Abstract
Antarctic environments are characterized by polar climate, making it difficult for the development of any form of life. The biogeochemical cycles and food web in such restrictive environments may be exclusively formed by microorganisms. Polar mycological studies are recent and there is much to know about the diversity and genetic resources of these microorganisms. In this sense, the molecular taxonomic approach was applied to identify 129 fungal isolates from marine and terrestrial samples collected from the King George Island (South Shetland Islands, Maritime Antarctic). Additionally, the production of cold-adapted enzymes by these microorganisms was evaluated. Among the 129 isolates, 69.0% were identified by ITS-sequencing and affiliated into 12 genera. Cadophora, Geomyces, Penicillium, Cosmospora, and Cladosporium were the most abundant genera. Representatives of Cosmospora were isolated only from terrestrial samples, while representatives of the others genera were recovered from marine and terrestrial samples. A total of 29, 19, and 74 isolates were able to produce ligninolytic enzymes, xylanase, and l-asparaginase, respectively. Representatives of Cadophora showed great ability to produce lignin peroxidase (LiP) and laccase at 15.0 °C in liquid medium, while representatives of Penicillium and non-identified fungi were the best producers of xylanase and l-asparaginase at 20.0 °C. The high number of fungi able to produce enzymes at moderate temperature reveals their potential for industrial production and biotechnological applications. The present study broadens the knowledge of fungal diversity associated with the southern polar region. Additionally, data from molecular taxonomy suggest that two filamentous fungi may be considered as potential new species.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The Antarctic environment, including the maritime and continental areas, is considered the coldest, driest, and most inhospitable place on Earth, covered by ice and surrounded by frozen sea (Tindall 2004). As a restrictive environment, diversity in general tends to be lower, and in some systems, biogeochemical cycles and food chains are exclusively formed by microorganisms (Vincent 2000).
The majority of the filamentous fungi that occur in the Antarctic environments are found on other continents, while few species are known to be endemic. Considering the growth temperature, fungi that live in these environments are classified as psychrophilic, psychrotolerant (psychrotrophic), and mesophilic-psychrotolerant. The first ones are able to grow at 0.0 °C with optimum growth at temperatures ≤ 15.0 °C, and maximum growth at temperatures ≤ 20.0 °C. Psychrotolerant fungi can grow at 0.0 °C, but their maximum growth temperature is between 15.0 and 25.0 °C, while the mesophilic-psychrotolerant have the ability to grow at low temperatures, with optimum growth at temperatures > 25.0 and ≤ 40.0 °C (Ruisi et al. 2007; Pesciaroli et al. 2012). According to Godinho et al. (2013), most of the filamentous fungi recovered from Antarctic samples are psychrotolerant, and few are psychrophilic.
The production of enzymes active at low temperatures is one of several survival strategies developed by microbial communities that inhabit Antarctica (Loperena et al. 2012; Duarte et al. 2018). Cold-adapted enzymes have properties that could be interesting for many industrial and/or environmental sectors (Siddiqui and Cavicchioli 2006). According to Feller and Gerday (2003), cold-adapted enzymes are up to tenfold more active at low/moderate temperatures than their mesophilic homologs. Additionally, they can be inactivated by temperature before the unfolding of the protein structure and their heat-lability (mainly in psychrophilic and psychrotolerant microorganisms) can propitiate selective inactivation in complex mixtures. In this context, in the present study, we investigated the production of ligninases, xylanases, and l-asparaginases by filamentous fungi from the Antarctic.
Cold-adapted ligninases and xylanases can be applied in the degradation of lignocellulosic material for biofuel production and in bioremediation of cold environments or processes (Margesin et al. 2002; Del-Cid et al. 2014). Cold-active xylanases are also useful for the generation of chemicals from lignocellulose (Del-Cid et al. 2014). Additionally, the ability of filamentous fungi from Antarctic environments to produce ligninases, xylanases, and/or l-ASNase at moderate temperatures can offer economic benefits by reducing energy costs in production processes conducted at moderate temperatures (25.0–40.0 °C) (Feller and Gerday 2003).
Considering the biotechnological potential of extremophile microorganisms from cold environments, in this study, we assessed filamentous fungi recovered from different samples collected on King George Island (Maritime Antarctica) and also evaluated their ability of producing of cold-adapted enzymes.
Materials and methods
Sampling and isolation
The fungal isolates (n = 129) were recovered from different marine and terrestrial substrates collected from the King George Island (Maritime Antarctica) by the Brazilian Antarctic Program team during an expedition in the southern polar summer of 2010. Sample description and GPS data are listed in Table 1.
Filamentous fungi were isolated on potato dextrose agar medium (in g L−1: 200.0 potato extract, 20.0 dextrose, and 20.0 agar) supplemented with rifampicin (300.0 µg mL−1) and incubated at 15.0 °C. Cultivation and purification of the isolates were performed on the same agar medium. All isolates are maintained in two different preservation methods (cryopreservation at − 80.0 °C in 10% glycerol and in water at 4.0 °C—Castellani) at the Brazilian Collection of Environmental and Industrial Microorganisms—CBMAI (UNICAMP, SP, Brazil) and at the UNESP Central of Microbial Resources—CRM-UNESP (UNESP, SP, Brazil).
Taxonomic identification
Filamentous fungi identification was carried out by ITS barcode sequencing and phylogenetic analysis. The ITS1-5.8S-ITS2 region of the ribosomal DNA was amplified from the genomic DNA extracted from fungal isolates. DNA extraction was carried out following a combined protocol from Moller et al. (1992) and Gerardo et al. (2004) using freshly grown mycelia. ITS4 and ITS5 primers (White et al. 1990) were used for the amplification. PCR conditions were the following: initial denaturation at 94.0 °C for 3 min, followed by 35 cycles at 94.0 °C for 1 min, 55.0 °C for 1 min, and 72.0 °C for 2 min. Amplicons were cleaned up with the Wizard® SV Gel mini kit and PCR Clean-Up System (Promega) and quantified in NanoDrop® (Thermo Scientific). Cycle sequencing reactions were carried out using the BigDye Terminator v. 3.1 Cycle Sequencing Kit (Life Technologies) following the manufacturer’s instructions. The products were sequenced on ABI 3330xl (Life Technologies). Forward and reverse sequences were compiled into contigs in BioEdit v. 7.0.5.3 (Hall 1999). Contigs were queried using the NCBI-GenBank (http://www.ncbi.nml.nih.gov) database to find the closest known sequences.
Sequences of the ITS region were aligned with sequences from the closest relatives found in the database in MAFFT v.7.110. (Katoh and Standley 2013) and manually inspected in Bioedit. The trees were inferred using MEGA v. 6.0 (Tamura et al. 2011) by the neighbor-joining algorithm, Kimura 2-parameters as the nucleotide substitution model, and 1000 bootstrap pseudo-replicates. Sites containing gaps were excluded from the analysis.
Screening of cold-adapted enzymes
Ligninolytic enzymes
The isolates were submitted to a screening on the solid medium using the method described by Verma et al. (2010). Therefore, fungal strains were cultivated at 15.0 °C in B&K medium containing (in g L−1): 10.0 glucose, 2.0 peptone, 1.0 yeast extract, 20.0 agar, and 4.0 mM guaiacol. The presence of brown coloring (guaiacoquinone) on and around the mycelium in the medium supplemented with guaiacol indicated that ligninolytic enzymes are present.
Xylanase
Antarctic-derived fungi were inoculated in tubes containing 1.0 mL of YNB (Yeast Nitrogen Base) medium with xylan as the carbon source. The cultures were incubated at 20.0 °C and at 120 rpm for 7 days. After this period, the enzymatic broth (supernatant) was obtained by centrifugation (15 min at 3000 rpm). Aliquots of 25.0 μL were transferred to sterilized straws arranged into Petri dishes containing agar-xylan medium. Plates were incubated for 48 h at 20.0 °C. After the incubation time, the straws were removed and the culture medium was colored with lugol. The presence of a degradation halo in the culture medium indicated that enzymes are present (Costa 2014).
l-Asparaginase
Screening was performed using Czapek Dox medium (Gulati et al. 1997) containing (in g L−1): 2.0 glucose, 10.0 l-asparagine, 1.52 KH2PO4, 0.52 KCl, 0.52 MgSO4·7H2O, 0.001 CuNO3·3H2O, 0.001 ZnSO4·7H2O, and 0.001 FeSO4·7H2O, pH 6.2, added with phenol red (0.009% final concentration). Plates were incubated for 7 days at 20.0 °C (l-ASNase activity was identified by the formation of a pink zone around colonies).
Assay of enzymatic activities
Ligninolytic enzymes
Fungal strains were cultivated in 2% (w/v) malt extract agar (MA2). Mycelium fragments (0.5 cm in diameter) taken from the edge of the colonies (n = 3) were transferred to 150-mL Erlenmeyer flasks containing 50 mL of MA2%. The assays were incubated at 15.0 °C and 25.0 °C under agitation at 150 rpm for 7 days, in duplicate. The extracts were obtained by centrifuging the liquid culture medium at 10,000 rpm for 30 min at 4.0 °C. Enzymatic activity was determined in triplicate using different enzyme substrates according to the methodology described below. One enzyme unit was defined as 1.0 μmol of product formed per minute under the assay conditions, obtained from calculations performed from Eq. 1, which was derived from the Beer–Lambert Law:
where ∆A is the difference between the final and initial absorbance, V is the reaction volume (0.001 L in all cases), 106 is the conversion of moles from ɛ to μmols, ɛ is the extinction coefficient (M−1 cm−1), R is the amount of enzyme in the broth (L), and T is the reaction time (min).
Laccase activity
Laccase activity was determined using 2.2′-azinobis-(3-ethylbenzothiazoline (ABTS) as previously described by Buswell et al. (1995). The mixture was composed of 0.3 mL sodium acetate buffer (0.1 M pH 5.0), 0.1 mL ABTS solution (0.03% w/v), and 0.6 mL enzyme solution. ABTS oxidation was measured by monitoring the increase in absorbance at 420 nm (ε = 36,000 M−1 cm−1) at 37.0 °C for 10 min.
Lignin peroxidase activity
LiP activity was determined by the oxidation of veratryl alcohol as previously described by Tien and Kirk (1984). The mixture reaction was composed of 500.0 μL enzyme extract, 1.0 mL sodium tartrate buffer (125.0 mM pH 3.0), 500.0 μL veratryl alcohol (10.0 mM), and 500.0 μL hydrogen peroxide (2.0 mM). The reaction was initiated with hydrogen peroxide and the appearance of veratraldehyde was measured at 310 nm (ε = 9300 M−1cm−1).
Xylanase
Fungal strains were cultivated in 2% (w/v) malt extract agar (MA2) for 7 days at 20.0 °C. After this period, three fungal plugs (0.5 cm in diameter) taken from the edge of the colony were transferred to 200-mL Erlenmeyer flasks containing 50.0 mL of Mandels and Sternberg medium (MS), consisting of (in g L−1) 1.0 peptone, 1.4 (NH4)2SO4, 2.0 KH2PO4, 0.3 urea, 0.3 CaCl2, 0.3 MgSO4·7H2O, 0.005 FeSO4·7H2O, 0.001 MnSO4·H2O, 0.001 ZnSO4·7H2O, and 0.002 CoCl2, pH 6.0, with 10.0 birchwood xylan (Sigma-Aldrich) as the carbon source. Assays were incubated for 7 days at 140 rpm and 20.0 °C. Cultures were harvested by centrifugation at 12,074×g for 30 min and the supernatant was used for the enzymatic quantification.
Xylanase activity
Xylanase activity was quantified by determining the amount of reducing sugar released from xylan derived from birchwood according to Bailey et al. (1992). The xylanase activity assay was performed by adding 20.0 μL of enzymatic broth into 50.0 mM citrate buffer (pH 5.0) with 1% (w/v) birchwood xylan (Sigma) at 40.0 °C for 5 min. The generated reducing sugar was measured by using dinitrosalicylic acid (DNS) (Miller 1959). One unit of enzyme activity was defined as the amount of enzyme required to release 1 μmol of product equivalent per min in assay conditions.
l-Asparaginase
Fungal strains were cultivated in PDA at 20.0 °C for 7 days. After this period, three mycelium fragments (0.5 cm in diameter) taken from the edge of the colonies were transferred to 200 mL-Erlenmeyer flasks containing 50.0 mL of the modified Czapek Dox medium consisting of 10.0 g L−1 l-proline as the inducer of l-ASNase, and incubated at 20.0 °C for 7 days. Cultures were harvested by centrifugation at 12,074 g for 30 min and the supernatant was used for enzymatic quantification.
l-Asparaginase activity
l-ASNase activity was measured by the ammonia produced during the hydrolysis of l-asparagine (Sigma–Aldrich/USA) catalyzed by the enzyme through Nessler’s reagent (Merck–Millipore). The method is divided into two steps. The first step was performed by adding 0.1 mL of enzymatic broth, 1.0 mL Tris buffer (50 mM) pH 8.6, asparagine 0.1 mL (189.0 mM), and 0.9 mL of Milli-Q water. The mixture was incubated at 37.0 °C for 30 min. After this period, 0.1 mL of trichloroacetic acid (TCA) 1.5 M was added to stop the reaction. In the second step, the mixture reaction was composed of 0.2 mL supernatant, 4.3 mL milli-Q water, and 0.5 mL Nessler’s reagent. The reading was conducted spectrophotometrically at 436 nm. The concentration of ammonia produced in the reaction was determined based on a standard curve obtained using ammonium sulfate as standard (0.0 to 20.0 μmol mL−1). One l-ASNase unit was considered as the amount of enzyme that catalyzes the formation of 1.0 μmol of ammonia per min at 37.0 °C and was expressed (Gulati et al. 1997; Theantana et al. 2007).
Results
Taxonomic evaluation
From the total number of isolates (n = 129), 89 (69%) were identified based on molecular taxonomy, the other fungal isolates (31%) could not be identified due to PCR amplification failures or low-quality sequences. Data derived from sequencing (Table 2) (Online Resource 1, Table S1) and phylogenetic analyses (Online Resource 1, Fig. S1) revealed that the Antarctic filamentous fungi recovered from the terrestrial and marine samples belong to 12 ascomycetous genera: Acremonium, Cadophora, Cercospora, Cladosporium, Cosmospora, Geomyces, Hypocrea, Oidiodendron, Penicillium, Pseudeurotium, Pseudogymnoascus, and Thelebolus.
Representatives of the genus Cadophora (n = 29, 32.5%) were isolated in greater abundance, followed by Geomyces (n = 18, 20.4%), Penicillium (n = 12, 13.6%), Cosmospora (n = 8, 9%), Cladosporium (n = 7, 9%), and the genera Thelebolus, Oidiodendron, Acremonium, Cercospora, Hypocrea, and Pseudeurotium, which were isolated in low abundances.
Fungal isolates representing the three most abundant genera were recovered from marine and terrestrial samples. Filamentous fungi affiliated to the Cadophora genus were recovered from wood (n = 11), marine sediment (n = 5), lichen (n = 4), ascidian (n = 3), sea star (n = 2), sea urchin (n = 1), Salpa sp. (n = 1), and whale bones (n = 1). Representatives of three different taxa of Cadophora were identified: Cadophora malorum, Cadophora luteo-olivacea, and Cadophora sp. The genus Geomyces was the second in abundance and was recovered from eight different samples (Table 2). The majority of isolates was recovered from lichen samples (n = 8) and the other ones from Salpa sp. (n = 4), amphipod (n = 2), wood (n = 1), isopod (n = 1), sea star (n = 1), and marine sediment (n = 1). Penicillium isolates were recovered from seven different samples, including sea star (n = 3), lichen (n = 2), wood (n = 2), Nacella sp. (n = 2), and isopod, whale bones, and Salpa sp. (n = 1, each one).
The genus Cosmospora was recovered only from terrestrial samples, including whale bones (n = 7) and ornithogenic soil (n = 1). Representatives of Cladosporium were isolated from marine and terrestrial samples, one isolate was recovered from isopod identified as C. halotolerans, and the others identified as Cladosporium sp. and recovered from ornithogenic soil (n = 2), wood (n = 1), amphipod (n = 1), Salpa sp. (n = 1), and macroalgae (n = 1). Additional genera isolated in less abundance were Thelebolus (sea star and ornithogenic soil), Oidiodendron (lichen and macroalgae), Acremonium (lichen), Cercospora (sea urchin), Hypocrea (lichen), and Pseudeurotium (wood and marine sediment).
Cold-adapted enzymes
Ligninolytic enzymes
A total of 29 (22.4%) filamentous fungi were selected based on the presence of brown coloring in solid medium containing guaiacol, which indicates the presence of ligninolytic enzymes. Some fungi showed a very intense brown color around the mycelium, while others had a brown color a bit lighter, which may be associated with the enzymatic potential of each isolate.
Antarctic filamentous fungi considered positive for ligninolytic enzymes were isolated from terrestrial (n = 17) and marine samples (n = 12). The majority of ligninolytic fungi were recovered from wood (n = 13), others were isolated from sea star (n = 3), marine sediment (n = 2), lichens (n = 2), whale bones (n = 2), macroalgae (n = 1), amphipod (n = 1), isopod (n = 1), Nacella sp. (n = 1), sea urchin (n = 1), and Salpa sp. (n = 1).
Most of the ligninolytic fungi were identified as belonging to the genus Cadophora (n = 12), ten isolates were representatives of Cadophora malorum and two Cadophora luteo-olivacea. Other fungi selected based on the potential to produce ligninolytic enzymes were identified as representatives of the genera Penicillium, Oidiodendron, Cosmospora, Geomyces, and Cladosporium. A total of four fungal isolates could not be identified by molecular taxonomy.
Among the selected fungi (n = 29), Cadophora luteo-olivacea P1 and Cadophora malorum A2B showed the highest laccase activity at 15.0 °C (Fig. 1a), both recovered from marine samples: marine sediment and sea star, respectively. Cadophora malorum AS-2A, also recovered from the marine sample (ascidian), produced the highest amount of laccase at 25.0 °C (Fig. 1a). Other isolates were able to produce laccase in very low amounts. On the other hand, the best production of LiP at 15.0 °C and 25.0 °C (Fig. 1b) was achieved by fungi recovered from terrestrial samples: Cosmospora sp. OB4B, which was recovered from whale bones and Cadophora malorum M7, recovered from the wood sample.
A higher number of fungi from marine origin were able to produce laccase and LiP at 25.0 °C in comparison to the number of isolates capable of producing these enzymes at 15.0 °C. On the other hand, a higher number of fungi recovered from terrestrial samples were able to produce these enzymes at 15.0 °C (Fig. 2).
Xylanase
A total of 19 (14.7%) Antarctic filamentous fungi were able to degrade xylan in the solid medium, seven were isolated from terrestrial samples, and 12 from marine samples: sea star (n = 3), amphipod, Nacella sp. (n = 2); marine sediment, snail, Salpa sp., ascidian (n = 1), wood (n = 3), lichens, whale bones, ornithogenic soil (n = 1). Among the positive isolates, four were identified as Penicillium sp. and three as Cadophora luteo-olivacea.
The results of enzymatic activity revealed that seven filamentous fungi were able to produce more than 10 U mL−1 of xylanase (Fig. 3). Three of them were identified as Penicillium sp. (E2B, N5, and E2-1) and four could not be identified based on barcode sequences (N2C, C2, L1-4, and E2C). The highest number of fungi able to produce xylanase was recovered from marine sources. The unidentified fungus N2C (recovered from Nacella sp.) and Penicillium sp. E2B (recovered from sea star) showed the best xylanase activity.
l-asparaginase
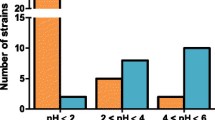
A total of 74 (57%) Antarctic filamentous fungi were positive for l-ASNase based on the formation of a pink zone in solid medium as a result of the conversion of the phenol red from yellow (acidic condition) to pink (alkaline condition). Among the positive strains, 36 and 38 were isolated from terrestrial and marine samples, respectively: Salpa sp. (n = 12), sea star (n = 8), marine sediment (n = 7), isopod (n = 7), Nacella sp. (n = 2), ascidian (n = 2), lichens (n = 17), whale bones (n = 10), ornithogenic soil (n = 5), and wood (n = 4).
A total of 42 filamentous fungi able to produce l-ASNase in solid medium were identified using molecular taxonomy: Geomyces sp. (n = 11), Pseudogymnoascus sp. (n = 1), Cosmospora sp. (n = 7), Hypocrea sp. (n = 2), Cadophora luteo-olivacea (n = 3), Cadophora malorum (n = 6), Cladosporium sp. (n = 4), Penicillium sp. (n = 5), Pseudeurotium sp., Oidiodendron sp., and Acremonium sp. (n = 1).
Among the total of filamentous fungi that exhibited l-ASNase activity in liquid medium (n = 8), three were identified as Cosmospora sp. (all isolated from whale bones: OB4B, OB1B, and OB2), two as Penicillium sp. (recovered from sea star: C2 and E2B), and one as Geomyces sp. (recovered from Salpa sp.: S2B) (Fig. 4). The isolates OB7A (recovered from whale bones) and L2-16A (recovered from lichen) could not be identified using molecular taxonomy. Penicillium sp. C2 and the unidentified fungus OB7A showed the highest asparaginase activities (Fig. 4). Penicillium sp. isolates C2 and E2B were also able to produce xylanase.
Discussion
Taxonomic evaluation
All filamentous fungi identified in the present study belong to the phylum Ascomycota. Previous studies reported the prevalence of Ascomycota among the filamentous fungi found in the Antarctic environments (Arenz et al. 2006; Arenz and Blanchette 2009; Santiago et al. 2016), including studies related to next-generation DNA sequencing (Dreesens et al. 2014; Pudasaini et al. 2017). Filamentous fungi belonging to the phylum Basidiomycota are rarely found in Antarctica (Arenz and Blanchette 2009), the representatives of this phylum were mainly composed of yeasts (Ludley and Robinson 2008; Duarte et al. 2013, 2016). However, the predominance of ascomycetes observed in the present study may be due to the culture medium used for fungal isolation, since representatives of ascomycetes show faster growth on PDA than basidiomycetes.
Based on sequencing and phylogenetic analyses, 12 different genera were identified, all of them previously reported in the Antarctic environments. The most abundant genus was Cadophora, followed by Geomyces, Penicillium, Cosmospora, and Cladosporium. With the exception of the genus Cosmospora, whose representatives were recovered only from terrestrial samples, all the other representatives of the most abundant genera were isolated from terrestrial and marine samples, suggesting they are widespread and adapted to the Antarctic extreme conditions.
According to Arenz et al. (2006), representatives of Cadophora have been reported as wood decomposers in the Antarctic environments, and in the present study, eight Cadophora isolates were recovered from wood samples. Cadophora is commonly found in Antarctic soils and has a broad growth temperature ranging from − 3.0 to 45.0 °C (Onofri et al. 2004). The predominance of this genus may suggest that it is an important microorganism in nutrient cycling in the polar regions.
The genus Geomyces presents psychrophilic features and halotolerance (Godinho et al. 2013). Species belonging to this genus have been reported by Zhdanova et al. (2000) to grow in high levels of radiation (greater than 220 mR h−1, measured as the level of gamma radiation). These properties can explain their high abundance in Antarctic samples.
Many species in the genera Penicillium and Cladosporium are psychrotolerant, often found in different environments, including the Antarctic and Arctic polar regions (Mcrae et al. 1999; Held et al. 2006; Sonjak et al. 2006; Gonçalves et al. 2013, 2015). According to Onofri et al. (2004), the species Cladosporium cladosporioides can grow in temperatures ranging from 0.0 to 32.0 °C with optimum temperatures ranging from 20.0 to 28.0 °C. Additionally, representatives of the genus Penicillium have a strong capacity to form spores and to tolerate saline conditions (Menezes et al. 2010).
Despite this, in the present study, the genus Cosmospora were recovered only from terrestrial Antarctic samples. Representatives of this genus were reported from samples from Antarctic lakes (Gonçalves et al. 2012). Additionally, they were also found in the rhizosphere soil of Deschampsia antarctica (Gonçalves et al. 2015), one of the two vascular plants that inhabit the maritime Antarctic. The majority of Cosmospora species is the sexual stage of Acremonium-like or Fusarium-like fungi (Gräfenhan et al. 2011). Representatives of these genera are psychrotrophic and are reported also in Arctic soil (Ali et al. 2013).
Most fungi reported from Antarctica are considered asexual. This fact can be explained by a survival strategy, since asexual reproduction can be completed in a shorter time and without high metabolic costs. Some exceptions of sexual fungi have been reported, such as Thelebolus, frequently isolated from Arctic and Antarctic environments (Hoog et al. 2005; Ruisi et al. 2007), and Cosmospora.
Among all fungi identified in this study, a large amount came from terrestrial samples, the majority were isolated from wood and lichen. Antarctic lichens have been reported as a natural microhabitat with favorable conditions for microbial survival. Santiago et al. (2016) identified 74 taxa from 21 genera of filamentous fungi and yeasts recovered from Antarctic lichens, Pseudogymnoascus, Thelebolus, and Antarctomyces are the most abundant genera. In addition, Duarte et al. (2016) recovered 200 isolates of yeasts from Antarctic lichen samples collected from different islands (South Shetland Archipelago), six of them are considered potential new species.
Many new fungal species recovered from Antarctic environments were recently described (Laich et al. 2013, 2014; Zhang et al. 2014; Turchetti et al. 2015; de Menezes et al. 2017), highlighting the relevance of the studies related to the diversity and exploitation of Antarctic-derived fungi. In the present study, the isolates Pseudeurotium sp. M14 and Oidiodendron sp. A11-M1 showed low similarity with sequences deposited in GenBank. In this sense, they may represent new species. However, further taxonomic studies should be done in order to confirm this hypothesis.
Cold-adapted enzymes
Considering the enzyme-producing ability of microorganisms, the properties of cold-adapted enzymes, and that fungi from maritime Antarctica are mainly classified as psychrotolerant, the selection approach applied in the present study was carried out at 15.0, 20.0, and 25.0 °C.
Different groups of fungi have been reported as producers of ligninolytic enzymes. Among them, the basidiomycetes (white rot fungi) are considered the most efficient in the breakdown of lignocellulosic materials (Bonugli-Santos et al. 2010a). In the present study, a higher number of fungi from the terrestrial Antarctic samples showed the ability to produce laccase and LiP at 15.0 and 25.0 °C. However, the number of marine-derived fungi able to produce these enzymes was also significant.
Despite the relevance of the ligninolytic enzymes for bioremediation of environmental pollutants and industrial applications (Bonugli-Santos et al. 2010b), studies related to the production of these enzymes by Antarctic fungi are scarce (Duarte et al. 2018). The ability of fungi from Antarctic origin to degrade environmental pollutants was reported by Gerginova et al. (2013) and Litova et al. (2014). Some of the genera identified in the present study (e.g., Geomyces and Thelebolus) were isolated from Antarctic soils contaminated with oil (Kerry 1990), highlighting their potential for oil tolerance/degradation. Additionally, the fungus Geomyces sp. M22 showed the ability to produce ligninolytic enzymes, which are one of the microbial pathways for pollutant degradation.
The largest number of fungi able to produce xylanase was recovered from marine sources. Bugni and Ireland (2004) highlighted the biotechnological potential of the marine microorganisms, based on their adaptation to the saline and pressure conditions that can lead to the production of natural products different from those produced by their terrestrial counterparts.
Xylanase have many industrial applications, including bread-making to help the dough rise (Courtin and Delcour 2002), pulp pre-bleaching in paper production (Bajpai 2004), as an animal feed additive to improve digestibility (Cowieson et al. 2006), production of ethanol (Polizeli et al. 2005), and in the synthesis of xylitol (Granström et al. 2007). However, there are few studies reported in the literature related to the production of this enzyme by Antarctic fungi (Duarte et al. 2018). According to Bradner et al. (1999) and Del-Cid et al. (2014), filamentous fungi from Antarctic ornithogenic soil and from marine sponges were reported as xylanase producers, including representatives of the genera Alternaria, Cladosporium, Penicillium, Phoma, and Trichoderma. In the study reported by Del-Cid et al. (2014), the fungus Cladosporium sp. recovered from Antarctic marine sponge was reported as a great producer of this enzyme at low temperatures when pure xylans from hardwoods (birchwood and beechwood) were used as carbon sources.
l-Asparaginase is a relevant medical enzyme, seeing that it is used as an anti-tumoral medicine in the treatment of lymphoblastic leukemia and lymphosarcoma. l-ASNase catalyzes the hydrolysis of l-asparagine into l-aspartic acid and ammonia. The depletion of l-asparagine in blood blocks the protein synthesis and leads to DNA and RNA synthesis inhibition, resulting in cellular apoptosis (Lopes et al. 2017). The high number of Antarctic filamentous fungi that were selected as l-ASNase producers (57%) in the first screening in solid medium, revealed the potential of these microorganisms. The microbial production of this enzyme is cost effective and eco-friendly, and the production of l-ASNase by an eukaryotic organism could avoid allergenic effects that are caused by l-ASNase industrially produced by prokaryotic organisms (Theantana et al. 2007).
Considering the potentially different properties of the microbial cold-adapted enzymes, the collection of Antarctic fungi structured from the marine and terrestrial samples can be considered relevant for biotechnological exploitation and is available for new studies. Additionally, fungal strains selected in the present study may produce enzymes with kinetic parameters important for specific biotechnological applications. In this sense, further studies related to low temperature activity, enzymatic optimization, and characterization should be performed.
References
Ali SH, Alias SA, Siang HY, Smykla J, Pang KL, Guo SY, Convey P (2013) Studies on diversity of soil microfungi in the Hornsund area, Spitsbergen. Pol Polar Res 34:39–54
Arenz BE, Blanchette RA (2009) Investigations of fungal diversity in wooden structures and soils at historic sites on the Antarctic Peninsula. Can J Microbiol 55:46–56
Arenz BE, Held BW, Jurges JA, Farrell RL, Blanchette RA (2006) Fungal diversity in soils and historic wood from the Ross sea region of Antarctica. Soil Biol Biochem 38:3057–3064
Bailey MJ, Biely P, Poutanen K (1992) Interlaboratory testing of methods for assay of xylanase activity. J Biotechnol 23:257–270
Bajpai P (2004) Biological bleaching of chemical pulps. Crit Rev Biotechnol 24:1–58
Bonugli-Santos RC, Durrant LR, Sette LD (2010a) Laccase activity and putative laccase genes in marine-derived basidiomycetes. Mycol Res 114:863–872
Bonugli-Santos RC, Durrant LR, Sette LD (2010b) Production of laccase, manganese peroxidase and lignin peroxidase by Brazilian marine-derived fungi. Enz Microb Technol 46:17–37
Bradner JR, Gillings M, Nevalainen KMH (1999) Qualitative assessment of hydrolytic activities in antarctic microfungi grown at different temperatures on solid media. World J Microbiol Biotechnol 15:131–132
Bugni TS, Ireland CM (2004) Marine-derived fungi: a chemically and biologically diverse group of microorganisms. Nat Prod Rep 21:143–163
Buswell JK, Cai YJ, Chang ST (1995) Effect of nutrient nitrogen on manganese peroxidase and laccase production by Lentinula (Lentinus) edodes. FEMS Microbiol Lett 128:81–88
Costa RR (2014) Perfil Enzimático e Potencial Biotecnológico de Fungos Isolados de Jardins de Fungo das Formigas Cortadeiras. Dissertation. Universidade Estadual Paulista “Julio de Mesquita Filho”. Rio Claro, São Paulo
Courtin CM, Delcour JA (2002) Arabinoxylans and endoxylanases in wheat flour bread-making. J Cereal Sci 35:225–243
Cowieson AJ, Hruby M, Pierson EE (2006) Evolving enzyme technology: impact on commercial poultry nutrition. Nutr Res Rev 19:90–103
De Menezes GC, Godinho VM, Porto BA, Gonçalves VN, Rosa LH (2017) Antarctomyces pellizariae sp. nov., a new, endemic, blue, snow resident psychrophilic ascomycete fungus from Antarctica. Extremophiles 21:259–269
Del-Cid A, Ubilla P, Rayanal MC, Medina E, Vaca I, Levicán G, Evzaquirre J, Chávez R (2014) Cold-active xylanase produced by fungi associated with Antarctic marine sponges. Appl Biochem Biotechnol 172:524–532
Dreesens LL, Lee CK, Cary SG (2014) The distribution and identity of edaphic fungi in the McMurdo Dry Valleys. Biology 3:466–483
Duarte AWF, Dayo-Owoyemi I, Nobre FS, Pagnocca FC, Chaud LSC, Pessoa JR, Felipe MGA, Sette LD (2013) Taxonomic assessment and enzymes production by yeasts isolated from marine and terrestrial Antarctic samples. Extremophiles 17:1023–1035
Duarte AWF, Passarini MR, Delforno TP, Pellizzari FM, Cipro CVZ, Montone RC, Petry MV, Putzke J, Rosa LH, Sette LD (2016) Yeasts from macroalgae and lichens that inhabit the South Shetland Islands, Antarctica. Environ Microbiol Rep 8:874–886
Duarte AWF, Santos Dos, Vianna MV, Vieira JMF, Mallagutti VH, Inforsato FJ, Wentzel LCP, Lario LD, Rodrigues A, Pagnocca FC, Pessoa JR, Sette LD (2018) Cold-adapted enzymes produced by fungi from terrestrial and marine Antarctic environments. Crit Rev Biotechnol 38:600–619
Feller G, Gerday C (2003) Psychrophilic enzymes: hot topics in cold adaptation. Nature 1:200–208
Gerardo NM, Currie CR, Price SL, Mueller UG (2004) Exploiting a mutualism: parasite specialization on cultivars within the fungus-growing ants symbiosis. Proc R Soc Lond B Bio Sci 271:1791–1798
Gerginova MG, Peneva NM, Krumova ET, Alexieva ZA (2013) Biodegradation ability of fungal strains isolated from Antarctic towards pah. In: Proceedings of the 13th international conference of environmental science and technology Athens, Greece, 5–7 Sept
Godinho VM, Furbino LE, Santiago IF, Pellizzari FM, Yokoya N, Pupo D, Alves TMA, Sales PA, Romanha AJ, Zani CL, Cantrell CL, Rosa CA, Rosa LH (2013) Diversity and bioprospecting of fungal communities associated with endemic and cold-adapted macroalgae in Antarctica. ISME J 7:1434–1451
Gonçalves VN, Vaz AB, Rosa CA, Rosa LH (2012) Diversity and distribution of fungal communities in lakes of Antarctica. FEMS Microbiol Ecol 82:459–471
Gonçalves VN, Campos LS, Melo IS, Pellizari VH, Rosa CA, Rosa LH (2013) Penicillium solitum: a mesophilic, psychrotolerant fungus present in marine sediments from Antarctica. Polar Biol 36:1823–1831
Gonçalves VN, Carvalho CR, Johann S, Mendes G, Alves TMA, Zani CL, Junior PAS, Murta SMF, Romanha AJ, Cantrell CL, Rosa CA, Rosa LH (2015) Antibacterial, antifungal and antiprotozoal activities of fungal communities present in different substrates from Antarctica. Polar Biol 38:1143–1152
Gräfenhan T, Schroers JH, Nirenberg HI, Seifert KA (2011) An overview of the taxonomy, phylogeny, and typification of nectriaceous fungi in Cosmospora, Acremonium, Fusarium, Stilbella, and Volutella. Stud Mycol 68:79–113
Granström TB, Izumori K, Leisola M (2007) A rare sugar xylitol. Part II: biotechnological production and future applications of xylitol. Appl Microbiol Biotechnol 74:273–276
Gulati R, Saxena R, Gupta RA (1997) Rapid plate assay for screening L-asparaginase producing microorganisms. Lett Appl Microbiol 24:23–26
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Held BW, Jurgens JA, Duncan SM, Farrell RL, Blanchette RA (2006) Assessment of fungal diversity and deterioration in a wooden structure at New Harbor, Antarctica. Polar Biol 29:526–531
Hoog GS, Gottlich E, Platas G, Genilloud O, Leotta G, Brummelen J (2005) Evolution, taxonomy and ecology of the genus Thelebolus in Antarctica. Stud Mycol 51:33–76
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Bio Evol 30:772–780
Kerry E (1990) Microorganisms colonizing plants and soil subjected to different degrees of human activity, including petroleum contamination, in the Vestfoid Hills and MacRobertson Land, Antarctica. Polar Biol 10:423–430
Laich F, Vaca I, Chávez R (2013) Rhodotorula portillonensis sp. nov., a bassidiomycetous yeast isolated from Antarctic shallow-water marine sediment. Int J Syst Evol Microbiol 63:3884–3891
Laich F, Chávez R, Vaca I (2014) Leucosporidium escuderoi f.a., sp. nov., a basidiomycetous yeast associated with an Antarctic marine sponge. Antonie Van Leeuwenhoek 105:593–601
Litova K, Gerginova M, Peneva N, Manasiev J, Alexieva Z (2014) Growth of Antarctic fungal strains on phenol at low temperatures. J BioSci Biotech 1:43–46
Loperena L, Sonia V, Varela H, Lupo S, Bergalli A, Guigou M, Pellegrino A, Bernardo A, Calviño A, Rivas F, Batista S (2012) Extracellular enzymes produced by microorganisms isolated from maritime Antarctica. World J Microbiol Biotechnol 28:2249–2256
Lopes AM, Oliveira-Nascimento L, Ribeiro A, Tairum-Jr AC, Breyer CA, Oliveira MA, Monteiro G, Souza-Motta CM, Magalhães PO, Avendaño JGF, Cavaco-Paulo AM, Mazzola PG, Rangel-Yagui CO, Sette LD, Converti A, Pessoa A (2017) Therapeutic l-asparaginase: upstream, downstream and beyond. Crit Rev Biotechnol 37:82–99
Ludley KE, Robinson CH (2008) ‘Decomposer’ basidiomycota in Arctic and Antarctic ecosystems. Soil Biol Biochem 40:11–29
Margesin R, Feller G, Gerday C, Russell N (2002) Cold-adapted microorganisms: adaptation strategies and biotechnological potential. In: Bitton (ed) The encyclopedia of environmental microbiology. Wiley, New York, pp 871–885
Mcrae CF, Hocking AD, Seppelt RD (1999) Penicillium species from terrestrial habitats in the Windmill Islands, East Antarctica, including a new species, Penicillium antarcticum. Polar Biol 21:97–111
Menezes CB, Bonugli-Santos RC, Miqueletto PB, Passarini MRZ, Silva CHD, Justo MR, Leal RR, Fantinatti-Garboggini F, Oliveira VM, Berlinck RGS, Sette LD (2010) Microbial diversity associated with algae, ascidians and sponges from the north coast of São Paulo state, Brazil. Microbiol Res 165:466–482
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31:426–428
Moller EM, Bahnweg G, Sandermann H, Geiger HH (1992) A simple and efficient protocol for isolation of high molecular weight DNA from filamentous fungi fruit bodies and infected plant tissues. Nucleic Acids Res 20:6115e6116
Onofri S, Selbmann L, Zucconi SP (2004) Antarctic microfungi as models for exobiology. Planet Space Sci 52:229–237
Pesciaroli C, Cupini F, Selbmann L, Barghini P, Fenice M (2012) Temperature preferences of bacteria isolated from seawater collected in Kandalaksha Bay, White Sea, Russia. Polar Biol 35:435–445
Polizeli ML, Rizzatti AC, Monti R, Terenzi HF, Jorge JA, Amorim DS (2005) Xylanases from fungi: properties and industrial applications. Appl Microbiol Biotechnol 67:577–591
Pudasaini S, Wilson J, Ji M, van Dorst J, Snape I, Palmer AS, Burns BP, Ferrari (2017) Microbial diversity of browning peninsula, Eastern Antarctica revealed using molecular and cultivation methods. Front Microbiol 8:591
Ruisi S, Barreca D, Selbmann L, Zucconi L, Onofri S (2007) Fungi in Antarctica. Rev Environ Sci Biotechnol 6:127–141
Santiago IF, Soares MA, Rosa CA, Rosa LA (2016) Lichensphere: a protected natural microhabitat of the non-lichenised fungal communities living in extreme environments of Antarctica. Extremophiles 19:1087–1097
Siddiqui KS, Cavicchioli R (2006) Cold-adpated enzymes. Annu Rev Biochem 75:403–433
Sonjak S, Frisvad JC, Gunde-Cimerman N (2006) Penicillium mycobiota in Arctic subglacial ice. Microbial Ecol 52:207–216
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Theantana T, Hyde KD, Lumyong S (2007) Asparaginase production by Endophytic fungi isolated from some Thai medicinal plants. KMITL Sci Tech 7:13-1
Tien M, Kirk TK (1984) Lignin-degrading enzyme from Phanerochaete chrysosporium: purification, characterization and catalytic properties of unique H2O2 requiring oxygenase. Proc Natl Aca Sci USA 81:2280–2284
Tindall BJ (2004) Prokaryotic diversity in the Antarctic: the tip of the iceberg. Microb Ecol 47:271–283
Turchetti B, Selbmann L, Blanchette RA, Di Mauro S, Marchegiani E, Zucconi L, Arenz BE, Buzzini P (2015) Cryptococcus vaughanmartiniae sp. nov. and Cryptococcus onofrii sp. nov.: two new species isolated from worldwide cold environments. Extremophiles 19:149–159
Verma AK, Ragggukkumar C, Verma P, Shouche YS, Nail CG (2010) Four marine-derived fungi for bioremediation of raw textile mill effluents. Biodegradation 21:217–233
Vincent WF (2000) Evolutionary origins of Antarctic microbiota: invasion, selection and endemism. Antarct Sci 12:374–385
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Shinsky TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, New York, pp 315–322
Zhang T, Zhang YQ, Liu HY, Zhao LX, Yu LY (2014) Cryptococcus fildesensis sp. nov., a psycrophilic basidiomycetous yeast isolated from Antarctic moss. Int J Syst Evol Microbiol 64:675–679
Zhdanova NN, Zakaarchenko VA, Vember VV, Nakonechnaya LT (2000) Fungi from Chernobyl: mycobiota of the inner regions of the containment structures of the damaged nuclear reactor. Mycol Res 104:1421–1426
Acknowledgements
This study was financed by Fundação de Amparo à Pesquisa do Estado de São Paulo (Grants 2013/19486-0, 2016/07957-7). MBB and JAS thank the Coordenação de Aperfeiçoamento Pessoal de Nível Superior (CAPES) for their scholarships. LDS thanks the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq) for the Productivity Fellowship (303145/2016-1) and the Brazilian Antarctic Program (PROANTAR). The authors thank Professor Eduardo C. M. Hajdu and Dr. Itamar S. de Melo for the samplings of marine invertebrates and ornithogenic soil, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
300_2018_2387_MOESM1_ESM.pdf
Supplementary material 1 (PDF 2418 kb) Electronic Supplementary Material: Online Resource 1. Table S1 Data related to the molecular identification of filamentous fungi from King George Island: numbers of pb analyzed and GenBank accession numbers; Fig. S1 Phylogenetic analysis of the sequences of fungi from King George Island, compared with type and closest sequences of the species following BLAST analysis. (A) Acremonium; (B) Cadophora; (C) Cercospora; (D) Cladosporium; (E) Cosmospora; (F) Geomyces and Pseudogymnoascus; (G) Oidiodendron; (H) Pseudeurotium; (I) Thelebolus. The trees were constructed based on the ITS region using MEGA v. 6.0 by the neighbor-joining algorithm, Kimura 2-parameters as the nucleotide substitution model and 1000 bootstrap pseudo-replicates (values lower than 50% are not shown). Sites containing gaps were excluded from the analysis. The scale bar indicates substitutions per site. All sequences were retrieved from GenBank (accessions are shown in parentheses). CBS (Centraalbureau voor Shimmelcutures), ATCC (American Type Culture Collection), IMI (International Mycological Institute), UAMH (University of Alberta Microfungus Collection and Herbarium) and MUCL (Mycothèque de l´Université Catholique de Louvain) voucher accessions are also shown. T: type strain; NT: neotype strain.
Rights and permissions
About this article
Cite this article
Duarte, A.W.F., Barato, M.B., Nobre, F.S. et al. Production of cold-adapted enzymes by filamentous fungi from King George Island, Antarctica. Polar Biol 41, 2511–2521 (2018). https://doi.org/10.1007/s00300-018-2387-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-018-2387-1