Abstract
Objectives
To evaluate whether a deep learning (DL) model using both three-dimensional (3D) black-blood (BB) imaging and 3D gradient echo (GRE) imaging may improve the detection and segmentation performance of brain metastases compared to that using only 3D GRE imaging.
Methods
A total of 188 patients with brain metastases (917 lesions) who underwent a brain metastasis MRI protocol including contrast-enhanced 3D BB and 3D GRE were included in the training set. DL models based on 3D U-net were constructed. The models were validated in the test set consisting of 45 patients with brain metastases (203 lesions) and 49 patients without brain metastases.
Results
The combined 3D BB and 3D GRE model yielded better performance than the 3D GRE model (sensitivities of 93.1% vs 76.8%, p < 0.001), and this effect was significantly stronger in subgroups with small metastases (p interaction < 0.001). For metastases < 3 mm, ≥ 3 mm and < 10 mm, and ≥ 10 mm, the sensitivities were 82.4%, 93.2%, and 100%, respectively. The combined 3D BB and 3D GRE model showed a false-positive per case of 0.59 in the test set. The combined 3D BB and 3D GRE model showed a Dice coefficient of 0.822, while 3D GRE model showed a lower Dice coefficient of 0.756.
Conclusions
The combined 3D BB and 3D GRE DL model may improve the detection and segmentation performance of brain metastases, especially in detecting small metastases.
Key Points
• The combined 3D BB and 3D GRE model yielded better performance for the detection of brain metastases than the 3D GRE model (p < 0.001), with sensitivities of 93.1% and 76.8%, respectively.
• The combined 3D BB and 3D GRE model showed a false-positive rate per case of 0.59 in the test set.
• The combined 3D BB and 3D GRE model showed a Dice coefficient of 0.822, while the 3D GRE model showed a lower Dice coefficient of 0.756.
Similar content being viewed by others

Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Brain metastases are the most common type of intracranial tumors in adults; they occur in 20–40% of patients with systemic cancer and are the major cause of morbidity and mortality [1, 2]. For managing patients with brain metastases, early and accurate diagnosis is crucial to determine the treatment strategy and patient prognosis. Surgical resection, stereotactic radiosurgery, and whole brain radiation therapy have been shown to increase survival in eligible patients compared with untreated ones [2,3,4,5]. However, manual detection and segmentation of brain metastases is not only laborious and time-consuming but also erroneous; interobserver variability in target volume delineation has been reported [6], and manual segmentation of all of the lesions may be challenging.
Several previous studies have proposed various deep learning (DL) methods for the detection and/or segmentation of brain metastases. These studies have applied convolutional neural networks (CNNs) [7,8,9,10], fully convolutional networks [11], or single-shot detector models [12]; however, most of the studies showed substantial numbers of false-positive (FP) findings with generally low segmentation performance (Dice coefficients lower than 0.8) [9, 10, 12]. Furthermore, the sensitivity was low in detecting small metastases (smaller than 6–7 mm), with a maximum sensitivity of 70% [9, 12]. Considering the fact that the presence of even small brain metastases may change the whole treatment plan [13], the detection of small brain metastases is crucial in the real-world implementation of DL algorithms.
The low performance and limited clinical feasibility of previous DL approaches may be partially attributed to the fact that most previous studies have focused on DL approaches using contrast-enhanced three-dimensional (3D) gradient echo (GRE) imaging. For the detection of brain metastases, either contrast-enhanced 3D GRE or spin echo (SE) imaging with black-blood (BB) imaging techniques, such as motion-sensitized driven-equilibrium (MSDE) or improved MSDE (iMSDE), is commonly used [14,15,16]. A previous meta-analysis has shown that brain metastases can be more easily detected in contrast-enhanced SE images than GRE images, especially small lesions (< 5 mm) [17]. Moreover, a recent consensus recommendation has stated that 3D SE imaging is preferable to 3D GRE imaging in a brain metastases imaging protocol [18]. Compared to 3D GRE images, the 3D SE with BB imaging technique can suppress blood vessel signals, which enables clearer delineation and better detection of small brain metastases [15]. We hypothesized that addition of 3D BB imaging to 3D GRE imaging in DL models may improve the detection and segmentation performance of brain metastases.
Thus, the purpose of this study was to evaluate whether a DL model using both 3D BB imaging and 3D GRE imaging may improve the detection and segmentation performance of brain metastases compared to that using only 3D GRE imaging.
Materials and methods
Patient population
The institutional review board waived the need for obtaining informed patient consent and ethical approval was obtained for this retrospective study. Between January 2018 and December 2019, 188 consecutive patients who followed our brain metastasis protocol including 3D BB imaging and were diagnosed with a newly developed brain metastasis were included in the training set. For the test set, between January 2020 and May 2020, 45 consecutive patients with a newly developed brain metastasis were included. Additionally, 49 patients without a brain metastasis were included after age and sex matching (detailed information of the exclusion criteria can be found in Supplementary Material S1). Brain metastases were determined based on the review of both initial and follow-up images by two radiologists (4 and 9 years of experience, respectively). In the rare case of ambiguity, a senior radiologist (with 15 years of experience) was consulted for the final decision.
MRI protocol
MRI was performed using various 3.0-T MRI scanners (Achieva/Ingenia/Ingenia CX/Ingenia Elition X, Philips Medical Systems) with an eight-channel sensitivity-encoding head coil. Five minutes after administering a gadolinium-based contrast (0.1 mL/kg gadobutrol; Gadovist, Bayer Schering Pharma), a 3D fast SE sequence with the iMSDE (BB imaging) was performed, followed by 3D GRE imaging (Supplementary Material S2).
Image pre-processing and segmentation
Image resampling to 1-mm isovoxels, low-frequency intensity non-uniformity correction by the N4 bias algorithm and coregistration of 3D GRE images to 3D BB images were performed using Advanced Normalization Tools (ANTs) (version 2.3.4., http://stnava.github.io/ANTs/). Skull stripping was performed using the Multi-cONtrast brain STRipping (MONSTR) [19]. Signal intensity was normalized using the z-score.
A junior radiologist (with 4 years of experience) independently segmented the brain metastases on the 3D BB images while referring to 3D GRE images, while another neuroradiologist (with 9 years of experience) modified the segmented lesions if necessary or confirmed the segmented lesions. In the rare case of ambiguity, a senior radiologist (with 16 years of experience) modified and confirmed the segmented lesions.
DL architecture
To segment the brain metastasis, 3D U-net-based DL models were used. The overall architecture of our DL model is presented in Fig. 1. The input of the model was 3D GRE, 3D BB, or combined 3D GRE and 3D BB images, and the output was the segmentation maps. The size of the inputs and outputs of the model was 184 × 184 × 152 by cropping the background areas of the brain. Three DL models were constructed depending on the input images (combined 3D BB and 3D GRE model, 3D BB model, and 3D GRE model). For a combined 3D BB and 3D GRE model, two 3D images were concatenated along a channel dimension (184 × 184 × 152 × 2). The model was trained with a fivefold cross-validation.
The original U-net consists of (1) an encoder that extracts the features of the input images using several convolutional blocks and max-pooling layers and (2) a decoder (“segmentation decoder”) that reconstructs the desired segmentation maps using the extracted features, convolutional blocks, and up-scale layers [20]. To regularize the encoder, an additional decoder (“reconstruction decoder”) that reconstructed the input images from the extracted features was added at the end of the encoder, inspired by an autoencoder network [21]. The model reconstructed both the segmentation maps and the input images using the shared encoder. The reconstruction decoder was used only during the training phase and not used during the inference phase.
The encoder and decoders had several convolutional blocks, and each convolutional block consisted of a 3D convolutional layer, leaky rectified linear unit (ReLU), and group normalization layer (detailed information of the DL architecture in Supplementary Material S3) [22, 23].
The DL model was implemented using the Python Keras library with a TensorFlow backend [24]. It was trained using the Adam optimizer with β1 = 0.9 and β2 = 0.999 for 500 epochs with a learning rate of 0.0001 [25]. The training took approximately 48 h using an Intel i9-9820X central processing unit (CPU) and an Nvidia TITAN RTX graphics processing unit (GPU) with CUDA version 10.0. In addition, training data augmentation and test-time augmentation were performed during the training and inference phase, respectively (detailed information of data augmentation in Supplementary Material S3).
Statistical analysis
The performances of the three models, including the models trained with both 3D BB and 3D GRE images, and with 3D BB only or 3D GRE only, were assessed by using lesion-based sensitivity, precision, and Dice coefficient (definitions are shown in Supplementary Material S4). The sensitivities were also evaluated with respect to the size of the metastases (< 3 mm, ≥ 3 mm and < 10 mm, and ≥ 10 mm). For patient-by-patient analysis, area under the receiver operating characteristic curve, sensitivity, and specificity were obtained. In addition, the detection performance was also evaluated by calculating the number of FPs per patient. Because all MRI scans from test patients were integrated as one data set, the within-patient correlation for patients with multiple lesions was not considered.
The sensitivities of the combined 3D BB and 3D GRE model and the other two models were compared pairwise using a logistic regression analysis with the generalized estimating equation in a per-lesion analysis [26]. We also analyzed the interaction between the subgroups with different sizes of metastases (< 3 mm, ≥ 3 mm and < 10 mm, and ≥ 10 mm) and DL models in the logistic regression analysis to assess whether the sensitivities differed based on the different subgroups on the DL models. Identical analysis was performed in subgroups with different numbers of metastases (1 to 3, 4 to 10, > 10). The FPs per patient among DL models were compared by using Poisson regression with a generalized estimating equation in a per-patient analysis [27].
Results
Patient characteristics and characteristics of the brain metastases
A total of 282 patients were included in our study (mean age, 61.7 ± 13.1; 129 females and 153 males). The total number of brain metastases was 1120 (917 and 203 in the training and test sets, respectively). The distributions of the number and sizes of the metastases across patients are shown in Fig. 2. Clinical characteristics and characteristics of the brain metastases (mean volume, number, and size) are shown in Table 1.
Detection performance of the DL models
The detection sensitivities and precisions for the DL models in the patients with brain metastases are summarized in Tables 2 and 3. The combined 3D BB and 3D GRE model achieved an overall detection sensitivity of 93.1% and precision of 84.8% for the patients with brain metastases in the test set. Specifically, for metastases < 3 mm, ≥ 3 mm and < 10 mm, and ≥ 10 mm, the sensitivities were 82.4%, 93.2%, and 100%, respectively. Examples of true-positive inferences in various sizes that were detected on all three models are shown in Supplementary Figure 1.
The overall sensitivity of the combined 3D GRE and 3D BB model was significantly higher than that of the 3D GRE model (93.1% vs 76.8%, p < 0.001) (Fig. 3), and this effect was significantly stronger in subgroups with small metastases, but not in subgroups according to number of metastases (p interaction < 0.001 and p interaction = 0.686, respectively). Specifically, the sensitivities of the combined 3D GRE and 3D BB model were significantly higher than those for the 3D GRE model in the subgroup with metastases < 3 mm (p < 0.001) as well as the subgroup with metastases ≥ 3 mm and < 10 mm (p = 0.014). The sensitivities of the combined 3D GRE and 3D BB model were significantly higher than those for the 3D GRE model in the subgroup with 4 to 10 metastases (p = 0.014) as well as the subgroup with > 10 metastases (p < 0.001). On the other hand, the overall sensitivity of the combined 3D GRE and 3D BB model was similar to that of the 3D BB model (93.1% vs 92.6%, p = 0.847).
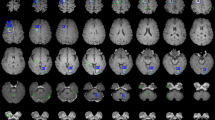
Examples of true-positive inferences that were detected on the combined three-dimensional (3D) black-blood (BB) and 3D gradient echo (GRE) model on representative test patients. a A 4.4-mm metastasis and b a 1.7-mm metastasis that were missed on the 3D GRE model but detected on the 3D BB model as well as the combined 3D BB and 3D GRE model
The overall area under the curve, sensitivities, and specificities for the DL models in patient-by-patient analysis are summarized in Supplementary Table 1. The overall sensitivity and specificity for the combined 3D BB and 3D GRE model in detecting brain metastases in the patient-by-patient analysis were 100% and 69.4%, respectively, in the entire test set (including both patients with and without brain metastases).
The number of FP lesions for the DL models is shown in Table 4. The FP per patient was 0.59 in the test set. The majority of FP lesions were insufficiently suppressed blood signals from peripheral vessels (48 of 55 [87.3%] FP lesions) as shown in Fig. 4 (detailed information of FP lesions in Supplementary Material S5). The number of FP lesions per patient was significantly lower in the combined 3D BB and 3D GRE model compared with that in the 3D BB model (p = 0.004). However, the number of FP lesions per patient was significantly higher in the combined 3D BB and 3D GRE model compared with that in the 3D GRE model (p < 0.001).
Examples of false-positive (FP) inferences. a An FP inference of an insufficiently suppressed blood signal from the peripheral vessel that was misclassified as a brain metastasis on the three-dimensional (3D) black-blood (BB) model but not on the combined 3D BB and 3D gradient echo (GRE) model or the 3D GRE model. b An FP inference of an insufficiently suppressed blood signal from the peripheral vessel that was misclassified as a brain metastasis on the 3D BB model as well as the combined 3D BB and 3D GRE model but not on the 3D GRE model
Segmentation performance of the DL models
The Dice coefficients for the DL models for the patients with brain metastases are summarized in Table 5. Compared to the combined 3D BB and 3D GRE model with a Dice coefficient of 0.822, the 3D BB and 3D GRE models showed similar or lower Dice coefficients of 0.827 and 0.756, respectively.
Discussion
In our study, the combined 3D BB and 3D GRE model showed promising results for detecting and segmenting brain metastases. The DL model was able to detect 189 of 203 (93.1%) brain metastases in the test set, with a Dice coefficient of 0.822. The Response Assessment in Neuro-Oncology Brain Metastases (RANO BM) suggests a target lesion size criterion of 10 mm [28]. Therefore, our model showing a sensitivity of 100% and Dice coefficient of 0.822 in detecting and segmenting brain metastases ≥ 10 mm may be suitable for response assessment of brain metastases. Furthermore, compared with previously reported DL studies for detecting and segmenting brain metastases, there were three major findings in the current study. First, we combined both 3D BB and 3D GRE imaging in our DL algorithm, resulting in a high sensitivity of 82.4% for detecting small metastases (smaller than 3 mm), which is the highest reported sensitivity, to the best of our knowledge. Second, we demonstrated that the combined 3D BB and 3D GRE model shows superior performance over the 3D GRE models. Third, the Dice coefficient of 0.822 shows the highest performance reported in segmenting brain metastases [8, 9, 11].
Several studies have attempted to identify methods and algorithms for the detection and/or segmentation of brain metastases [8,9,10,11,12, 29], but a majority of the previous DL models on the detection and segmentation of brain tumors were focused on gliomas [30,31,32,33]. Compared with gliomas [34], brain metastases present several unique challenges. Brain metastases may be smaller than gliomas; however, it is important to note that the detection of even small lesions is essential for treatment planning [35]. Moreover, brain metastases near the peripheral vessels may be missed or, conversely, peripheral vessels may be mislabeled as metastases [14, 36, 37]. Our combined 3D BB and 3D GRE model showed a sensitivity of 93.1% and precision of 84.8% in detecting brain metastases, indicating that this model is one of the best performing models among previously reported ones [8,9,10,11,12, 29]. Although a previous DL model showed a slightly higher sensitivity of 96% [10] in detecting brain metastases, it also showed a higher number of FP lesions per patient of 19.9 [10]; however, we showed a substantially lower number of FP lesions (0.59) per patient in the test set. Considering the trade-off between sensitivity and precision, our model may be more optimal for clinical application. Compared to previous studies that used 2D- or 2.5D-based detection models [9, 10, 12], our model was based on 3D U-net, which utilizes 3D information for detecting and segmenting brain metastases. In our model, deep supervision loss, which was computed at the low-resolution feature maps, and an additional reconstruction decoder could guide the final segmentation of metastases. In addition, by using test-time augmentation as well as data augmentation, it was possible to increase the sensitivity in detecting metastases and reduce the number of FP lesions.
In our study, a substantial proportion of brain metastases were small in size (< 3 mm). Specifically, the overall proportion of small metastases (< 3 mm) in our dataset was 27.4% (307 of 1122 lesions), whereas previous studies have either reported a lower proportion of small metastases (< 3 mm) (11.1%, 127 of 1147 lesions) [12] or did not report on the proportion of small metastases [8,9,10,11]. A previous meta-analysis has shown that BB images show superior performance for detecting small lesions (< 5 mm) [17]. In accordance with this, the 3D GRE model showed a substantially lower performance of 23.5% in detecting metastases < 3 mm than the 3D BB model or the combined 3D BB and 3D GRE model. Because 3D GRE imaging does not suppress blood signals and has a lower contrast-to-noise ratio compared to 3D BB imaging, brain metastases that are small in size may be easily missed in the DL model [17]. When 3D BB was added into the model, the sensitivity in detecting metastases < 3 mm increased to 82.4%, highlighting the importance of 3D BB imaging in detecting small metastases. Our reported sensitivity of detecting small metastases is by far the highest among previous studies, some of which reported sensitivities of 15% for identifying brain metastases < 3 mm [12] and 50% for identifying lesions smaller than < 7 mm [9]. On the other hand, the sensitivities of the combined 3D BB and 3D GRE model as well as the 3D BB model for small metastases (< 3 mm) were similar, but the number of FPs was significantly lower when the former model was used. This may be due to the fact that, when only 3D BB imaging is used, some blood vessels that are incompletely suppressed may mimic brain metastases [36], leading to an increased number of FPs. A previous study implemented multiparametric MRI, which included 3D BB, 3D GRE, and 3D fluid-attenuation inversion recovery imaging, for the detection and segmentation of brain metastases [9]; however, the overall performance was lower, with a sensitivity of 83% and FP lesions per patient of 8.3; furthermore, the model’s performance with and without 3D BB was not compared.
Our study showed a Dice coefficient of 0.822 in segmenting brain metastases, which suggests that our model performed well. Previous studies have reported Dice coefficients of 0.77 [8], 0.79 [9], and 0.85 [11] in segmenting brain metastases, respectively. However, the previous study that had reported a Dice coefficient of 0.85 had not included brain metastases smaller than 5 mm for this calculation [11]. Thus, we speculate that the Dice coefficient in their study might have been lowered if small metastases had been included, because smaller lesions have shown to be associated with lower Dice coefficients [38, 39]
In the test set, we included not only patients with brain metastases but also those without brain metastases to evaluate specificity. Previous studies on DL models of brain metastases did not include patients without brain metastases, which may limit the evaluation of true performance of the DL model. The reported incidence of brain metastases in patients during the diagnosis of primary cancer is low [40], and many patients undergoing an MRI for brain metastases may not have brain metastases. Our results show that there are FP lesions in patients without brain metastases; most of these are insufficiently suppressed blood signals. Thus, the DL results should be reviewed by the radiologist with caution.
Our study has several limitations. First, this was a single center, retrospective study with a relatively small data size. Evaluations of patients at external locations with different hardware implementations are necessary to validate the model’s robustness. Second, our approach implemented both 3D BB and 3D GRE imaging, which may have limited availability in certain MRI scanners. Although 3D BB imaging is ideally recommended in MRI protocols for brain metastases [18], it is not technically available in all MRI instruments sold by all vendors; furthermore, it may require additional scan time. Future DL models implementing synthetic 3D BB imaging without additional scan time may be an alternative approach [41]. Third, because our model was mostly trained and tested on patients with brain metastases, our results may be unlikely to represent true systematic model inaccuracies. Nonetheless, we believe that our results could establish a foundation for future prospective research for the verification of our DL model in actual clinical practice. Fourth, segmentation of the lesions was performed in consensus rather than by multi-readers, which may limit the robustness and reliability of deep features.
In conclusion, the combined 3D BB and 3D GRE DL model may improve the detection and segmentation performance of brain metastases, especially for the detection of small metastases.
Abbreviations
- 3D:
-
Three-dimensional
- BB:
-
Black blood
- CNN:
-
Convolutional neural network
- DL:
-
Deep learning
- FP:
-
False positive
- GRE:
-
Gradient echo
- iMSDE:
-
Improved motion-sensitized driven-equilibrium
- SE:
-
Spin echo
- TE:
-
Echo time
- TR:
-
Repetition time
References
Bradley KA, Mehta MP (2004) Management of brain metastases. Semin Oncol 31(5). WB Saunders, 2004
Loeffler J, Patchell R, Sawaya R (1997) Metastatic brain cancer. Cancer 2523
Kondziolka D, Patel A, Lunsford LD, Kassam A, Flickinger JC (1999) Stereotactic radiosurgery plus whole brain radiotherapy versus radiotherapy alone for patients with multiple brain metastases. Int J Radiat Oncol Biol Phys 45:427–434
Patchell RA, Tibbs PA, Walsh JW et al (1990) A randomized trial of surgery in the treatment of single metastases to the brain. N Engl J Med 322:494–500
Mehta MP, Rodrigus P, Terhaard C et al (2003) Survival and neurologic outcomes in a randomized trial of motexafin gadolinium and whole-brain radiation therapy in brain metastases. J Clin Oncol 21:2529–2536
Growcott S, Dembrey T, Patel R, Eaton D, Cameron A (2020) Inter-observer variability in target volume delineations of benign and metastatic brain tumours for stereotactic radiosurgery: results of a national quality assurance programme. Clin Oncol (R Coll Radiol) 32:13–25
Kamnitsas K, Ledig C, Newcombe VFJ et al (2017) Efficient multi-scale 3D CNN with fully connected CRF for accurate brain lesion segmentation. Med Image Anal 36:61–78
Charron O, Lallement A, Jarnet D, Noblet V, Clavier JB, Meyer P (2018) Automatic detection and segmentation of brain metastases on multimodal MR images with a deep convolutional neural network. Comput Biol Med 95:43–54
Grøvik E, Yi D, Iv M, Tong E, Rubin D, Zaharchuk G (2020) Deep learning enables automatic detection and segmentation of brain metastases on multisequence MRI. J Magn Reson Imaging 51:175–182
Zhang M, Young GS, Chen H et al (2020) Deep-learning detection of cancer metastases to the brain on MRI. J Magn Reson Imaging. https://doi.org/10.1002/jmri.27129
Xue J, Wang B, Ming Y et al (2020) Deep learning-based detection and segmentation-assisted management of brain metastases. Neuro Oncol 22:505–514
Zhou Z, Sanders JW, Johnson JM et al (2020) Computer-aided detection of brain metastases in T1-weighted MRI for stereotactic radiosurgery using deep learning single-shot detectors. Radiology 295:407–415
Lin X, DeAngelis LM (2015) Treatment of brain metastases. J Clin Oncol 33:3475–3484
Park J, Kim J, Yoo E, Lee H, Chang J-H, Kim EY (2012) Detection of small metastatic brain tumors: comparison of 3D contrast-enhanced whole-brain black-blood imaging and MP-RAGE imaging. Invest Radiol 47:136–141
Park J, Kim EY (2010) Contrast-enhanced, three-dimensional, whole-brain, black-blood imaging: application to small brain metastases. Magn Reson Med 63:553–561
Park YW, Ahn SJ (2018) Comparison of contrast-enhanced T2 FLAIR and 3D T1 black-blood fast spin-echo for detection of leptomeningeal metastases. Investig Magn Reson Imaging 22:86–93
Suh CH, Jung SC, Kim KW, Pyo J (2016) The detectability of brain metastases using contrast-enhanced spin-echo or gradient-echo images: a systematic review and meta-analysis. J Neurooncol 129:363–371
Kaufmann TJ, Smits M, Boxerman J et al (2020) Consensus recommendations for a standardized brain tumor imaging protocol for clinical trials in brain metastases. Neuro Oncol 22:757–772
Roy S, Butman JA, Pham DL (2017) Robust skull stripping using multiple MR image contrasts insensitive to pathology. Neuroimage 146:132–147
Ronneberger O, Fischer P, Brox T (2015) U-net: Convolutional networks for biomedical image segmentation. International conference on medical image computing and computer-assisted intervention. Springer, Cham
Myronenko A (2018) 3D MRI brain tumor segmentation using autoencoder regularization. International MICCAI Brain Lesion Workshop. Springer, Cham
Maas AL, Hannun AY, Ng AY (2013) Rectifier nonlinearities improve neural network acoustic models. Proc icml 3(1)
Wu Y, He K (2018) Group normalization. Proceedings of the European conference on computer vision (ECCV)
Abadi M, Barham P, Chen J et al (2016) TensorFlow: a system for large-scale machine learning. 12th {USENIX} symposium on operating systems design and implementation ({OSDI} 16)
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. arXiv preprint arXiv:14126980 22
Zeger SL, Liang KY, Albert PS (1988) Models for longitudinal data: a generalized estimating equation approach. Biometrics 44:1049–1060
Consul P, Famoye F (1992) Generalized Poisson regression model. Commun Stat Theory Methods 21:89–109
Lin NU, Lee EQ, Aoyama H et al (2015) Response assessment criteria for brain metastases: proposal from the RANO group. Lancet Oncol 16:e270–e278
Sunwoo L, Kim YJ, Choi SH et al (2017) Computer-aided detection of brain metastasis on 3D MR imaging: observer performance study. PLoS One 12:e0178265
Chang K, Beers AL, Bai HX et al (2019) Automatic assessment of glioma burden: a deep learning algorithm for fully automated volumetric and bidimensional measurement. Neuro Oncol 21:1412–1422
Kickingereder P, Isensee F, Tursunova I et al (2019) Automated quantitative tumour response assessment of MRI in neuro-oncology with artificial neural networks: a multicentre, retrospective study. Lancet Oncol 20:728–740
Bakas S, Reyes M, Jakab A et al (2018) Identifying the best machine learning algorithms for brain tumor segmentation, progression assessment, and overall survival prediction in the BRATS challenge. arXiv preprint arXiv:181102629
Weninger L, Rippel O, Koppers S, Merhof D (2018) Segmentation of brain tumors and patient survival prediction: methods for the BraTS 2018 challenge. International MICCAI Brain Lesion Workshop. Springer, Cham
Park YW, Han K, Ahn SS et al (2018) Prediction of IDH1-mutation and 1p/19q-codeletion status using preoperative MR imaging phenotypes in lower grade gliomas. AJNR Am J Neuroradiol 39:37–42
Anzalone N, Essig M, Lee SK et al (2013) Optimizing contrast-enhanced magnetic resonance imaging characterization of brain metastases: relevance to stereotactic radiosurgery. Neurosurgery 72:691–701
Nagao E, Yoshiura T, Hiwatashi A et al (2011) 3D turbo spin-echo sequence with motion-sensitized driven-equilibrium preparation for detection of brain metastases on 3T MR imaging. AJNR Am J Neuroradiol 32:664–670
Kato Y, Higano S, Tamura H et al (2009) Usefulness of contrast-enhanced T1-weighted sampling perfection with application-optimized contrasts by using different flip angle evolutions in detection of small brain metastasis at 3T MR imaging: comparison with magnetization-prepared rapid acquisition of gradient echo imaging. AJNR Am J Neuroradiol 30:923–929
Woo I, Lee A, Jung SC et al (2019) Fully automatic segmentation of acute ischemic lesions on diffusion-weighted imaging using convolutional neural networks: comparison with conventional algorithms. Korean J Radiol 20:1275–1284
Xue Y, Farhat FG, Boukrina O et al (2020) A multi-path 2.5 dimensional convolutional neural network system for segmenting stroke lesions in brain MRI images. Neuroimage Clin 25:102118
Cagney DN, Martin AM, Catalano PJ et al (2017) Incidence and prognosis of patients with brain metastases at diagnosis of systemic malignancy: a population-based study. Neuro Oncol 19:1511–1521
Jun Y, Eo T, Kim T et al (2018) Deep-learned 3D black-blood imaging using automatic labelling technique and 3D convolutional neural networks for detecting metastatic brain tumors. Sci Rep 8:1–11
Funding
This research received funding from the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, Information and Communication Technologies & Future Planning (2020R1A2C1003886). This research was also supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2020R1I1A1A01071648). This study was financially supported by the Faculty Research Grant of Yonsei University College of Medicine (6-2020-0149). This research was supported by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science and ICT (2019R1A2B5B01070488), Brain Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (2018M3C7A1024734), and Y-BASE R&E Institute a Brain Korea 21, Yonsei University.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Professor Seung-Koo Lee, MD, PhD, from Yonsei University College of Medicine (slee@yuhs.ac).
Conflict of interest
The authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
One of the authors has significant statistical expertise (K.H, a biostatistician with 10 years of experience in biostatistics).
Informed consent
The institutional review board waived the requirement to obtain informed patient consent for this retrospective study.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 335 kb)
Rights and permissions
About this article
Cite this article
Park, Y.W., Jun, Y., Lee, Y. et al. Robust performance of deep learning for automatic detection and segmentation of brain metastases using three-dimensional black-blood and three-dimensional gradient echo imaging. Eur Radiol 31, 6686–6695 (2021). https://doi.org/10.1007/s00330-021-07783-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-021-07783-3