Abstract
Objectives
To establish and validate a radiomics nomogram for prediction of induction chemotherapy (IC) response and survival in nasopharyngeal carcinoma (NPC) patients.
Methods
One hundred twenty-three NPC patients (100 in training and 23 in validation cohort) with multi-MR images were enrolled. A radiomics nomogram was established by integrating the clinical data and radiomics signature generated by support vector machine.
Results
The radiomics signature consisting of 19 selected features from the joint T1-weighted (T1-WI), T2-weighted (T2-WI), and contrast-enhanced T1-weighted MRI images (T1-C) showed good prognostic performance in terms of evaluating IC response in two cohorts. The radiomics nomogram established by integrating the radiomics signature with clinical data outperformed clinical nomogram alone (C-index in validation cohort, 0.863 vs 0.549; p < 0.01). Decision curve analysis demonstrated the clinical utility of the radiomics nomogram. Survival analysis showed that IC responders had significant better PFS (progression-free survival) than non-responders (3-year PFS 84.81% vs 39.75%, p < 0.001). Low-risk groups defined by radiomics signature had significant better PFS than high-risk groups (3-year PFS 76.24% vs 48.04%, p < 0.05).
Conclusions
Multiparametric MRI-based radiomics could be helpful for personalized risk stratification and treatment in NPC patients receiving IC.
Key Points
• MRI Radiomics can predict IC response and survival in non-endemic NPC.
• Radiomics signature in combination with clinical data showed excellent predictive performance.
• Radiomics signature could separate patients into two groups with different prognosis.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Nasopharyngeal carcinoma (NPC) is the most common head and neck cancer in Southeast Asia. Previous studies indicated that NPC had different characteristics in endemic vs non-endemic areas [1, 2]. Concurrent chemoradiation (CCRT) is a standard of care for locally advanced NPC [3]. With the development of intensity-modulated radiotherapy (IMRT), despite a favorable 5-year local rate (> 90%), distant metastases are the predominant risk [2, 4, 5], suggesting a need for more systemic therapies. Induction chemotherapy (IC) is better tolerated than adjuvant chemotherapy, and the achieved dose intensity is greater. A meta-analysis and one phase III trial both indicated that IC could improve survival compared with CCRT alone in locoregionally advanced NPC [6, 7]. We suggested that IC-CCRT was safe and effective in the treatment of NPC from non-endemic regions [8].
Tumor response to IC is an independent prognostic factor for survival after IMRT in NPC [9]. However, not all patients respond well to IC. Pretreatment identification of non-responders would allow more personalized treatment selection, avoiding toxicity and unnecessary costs for non-responders. However, no ideal clinical features or biological biomarker is available today.
Modalities like 18F-fluorodeoxyglucose positron emission tomography and MRI-DWI have been proposed as potential imaging biomarkers for the prediction of response to IC in NPC [10, 11]. Radiomic analysis is promising in cancer diagnosis, prognosis, and prediction of response to treatment [12, 13]. Recent studies showed that texture parameter of pretreatment MRI or MRI-based radiomics nomogram could predict PFS in endemic NPC [14, 15]. Pretreatment MRI signature could also predict early IC response in endemic NPC patients, although there was no independent validation [16]. However, no study has reported the role of MRI-based radiomics for the prediction of IC response in non-endemic region. Its role in predicting both IC response and survival is still unclear. The current study aims to investigate the role of a MRI-based radiomics nomogram in predicting the response to IC in locally advanced NPC patients of Northwest China. Also, we sought to correlate it with clinical parameters and survival.
Materials and methods
Patients
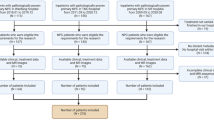
A total of 200 patients with histologically confirmed NPC treated with IC and CCRT between January 2012 and December 2016 in Xijing Hospital were enrolled in the study. T1-WI, T2-WI, and T1-C (T1 contrast) images were collected before treatment (3-T GE or Siemens MRI scanners). All patients were staged according to the 7th Edition of American Joint Committee on Cancer (AJCC) TNM classification. A total of 123 patients were enrolled and were allocated to the training and validation cohorts randomly. One hundred and 23 patients were allocated to the training cohort (28 responders vs 72 non-responders) and validation cohort (6 responders vs 17 non-responders), respectively. The details of chemotherapy are shown in supplemental materials.
Endpoints and follow-up
The clinical treatment response after IC was defined as stable disease (SD), progressive disease (PD), partial response (PR), or complete response (CR) based on the Response Evaluation Criteria in Solid Tumors 1.1 (RECIST) criteria [17]. Patients were categorized into IC responders (CR/PR) and non-responders (SD/PD). All patients were followed up weekly during radiotherapy treatment and every 2–3 months during the first 2 years, then every 3–4 months during the year after radiotherapy treatment 3–5, and annually thereafter.
Tumor segmentation, radiomics feature extraction, and development of multiparametric MRI-based radiomics model
The details of tumor segmentation (Fig. 1a) are shown in supplemental methods. Imaging features calculated for each patient from a unimodal MR image using MATLAB procedure algorithm (Fig. 1b), which was described in the supplemental material. Features from T1-C, T1-WI, and T2-WI or the combination were used for subsequent analysis, respectively. For the feature selection step, a filter method based on the selection frequency was employed to obtain a ranking of features with the most discriminative power. The frequency of each feature selected by the two-step feature selection methods across 100 training partitions of the leave-one-out cross-validation (LOOCV) was calculated to assess the variable importance (> 50% selection probability across partitions). The two-step feature selection method consisted the two-sample t test between responders and non-responder in the training cohort and the logistic regression model with LASSO (the least absolute shrinkage and selection operator) (Fig. 1c) [18,19,20]. A tuning parameter λ which controls the strength of regularization was chosen using tenfold cross-validation repeated 1000 times via minimum criteria. All important features were ranked by the selection frequency.
A general schema for predictive nomogram establishment. a Three-dimensional manual segmentation. b Four types of features were finally calculated for each patient from single segmentation. c The two-sample t test and the logistic regression model with LASSO based on LOOCV were used to select useful features. d Radiomics signature was established with support vector machine. e Multivariable backward stepwise logistic regression model was used to develop a radiomics nomogram
The ranked 29 features (> 50% selection probability) were used to generate different subsets and train the SVM (support vector machine) model. For example, the first subset consisted of the first two features, the second one consisted of the first three features, and so on. Twenty-eight feature subsets were generated in total. The support vector machine (SVM) with linear kernel was developed to evaluate the role of each feature subset in predicting the response to IC (Fig. 1d). Tenfold cross-validation was applied to select the regularization parameter C from 0.01 to 1 with a step size 0.01 based on AUC maximum criteria. In this study, fivefold cross-validation was used to estimate group classification and prediction accuracies and repeated 1000 times in the training cohort. Furthermore, the feature subset with the best performance was used to train and validate the final SVM model in the training and validation cohort respectively. The optimal cutoff value of the radscore were based on the score test from the Cox regression model and calculated from the “cutp” function of the “survMisc” R package [21]. Random forest (RF) was used as a second machine learning algorithm to validate the previous results (supplemental methods).
Accuracy (ACC), positive predictive value (PPV), negative predictive value (NPV), and the area under the receiver operating characteristic curve (AUC) are used for evaluating the model performance. ROC curves were drawn to display and compare the performance of different models [22]. Identification of the role of radiomics signature in the prediction of survival is shown in supplemental materials.
Development of the clinical and radiomics nomogram
Multivariable logistic regression model–based nomograms are shown in Fig. 1e. The clinical parameters used in the clinical model include the following: age; gender; T, N, and clinical stage; EB-DNA; histology; IC cycles and regimens; the diameter and volume of lymph nodes and nasopharyngeal before IC; and the level of leukocytes, platelets, lymphocytes, and albumin before IC. Radscore from the joint multimodal images was enrolled in the establishment of radiomics nomogram. The likelihood ratio test with Akaike’s information criterion was applied as the stopping rule for backward stepwise logistic regression analysis. Harrell’s C-index, AUC, ACC, PPV, and NPV were used to quantify the discriminative performance. Decision curve analysis was conducted to estimate the clinical usefulness of the models. The details of statistical analysis are described in the supplemental material.
Results
Clinical characteristics of the patients
The characteristics of all NPC patients in both the training and validation cohorts are shown in Table 1 and Supplemental Table 1. There was no significant difference between the two cohorts in terms of IC response rate (28% and 26.1% in the training and validation cohorts, respectively, p = 0.853). Also, no significant difference was observed between IC responders and non-responders, except that there were more females and more patients with ≥ 5000 copies/ml EBV DNA level in the responders group of the training cohort.
Establishment of multiparametric MRI-based radiomics signature
A total of 4503 imaging features were finally calculated for each patient from the extracted tumor region of a unimodal MR image. Twenty-nine features (> 50% selection probability) selected from the combination of the three modalities were ranked by the selection frequency and the top 19 features consisting of 5 from enhanced T1-C, 4 from T1-WI, and 10 from T2-WI images showed the best performance in training the models (Table 2, Fig. 2). The classification accuracy and the AUC value using the combination of three modalities by repeated fivefold cross-validation were 88.58% and 0.8866, respectively (Supplemental Table 3). Multi-modality images showed better prediction performance than unimodal MR image alone (Supplemental Table 3) and RF model generated similar results to SVM model (Supplemental Table 7). The distributions of radiomics signatures for each patient in both the training and validation cohorts are shown in Fig. 3 a and b. There was a significant difference in radiomics scores between responders and non-responders in both the training cohort (p < 0.001) and the validation cohort (p = 0.002; Table 1). The AUC, ACC, PPV, and NPV of the SVM model in the training cohort were 0.9494, 95.00%, 96.00%, and 94.67%, respectively, and in the validation cohort were 0.8725, 86.96%, 71.43%, and 93.75%, respectively (Table 3). ROC curve is shown in Fig. 3c.
One thousand times of fivefold cross-validation were used to examine different subsets of features and evaluate the SVM models. Features of the feature subset with the highest AUC value were the most important and the feature subset was used to train and validate the final SVM model in the training and validation cohort respectively
Establishment of clinical and radiomics nomogram
Sex and the level of leukocytes before treatment were identified as independent predictors with significance for IC response (p < 0.05). The clinical nomogram based on these two factors yielded a C-index of 0.708 (95% CI, 0.588–0.827) in the training cohort and 0.549 (95% CI, 0.197–0.705) in the validation cohort. The AUC, ACC, PPV, and NPV of clinical nomogram in the training cohort were 0.7076, 76%, 62.50%, and 78.57%, respectively, and in the validation cohort were 0.5490, 65.22%, 0%, and 71.43%, respectively (Table 3). ROC curve is shown in Fig. 4a. We next developed a radiomics nomogram that integrated the radiomics signature (p < 0.0001) from the joint T1-C, T1-WI, and T2-WI with all the independent clinical factors, which provided a higher C-index of 0.952 (95% CI, 0.895–1) in the training cohort and 0.863 (95%CI: 0.706–1) in the validation cohort (Fig. 4c). The AUC, ACC, PPV, and NPV of the radiomics nomogram in the training cohort were 0.9524, 95.00%, 92.59%, and 95.89%, respectively, and in the validation cohort were 0.8627, 82.61%, 100%, and 80.95%, respectively (Table 3). The ROC curves are shown in Fig. 4b. The calibration curves of the radiomics nomogram showed good agreement between the observed actual probabilities of the response and nomogram predicted probabilities in the training and validation cohorts shown in Fig. 5 a and b, respectively. The decision curve analysis for the radiomics nomogram is presented in Fig. 5c, showing that the radiomics nomogram provided better performance.
The radiomics nomogram and the ROC curves in both the training and validation cohorts. a ROC curves for the clinical nomogram in the training and validation cohorts. b ROC curves for the radiomics nomogram in the training and validation cohorts. c Radiomics nomogram integrating the radiomics signature from the joint T1-C, T1-W, and T2-W with all the independent clinical factors. d Representative T2-WI image of NPC patient as IC responder before and after IC. e Representative T2-WI image of NPC patient as IC non-responder before and after IC
The calibration and decision curves. a Calibration curves for the radiomics nomogram in the training cohort. The dashed line indicated ideal reference line where predicted probabilities would match the observed proportions. The solid line represented the performance of the radiomics nomogram. b Calibration curves for the radiomics nomogram in the validation cohort. c Decision curve analysis for the radiomics nomogram. The x-axis represented the probability of the response to IC ranging from 0 to 100%. The y-axis measured the net benefit. The black line represented the scheme in which all patients were assumed to be responders. The gray line represented the assumption that all patients were non-responders
The role of radiomics signature in the prediction of survival
Survival analysis showed that responders to IC had significantly better PFS than non-responders (3-year PFS 84.81% vs 39.75%, p < 0.001; Fig. 6a). Log-rank test was used to select the radscore with significant significance and radscore value (− 0.317) was used to divide patients into high- and low-risk groups for different PFS. Low-risk groups with higher radiomics signature had significantly better PFS than higher-risk groups (3-year PFS 76.24% vs 48.04%, p < 0.05; Fig. 6b).
Discussion
In this study, we established a multiparametric MRI-based radiomics nomogram to predict IC response before treatment in locally advanced non-endemic NPC. The radiomics signatures from combination of T1-WI, T2-WI, and enhanced T1-WI images demonstrated excellent prediction value. The nomogram established by addition of MRI radiomics to clinical parameters outperformed that based on clinical data alone. Also, the radiomics signature could separate patients into high- and low-risk groups based on significantly different PFS rate.
Anatomical MRI has been widely used to determine the treatment response in NPC [23]. Currently, there are no ideal clinical features or biomarkers to predict IC response in NPC. One study found NPC patients with low ADCs before treatment responded better to IC [10]. In a retrospective study consisting of 120 endemic NPC patients, pretreatment MRI radiomics signature could predict early IC response [16]. However, there was no validation by an independent cohort. One study showed that MRI-based texture analysis had the potential to predict chemoradiotherapy response with a high accuracy, but this was in a relatively small number of NPC patients [24]. In another study, T2-weighted and contrast-enhanced T1-weighted MRI radiomics predicted progression of endemic NPC in both the training and validation groups [25].
However, several key points need to be considered in MRI radiomics prediction model establishment. Firstly, the majority of MRI radiomics studies were performed in endemic NPC patients and the predictive model cannot be directly applied to non-endemic NPC, given that characteristics differentiate endemic and non-endemic tumors. Our previous studies indicated that WHO type II histology represented a higher proportion of cases diagnosed in northwest China versus endemic regions (> 25% vs < 10%) and WHO type II histology was associated with poor outcomes. Furthermore, less than 15% patients had detectable EBV DNA, while over 90% patients in endemic regions have detectable EBV DNA [1, 2]. Here, we established and validated a radiomics nomogram for non-endemic NPC patients and more studies need to be done to study the difference of radiomics between endemic and non-endemic NPC.
Secondly, the role of clinical factors or the combination of clinical factors and radiomics needs to be clarified. One recent study extracted 970 radiomics features from T2-WI and contrast-enhanced T1-WI MRI [15]. The radiomics nomogram was established by combining radiomics signature with TNM and showed a significant improvement in PFS prediction of NPC when compared with TNM alone. Our study developed a radiomics nomogram that integrated radiomics signature from the joint T1-C, T1-WI, and T2-WI with all the independent clinical factors. This provided a higher C-index in both the training and validation cohorts, indicating that radiomics was more accurate than the clinical nomogram alone in the prediction of IC response. Accordingly, radiomics features seems to be more sensitively and closely correlated with the clinical endpoints we defined (clinical treatment response after IC was the endpoint in this study) when compared with diagnostic radiologic, or clinical information.
Lastly, the role of multiple versus single MRI sequences needs to be studied. Here, we used three MRI sequences to extract image radiomics features, and 4503 imaging features were calculated for each patient from single modality. Of these, 19 features consisting of 5 from enhanced T1-C, 4 from T1-WI, 10 from T2-WI images were input to the SVM model. Our results showed that features extracted from several sequences performed better than when acquired from a single sequence, as shown by both fivefold cross-validation in the training cohort and independent validation in the validation cohort. In order to overcome the limitation of SVM, we also used the random forest (RF) as a second machine learning algorithm and the results were similar. Image information from different sequences could be a more meaningful representation of the radiomics diversity within the tumor.
Given the capability to predict IC response in NPC, we wondered whether this model could also predict survival, which is a very important clinical endpoint. Some reports have showed that tumor response to IC was an independent prognostic factor for survival after radiation therapy in NPC [9]. In this study, we also found that IC responders had significantly better PFS than those non-responders (3-year PFS 84.81% vs 39.75%, p < 0.001). Also, we divided patients into high-risk and low-risk groups by radiomics signature. The low-risk group had a significantly better PFS. These results were in accordance with a previous study showing a significant improvement in PFS prediction by combination of radiomics and TNM than by TNM alone [15].
It has long been demonstrated that radiomics features were associated with tumor biology. Adding radiomics features to molecular pathology is an important extension of radiomics [13]. Panth KM et al found radiomics was able to identify early effects of changed gene expression after radiation treatment in tumors with similar volumes [26]. “Radio-genomics” combining radiomics with genomics data is another emerging prognostic tool [13]. One study indicated that neuroimaging and DNA microarray analysis could create a multidimensional map of gene expression patterns in glioblastoma multiforme [27]. Another study indicated that measures of spatial diversity from MRI images were associated with both tumor subtype status and survival status [28]. In NLCLC, a study established a radiogenomics map and found a metagene from EGF pathway was significantly correlated with ground-glass opacity and irregular nodules or nodules with poorly defined margins [29]. NRG Oncology HN001 trial (NCT02135042) is ongoing with the aim to identify NPC patients who would more likely benefit from additional systemic chemotherapy based on EBV DNA levels. Therefore, radiogenomics in prediction of NPC treatment response and prognosis is an attractive future direction.
Because of its retrospective nature, our study has several limitations. Firstly, due to the critical inclusion criteria of patients included, we had a relatively small number of patients that may affect the final conclusion of the study. Secondly, the single institutional nature of the study may also limit the applicability of our findings for patients from other geographical regions and institutions. Lastly, as pathologic examination of the treatment response was not possible in NPC patients who are mainly treated by radiotherapy, IC response evaluation only based on anatomical MRI imaging might be less accurate. Furthermore, other MRI technique, especially DCE-MRI-based radiomics, has been used in the prediction of treatment response for breast cancer and its role in NPC needs to be further explored [30].
In conclusion, the current study established a multiparametric MRI-based radiomics nomogram to predict the IC response before treatment in locally advanced NPC in non-endemic areas. The results might be more easily generalizable in other countries, due to the non-endemic nature of this cohort. MRI-based radiomics could be especially helpful for personalized risk stratification and treatment in NPC patients receiving IC.
Abbreviations
- CCRT:
-
Concurrent chemoradiation
- IC:
-
Induction chemotherapy
- IMRT:
-
Intensity-modulated radiotherapy
- LASSO:
-
Least absolute shrinkage and selection operator
- NPC:
-
Nasopharyngeal carcinoma
- PFS:
-
Progression-free survival
- RF:
-
Random forest
- SVM:
-
Support vector machine
- T1-C:
-
T1 contrast
References
Zang J, Li C, Zhao LN et al (2016) Prognostic model of death and distant metastasis for nasopharyngeal carcinoma patients receiving 3DCRT/IMRT in nonendemic area of China. Medicine (Baltimore) 95:e3794
Zhao LN, Zhou B, Shi M et al (2012) Clinical outcome for nasopharyngeal carcinoma with predominantly WHO II histology treated with intensity-modulated radiation therapy in non-endemic region of China. Oral Oncol 48:864–869
Al-Sarraf M, LeBlanc M, Giri PG et al (1998) Chemoradiotherapy versus radiotherapy in patients with advanced nasopharyngeal cancer: phase III randomized intergroup study 0099. J Clin Oncol 16:1310–1317
Wang J, Shi M, Hsia Y et al (2012) Failure patterns and survival in patients with nasopharyngeal carcinoma treated with intensity modulated radiation in Northwest China: a pilot study. Radiat Oncol 7:2
Wee CW, Keam B, Heo DS, Sung MW, Won TB, Wu HG (2015) Locoregionally advanced nasopharyngeal carcinoma treated with intensity-modulated radiotherapy plus concurrent weekly cisplatin with or without neoadjuvant chemotherapy. Radiat Oncol J 33:98–108
Blanchard P, Lee A, Marguet S et al (2015) Chemotherapy and radiotherapy in nasopharyngeal carcinoma: an update of the MAC-NPC meta-analysis. Lancet Oncol 16:645–655
Cao SM, Yang Q, Guo L et al (2017) Neoadjuvant chemotherapy followed by concurrent chemoradiotherapy versus concurrent chemoradiotherapy alone in locoregionally advanced nasopharyngeal carcinoma: a phase III multicentre randomised controlled trial. Eur J Cancer 75:14–23
Zhao L, Xu M, Jiang W et al (2017) Induction chemotherapy for the treatment of non-endemic locally advanced nasopharyngeal carcinoma. Oncotarget 8:6763–6774
Peng H, Chen L, Zhang Y et al (2016) The tumour response to induction chemotherapy has prognostic value for long-term survival outcomes after intensity-modulated radiation therapy in nasopharyngeal carcinoma. Sci Rep 6:24835
Zhang GY, Wang YJ, Liu JP et al (2015) Pretreatment diffusion-weighted MRI can predict the response to neoadjuvant chemotherapy in patients with nasopharyngeal carcinoma. Biomed Res Int 2015:307943
Yen RF, Chen TH, Ting LL, Tzen KY, Pan MH, Hong RL (2005) Early restaging whole-body (18)F-FDG PET during induction chemotherapy predicts clinical outcome in patients with locoregionally advanced nasopharyngeal carcinoma. Eur J Nucl Med Mol Imaging 32:1152–1159
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577
Verma V, Simone CB 2nd, Krishnan S, Lin SH, Yang J, Hahn SM (2017) The rise of radiomics and implications for oncologic management. J Natl Cancer Inst 109
Mao J, Fang J, Duan X et al (2019) Predictive value of pretreatment MRI texture analysis in patients with primary nasopharyngeal carcinoma. Eur Radiol. https://doi.org/10.1007/s00330-018-5961-6
Zhang B, Tian J, Dong D et al (2017) Radiomics features of multiparametric MRI as novel prognostic factors in advanced nasopharyngeal carcinoma. Clin Cancer Res. https://doi.org/10.1158/1078-0432.CCR-16-2910
Wang G, He L, Yuan C, Huang Y, Liu Z, Liang C (2018) Pretreatment MR imaging radiomics signatures for response prediction to induction chemotherapy in patients with nasopharyngeal carcinoma. Eur J Radiol 98:100–106
Eisenhauer EA, Therasse P, Bogaerts J et al (2009) New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 45:228–247
Sauerbrei W, Royston P, Binder H (2007) Selection of important variables and determination of functional form for continuous predictors in multivariable model building. Stat Med 26:5512–5528
Smyser CD, Dosenbach NU, Smyser TA et al (2016) Prediction of brain maturity in infants using machine-learning algorithms. Neuroimage 136:1–9
Liu Z, Zhang XY, Shi YJ et al (2017) Radiomics analysis for evaluation of pathological complete response to neoadjuvant chemoradiotherapy in locally advanced rectal cancer. Clin Cancer Res 23:7253–7262
Contal COQJ (1999) An application of changepoint methods in studying the effect of age on survival in breast cancer. Comput Stat Data Anal 30:253–270
Cui Z, Xia Z, Su M, Shu H, Gong G (2016) Disrupted white matter connectivity underlying developmental dyslexia: a machine learning approach. Hum Brain Mapp 37:1443–1458
Liao XB, Mao YP, Liu LZ et al (2008) How does magnetic resonance imaging influence staging according to AJCC staging system for nasopharyngeal carcinoma compared with computed tomography? Int J Radiat Oncol Biol Phys 72:1368–1377
Liu J, Mao Y, Li Z et al (2016) Use of texture analysis based on contrast-enhanced MRI to predict treatment response to chemoradiotherapy in nasopharyngeal carcinoma. J Magn Reson Imaging 44:445–455
Zhang B, Ouyang F, Gu D et al (2017) Advanced nasopharyngeal carcinoma: pre-treatment prediction of progression based on multi-parametric MRI radiomics. Oncotarget 8:72457–72465
Panth KM, Leijenaar RT, Carvalho S et al (2015) Is there a causal relationship between genetic changes and radiomics-based image features? An in vivo preclinical experiment with doxycycline inducible GADD34 tumor cells. Radiother Oncol 116:462–466
Diehn M, Nardini C, Wang DS et al (2008) Identification of noninvasive imaging surrogates for brain tumor gene-expression modules. Proc Natl Acad Sci U S A 105:5213–5218
Lee J, Narang S, Martinez J, Rao G, Rao A (2015) Spatial habitat features derived from multiparametric magnetic resonance imaging data are associated with molecular subtype and 12-month survival status in glioblastoma multiforme. PLoS One 10:e0136557
Zhou M, Leung A, Echegaray S et al (2017) Non-small cell lung cancer radiogenomics map identifies relationships between molecular and imaging phenotypes with prognostic implications. Radiology. https://doi.org/10.1148/radiol.2017161845:161845
Mani S, Chen Y, Li X et al (2013) Machine learning for predicting the response of breast cancer to neoadjuvant chemotherapy. J Am Med Inform Assoc 20:688–695
Funding
This study has received funding by the National Natural Science Foundation of China Grants 81872699 and Key project of Shanxi Province 2017ZDXM-SF-043.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Guarantor
The scientific guarantor of this publication is Lina Zhao.
Conflict of interest
The authors of this manuscript declare no relationships with any companies, whose products or services may be related to the subject matter of the article.
Statistics and biometry
Yutian Yin, one of the authors, has significant statistical expertise.
Informed consent
Written informed consent was not required for this study because the retrospective nature of the study.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• retrospective
• diagnostic or prognostic study
• performed at one institution
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1217 kb)
Rights and permissions
About this article
Cite this article
Zhao, L., Gong, J., Xi, Y. et al. MRI-based radiomics nomogram may predict the response to induction chemotherapy and survival in locally advanced nasopharyngeal carcinoma. Eur Radiol 30, 537–546 (2020). https://doi.org/10.1007/s00330-019-06211-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-019-06211-x