesion and migration) pathway, and the extracellular matrix organization pathway.
Transcriptomics analysis. Volcano plots for oncogenetic pathways in the 21 sarcomas and (a) and 8 undifferentiated pleomorphic sarcomas (UPS, b). Main differentially expressed pathways of oncogenesis in UPS from the radiomics group B versus groups A and C (c). Synthesis of the activated oncogenetic pathways from the radiomics groups A and C and from the radiomics group B (d). Abbreviations: CE-T1-WI, contrast-enhanced T1-weighted imaging; MRI0 and MRI1, magnetic resonance imaging performed during the diagnostic interval and before the beginning of treatment. Dashed gray lines correspond to significance (p value < 0.05)
To verify whether the trends identified in the overall set of samples were conserved in the most frequent histotype, we reproduced the analysis in the UPS subgroup, which revealed 1466 differentially expressed genes. The enrichment analysis confirmed the trends found in the entire cohort with 162 enriched pathways (Fig. 5c, d, Supplementary Data S12-S13). In summary, typical oncogenesis functions like regulation of cell proliferation and apoptosis, and inhibition of the immune response were upregulated in the patients from the more aggressive B group versus the A+C pool.
Survival analysis
The median follow-up was 82.1 months. There were 24 metastatic relapses, 10 local relapses, and 16 deaths related to disease (Supplementary Data S14). No correlations were found between the delay from MRI0 and MRI1 and MFS, LFS, and OS in univariate Cox regression (p = 0.7680, 0.8230, and 0.3680, respectively). No significant differences were found between the MFS, LFS, and OS depending on the three radiomics groups (p = 0.6731, 0.0622, and 0.8970, respectively) (Table 3). However, none of the patients from the A group demonstrated local relapse while the local relapse rates from groups B and C were similar at 2 years (73.3%, 95%CI: 54–99.5, and 78.5%, 95%CI: 61.8–99.7). Thus, significant higher LFS were observed in the group A versus the B-C groups (p = 0.0277) (Fig. 6).
Discussion
Medical imaging enables longitudinal and non-invasive monitoring of the changes occurring in tumor and their surrounding tissues. Since STS are prone to long diagnostic intervals, which require updating the local staging, we proposed an original delta-radiomics-based methodology to quantify the “natural” shape and textural changes arising before any therapeutic intervention. We found three consistent patterns of natural evolution, which were radiologically and biologically explainable, and demonstrated distinct oncogenetic pathway activations.
We transposed bioinformatics tools to the radiomics field to obtain three reliable groups, according to resampling-based consensus hierarchical clustering performed on normally distributed delta RFs from two RF calculation methods. We believe this design counterbalanced the noise due to our small population. These three groups were comparable regarding the main histological types, MRI0-MRI1 delays, initial size, and change in LD from MRI0 to MRI1.
A frequent reproach to radiomics is the lack of explicability with human vocabulary. Thus, to further validate the delta-radiomics groups, we investigated whether they could be explainable by radiologists. Indeed, radiologically, a gradient of morphological evolution was noticed from the radiomics group A to the radiomics group B in terms of changes in the intra-tumoral aspect (increase in visual heterogeneity, complexity, and necrosis) and in the tumor periphery (increase in peritumoral edema, peritumoral enhancement, and more infiltrative margins). At baseline, these radiological features have been linked to higher grade and poorer survivals [6, 7, 25, 32]. Moreover, we observed that the delta-radiomics groups (evaluating the intra-tumoral texture and not the surrounding tissues) were associated with the changes in radiological features evaluating the peritumoral tissues, highlighting the collinearity of intra- and peritumoral evolutions.
Logically, if these features appear or amplify before beginning the treatment, this could have a negative impact on the patients’ outcome. Indeed, we found that no local relapse occurred in the A group compared with the two others, whereas the lowest LFS was seen in the B group. The lack of modification of the peritumoral tissue and tumor margins on the MRI of A group patients is in agreement with this better local control. Indeed, abnormal peritumoral MRI findings have been associated with satellite tumor cells spreading [33, 34].
Although a fundamental assumption behind radiomics is the relationship between gene expression and radiophenotype, no studies had directly investigated this link in STS. Since our delta-radiomics groups reflected the natural evolutive patterns before any intervention, we hypothesized that they would also reflect the unconstrained expression of the tumor genotype. Indeed, we identified differentially expressed genes from main oncogenetic pathways associated to the delta-radiomics groups. The B group, with the most aggressive aspect and the worst local outcome, was characterized by an overexpression of genes involved in the Hedgehog pathway and by a lower expression of genes belonging to the NK cell genesets. Our results are in agreement with previous studies, which demonstrated the important role of developmental pathways including Hedgehog and NK cells in the tumorigenesis and immune surveillance of sarcomas [35, 36].
Our findings could have practical applications. Regarding the poorer LFS in the B group, these patients could benefit from more aggressive locoregional treatments including larger surgical margins, higher dose of peri-operative radiotherapy, and closer local follow-up. Regarding systemic treatments, some of the identified differentially expressed genes and pathways may represent targetable therapeutic vulnerabilities. For instance, anti-angiogenic treatments or Bcl-2 inhibitors may be more effective in the B group. In the future, medical imaging may help to tailor the therapeutic strategy in STS patients, and to perform a “virtual” biopsy inferred from the tumor radiophenotype.
Our study has limitations. First, this was a retrospective single-center study, which implies heterogeneous MRI protocols and lack of frozen specimens for RNA-sequencing in a third of patients. Hence, our delta-radiomics analysis was achieved on FS-CE-T1-WI only. Second, dynamic contrast enhancement, spectroscopy and diffusion imaging, and positron emission tomography could have improved the assessment of tumor angiogenesis, cellularity, and metabolism [35, 37] but they are never performed outside reference centers. Third, the small population led to a lack of statistical power for transcriptomics analyses, which prompted us not to adjust the p values with standard method in order to detect biologically meaningful trends; however, our results were confirmed using an alternative method named SAM which is dedicated to very imbalanced datasets in terms of number of variables and number of samples [29]. Fourth, there were important variations in the delay between the two pre-treatment MRIs that could have influenced our findings though no association was found between this delay and patients’ outcome. Fifth, in order to limit the exclusion of several RFs before the calculation and selection step of delta RFs, we choose to apply a long post-processing pipeline in order to obtain comparable images between patients and between two evaluations per patient and, then, to make the VOIs adjusted and validated by the most experienced radiologists from our sarcoma reference center, this instead of filtering RFs at a single time point depending on their reproducibility over various voxel sizes, numbers of gray levels, or experience of the segmenting radiologist. Sixth, although the final agreement table between the consensus clusterings on LR and RC features was an additional security to avoid tumor misclassification, it led to the additional exclusion of 11 patients (17%) without agreement.
In conclusion, this exploratory study provides new research leads regarding the radiogenomics of STS. We have highlighted original correlations between meaningful delta-radiomics-based clusters quantifying the natural tumor changes before any treatment, and important oncogenetic pathways and local control of the disease after curative treatment, bridging the gap between quantitative imaging and cancer biology.
Abbreviations
- AC:
-
Absolute change
- CE:
-
Contrast-enhanced
- CI:
-
Confidence interval
- DGE:
-
Differential gene expression
- FNCLCC:
-
French “Fédération Nationale des Centres de Lutte Contre le Cancer”
- FS:
-
Fat sat
- HR:
-
Hazard ratio
- IQR:
-
Interquartile range
- IRB:
-
Institutional Review Board
- LD:
-
Longest diameter
- LFS:
-
Local relapse-free survival
- LR:
-
Log ratio
- MFS:
-
Metastatic relapse-free survival
- MRI:
-
Magnetic resonance imaging
- NK:
-
Natural killer
- OS:
-
Overall survival
- RF:
-
Radiomics feature
- RNA:
-
Ribonucleic acids
- SAM:
-
Significance analysis of microarrays
- SI:
-
Signal intensity
- STS:
-
Soft tissue sarcoma
- TSE:
-
Turbo spin echo
- TWAC:
-
Time-weighted absolute change
- TWRC:
-
Time-weighted relative change
- WI:
-
Weighted imaging
References
Fletcher CDMF, Bridge JA, Hogendoorn PCW, Martens F (2020) WHO classification of soft tissue and bone tumours, 5th edn. IARC Publications, Lyon, France
Soomers V, Husson O, Young R et al (2020) The sarcoma diagnostic interval: a systematic review on length, contributing factors and patient outcomes. ESMO Open 5:e000592
Gronchi A, Miah AB, Dei Tos AP et al (2021) Soft tissue and visceral sarcomas: ESMO-EURACAN-GENTURIS Clinical Practice Guidelines for diagnosis, treatment and follow-up☆. Ann Oncol S0923-7534(21):02184–0
Callegaro D, Miceli R, Bonvalot S et al (2016) Development and external validation of two nomograms to predict overall survival and occurrence of distant metastases in adults after surgical resection of localised soft-tissue sarcomas of the extremities: a retrospective analysis. Lancet Oncol 17:671–680
Coindre JM, Terrier P, Bui NB et al (1996) Prognostic factors in adult patients with locally controlled soft tissue sarcoma. A study of 546 patients from the French Federation of Cancer Centers Sarcoma Group. J Clin Oncol 14:869–877
Zhao F, Ahlawat S, Farahani SJ et al (2014) Can MR imaging be used to predict tumor grade in soft-tissue sarcoma? Radiology 272:192–201
Crombé A, Marcellin P-J, Buy X et al (2019) Soft-tissue sarcomas: assessment of MRI features correlating with histologic grade and patient outcome. Radiology 291:710–721
Peeken JC, Neumann J, Asadpour R et al (2021) Prognostic assessment in high-grade soft-tissue sarcoma patients: a comparison of semantic image analysis and radiomics. Cancers (Basel) 13:1929
Limkin EJ, Sun R, Dercle L et al (2017) Promises and challenges for the implementation of computational medical imaging (radiomics) in oncology. Ann Oncol 28:1191–1206
Lambin P, Leijenaar RTH, Deist TM, et al (2017) Radiomics: the bridge between medical imaging and personalized medicine. Nat Rev Clin Oncol. 2017;14:749–762
Corino VDA, Montin E, Messina A et al (2018) Radiomic analysis of soft tissues sarcomas can distinguish intermediate from high-grade lesions. J Magn Reson Imaging 47:829–840
Peeken JC, Spraker MB, Knebel C et al (2019) Tumor grading of soft tissue sarcomas using MRI-based radiomics. EBioMedicine 48:332–340
Yan R, Hao D, Li J et al (2021) Magnetic resonance imaging-based radiomics nomogram for prediction of the histopathological grade of soft tissue sarcomas: a two-center study. J Magn Reson Imaging 53:1683–1696
Wang H, Chen H, Duan S et al (2020) Radiomics and machine learning with multiparametric preoperative MRI may accurately predict the histopathological grades of soft tissue sarcomas. J Magn Reson Imaging 51:791–797
Crombé A, Fadli D, Buy X et al (2020) High-grade soft-tissue sarcomas: can optimizing dynamic contrast-enhanced MRI postprocessing improve prognostic radiomics models? J Magn Reson Imaging 52:282–297
Crombé A, Le Loarer F, Sitbon M et al (2020) Can radiomics improve the prediction of metastatic relapse of myxoid/round cell liposarcomas? Eur Radiol 30:2413–2424
Spraker MB, Wootton LS, Hippe DS et al (2019) MRI radiomic features are independently associated with overall survival in soft tissue sarcoma. Adv Radiat Oncol 4:413–421
Crombé A, Périer C, Kind M et al (2019) T2 -based MRI delta-radiomics improve response prediction in soft-tissue sarcomas treated by neoadjuvant chemotherapy. J Magn Reson Imaging 50:497–510
Peeken JC, Asadpour R, Specht K et al (2021) MRI-based delta-radiomics predicts pathologic complete response in high-grade soft-tissue sarcoma patients treated with neoadjuvant therapy. Radiother Oncol 164:73–82
Chen S, Li N, Tang Y et al (2021) Radiomics analysis of fat-saturated T2-weighted MRI sequences for the prediction of prognosis in soft tissue sarcoma of the extremities and trunk treated with neoadjuvant radiotherapy. Front Oncol 11:710649
Fadli D, Kind M, Michot A et al (2021) Natural changes in radiological and radiomics features on MRIs of soft-tissue sarcomas naïve of treatment: correlations with histology and patients’ outcomes. J Magn Reson Imaging. https://doi.org/10.1002/jmri.28021
Trojani M, Contesso G, Coindre JM et al (1984) Soft-tissue sarcomas of adults; study of pathological prognostic variables and definition of a histopathological grading system. Int J Cancer 33:37–42
McCormick M, Liu X, Jomier J et al (2014) ITK: enabling reproducible research and open science. Front Neuroinform 8:13
Tustison NJ, Avants BB, Cook PA et al (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29:1310–1320
Nyúl LG, Udupa JK (1999) On standardizing the MR image intensity scale. Magn Reson Med 42:1072–1081
Nakamura T, Matsumine A, Matsubara T et al (2017) Infiltrative tumor growth patterns on magnetic resonance imaging associated with systemic inflammation and oncological outcome in patients with high-grade soft-tissue sarcoma. PLoS One 12:e0181787
Law CW, Chen Y, Shi W, Smyth GK (2014) voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol 15:R29.
Wilkerson MD, Hayes DN (2010) ConsensusClusterPlus: a class discovery tool with confidence assessments and item tracking. Bioinformatics 26:1572–1573
Tusher VG, Tibshirani R, Chu G (2001) Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci U S A 98:5116–5121
Subramanian A, Tamayo P, Mootha VK et al (2005) Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci U S A 102:15545–15550
Newman AM, Liu CL, Green MR et al (2015) Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12:453–457
Hanna SL, Fletcher BD, Parham DM, Bugg MF (1991) Muscle edema in musculoskeletal tumors: MR imaging characteristics and clinical significance. J Magn Reson Imaging 1:441–449
Crombé A, Le Loarer F, Stoeckle E et al (2018) MRI assessment of surrounding tissues in soft-tissue sarcoma during neoadjuvant chemotherapy can help predicting response and prognosis. Eur J Radiol 109:178–187
White LM, Wunder JS, Bell RS et al (2005) Histologic assessment of peritumoral edema in soft tissue sarcoma. Int J Radiat Oncol Biol Phys 61:1439–1445
Fortes-Andrade T, Almeida JS, Sousa LM et al (2021) The role of natural killer cells in soft tissue sarcoma: prospects for immunotherapy. Cancers (Basel) 13:3865
Kelleher FC, Cain JE, Healy JM et al (2012) Prevailing importance of the hedgehog signaling pathway and the potential for treatment advancement in sarcoma. Pharmacol Ther 136:153–168
Ahlawat S, Fritz J, Morris CD, Fayad LM (2019) Magnetic resonance imaging biomarkers in musculoskeletal soft tissue tumors: review of conventional features and focus on nonmorphologic imaging. J Magn Reson Imaging 50:11–27
Funding
The authors state that this work has not received any funding.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Prof. Antoine Italiano.
Conflict of interest
The authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
Three of the authors have significant statistical expertise (C.L. and F.B. are bioinformaticians; A.C. has a PhD in applied mathematics).
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Study subjects or cohorts overlap
Some study subjects or cohorts have been previously reported in the study by Fadli et al [20].
Methodology
• retrospective
• diagnostic or prognostic study/observational/experimental
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Crombé, A., Bertolo, F., Fadli, D. et al. Distinct patterns of the natural evolution of soft tissue sarcomas on pre-treatment MRIs captured with delta-radiomics correlate with gene expression profiles. Eur Radiol 33, 1205–1218 (2023). https://doi.org/10.1007/s00330-022-09104-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-022-09104-8