Abstract
Few studies have addressed the diversity of cultivable fungi from marine sediments, especially those from Antarctica. In the present study, we evaluated the presence and distribution of cultivable fungi in marine core sediments obtained from 100, 500, 700 and 1,100 m below the Antarctic Ocean surface. Fifty-two fungal isolates were identified as Penicillium solitum by their physiological and morphological characteristics, and the identity of 12 representative isolates was further confirmed by sequencing of the ITS1-5.8S-ITS2 and β-tubulin genes. P. solitum displayed high sequence similarity to Penicillium taxa that have been described from other marine habitats. Conidial germination of P. solitum occurred at low temperatures and high salinities. In addition, P. solitum displayed extracellular amylasic and esterasic activities. The isolation of P. solitum from marine sediments in Antarctica and its survival at low temperatures and high salt concentrations suggest that it is adapted to the cold and halophilic environment of the Antarctic oceans. Because P. solitum produces extracellular enzymes, it is an interesting eukaryotic model for the study of structure–function relationships during enzymatic biocatalysis and biotransformation under extreme conditions. Marine sediments from Antarctica may represent a unique source for obtaining extremophilic fungi. New studies using different culture media, temperatures ranges and pressure conditions as well as metagenomic techniques can assist in understanding the extremophilic fungal communities in marine sediments across the Antarctic Ocean.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Marine fungi are an ecologically important group of microorganisms that play key roles in the energy flow of oceans (Kohlmeyer and Kohlmeyer 1979; Hyde and Lee 1995). Fungi in marine environments are present on a variety of substrata such as decaying wood, leaves, macroalgae and seagrasses (Jones 2011). However, according to Bhadury et al. (2011), the diversity and functions of fungi in marine systems have been underestimated. Recent studies have suggested that marine fungi are saprobes, pathogens or symbionts forming an integral part of coastal and deep-sea environments (Hyde et al. 1998; Bass et al. 2007) that are present in open waters (pelagic) and in bottom (benthic) zones. Few studies have addressed the occurrence of fungi in marine sediments (Raghukumar and Raghukumar 1998; Edgcomb et al. 2011; Orsi et al. 2013); fungi thrive in habitats characterised by extreme conditions, including high hydrostatic pressures and low temperatures (Snelgrove et al. 1997).
Antarctica is one of the harshest, most pristine and most unique ecosystems in the world. During the last few decades, the archeal, bacterial, microalgal and fungal diversity in different Antarctic habitats has been studied. Mycological studies in Antarctica have included species living in lakes (Ellis-Evans 1985; Brunati et al. 2009; Gonçalves et al. 2012), soil, historic woodlands (Arenz et al. 2006; Fell et al. 2006) and on macroalgae (Loque et al. 2010) and plants (Bradner et al. 2000; Tosi et al. 2002; Rosa et al. 2009; Uspon et al. 2009). Only one uncultured, unidentified fungal taxon has been detected in Antarctic marine plankton via metagenomic sequencing (Lopez-Garcia et al. 2001).
Taken as a whole, marine sediments from Antarctica represent special habitats that are suitable for the exploration of fungal life under extreme conditions. The aims of the present study were to isolate and culture fungal species from sediments at various depths in the Antarctic Ocean and to classify these species using morphological, physiological and molecular methods.
Materials and methods
Study area and source of fungi
The samples were collected at different sites in the Bransfield Strait of Admiralty Bay, King George Island, Antarctica (Fig. 1). All sites were sampled during the austral summer season in February 2010 from on-board the Brazilian Navy Polar Ship Ary Rongel (H-44). Twelve sediment samples were obtained from depths of 100 (three samples), 500 (three samples), 700 (three samples) and 1,100 m (three samples) using a 50-cm3 USNEL-type box corer. Additionally, measures against contamination were maintained throughout the sample collection and microbial isolation phases, except for the air control on the navy deck and in the laboratory. However, as soon as the core had been brought on-board, samples were collected by inserting in sterile whirl-pack bags into the lower end of the core, into which 10-cm-long sediment samples were extruded to avoid aerial contamination. The bags were closed with rubber bands and transported within 24 h to the laminar flow hood in the laboratory, where a portion of the sediment from the centre of each subsample that had not been in contact with the walls of the core pipe was removed using an alcohol-/flame-sterilised spatula and placed in sterile containers for isolation of the fungi.
Maps showing (a) Antarctica, (b ) Antarctic Peninsula, (c) King George Island and (d) Admiralty Bay at King George Island. The positions of the sediment collection sites in Admiralty Bay are shown in 1 (62°05′6607″S, 58°21′2300″W), 2 (62°05′9003″S, 58°25′7822″W) and 3 (62°06′3411″S, 58°26′7762″W) at 100 m depth; 4 (62°09′8731″S, 58°23′7249″W), 5 (62°11′0334″S, 58°22′7174″W) and 6 (62°11′4137″S, 58°19′7172″W) at 500 m depth; 7 (62°16′2400″S, 58°21′7500″W), 8 (62°15′4962″S, 58°17′2124″W) and 9 (62°15′7010″S, 58°12′5956″W) at 700 m; 10 (62°17′4406″S, 58°19′8244″W), 11 (62°16′8044″S, 58°16′1651′W) and 12 (62°16′6460″S, 58°11′4731″W) at 1,100 m depth
Fungal isolation
To isolate fungi from different sediment samples, 1 g of each sample was re-suspended in 9 ml of Hanks’ buffer (Sigma, USA). Each suspension was diluted to 10−2, and 100 μl of each dilution was plated on Marine Agar (MA-Difco/USA), supplemented with 2 % glucose and 200 mg l−1 of chloramphenicol (Sigma, USA) and incubated at 15 °C for 30 days. All sediment samples were processed using a flame-sterilised spatula in a laminar flow hood to avoid environmental contamination. The fungal colonies present on each plate were counted, purified, maintained on marine agar slants and preserved at −80 °C for later identification. The fungal isolates were deposited at the Collection of Microorganisms and Cells of the Universidade Federal of Minas Gerais under code UFMGCB.
Molecular taxonomy
Fungal DNA extraction was performed as described by Rosa et al. (2009). The dried DNA was re-suspended in 100 μl of TE buffer (10 mM Tris, 10 mM Na-EDTA, pH 8.0) and 2.5 μl of RNase solution. The latter was prepared by adding 10 mg of pancreatic RNase (20 U mg−1) to 1 ml of Na acetate (0.01 M), heating at 100 °C for 15 min, cooling the solution slowly to room temperature and adjusting the pH to 7.4 with 100 μl of Tris-HCl. The sample was incubated at 37 °C for 30 min and then stored at −20 °C until use. The internal transcribed spacer region (ITS1-5.8S-ITS2) of the rRNA gene was amplified with the universal primers ITS1 and ITS4, as described by White et al. (1990). Amplification of the ITS region was performed as follows: 95 °C for 5 min, 40 cycles of 94 °C for 60 s, 50 °C for 60 s, 72 °C for 60 s, and a final extension at 72 °C for 5 min.
Amplification of the β-tubulin gene was performed with the Bt2a and Bt2b primers (Glass and Donaldson 1995). PCR assays were conducted in 50 μl reaction mixtures containing 1 μl of genomic DNA (10 ng μl−1), 5 μl of PCR buffer (100 mM Tris-HCl, 500 mM KCl, pH 8.8), 2 μl of dNTPs (10 mM) plus 3 μl of MgCl2 (25 mM), 1 μl of each primer (50 pmol μl−1), 1 μl of dimethyl sulphoxide (DMSO, Merck, USA), 2 μl of betaine (5 M), 0.2 μl of Taq polymerase (5 U μl−1 DNA) and 33.8 μl of ultrapure sterile water. PCR amplifications were performed with a Mastercycler pro (Eppendorf, Hamburg, Germany) that was programmed for an initial denaturation at 94° C for 5 min, followed by 35 cycles of 1 min of denaturation at 94 °C, primer annealing for 1 min at 59 °C and extension for 1.30 min at 72 °C, with a final 7 min elongation step at 72 °C. After amplification of the β-tubulin template, excess primers and dNTPs were removed from the reaction mixture using a commercial GFX column with the PCR DNA Purification kit (Amersham Bioscience, Roosendaal, Netherlands). Purified PCR fragments were re-suspended in 50 μl of TE buffer.
The amplified DNA was concentrated with a Wizard Plus SV Miniprep DNA Purification System (Promega, USA) and was sequenced with a ABI 3730xl DNA sequencer (Applied Biosystem®, USA). The sequences were analysed using SeqMan Π and Lasergene software (DNASTAR/Inc.), and consensus sequences were obtained with Bioedit v. 7.0.5.3 software. To make species identifications, the consensus sequences were aligned with all sequences of related species retrieved from the GenBank database using the Fasta 2.0 program (Altschul et al. 1997). The consensus sequences were deposited into GenBank under the accession numbers HM746658 (ITS1-5.8S-ITS2) and KC408929 (β-tubulin). The fungal sequences were used to determine phylogenetic relationships with the nearest species relatives, which were estimated with MEGA Version 5.0 (Tamura et al. 2011). The maximum composite likelihood model was used to estimate evolutionary distance with bootstrap values calculated from 1,000 replicate runs. The GenBank sequences of the fungi most closely related to the Antarctic marine fungus were used in the initial molecular analysis (Table 1). To complete the molecular identification, sequences from known types or reference fungal sequences in the worldwide culture collections were added to improve the accuracy of the phylogenetic analysis. The fungal taxonomy assigned follows the guidelines established by Kirk et al. (2011), the MycoBank database (www.mycobank.org) and the Index Fungorum database (www.indexfungorum.org).
Physiological and morphological taxonomy
Fungal macroscopic parameters (colony colour and texture, reverse colour, border type and radial growth rate) and colony diameters were observed on various media, including Czapek yeast extract agar (CYA), malt extract agar (MEA) and yeast extract sucrose agar (YES). Fungal isolates were inoculated into 3-point cultures and incubated for 7 days in the dark at 25 °C. A duplicate set of MEA plates was incubated in the dark at 25 °C to determine the microscopic parameters (hyphae, conidiophores and conidia) using slide cultures mounted in methyl blue in polyvinyl lactophenol.
For scanning electron microscopy (SEM), the MA plates were inoculated with a 5-mm mycelial disc from fresh fungal cultures. The cultures were incubated at 15 °C, and after 7 days, agar discs with fungi were examined by SEM. Specimens were fixed in 2 % glutaraldehyde in 0.1 M NaPO4 buffer and washed in buffered 1 % OsO4 for 2 h. The material was dehydrated using an ethanol series (10, 25, 40, 60, 75, 85, 95 and 100 %) for 15 min per concentration. The material was dried in a critical point drying apparatus, sputter-coated with gold and viewed with a field emission SEM (Leo 982, Zeiss + Leica).
Fungal growth responses to temperature and salinity
The fungi were grown on MA at 0, 10, 15, 20, 25 and 37 °C and on MEA with 1, 5, 10 and 15 % NaCl at 15 °C. The fungi were inoculated onto each medium by the transfer of blocks (4 mm2) from 10-day-old pre-cultures grown at 15 °C on MA. Plates were incubated in triplicate for seven days, and the colony diameter was measured in mm.
Extracellular enzymatic activity
Fungal isolates were screened for their ability to degrade starch, protein (casein), lipids (tributyrin and Tween-80), pectin and cellulose according to procedures described by Brizzio et al. (2007). A spot of spores was grown for 72 h on MA and then inoculated onto agar plates. Plates containing each substrate were incubated at 5, 10, 15, 20, 25 and 30 °C. Enzymatic activity was analysed after 10 days. The halo/colony (h/c) ratio was used as a semi-quantitative assessment of extracellular enzymatic activity. Sporobolomyces ruberrimus CRUB1141 (amylase), Leucosporidiella fragaria CRUB1211 (protease), Rhodotorula mucilaginosa CRUB138 (lipase and pectinase) and Trichosporon mycotoxinivorans UFMG-HB20 were used as positive controls for enzymatic activities. Data for both temperatures were statistically compared using Student’s t test.
Results and discussion
A total of 12 sediment samples were collected and analysed for fungal presence and colony forming units (CFUs). A total of 52 fungal isolates were obtained (13 isolates at 100 m, 19 at 500 m, 2 at 700 m and 18 at 1,100 m depth) using a selective marine agar, which exhibited the same macro-(colony colour and texture, reverse colour, border type and radial growth rate) and micromorphology (hyphae, conidiophores and conidia). One isolate per depth was selected for identification and physiological analysis. The fungal CFU g−1 densities were 1.3 × 104 at 100 m, 1.2 × 104 at 500 m, 0.2 × 104 at 700 m and 1.1 × 104 at 1,100 m depth.
Based on sequencing of the ITS regions and BLASTn comparisons with GenBank, the Antarctic marine sediment fungus displayed sequence similarities (97–99 %) with Penicillium species from a wide variety of habitats and substrates, including cave soil (Mexico), a stone chamber (Japan), mangrove soil (China), the thallus of red algae (Mediterranean Sea), wastewater sludge (Canada, China), wood (Antarctica) and soil (Antarctica) (Table 1). In addition, the marine Antarctic fungus shared 97 % sequence similarity with Penicillium mali (AM158219) (a synonymy of P. solitum) that was isolated from deep-sea sediment in the Pacific Ocean near China. Because the Antarctic marine fungus exhibited high sequence similarity to other Penicillium taxa living in a wide variety of habitats, especially those in deep-sea sediments, there may be a cosmopolitan distribution of or phylogenetic relationships among species present in extremophile environments.
The ITS sequence from the Antarctic marine Penicillium was compared with sequences from the species deposited in the GenBank database (Fig. 2a). However, the ITS region from terverticillate Penicillia alone cannot differentiate Penicillium species (Skouboe et al. 2000). For this reason, we also analysed the β-tubulin gene (Seifert and Louis-Seize 2000) for classifying the marine Antarctic Penicillium (Fig. 2b). In this analysis, the Antarctic Penicillium was 100 % identical to the species P. solitum (AF000934), but it was 98.85, 98 and 97.4 % similar to P. discolor (AY674349), P. chinulatum (AF003536) and P. cavernicola (AY674337), respectively.
To confirm the molecular taxonomy, we performed classic physiological and morphological analyses that revealed the following characteristics of the Antarctic marine Penicillium: colony diameters of 19.7, 24.25 and 33.92 mm after 7 days at 25 °C on CYA, MEA and YES, respectively; the absence of growth at 37 °C; the absence of synnemata or fasciculation and sclerotia; the presence of clear to light yellow exudate droplets on CYA; a velutinous colony texture that was green with a cream-coloured reverse and a green conidium on CYA; and an orange-yellow reverse on YES (Fig. 3f). After 7 days at 25 °C on MEA, the Antarctic marine Penicillium showed terverticillate conidiophores, appressed elements, subsurface hyphae, slightly rough-walled and globose conidia (4 μm), cylindrical phialides tapering to a distinct collulum (10 × 2.5 μm), cylindrical metulae (13 × 3 μm), cylindrical rami (15 × 3.5 μm) and rough-walled stipes (200 × 4 μm) (Fig. 3g–m). All of the physiological and morphological characteristics displayed by the marine Antarctic Penicillium were identical to those of P. solitum Westling (Ascomycota).
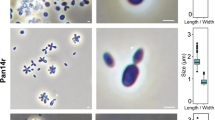
Morphology of the marine Antarctic Penicillium species. Seven-day-old colonies on a verse and b reverse CYA; c verse and d reverse MEA; and e verse and f reverse YES at 25 ± 2 °C. Optical micrographs of conidiophores (g, h) and conidia (i). Scanning electron micrographs of conidiophores (j, k), conidiogenic cells (l) and conidia (m)
Penicillium solitum is very closely related to P. commune and P. palitans, which cannot easily be differentiated by the morphology of their conidiogenous structures (Lund 1995). However, when P. solitum was grown on YES agar, it had a unique yellowish orange colour (Sùrensen et al. 1999) (Fig. 3f). Additionally, P. solitum had smooth to slightly rough-walled conidia (Fig. 3m), which contrast with the conspicuously dark green and rough-walled conidia of P. cavernicola, P. echinulatum and P. discolor in the series Solita (www.mycobank.org).
According to Raghukumar et al. (2010), the majority of fungi recovered from marine sediments are closely related to terrestrial species. Fungi from all known phyla have been isolated from marine sediments, including species of the genera Aspergillus, Candida, Cryptococcus, Cylindrocarpon, Glomerella, Golovinomyces, Hortaea, Lodderomyces, Malassezia, Metshnikowia, Nematoctonus, Penicillium, Phoma, Pichia, Rhodotorula, Rhizophlyctis and Trichosporon (Lai et al. 2007; Calvez et al. 2009; Nagano et al. 2010; Singh et al. 2011). Among the non-melanised fungi, organisms in the Penicillium genus have been isolated from alpine, tundra and permafrost soils (Gunde-Cimerman et al. 2003) and marine sediments (Nagano et al. 2010; Singh et al. 2010). Biddle et al. (2005) recovered Penicillium species from sediment collected 200 m below the sea floor in Peru and Takami (1999) isolated P. lagena from a depth of 10,500 m in the Mariana Trench in the Pacific Ocean. Damare et al. (2006) and Singh et al. (2010) found that Penicillium sp. were the dominant fungi in deep-sea sediments of the Central Indian Basin.
In Antarctica, fungi distribution is correlated with the presence of birds, invertebrates, humans and plants (Tosi et al. 2002). According to Moller and Dryfuss (1996), most of the known Antarctic fungi are cosmopolitan cold-tolerant species that have been selected by, or have adapted to, the prevailing low temperatures. In the deep-sea environment, fungi are some of the few eukaryotes in the aphotic zone between 250 and 3,000 m below the Antarctic polar front (Lopez-Garcia et al. 2001). In Antarctica, Penicillium species have been identified in soils (Azmi and Seppelt 1998), lakes (Ellis-Evans 1996), historic woodlands (Arenz et al. 2006) and macroalgal thalli (Loque et al. 2010). P. solitum is an extremophile that has been isolated from Arctic subglacial ice (Sonjak et al. 2006); however, our study is the first report of P. solitum in Antarctic marine sediments.
According to Raghukumar et al. (2010), the fungi present in marine sediments demonstrate mesophilic, psychrotolerant and/or halotolerant physiologies. P. solitum was tested for growth at different temperatures and salinities (Fig. 4a, b). P. solitum displayed an interesting temperature plasticity, mesophilic behaviour and psychrotolerant adaptability at 5, 10, 15, 20, 25 and 30 °C, but it did not grow at 37 °C. P. solitum also demonstrated halotolerance at 1, 5, 10 and 15 % NaCl.
Microbial proliferation at cold temperatures is predicated on the capacity to synthesise cold-adapted enzymes (Kuddus et al. 2011). Extremophilic microorganisms are able to produce specific enzymes that facilitate the degradation of polymeric substrates such as cellulose, xylan, pectin, chitin and starch (Kristjansson and Hreggvidsson 1995). While the role of fungi in marine sediments remains unknown, some species produce extracellular enzymes, such as alkaline phosphatase, protease and polygalacturonases that transform organic matter (Raghukumar et al. 2010). P. solitum isolates (one from each depth) obtained from the marine sediments of Antarctica exhibited amylolytic and esterasic activities. Amylase activity was detected at 10, 15, 25 and 30 °C after 10 days, and esterase activity occurred at 10, 15 and 25 °C (Table 2). No xylanasic, proleolytic or cellulolytic activities were detected.
According to Kuddus et al. (2011), cold amylases reduce energy consumption because they have low activation energies and demonstrate activity at low temperatures. Amylases can also depolymerise complex starch or glycogen structures from algae, animal cells or microorganisms into oligosaccharides and smaller sugars such as glucose and maltose (Bertoldo and Antranikan 2002). In addition, esterases catalyse the hydrolysis of short-chain triacylglycerols, and their wide substrate specificity allows them to process various carbon sources or to be involved in catabolic pathways (Aurilia et al. 2008). The amylase and esterase activities of P. solitum might allow this organism to use the scarce available nutrients in marine sediments, thus increasing its survival under extreme conditions. Additionally, P. solitum amylase and esterase enzymes could be promising tools for pharmaceutical, cosmetic or chemical applications at low temperatures. However, further enzymatic studies using quantitative methods will be necessary to determine the specific amylolytic and esterasic activities of P. solitum.
We identified P. solium in Antarctic marine sediments, which are a habitat previously unknown for this organism. P. solitum grew at low temperatures and high salinity, and it degraded polymeric substrates, suggesting that P. solitum has metabolic adaptations that allow it to survive in this cold and halophytic environment. Thus, P. solitum is an interesting eukaryotic model for the study of structure–function relationships during enzymatic biocatalysis and biotransformation reactions under extreme conditions. Our study was based on cultivable fungi present in marine sediments collected in the Antarctic Ocean, where we found just one fungal species. However, marine sediments from Antarctica may represent a unique source for obtaining extremophile fungi. For perspective, new studies using different culture media, ranges of temperatures and pressure conditions as well as metagenomic techniques might assist in understanding the extremophile fungal communities in marine sediments across the Antarctic region.
References
Altschul SF, Madden TL, Schaffer AA, Zhang JH, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
Arenz BE, Held BW, Jurgens JA, Farrell RL, Blanchette RA (2006) Fungal diversity in soils and historic wood from the Ross Sea Region of Antarctica. Soil Biol Biochem 38:3057–3064
Aurilia V, Parracino A, D’Auria S (2008) Microbial carbohydrate esterase in cold adapted environments. Gene 410:234–240
Azmi OR, Seppelt RD (1998) The broad-scale distribution of microfungi in the Windmill Islands region, continental Antarctica. Polar Biol 19:92–100
Bass D, Howie A, Brown N, Barton N, Demidoba M et al (2007) Yeast forms dominate fungal diversity in the deep oceans. Proc R Soc B 274:3069–3077
Bertoldo C, Antranikan G (2002) Starch-hydrolyzing enzymes from thermophilic archaea and bacteria. Curr Opin Chem Biol 6:151–160
Bhadury P, Bik H, Lambshead JD, Austen MC, Smerdon GR, Rogers AD (2011) Molecular diversity of fungal phylotypes co-amplified alongside nematodes from coastal and deep-sea marine environments. PLoS One 6:e264451–e264457
Biddle JF, House CH, Brenchley JE (2005) Microbial stratification in deeply buried marine sediment reflects change in sulfate/methane profiles. Geobiology 3:287–295
Bradner JR, Sidhu RK, Yee B, Skotnicki ML, Selkirk PM, Nevalainen KMH (2000) A new microfungal isolate, Embellisia sp., associated with the Antarctic moss Bryum argenteum. Polar Biol 23:730–732
Brizzio S, Turchetti B, de García V, Libkind D, Buzzini P, van Broock M (2007) Extracellular enzymatic activities of basidiomycetous yeasts isolated from glacial and subglacial waters of northwest Patagonia (Argentina). Can J Microbiol 53:519–525
Brunati M, Rojas JL, Sponga F, Ciciliato I, Losi D, Göttlich E, de Hoog S, Genilloud O, Marinelli F (2009) Diversity and pharmaceutical screening of fungi from benthic mats of Antarctic lakes. Mar Genomics 2:43–50
Calvez TL, Burgaud G, Mahé S, Barbier G, Vandenkoornhuyse P (2009) Fungal diversity in deep-sea hydrothermal ecosystems. Appl Environ Microbiol 75:6415–6421
Damare S, Raghukumar C, Raghukumar S (2006) Fungi in deep-sea sediments of the Central Indian Basin. Deep-Sea Res 53:14–27
Edgcomb VP, Beaudoin D, Gast R, Biddle JF, Teske A (2011) Marine subsurface eukaryotes: the fungal majority. Environ Microbiol 13:172–183
Ellis-Evans JC (1985) Fungi from maritime Antarctic freshwater environments. Br Antarct Surv Bull 68:37–45
Ellis-Evans JC (1996) Microbial diversity and function in Antarctic freshwater ecosystems. Biodivers Conserv 5:1395–1431
Fell JW, Scorzetti G, Connell L, Craig S (2006) Biodiversity of micro-eukaryotes in Antarctic Dry Valley soils with < 5% soil moisture. Soil Biol Biochem 38:3107–3119
Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl Environ Microbiol 61:1323–1330
Gonçalves VN, Vaz ABM, Rosa CA, Rosa LH (2012) Diversity and distribution of fungal communities in lakes of Antarctica. FEMS Microbiol Ecol 82:459–471
Gunde-Cimerman N, Sonjak S, Zalar P, Frisvad JC, Diderichsen B, Plemenita A (2003) Extremophilic fungi in arctic ice: a relationship between adaptation to low temperature and water activity. Phys Chem Earth 28:1273–1278
Hyde KD, Lee SY (1995) Ecology of mangrove fungi and their role in nutrient cycling: what gaps occur in our knowledge? Hydrobiologia 295:107–118
Hyde KD, Jones EBG, Leaño E, Pointing SB, Poonyth AD, Vrijmoed LLP (1998) Role of fungi in marine ecosystems. Biodivers Conserv 7:1147–1161
Jones EBG (2011) Fifty years of marine mycology. Fungal Div 50:73–112
Kirk PM, Cannon PF, David JC, Stalpers JA (2011) Ainsworth and Bisby’s Dictionary of the Fungi, 19th edn. CAB International, Wallingford
Kohlmeyer J, Kohlmeyer E (1979) Marine mycology, the higher fungi, Academic Press
Kristjansson JK, Hreggvidsson GO (1995) Ecology and habitats of extremophiles. World J Microbiol Biotechnol 11:17–25
Kuddus M, Arif JM, Ramteke PW (2011) An overview of cold-active microbial α–amylase: adaptation strategies and biotechnological potentials. Biotechnology 3:246–258
Lai X, Cao L, Tan H, Fang S, Huang Y, Zhou S (2007) Fungal communities from methane hydrate-bearing deep-sea marine sediments in South China Sea. ISME J 1:756–762
Lopez-Garcia P, Rodriguez-Valera F, Pedros-Allo C, Moreira D (2001) Unexpected diversity of small eukaryotes in deep-sea Antarctic plankton. Nature 409:603–607
Loque CP, Medeiros AO, Pellizzari FM, Oliveira EC, Rosa CA, Rosa LH (2010) Fungal community associated with marine macroalgae from Antarctica. Polar Biol 33:641–648
Lund F (1995) Diagnostic characterization of Penicillium palitans, P. commune and P. solitum. Lett Appl Microbiol 20:228–231
Moller C, Dreyfuss MM (1996) Microfungi from Antarctic lichens, mosses and vascular plants. Mycology 88:922–933
Nagano Y, Nagahama T, Hatada Y, Nunoura T, Takami H, Miyazaki J, Takai K, Horikoshi K (2010) Fungal diversity in deep-sea sediments - the presence of novel fungal groups. Fungal Ecol 3:316–325
Orsi W, Biddle JF, Edgcomb V (2013) Deep sequencing of subseafloor eukaryotic rRNA reveals active Fungi across marine subsurface provinces. PLoS ONE 2:e56335
Raghukumar C, Raghukumar S (1998) Barotolerance of fungi isolated from deep-sea sediments of the Indian Ocean. Aquat Microbial Ecol 15:153–163
Raghukumar C, Damare SR, Singh P (2010) A review on deep-sea fungi occurrence, diversity and adaptations. Bota Mar 53:479–492
Rosa LH, Vaz ABM, Caligiorne RB, Campolina S, Rosa CA (2009) Endophytic fungi associated with the Antarctic Grass Deschampsia antarctica Desv. (Poaceae). Polar Biol 32:161–167
Seifert KA, Louis-Seize G (2000) Phylogeny and species concepts in the Penicillium aurantiogriseum complex as inferred from partial β-tubulin gene DNA sequences. In: Samson RA, Pitt JI (eds) Integration of Modern Taxonomic Methods for Penicillium and Aspergillus Classification. Harwood Academic Publishers, Netherlands, pp 189–198
Singh P, Raghukumar C, Verma P, Shouche Y (2010) Phylogenetic diversity of culturable fungi from the deep-sea sediments of the Central Indian Basin and their growth characteristics. Fungal Div 40:89–102
Singh P, Raghukumar C, Verma P, Shouche Y (2011) Fungal community analysis in the deep-sea sediments of the central indian basin by culture-independent. Approach Microbiol Ecol 61:507–517
Skouboe P, Taylor JW, Frisvad JC, Lauritsen D, Larsen L, Albaek C, Boysen M, Rossen L (2000) Molecular methods for differentiation of closely related Penicillium species. In: Samson RA, Pitt JI (eds) Integration of modern taxonomic methods for Penicillium and Aspergillus classification. Harwood Academic Publishers, Netherlands, pp 179–188
Snelgrove PVR, Blackburn TH, Hutchings P, Alongi D, Grassle JF, Hummel H, King G, Koike I, Lambshead PJD, Ramsing NB, Solis-Weiss V, Freckman DW (1997) The importance of marine sediment biodiversity in ecosystem processes. Ambio 26:578–582
Sonjak S, Frisvad JC, Gunde-Cimerman N (2006) Penicillium mycobiota in Arctic subglacial ice. Microbiol Ecol 52:207–216
Sùrensen D, Larsen TO, Christophersen C, Nielsen PH, Anthoni U (1999) Solistatin, an aromatic compactin analogue from Penicillium solitum. Phytochemistry 51:1027–1029
Takami H (1999) Isolation and characterization of microorganisms from deep-sea mud. In: Horikoshi K, Tsujii K (eds) Extremophiles in deep-sea environments. Springer, Tokyo, pp 3–26
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tosi S, Casado B, Gerdol R, Caretta G (2002) Fungi isolated from Antarctic mosses. Polar Biol 25:262–268
Uspon R, Newsham KK, Bridge PD, Pearce DA, Read DJ (2009) Taxonomic affinities of dark septate root endophytes of Colobanthus quitensis and Deschampsia antarctica, the two native Antarctic vascular plant species. Fungal Ecol 2:184–196
White TJ, Bruns TD, Lee SB, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Gelfand D, Sninsky J (eds) Innis NA. A guide to methods and applications. Academic Press San Diego, PCR protocols, pp 315–322
Acknowledgments
This study had financial and logistic support from the Brazilian Antarctic Program, Marine of Brazil. This work is part of the API activity 403, contributes to Microbiological and Ecological Responses to Global Environmental Changes in Polar Regions, and INCT Criosfera. This work was financially supported by the Fundação de Amparo a Pesquisa do Estado de Minas Gerais (FAPEMIG) and Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). We thank the anonymous reviewers for their comments, which help the quality of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Gonçalves, V.N., Campos, L.S., Melo, I.S. et al. Penicillium solitum: a mesophilic, psychrotolerant fungus present in marine sediments from Antarctica. Polar Biol 36, 1823–1831 (2013). https://doi.org/10.1007/s00300-013-1403-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-013-1403-8