Abstract.
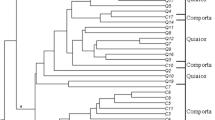
Eleven elite and popular Indian cultivars of Cymbopogon aromatic grasses of essential oil trade types – citronella, palmarosa and lemongrass – were characterized by means of RAPDs to discern the extent of diversity at the DNA level between and within the oil biotypes. Primary allelic variability and the genetic bases of the cultivated germplasm were computed through parameters of gene diversity, expected heterozygosity, allele number per locus, SENA and Shannon's information indices. The allelic diversity was found to be in the order: lemongrass > palmarosa > citronella. Lemongrasses displayed higher (1.89) allelic variability per locus than palmarosa (1.63) and citronella (1.40). Also, RAPDs of diagnostic and curatorial importance were discerned as 'stand along' molecular descriptors. Principal component analysis (PCA) resolved the cultivars into four clusters: one each of citronella and palmarosa, and two of lemongrasses (one of C. flexuosus and another of C. pendulus and its hybrid with C. khasianus). Proximity of the two species-groups of lemongrasses was also revealed as they shared the same dimension in the three-dimensional PCA. The molecular distinctions are discussed in relation to oil-chemotypic variations.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Author information
Authors and Affiliations
Additional information
Revision received: 16 March 2000
Electronic Publication
Rights and permissions
About this article
Cite this article
Sangwan, N., Yadav, U. & Sangwan, R. Molecular analysis of genetic diversity in elite Indian cultivars of essential oil trade types of aromatic grasses (Cymbopogon species). Plant Cell Rep 20, 437–444 (2001). https://doi.org/10.1007/s002990100324
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s002990100324