Abstract
Plants strive for phosphorus (P), which is an essential mineral for their life. Since P availability is limiting in most of the world’s soils, plants have evolved with a complex network of genes and their regulatory mechanisms to cope with soil P deficiency. Among them, purple acid phosphatases (PAPs) are predominantly associated with P remobilization within the plant and acquisition from the soil by hydrolyzing organic P compounds. P in such compounds remains otherwise unavailable to plants for assimilation. PAPs are ubiquitous in plants, and similar enzymes exist in bacteria, fungi, mammals, and unicellular eukaryotes, but having some differences in their catalytic center. In the recent past, PAPs’ roles have been extended to multiple plant processes like flowering, seed development, senescence, carbon metabolism, response to biotic and abiotic stresses, signaling, and root development. While new functions have been assigned to PAPs, the underlying mechanisms remained understood poorly. Here, we review the known functions of PAPs, the regulatory mechanisms, and their relevance in crop improvement for P-use-efficiency. We then discuss the mechanisms behind their functions and propose areas worthy of future research. Finally, we argue that PAPs could be a potential target for improving P utilization in crops. In turn, this is essential for sustainable agriculture.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
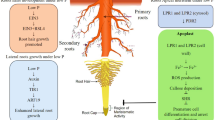
Phosphorus (P) is a crucial macronutrient and essential for life. Being the key constituent of nucleic acids, ATP, phospholipids, etc., P plays a critical role in cell structure and functions, including energy metabolism, metabolic pathways, and signal transduction (Raghothama 2005). Plants can absorb and assimilate only soluble inorganic phosphate (Pi; H2PO4−) which remains limited in the natural environment due to its reactive nature as it forms insoluble complexes with cationic minerals (Fe or Al in acidic soils; Ca or Mg in alkaline soils) (Oelkers and Valsami-Jones 2008; Arai and Sparks 2007). The P-rich biomolecules in decaying biological matter add organic phosphate (Po) to the soils. Po remains unavailable for root absorption unless hydrolyzed by enzymes to release Pi, which is acquired by Pi transporters (PTs) present in the plasma membrane of root epidermal cells (Dissanayanka et al. 2018). The availability of Pi in the soils remains at a suboptimal level (1–10 µM) which is many magnitudes lower than the Pi concentration in a plant cell (5–20 mM) (Shen et al. 2011). Hence, Pi uptake against the steep concentration gradients occurs at the expense of plant ATPs (Srivastava et al. 2018). Plants initiate an array of adaptive responses to combat Pi deficiency, broadly known as phosphate starvation response (PSR). These include (i) morphological alterations (root architecture, root to shoot ratio, and symbiotic associations with mycorrhizae), (ii) molecular adaptations (activation of Pi starvation-induced genes and associated regulators), (iii) physiochemical adaptations (enhanced internal Pi recycling, anthocyanin accumulation, and alterations in root exudates) (Fig. 1). The coordinated PSR results in increased Pi acquisition via roots and optimization in plant internal Pi utilization. The hydrolases (such as phospholipases, ribonucleases, acid phosphatases, etc.) and organic acids (citrate, malate, oxalate, etc.) in root exudates help liberate Pi from P complexes in the rhizosphere for roots acquisition (Raghothama 1999). The Pi uptake and transport are tightly regulated via high-affinity transporters present in the root epidermal cells and distributed to organelles/tissues by a specific Pi transporter networks (Poirier and Jung 2015; Gu et al. 2016). These coordinated actions ensure the optimum Pi uptake and utilization in plants.
Schematic depiction of the different phosphate starvation responses and functions of APases. Under Pi deprived conditions, plants adaptations include morphological, molecular, and physiochemical changes. The morphological (such as root system architecture) and physiochemical (organic acid and APases secretion) alterations result in enhanced availability of Pi in soil. The molecular activities drive the acquisition and translocation of Pi in the above-ground parts of plants
Acid phosphatases (APases) are a family of hydrolases ubiquitously present in living organisms and hydrolyze orthophosphate monoesters (Po) to release Pi at acidic pH (Fig. 2) (Olczak et al. 2003). The increase in APases expression and activity is a characteristic response of Pi deprived plants (Tran et al. 2010a; Zhang et al. 2014a; Secco et al. 2013; Mehra et al. 2016). Purple acid phosphatases (PAPs), a distinct group of APases, are widely studied for their roles in plant acclimation to Pi deficiency. PAPs are broadly involved in liberating Pi from phosphomonoesters in soil and senescing tissues, thus help in Pi uptake and distribution in the plant (Olczak et al. 2003; Tran et al. 2010a; López-Arredondo et al. 2014; Scheible and Rojas-Triana 2015; Gu et al. 2016). Besides, a few PAPs are also known to exhibit peroxidase activity in addition to the phosphatase activity (Klabunde et al. 1995; Del Pozo et al. 1999; Bozzo et al. 2002). Recent reports highlight their unanticipated roles in multiple stress responses and plant development (flowering, seed germination, etc.). This review aims to explore the recent advances made on the understanding PAPs’ role in Pi utilization and new emerging roles. We also discuss PAPs’ role in Pi signaling pathways, their regulation and utilization in crop improvement for Pi-use efficiency.

adapted from Schenk et al., 2008). B Schematic illustrations showing domain architecture/motifs present in PAPs. (i) The amino acid sequence of PAPs contain metallophos domain across the species and five conserved motifs with seven invariant amino acids. (ii) Few PAPs have FN3 domain (fibronectin domain) at N-terminal along with metallophos domain and might have a role in substrate selectivity. (iii) PAPs with phytase activity have REKA (Arg, Glu, Lys, Met, Ala) motif at C-terminal. C PAPs hydrolyze phosphate monoesters in the presence of a water molecule to release alcohol and inorganic phosphate
Schematic representation of active site, domain architecture and reaction mechanism of PAPs. A The active site of red kidney bean purple acid phosphatase (rkbPAP), a representative binuclear metallohydrolase with Zn and Fe ions (
Purple acid phosphatases (PAPs): ubiquitous family of APases
PAPs belong to the metallophosphatase superfamily proteins carrying a metallophos domain and bimetallic reaction center at their active site (Fig. 2) (Schenk et al. 2013). These enzymes are broad substrate-specific, function at pH 4–6, and temperature range of 25–60 °C (Tian and Liao 2015). PAPs give pink or purple color in an aqueous solution due to the electron transfer reaction from tyrosine to Fe(III) at 560 nm (Olczak et al. 2003). The electron transfer at catalytic center results in the inactivation of the enzyme (purple color), and the redox reaction (Fe3+–Fe2+) changes it to an active form (pink color) (Olczak et al. 2003). PAPs are also known as TRAPs (Tartrate-Resistant Acid Phosphatases) due to their resistance to tartrate-mediated inhibition of catalytic activity. The term “TRAPs” is mostly used for mammalian PAPs which are functional in different mammalian tissues such as the bovine spleen, hairy leukaemia cells, porcine uterine fluid, bone, lungs, and placenta (Sundararajan and Sarma 1954; Chen et al. 1975; Efstratiadis and Moss 1985; Ketcham et al. 1985, 1989). Their functions include bone resorption, antigen presentation, generation of ROS, and iron transport (Hayman et al. 1996; Halleen et al. 2003). Other than mammals, PAPs have been identified in plants, bacteria, and fungi. Different Aspergillus species have sequences closely related to plant PAPs and one such PAP (85 kDa) from Aspergillus ficuum has been characterized (Ullah and Cummins 1988; Schenk et al. 2000a). A few bacteria also have genes encoding PAPs, i.e., cyanobacterium Synechocystis sp., Mycobacterium tuberculosis, Mycobacterium leprae, Burkholderia cenocepacia, and Burkholderia pyrrocinia (Schenk et al. 2000a; Yeung et al. 2009; Zhu et al. 2019). Bioinformatic analysis also revealed 57 PAP-like sequences in 43 bacterial genomes and 4 in cyanobacteria (Yeung et al. 2009; Zhu et al. 2019). Surprisingly, a phospholipase D from Streptomyces chromofuscus (scPLD) also showed structural and mechanistic similarities to iron-dependent PAPs (Zambonelli and Roberts 2003). Although, PAPs are reported in a limited number of microorganisms; however, structural similarities suggest their catalytic roles similar to plants’ PAPs (Schenk et al. 2000a).
PAPs have been functionally characterized from several plants, including Arabidopsis, rice, tomato, kidney bean, tobacco, sweet potato, soybean, duckweed, potato, chickpea, and yellow lupin (Table 1). Recently PAPs have also been explored in model unicellular organisms, i.e., Phaeodactylum tricornutum (diatom) and Chlamydomonas reinhardtii (green algae), performing a similar function and providing new insights into algal P metabolism (Rivera-Solís et al. 2014; Wang et al. 2021). Among plants, PAPs exist as multigene family in Arabidopsis (29 AtPAPs), rice (26 OsPAPs); maize (33 ZmPAPs); soybean (35 GmPAPs), chickpea (25 CaPAPs); Jatropha curcas (25 JCrPAPs), Camellia sinensis (19 CsPAPs), and tomato (25 SlPAPs) (Li et al. 2002, 2012a; Zhang et al. 2011; Bhadouria et al. 2017; González-Muñoz et al. 2015; Venkidasamy et al. 2019; Yin et al. 2019; Srivastava et al. 2020). PAPs were also recently reported in Brassicaceae, Solanaceae, and Cucurbitaceae (Xie and Shang 2018). A subfamily of PAPs also exists, namely phosphodiesterase (PPD) in legumes, hydrolyze phosphodiesters, monoesters, and anhydrides (Olczak et al. 2009; Wang et al. 2015). However, there are still many organisms yet to be explored for the presence of PAPs. Nevertheless, the existence of PAPs multigene families in many plants suggests their conserved and diverse roles.
Structural and functional conservation among PAPs
Different phylogenetic studies divided the members of PAPs multigene family into groups, and the members of the same group largely have similar properties such as molecular weight, biochemical activity, subcellular localization, etc. (Li et al. 2002, 2012a; Tian and Liao 2015; Bhadouria et al. 2017; Srivastava et al. 2020). The functional conservation among the clade members thus aids in assigning a putative function to the new PAP identified in a clade. However, PAPs’ evolutionary history is not very clear; only a recent phylogenetic study proposed that functional diversification of plant PAPs occurred before the existence of early angiosperm (Xie and Shang 2018).
Further, the amino acid sequence analysis of mammals and plants PAPs revealed the presence of five conserved domains/motifs (DxG–GDXXY–GNH (D/E)–VXXH–GHXH) with seven invariant amino acid residues (bold letters) (Fig. 2) (Koonin 1994; Zhuo et al. 1994; Klabunde et al. 1995). However, variations in the number, position, and amino acid residues at the conserved motifs have been reported for different PAPs (Li et al. 2002, 2012a; Zhang et al. 2011; Bhadouria et al. 2017). These invariant residues contribute to the enzyme activity as a mutation in them either eliminates or reduces the catalytic activity (Koonin 1994; Zhuo et al. 1994; Funhoff et al. 2001; Mitić et al. 2009). The metallic center consists of invariable Fe3+ and a varying second metal ion M2+ (M = Fe/Zn/Mn) (Fig. 2) (Klabunde et al. 1995; Schenk et al. 1999; 2008; Boudalis et al. 2007). The in-vitro experiments show catalytic activity reconstitution with varied metal ions, but PAPs are more specific towards specific metal ions when studied in-vivo (Beck et al. 1986; Mitić et al. 2006, 2009). The ferrous ion produces hydroxyl radicals on reacting with hydrogen peroxide, thus imparts alternative function to TRAPs, i.e., generating reactive oxygen species (ROS). Similar to TRAPs, some plant PAPs also catalyze a peroxidation reaction in addition to dephosphorylation (Del Pozo et al. 1999; Bozzo et al. 2002, 2004). Both activities are independent of each other as phosphatase activity inhibitors such as molybdate, do not interfere with peroxidase activity. These enzymatic activities also have different pH optima with phosphatase activity being optimal at acidic pH (~ pH 5), whereas peroxidation occurs at alkaline pH (~ pH 8.5) (Bozzo et al. 2002, 2004). Only a few plants’ PAPs show peroxidase activity which is intriguing, and its relevance should be explored further.
PAPs are broadly divided based on their protein size as small (~ 35 kDa) and large (~ 80 kDa) molecular weight PAPs. The small PAPs were initially identified in mammals and large ones were identified in plants. Later, both types of PAPs were reported in mammals and plants (Schenk et al. 2000a, b; Flanagan et al. 2006). PAPs are reported to exist as monomer, dimer, tetramers, and oligomers (Table 1). Although, the presence of metallophos domain is a characteristic feature of PAPs; lately, other domains are also reported in some PAPs (Fig. 2). For instance, a structure similar to sterol desaturases at C-terminal is present in a PAP from Lupinus albus (Miller et al. 2001). However, the relevance of such a domain remains unexplored. A transmembrane domain at C-terminal is also present in a few plant PAPs, e.g., GmPAP35 in soybean, AtPAP2 and AtPAP9 in Arabidopsis, OsPAP9a in rice (Li et al. 2002, 2012a; Sun et al. 2012a; Zhang et al. 2011). The presence of only one such member in each plant except for Arabidopsis (Li et al. 2002, 2012a; Sun et al. 2012a; Zhang et al. 2011) is intriguing. It also remains to see if all these PAPs localize to the membrane. The physiological impact of this finding is yet to be studied in detail.
Kidney bean PAP (KbPAP) has been widely explored for protein structure (Sträter et al. 1995). Later, structure of human, pig, rat, sweet potato, and yellow lupin PAPs has also been solved (Sträter et al. 1995; Klabunde et al. 1996; Lindqvist et al. 1999; Guddat et al. 1999; Uppenberg et al. 1999; Schenk et al. 2008; Antonyuk et al. 2014). Broadly, PAPs’ protein structure consists of two domains (i.e., N-terminal and C-terminal) joined by the disulfide bridge. PAPs’ catalytic site is present at C-terminal domain, whereas the N-terminal domain (~ 120 residues) does not have any known function (Sträter et al. 1995). Mammalian PAPs have only one domain similar to the C-terminal domain (Sträter et al. 1995; Guddat et al. 1999). The C-terminal domain consists of two β sheets alternating with α helix, conserved in both animals and plants. The metal ions are in contact with carboxyl ends of the β strands (Sträter et al. 1995; Guddat et al. 1999). The N-terminal subdomain consists of β sheets and does not participate in catalysis. Later, the structural study of PPD1 (PAP from yellow lupin) revealed that its N-terminal has a unique domain called fibronectin III (FN3) like domain (two β-sheets making up a seven-stranded β barrel) spanning the whole N-terminal (1–150 residues) (Fig. 2). However, an FN3 domain (150–250 residues) is also present in other PAPs including red kidney PAP and sweet potato PAP (Sträter et al. 1995; Tsyguelnaia and Doolittle 1998). This domain is proposed to participate in substrate selectivity (Antonyuk et al. 2014). The FN3 domains in animals are known to mediate protein–protein interactions or cell adhesion (Craig et al. 2004), and might also have similar functions in plants. PAPs’ catalytic mechanism will not be discussed in detail here as an excellent review is available (Schenk et al. 2013). Briefly, the non-esterified oxygen atoms in the phosphate group attack one of the metal ion (M2+: Fe/Mn/Zn) and the other oxygen atom attacks Fe3+ ion. Thus, a transition state is formed, changing the position of the phosphate group. The nucleophilic attack by the hydroxyl group present between two ions releases the leaving group (ROH). The water molecules attack this transition state that results in the release of the phosphate group (Schenk et al. 2008, 2013). Recently, the crystal structure of rkbPAP in complex with vanadate derivative of ATP has been identified, which mimics the natural enzyme–substrate complex. This structure facilitates viewing the trajectory of substrate binding and targets release from the catalytic site, confirming the earlier predicted mechanism (Feder et al. 2020).
PAPs localize to various subcellular organelles
PAPs are commonly classified as intracellular and secreted based on their functioning in cytoplasm or in extracellular spaces, respectively. The intracellular PAPs are localized in different cell compartments like cytoplasm, plasma membrane, apoplast, tonoplast, mitochondria, chloroplast, golgi bodies, and endoplasmic reticulum (Table 1). The secreted PAPs are either associated with root surface or secreted out in the soil, and aid in Pi solubilization in the rhizosphere. Some PAPs even have dual localization, which in some cases, changes based on prevailing condition or stress. For instance, AtPAP26 exist as both vacuolar and secreted protein (Veljanovski et al. 2006; Hurley et al. 2010). AtPAP2 localizes to the outer envelope of chloroplast, mitochondria, and plasma membrane (Sun et al. 2012b, 2018). AtPAP9 is localized to the plasma membrane and cell wall (Zamani et al. 2012). Both AtPAP2 and AtPAP9 have a transmembrane domain at C-terminal (Sun et al. 2012b; Zamani et al. 2012). AtPAP18 is also a dual-targeted protein, as its transient expression in onion epidermal cells resulted in localization to apoplast after 24 h and vacuoles after 72 h (Zamani et al. 2012). Vacuoles are the cellular reservoir of P and they behave as a source or sink in a condition-dependent manner. Therefore, a wide localization profile of PAPs may help plants in maintaining organelles' P balance. However, it remains to be explored what controls the localization of PAPs in different cellular chambers.
As far as the synthesis of PAPs is considered, induction of PAPs occurs first in roots compared to shoot on Pi deprivation (Bozzo et al. 2004, 2006; Secco et al. 2013). Reports suggest that the synthesis of intracellular APases occurs prior to secreted APases under Pi deprivation condition, as shown in tomato suspension cells (Bozzo et al. 2004, 2006). It appears that plants primarily rely on cytosolic P and activate secreted PAPs later, to acquire P from the rhizosphere.
Complex molecular regulation of PAPs expression and activity
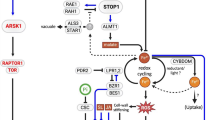
Most PAPs are reported to be induced transcriptionally in response to Pi deficiency (Zhang et al. 2011; Li et al. 2012a; González-Muñoz et al. 2015; Mehra et al. 2016). The MYB domain transcription factor, PHR1 and its homologue (PHL1, PHL2, and PHL3) are involved in transcriptional regulation of various PSR genes, including PAPs such as AtPAP10, OsPAP21b (Sun et al. 2016; Mehra et al. 2017). The PHR1 or its homologues positively regulate the expression of APases including PAPs, as PHR1 overexpression enhances APase activity and mutants displayed a reduced APase activity (Rubio et al. 2001; Zhang et al. 2011; Yao et al. 2014a; Sun et al. 2016; Mehra et al. 2017). Most of the PAPs have PHR1 binding site, i.e., P1BS elements in their promoter, but not all, yet most are responsive to Pi deficiency (Bhadouria et al. 2017). Moreover, only a few studies have shown the physical binding of PHRs at the PAP’s promoter (Mehra et al. 2017). Other cis-regulatory elements such as NIT2, PHO, helix-loop-helix, TC elements, TATA box-like, etc., are present in the promoter of many Pi responsive gene suggesting an additional regulation (Mukatira et al. 2001, Hammond et al. 2004). WRKY75 and ZAT6 are other transcription factors that are induced under Pi deprivation and positively regulate the expression of PSI genes such as phosphatases in Arabidopsis (Devaiah et al. 2007a, b). Besides, altered expression of PAPs is also reported in overexpression/mutant lines of other transcription factors such as OsMYB2P-1, StMYB44, and AP2/ERF, implying that other members of this regulatory pathway need to be identified for having clearer information on the regulation of PAPs (Dai et al. 2012; Chen et al. 2018; Zhou et al. 2017). Intriguingly, the expression of OsPAP10 was enhanced in the mutants of SPX domain-containing transporters, OsSPX-MFS1 (Wang et al. 2012). Further, the nuclear translocation of PHR1 is facilitated by SPX protein in Pi dependent manner (Wang et al. 2014a; Puga et al. 2014). Under Pi sufficient condition, SPXs remain bound to PHR1; therefore, inhibit its binding to promoter of the target PSR genes. In Pi deficient condition, the affinity of SPX towards PHRs decreases leading to the liberation of PHRs for the induction of PSR genes (Wang et al. 2014a; Puga et al. 2014). The SPX dependent regulation varies in different crops as PvSPX1, GmSPX3, and AtSPX1 positively regulate the expression of PAPs (PvPAP3, GmPAP21, AtPAP17, AtPAP2) in Phaseolus vulgaris, soybean, and Arabidopsis, whereas rice SPX1-5 negatively regulates PAPs expression (Duan et al. 2008; Liu et al. 2010a; Shi et al. 2014; Yao et al. 2014a, b; Wang et al. 2014a; Qi et al. 2017). In another study, the overexpression of BnA2SPX1 in Arabidopsis resulted in the downregulation of AtPAP10 expression (Du et al. 2017). These results imply that a complex regulatory mechanism underlying SPX members exists in different plants (Fig. 3) (Lv et al. 2014; Liu et al. 2010a; Shi et al. 2014).
Molecular regulation of PAPs. Under Pi deficient conditions, SPX proteins do not bind to PHR/PHL transcription factor, thus activating PAPs expression (monocots), whereas in dicots SPX1 positively regulates PHR1. PHR controls the miR399-PHO2 pathway, which is also affected by sucrose. OsSPX-MFS1 and OsMYB2P-1 affect the expression of PAPs negatively and positively, respectively. The post-translational modifications such as glycosylation affect the activity and targeting of PAPs. Serine proteases are involved in feedback inhibition of PAPs’ activity by Pi. Auxin signaling also affects Pi signaling through OsARF12. Cytokinin level and PAPs activity are inversely related. Ethylene, as local sensing molecule provides stability to PAPs at protein level. Arrows indicate activation, perpendicular lines show inhibition and dotted lines show indirect effect
Plant hormones are known to influence the expression of many Pi deficiency responsive genes, including PAPs. Auxins usually participate positively in Pi deficiency signaling. However, rice mutants of auxin-responsive transcription factor, osarf12, had an elevated expression of PAPs like, OsPAP9b, OsPAP10a, OsPAP10c, and OsPAP27a, suggesting a negative regulation in WT (Wang et al. 2014b). On the other hand, cytokinins regulate PAPs (AtPAP17 or OsPAP10) expression, negatively exhibited by the exogenous cytokinins application (Fang et al. 2009). Ethylene is a component of local cellular signaling (in response to Pi levels in rhizosphere) and influences the AtPAP10 regulation in Pi dependent manner. The AtPAP10 expression, protein accumulation, and activity increase under P sufficient condition in the presence of ACC (ethylene precursor) (Fig. 3) (Zhang et al. 2014a). These studies suggest the involvement of local Pi signaling in PAPs regulation.
Moreover, systemic signaling components (in response to cellular Pi levels) also influence PAPs behavior. Sucrose, a component of systemic signaling is required for the transcription of AtPAP10 (Zhang et al. 2014a). The presence of sucrose also affects the miRNA399 expression, a positive regulator of PSR (Liu et al. 2010b). The miR399 functions downstream to PHR1 (Bari et al. 2006). PHO2 (UBC24) is a member of the E2 ubiquitin conjugase family which assists the degradation of Pi transporters in sufficient conditions. PHO2 expression is suppressed by miR399s under Pi deficiency, thus enhancing Pi acquisition (Wu et al. 2013). The miRNA399 overexpression or mutation in PHO2 in Arabidopsis or rice results in an increased root-associated APase activity than WT (Zhang et al. 2011; Tao et al. 2016). This suggests that miRNA399-PHO2 pathway also affects root-associated APase activity, most likely indirectly.
Furthermore, PAPs are also regulated at the post-transcriptional level under Pi deficiency. It is reported that a high Pi concentration inhibits the PAPs activity, possibly through a feedback mechanism (Zhang et al. 2014b; Mehra et al. 2017). The P resupply to the Pi deficient tomato suspension cells also resulted in the degradation of APases by enzymes such as serine proteases (Bozzo et al. 2004, 2006). Transcripts profiling further revealed a reduction in the expression of different PAPs such as OsPAP3b, OsPAP21b, OsPAP23, on Pi resupply (Secco et al. 2013). On the other hand, few PAPs such as OsPAP10d, OsPAP7, OsPAP20b, and OsPAP21a show constitutive expression under P sufficient or starvation, but their transcripts decrease on P resupply (Secco et al. 2013). Other PAPs like AtPAP10, StPAP1, OsPAP26 or AtPAP26 do not show transcript induction under Pi deprivation; instead respond post-transcriptionally (Zimmermann et al. 2004; Veljanovski et al. 2006; Tran et al. 2010b; Wang et al. 2011; Gao et al. 2017). The constitutive mRNA level of these PAPs may help the cellular machinery cope quickly under stress conditions (Veljanovski et al. 2006; Wang et al. 2011). Since PAPs form a multigene family in plants, sub-functionalization or neo-functionalization may be defining their varied behavior in response to Pi deficiency.
Post-translational modification (PTM) has been reported to influence PAPs’ activity. Glycosylation affects protein structure, folding, enzyme activity, trafficking, interaction, and localization (Spiro 2002). Glycosylation also plays an important role in PAPs functioning (Fig. 3). PAPs exist as glycosylated protein, which may assist in their targeting into the different cellular organelles and enzymatic activities. The majority of PAPs carry xylosylated or fucosylated amino acids, while others have mannose as sugar moiety (Olczak et al. 2003). Plant PAPs consist of almost 10% sugar and have three to five glycosylation sites (Plaxton and Shane 2015). Two PAPs present in P. vulgaris and L. luteus have been characterized, and glycosylation at C-terminal was found crucial for the secretion of the enzyme (Olczak and Olczak 2007). Notably, two glycoforms of AtPAP26, i.e., S1 and S2 were identified differing in glycosylation pattern and both are secreted (Veljanovski et al. 2006). It was further revealed that their differential glycosylation pattern affects substrate specificities, as AtPAP26-S2 can hydrolyze phenyl-P substrates, but AtPAP26-S1 cannot (Tran et al. 2010b). AtPAP26-S2 is a high mannose glycoform compared to AtPAP26-S1. AtPAP26-S2 was found to interact with AtGAL1 (Galanthus nivalis agglutinin‐related and apple domain lectin‐1) due to high mannose glycans, enhancing its catalytic activity and physiochemical properties (Ghahremani et al. 2019). This explains the role of glycosylation in the regulation of enzymatic activity and substrate specificities. Understanding the glycosylation pattern of PAPs and other PTMs can help reveal novel functions of PAPs and their subcellular localization. Most of the information on PAPs regulation has been generated in Arabidopsis or rice. Whether there is a major difference in PAPs’ regulation in other plants is a subject of future research.
The diverse functions of PAPs
PAPs’ functions are mainly linked to Pi homeostasis. However, PAPs transcripts are also reported to be induced by other abiotic (N or K deficiency, salt stress) and biotic stresses (pathogen attack) (Bhadouria et al. 2017; Deng et al. 2014; Ravichandran et al. 2013). Additionally, PAPs are linked with root growth, symbiotic association, carbon metabolism, phospholipid hydrolysis, defense response, and cellular signaling (Kaida et al. 2009; Sun et al. 2012b; Ravichandran et al. 2015; Del Vecchio et al. 2014; Mehra et al. 2017; Xue et al. 2018; Li et al. 2019). PAPs’ functional horizon is expanding; it is now evident that there is more to reveal about them besides their roles in Pi deficiency adaptation.
PAPs’ roles in mitigating P deficiency stress
PAPs’ broad substrate specificity helps plants survive Pi deficiency by hydrolyzing various organic P compounds for Pi supply. Hence, overexpression of PAP encoding genes is a common approach for enhancing plant Pi deficiency tolerance (Table 1). Usually, the PAPs overexpression improves performance in terms of biomass and total P accumulation, when plants grown on media supplied with an external Po (ATP, dNTPs, DNA) or in soil with organic manure (Table 1). For instance, OsPAP10a, OsPAP10c, and OsPAP21b overexpression helped plants hydrolyze the externally supplied ATP and other Po (Tian et al. 2012; Lu et al. 2016; Mehra et al. 2017; Deng et al. 2020). Kidney bean (PvPAP1, PvPAP3) and Stylosanthes spp. (SgPAP7, 10, 26) PAPs could also utilize dNTPs as Pi source; reflected by an increase in total P content and biomass on their overexpression in hairy roots (Liang et al. 2012; Lu et al. 2016). In Arabidopsis, AtPAP12 and AtPAP26 are prominent PAPs involved in Pi utilization, as atpap12/atpap26 mutant shows an inability to utilize the herring sperm DNA or glycerol-3-phosphate, which otherwise were hydrolyzed by WT Arabidopsis (Robinson et al. 2012). Expectedly, overexpression of AtPAP12 or 26 resulted in better growth on Pi deficient or ADP/ Fru-6P supplied media; however, there was no difference in total P content compared to WT (Wang et al. 2014c). Such PAPs might be involved in enhancing internal Pi recycling from P-rich biomolecules. Thus, both secreted and intracellular PAPs aid the plant performance in Pi limiting conditions.
Besides conventional APase activity, few PAPs such as LlPPD1 (Lupinus luteus) and AsPPD1 (Astragalus sinicus), can also hydrolyze phosphodiesters (Olczak et al. 2000, 2009; Wang et al. 2015). Moreover, a few PAPs display phytase activity, i.e., hydrolyzing phytates. Phytate is a complex of Ca or Mg with myo-inositol and constitutes 70–80% of organic P present in the soil. PAPs with phytase activity differ from other phytases in their catalytic mechanism and also have REKA motif at C-terminal (AtPAP15, NtPAP, GmPHY, and OsPHY), as identified, recently (Fig. 2) (Yao et al. 2012; Feder et al. 2020). The REKA motif at C-terminal is responsible for phytase activity, as identified by the study of NtPAP–phytate complex using homology modeling (Feder et al. 2020). These PAPs are categorized as phytases because of their higher affinity towards phytates (Hegeman and Grabau 2001; Jeong et al. 2017; Kuang et al. 2009; Dionisio et al. 2011; Liu et al. 2018). A handful of PAPs having phytase activity are secreted, whereas others are mostly active during seed germination (Hegeman and Grabau 2001; Kuang et al. 2009; Bhadouria et al. 2017). Secreted PAPs with phytase activity are NtPAP, AtPAP15, MtPAP1, MtPHY1, GmPAP4, and GmPAP14 (Table 1). CaPAP7, SlPAP1, OsPAP23, and GmPHY (GmpPAP29) are intracellular PAPs exhibiting phytase activity (Hegeman and Grabau 2001; Li et al. 2012b; Suen et al. 2015; Bhadouria et al. 2017). The exogenous application of CaPAP7 purified protein leads to an increase in biomass and Pi content of Arabidopsis seedlings grown on media containing phytate as the sole P source (Bhadouria et al. 2017). The overexpression of PAPs with phytase activity (MtPAP1, MtPHY1, OsPAP23, GmPAP14, etc.) led to better growth and higher P content when phytate was provided as the sole P source (Xiao et al. 2005, 2006; Ma et al. 2009; Li et al. 2012b; Kong et al. 2014, 2018). Thus, PAPs with phytase activity increase the substrate profile and strengthen the PAPs utility in improving crop P-use-efficiency.
The developing seeds store P in the form of phytate, which becomes the P source during germination when roots are not fully established. Seed germination involves phytates hydrolysis by phytases and simultaneous induction of APase activity (Gibson and Ullah 1988). The PAPs with phytase activity might be critical for seed germination, as many of them were isolated from germinating seeds (Hegeman and Grabau 2001; Li et al. 2012b). Such PAPs might be involved in seed size regulation, as suggested by CaPAP7 (chickpea PAP with phytase activity) higher expression in genotypes with lower seed weight or phytate content. Since seed phytate levels are directly proportional to seed weight, CaPAP7 might influence seed phytate content, thus seed weight (Bhadouria et al. 2017). PAPs with phytase activity help in seed germination; however, it would be interesting to investigate whether they also assist in P mobilization towards developing seeds, thus influencing crop yield.
PAPs with dual functionality
Some of the PAPs (KbPAP, AtPAP17, LeSAP1, and LeSAP2) consist of two catalytic sites displaying phosphatase and peroxidase activity (Klabunde et al. 1995; Del Pozo et al. 1999; Bozzo et al. 2002). The phosphatase activity is involved in Pi mobilization during its deprivation and tissue senescence (Del Pozo et al. 1999). The peroxidase activity is involved in ROS generation or scavenging, as shown in the case of mammalian PAPs (Del Pozo et al. 1999; Schenk et al. 2013). An increased ROS production is often reported under Pi deficiency that induces the antioxidant pathway genes (Pandey et al. 2013). Klabunde and co-workers had also reported that KbPAP acts as an antioxidant, thus reducing the ROS in seed (Klabunde et al. 1995). Both the activities appear to be independent of each other as APase inhibitors do not affect the peroxidase activity (Bozzo et al. 2002).
PAPs’ peroxidase activity appears to be very appealing as it may help provide tolerance against both Pi deficiency and oxidative stress. Tomato PAPs, LeSAP1, and LeSAP2 are predicted to be involved in ROS production and associated with oxidative burst accompanying pathogen attack. Additionally, various APases are also induced on pathogen infection (Bozzo et al. 2002). There are also few reports which assert the role of PAPs in improving resistance against biotic stress. AtPAP5, a low Pi inducible PAP is required for basal resistance against Pseudomonas syringae pv. Tomato DC3000 (Pst DC3000) (Ravichandran et al. 2013). The higher susceptibility of atpap5 mutants to Pst DC3000 complements the earlier results. Another PAP, AtPAP9 transcripts get induced to fungal infection in stipule and vascular tissues and, is surprisingly not responsive to Pi deficiency (Zamani et al. 2014). Lastly, a study identified the PAP localized in the QTL associated with FHB (Fusarium head blight) resistance in wheat (Liu et al. 2019). Plants have extensive antioxidant defense machinery; the evolution of a few PAPs with peroxidase activity might help plant tolerate oxidative stress that co-exists with Pi deficiency. Although the precise function and mechanism of peroxidase activity in PAPs remain obscure and need further investigation.
P remobilization from senescing tissues and flag leaf
It has been hypothesized that the plant remobilizes Pi from older tissues in response to senescence or cellular P levels dependent manner, i.e., Pi deficiency triggers the Pi remobilization from older tissues. Plants such as Banksia serrata and Hakea prostrata grown in severely Pi depleted soils, and soybean may recycle up to 85% and 50% of total P from senescing leaves, respectively (Crafts‐Brandner 1992; Lambers et al. 2015). Shane and co-workers have isolated two senescence induced intracellular PAP isoforms (HpPAP1 and HpPAP2) functioning in senescing leaves of H. prostrate (Shane et al. 2014). Similarly, OsPAP26 activity increases in rice during senescence, and the Arabidopsis atpap26 mutant showed delayed senescence (Robinson et al. 2012; Gao et al. 2017). Furthermore, Gregerson and Holm (2007) had reported the upregulation of one PAP in the senescing flag leaf of wheat. Rice grains receive most P by the mobilization from plant tissues via flag leaf (Wang et al. 2016). Also, PAPs mediated P mobilization towards developing seeds has been uncovered recently (Gregersen and Holm 2007; Jeong et al. 2017). Lastly, many Pi starvation-induced genes were also reported to be induced in the flag leaf at 15 DAA (days after anthesis), including four PAPs (OsPAP3, OsPAP9b, OsPAP10a, and OsPAP26 thus, emphasizing their role in seed P loading (Jeong et al. 2017). Although, there seems to be functional conservation among the PAPs; however, it remains to be distinguished the PAPs’ roles between senescence and Pi deficiency driven Pi remobilization while the former represents the developmental necessity; the latter is a survival adaptation.
PAPs involvement in response to other abiotic stresses
Emerging evidence shows that PAPs can also help plants tolerate salt and osmotic stress. The overexpression of AtPAP15 provides tolerance to salt and osmotic stress in addition to reduced phytate content in the seeds (Zhang et al. 2008). AtPAP17 is also responsive to salt, ABA and low Pi (Del Pozo et al. 1999). Similarly, GmPAP3 overexpression reduces the damaging effects of salt stress by decreasing oxidative damage (Liao et al. 2003; Deng et al. 2014). Similarly, in Pennisetum glaucum, PgPAP18 like (purple acid phosphatase 18 like) confers tolerance to multiple environmental stress like heat, salt stress, PEG, and CuSO4 in E. coli (Reddy et al. 2017). Such tolerance to oxidative stress may be due to the peroxidase activity of PAPs, as in the case of GmPAP3 or AtPAP17, but it needs experimental validation.
PAPs’ role in root growth modulation
The change in root system architecture (RSA) is one of the primary responses plants exhibit in response to Pi deficiency. In a few cases, PAPs are associated with a change in root growth patterns in transgenics. For instance, AtPAP10 overexpression resulted in increased root growth, such as primary and lateral root (LR) length and density under Pi deficiency (Wang et al. 2011). The overexpression of AtPAP12 or AtPAP26 also had similar results. Expectedly, the atpap10 or atpap12/atpap26 mutants resulted in attenuated root growth (Wang et al. 2011; Tran et al. 2010b). An increase in root growth is also reported on the overexpression of AtPAP15 and AtPAP18 in soybean or tobacco, respectively (Wang et al. 2009; Zamani et al. 2012). In rice also, OsPAP21b overexpression resulted in longer primary and lateral roots, whereas root biomass decreased in knockdown lines (Mehra et al. 2017). The GmPAP4 overexpression in Arabidopsis affected lateral root length and number positively (Kong et al. 2014). The overexpression of JcERF035 (an AP2/ERF transcription factor in Jatropha curcas L.) in Arabidopsis resulted in the suppression of AtPAP17 expression and lateral root formation (Chen et al. 2018). This transcription factor could be a missing link between PAPs expression and increased lateral root formation, but it needs further validation. A few genes like Arabidopsis LPR1, PDR2, and SHR are known to affect both low Pi sensing and root architecture modulation (Pandey et al. 2013). It is also proposed that PAP may alter root development via cell wall modulation (Mehra et al. 2017; Deng et al. 2020). However, there are no reports explaining PAPs’ direct roles in root architecture alteration and the mechanism behind it. It could be possible that their overexpression leads to the induction of Pi signaling, which in turn modulates root architecture and growth.
PAPs role in microbial associations
Symbiotic associations with bacteria or fungi are an inseparable part of plant’s life, and PAPs are also found to affect such associations. In legumes, root nodules are the site for nitrogen fixation in the soil, and PAPs such as GmPAP21, AsPPD1 are expressed in root nodules under Pi deficiency (Wang et al. 2015; Li et al. 2017). The overexpression of such PAPs negatively affected nodule formation. Whereas the silencing of AsPPD1 resulted in early senescence in nodules and reduced nitrogenase activity (Wang et al. 2015). Recently, an RNA-seq analysis also indicated the role of 16 PAPs in maintaining Pi homeostasis in soybean nodules (Xue et al. 2018). PAPs (e.g., AsPPD1, GmPAP21) are involved in maintaining ATP/ADP ratio and energy charge, affecting the nodule formation and nitrogenase activity (Wang et al. 2015; Li et al. 2017). Pi demand for root nodules is high; thus, PAPs activity may help supply much-needed Pi and maintain nitrogenase activity.
Besides, PAPs are also involved in regulating another important symbiotic association, i.e., mycorrhiza. GmPAP33 has been reported to play an important role in arbuscular mycorrhizal symbiosis. GmPAP33 gets induced on arbuscular mycorrhizal inoculation independent of P status (Li et al. 2019). The overexpression of GmPAP33 resulted in improved yield, high P content and large arbuscules, whereas knockdown resulted in an increase in phospholipid concentration and smaller arbuscules. This study further revealed that the role of GmPAP33 in arbuscule degeneration occurs through phospholipid hydrolysis (Li et al. 2019). However, PAPs’ precise mechanism controlling the size, shape, or number of nodules/arbuscule remained unexplored.
Carbon metabolism
Recently, carbon metabolism was linked to further improvement of yield in high yielding green revolution rice varieties (Li et al. 2018). Some of the PAPs, like AtPAP2, are reported to be involved in carbon metabolism (Sun et al. 2012a). In Arabidopsis, AtPAP2 does not respond to Pi starvation; instead, it is involved in carbon metabolism as AtPAP2 overexpression in Arabidopsis resulted in the enhanced biomass, seed yield and higher sugar content. Further, overexpression lines showed induction of sucrose phosphate synthase (Sun et al. 2012a, 2018). Similar results were also obtained on overexpressing AtPAP2 in Camelina sativa, i.e., improved growth rate and seed yield, thus enhancing oil yield/unit area (Zhang et al. 2012). Similarly, overexpression of NtPAP12, a cell wall-bound phosphatase localized in apoplast, activates callose and cellulose synthases by dephosphorylation/photophosphorylation, thereby influencing cell wall synthesis (Kaida et al. 2009). Recently, overexpression of PtPAP1 in P. tricornutum cells (diatoms) showed an increase in carbon content and led to the reallocation of this carbon flux to lipogenesis as depicted by higher lipid content. Furthermore, the PtPAP1 overexpression showed higher photosynthetic efficiency and expression of genes involved in photosynthesis (Wang et al. 2021). These results assert the involvement of PAPs in carbon metabolism, and open up the new avenue for enhanced biomass production using PAPs.
PAPs’ role in reproductive development
Interestingly, many PAPs showed high expression in reproductive tissues (Wang et al. 2011; Zhang et al. 2011; Mehra et al. 2015; Bhadouria et al. 2017). Zhu et al. (2005) reported that 28 AtPAPs showed expression in floral organs with varied intensity. Of these, seven members (AtPAP 6, 11, 14, 19, 23, 24, and 25) were expressed mainly in flowers. Even in chickpea, most of the CaPAPs are expressed in flower buds (Bhadouria et al. 2017). The detailed study of AtPAP23 expression showed its presence in the floral apical meristem, immature flower, petals, and anthers in Arabidopsis (Zhu et al. 2005). Unexpectedly, AtPAP23 transgenics (overexpression and knockdown lines) did not show any change in flower development. On the other hand, the overexpression of OsPAP10a and OsPAP21b resulted in early flowering (Tian et al. 2012; Mehra et al. 2017). These transgenics’ early flowering phenotype can be correlated with improved P levels, as Pi deficiency is known to delay the flowering. However, a direct mechanism linking the activity of PAPs and flowering is yet to be unravelled.
PAPs expression modulates cellular signaling
The PAPs overexpressing plants had only a marginally increased APase activity over the untransformed plants but displayed a marked increase in organic P utilization. PAPs (such as AtPAP25, OsPAP21b) can dephosphorylate a wide range of substrates, including phosphoserine, phosphotyrosine, and phosphothreonine. These metabolites play an important role in multiple cellular signaling processes (Del Vecchio et al. 2014; Mehra et al. 2017). Thus, besides working as phosphatase, PAPs may also influence cellular signaling. Tobacco PAP, NtPAP12 localizes to the apoplast and is involved in dephosphorylation of a-xylosidase and b-glucosidase. Its overexpression leads to decreased glycosidases activity and a higher level of xyloglucan and cello- oligosaccharides (Kaida et al. 2010). AtPAP25 and NtPAP12, both are cell wall localized protein and can be studied for their role in signaling or altering cell wall phosphoproteome under Pi deprivation. The signaling role of PAPs has not been explored and can be an interesting area for future research. The PAPs like OsPAP21b also forms higher molecular weight complexes while interacting with other unknown proteins (Tran et al. 2010b; Mehra et al. 2017). It is possible that such a big protein complex might be linked with some of the PAPs’ signaling roles.
Future perspectives
PAPs are central to plants response to Pi deficiency and adaptation. The emerging roles of PAPs in plant development processes like symbiotic/non-symbiotic association, flowering, seed germination, carbon metabolism, maintaining redox balance, nodule formation, oxidative stress, and defense mechanism suggest that they play greater roles in addition to Pi deficiency. The availability of whole-genome sequence of an ever-increasing number of plants has aided the identification of PAPs multigene family in diverse plants. Functional redundancy is expected among the members of a multigene family. However, emerging evidence reveals some level of sub- or neo-functionalization among PAPs and future research will strengthen PAPs’ wider role besides Pi mobilization. The recent advancements in cellular transcriptomics would greatly help assign more specific roles to PAPs. Moreover, the proteome profiling, especially phosphoproteome of P deficient or plants subjected to other stresses in spatiotemporal manner will further aid in unraveling the mechanistic of PAPs functions.
Overexpression of PAPs has been a common strategy to understand their functions. The utilization of CRISPR/cas9 to generate PAPs knockout will provide more precise information on their roles in plants. Most studies used a constitutive promoter for overexpression, which sometimes results in undesired effects in P sufficient conditions. As demonstrated recently, using a native promoter can be a better way to study their roles (Deng et al. 2020). Only a handful of studies have tested PAPs overexpression transgenics in soil using organic-P as a sole P source (Mehra et al. 2017; Deng et al. 2020). The transgenics overexpressing PAPs should be assessed for agronomic traits using organic fertilizers in fields to evaluate their real potential for improving crop P-use-efficiency. This will help figure out the best candidate to be used for developing P efficient crops. To develop Pi efficient crops, the breeding programme often utilizes QTL approach. Although there are few reports where PAPs have been identified falling in QTLs associated with low Pi tolerance, such as HvPAPa is associated with mqphy QTL (associated with phytase activity), few ZmPAPs also co-localize with low Pi responsive cQTLs in a meta-analysis (Dai et al. 2011; Zhang et al. 2014b; González-Muñoz et al. 2015). However, the information on PAPs association with P responsive QTL is at a nascent stage, and no PAP has been identified as a candidate gene so far. However, initial reports are encouraging and given their key role in P homeostasis, PAPs may be identified as an important candidate in future for molecular breeding to develop Pi efficient crops.
There remain many unanswered questions on PAPs biochemistry and new roles. What is the role of PAPs’ N-terminal domain? Is it responsible for protein–protein interactions leading to oligomerization as reported in TRAPs? How post-translation modifications influence their catalytic behavior and how are they regulated? Answering these questions should be a target of future research. It should also be investigated how PAPs having roles other than P utilization are regulated at molecular levels? Lastly, PAPs are degraded on sufficient P supply (Mehra et al. 2017) losing the advantages to plants growing in P supplied soils. Pinning down amino acids responsible for their interaction with proteasome machinery and mutating them using gene-editing may help further improve P-use-efficiency. Considering the advantages provided by the PAPs to the crops (e.g., improved P-use-efficiency, hydrolyzing organic P, better yield), they could be potential targets for improving low P tolerance using molecular breeding and transgenesis.
References
Antonyuk SV, Olczak M, Olczak T, Ciuraszkiewicz J, Strange RW (2014) The structure of a purple acid phosphatase involved in plant growth and pathogen defence exhibits a novel immunoglobulin-like fold. IUCrJ 1:101–109
Arai Y, Sparks DL (2007) Phosphate reaction dynamics in soils and soil components: a multiscale approach. Adv Agron 94:135–179
Bari R, Datt Pant B, Stitt M, Scheible WR (2006) PHO2, microRNA399, and PHR1 define a phosphate-signaling pathway in plants. Plant Physiol 141:988–999
Beck JL, McConachie LA, Summors AC, Arnold WN, De Jersey J, Zerner B (1986) Properties of a purple phosphatase from red kidney bean: a zinc-iron metalloenzyme. Biochim Biophys Acta 869:61–68
Bhadouria J, Singh AP, Mehra P, Verma L, Srivastawa R, Parida SK, Giri J (2017) Identification of purple acid phosphatases in chickpea and potential roles of CaPAP7 in seed phytate accumulation. Sci Rep 7:1–12
Boudalis AK, Aston RE, Smith SJ, Mirams RE, Riley MJ, Schenk G, Blackman AG, Hanton LR, Gahan LR (2007) Synthesis and characterization of the tetranuclear iron (III) complex of a new asymmetric multidentate ligand. A structural model for purple acid phosphatases. Dalton T 44:5132–5139
Bozzo GG, Plaxton WC (2008) The role of intracellular and secreted purple acid phosphatases in tomato phosphate nutrition. In: Preddy V, Watson R (eds) Tomatoes and tomato products. Science Publishers, Enfield, pp 216–233
Bozzo GG, Raghothama KG, Plaxton WC (2002) Purification and characterization of two secreted purple acid phosphatase isozymes from phosphate-starved tomato (Lycopersicon esculentum) cell cultures. Eur J Biochem 269:6278–6286
Bozzo GG, Raghothama KG, Plaxton WC (2004) Structural and kinetic properties of a novel purple acid phosphatase from phosphate-starved tomato (Lycopersicon esculentum) cell cultures. Biochem J 377:419–428
Bozzo GG, Dunn EL, Plaxton WC (2006) Differential synthesis of phosphate-starvation inducible purple acid phosphatase isozymes in tomato (Lycopersicon esculentum) suspension cells and seedlings. Plant Cell Environ 29:303–313
Cashikar AG, Kumaresan R, Madhusudhana Rao N (1997) Biochemical characterization and subcellular localization of the red kidney bean purple acid phosphatase. Plant Physiol 114:907–915
Chen TT, Bazer FW, Gebhardt BM, Roberts RM (1975) Uterine secretion in mammals: synthesis and placental transport of a purple acid phosphatase in pigs. Biol Reprod 13:304–313
Chen Y, Wu P, Zhao Q, Tang Y, Chen Y, Li M, Jiang H, Wu G (2018) Overexpression of a phosphate starvation response AP2/ERF gene from physic nut in Arabidopsis alters root morphological traits and phosphate starvation-induced anthocyanin accumulation. Front Plant Sci 9:1186
Crafts-Brandner SJ (1992) Significance of leaf phosphorus remobilization in yield production in soybean. Crop Sci 32:420–424
Craig D, Gao M, Schulten K, Vogel V (2004) Tuning the mechanical stability of fibronectin type III modules through sequence variations. Structure 12:21–30
Dai F, Qiu L, Ye L, Wu D, Zhou M, Zhang G (2011) Identification of a phytase gene in barley (Hordeum vulgare L.). PLoS ONE 6:e18829
Dai X, Wang Y, Yang A, Zhang WH (2012) OsMYB2P-1, an R2R3 MYB transcription factor, is involved in the regulation of phosphate-starvation responses and root architecture in rice. Plant Physiol 159:169–183
Del Pozo JC, Allona I, Rubio V, Leyva A, De La Peña A, Aragoncillo C, Paz-Ares J (1999) A type 5 acid phosphatase gene from Arabidopsis thaliana is induced by phosphate starvation and by some other types of phosphate mobilising/oxidative stress conditions. Plant J 19:579–589
Del Vecchio HA, Ying S, Park J, Knowles VL, Kanno S, Tanoi K, She YM, Plaxton WC (2014) The cell wall-targeted purple acid phosphatase AtPAP25 is critical for acclimation of Arabidopsis thaliana to nutritional phosphorus deprivation. Plant J 80(569):581
Deng L, Chen F, Jiang L, Lam HM, Xiao G (2014) Ectopic expression of GmPAP3 enhances salt tolerance in rice by alleviating oxidative damage. Plant Breed 133:348–355
Deng S, Lu L, Li J, Du Z, Liu T, Li W, Xu F, Shi L, Shou H, Wang C (2020) Purple acid phosphatase 10c (OsPAP10c) encodes a major acid phosphatase and regulates the plant growth under phosphate deficient condition in rice. J Exp Bot 71:4321–4332
Devaiah BN, Karthikeyan AS, Raghothama KG (2007a) WRKY75 transcription factor is a modulator of phosphate acquisition and root development in Arabidopsis. Plant Physiol 143:1789–1801
Devaiah BN, Nagarajan VK, Raghothama KG (2007b) Phosphate homeostasis and root development in Arabidopsis are synchronized by the zinc finger transcription factor ZAT6. Plant Physiol 145:147–159
Dionisio G, Madsen CK, Holm PB, Welinder KG, Jørgensen M, Stoger E, Arcalis E, Brinch-Pedersen H (2011) Cloning and characterization of purple acid phosphatase phytases from wheat, barley, maize, and rice. Plant Physiol 156:1087–1100
Dissanayaka DMSB, Plaxton WC, Lambers H, Siebers M, Marambe B, Wasaki J (2018) Molecular mechanisms underpinning phosphorus-use efficiency in rice. Plant Cell Environ 41:1483–1496
Du H, Yang C, Ding G, Shi L, Xu F (2017) Genome-wide identification and characterization of SPX domain-containing members and their responses to phosphate deficiency in Brassica napus. Front Plant Sci 8:35
Duan K, Yi K, Dang L, Huang H, Wu W, Wu P (2008) Characterization of a sub-family of Arabidopsis genes with the SPX domain reveals their diverse functions in plant tolerance to phosphorus starvation. Plant J 54:965–975
Efstratiadis T, Moss DW (1985) Tartrate-resistant acid phosphatase of human lung: apparent identity with osteoclastic acid phosphatase. Enzyme 33:34–40
Fang Z, Shao C, Meng Y, Wu P, Chen M (2009) Phosphate signaling in Arabidopsis and Oryza sativa. Plant Sci 176:170–180
Farhadi S, Sabet MS, Malboobi MA, Moieni A (2020) The critical role of AtPAP17 and AtPAP26 genes in Arabidopsis phosphate compensation network. Front Plant Sci 11:565865
Feder D, McGeary RP, Mitić N, Lonhienne T, Furtado A, Schulz BL, Henry RJ, Schmidt S, Guddat LW, Schenk G (2020) Structural elements that modulate the substrate specificity of plant purple acid phosphatases: avenues for improved phosphorus acquisition in crops. Plant Sci 294:110445
Flanagan JU, Cassady AI, Schenk G, Guddat LW, Hume DA (2006) Identification and molecular modeling of a novel, plant-like, human purple acid phosphatase. Gene 377:12–20
Funhoff EG, Ljusberg J, Wang Y, Andersson G, Averill BA (2001) Mutational analysis of the interaction between active site residues and the loop region in mammalian purple acid phosphatases. Biochemistry 40:11614–11622
Gao W, Lu L, Qiu W, Wang C, Shou H (2017) OsPAP26 encodes a major purple acid phosphatase and regulates phosphate remobilization in rice. Plant Cell Physiol 58:885–892
Ghahremani M, Park J, Anderson EM, Marty-Howard NJ, Mullen RT, Plaxton WC (2019) Lectin AtGAL1 interacts with high-mannose glycoform of the purple acid phosphatase AtPAP26 secreted by phosphate-starved Arabidopsis. Plant Cell Environ 42:1158–1166
Gibson DM, Ullah AHJ (1988) Purification and characterization of phytase from cotyledons of germinating soybean seeds. Arch Biochem Biophys 260:503–513
González-Muñoz E, Avendaño-Vázquez AO, Montes RA, de Folter S, Andrés-Hernández L, Abreu-Goodger C, Sawers RJ (2015) The maize (Zea mays ssp. mays var. B73) genome encodes 33 members of the purple acid phosphatase family. Front Plant Sci 6:341
Gregersen PL, Holm PB (2007) Transcriptome analysis of senescence in the flag leaf of wheat (Triticum aestivum L.). Plant Biotechnol J 5:192–206
Gu M, Chen A, Sun S, Xu G (2016) Complex regulation of plant phosphate transporters and the gap between molecular mechanisms and practical application: what is missing? Mol Plant 9:396–416
Guddat LW, McAlpine AS, Hume D, Hamilton S, de Jersey J, Martin JL (1999) Crystal structure of mammalian purple acid phosphatase. Structure 7:757–767
Halleen JM, Räisänen SR, Alatalo SL, Väänänen HK (2003) Potential function for the ROS-generating activity of TRACP. J Bone Miner Res 18:1908–1911
Hammond JP, Broadley MRWPJ (2004) Genetic responses to phosphorus deficiency. Ann Bot 94:323–332
Hayman AR, Jones SJ, Boyde A, Foster D, Colledge WH, Carlton MB, Evans MJ, Cox TM (1996) Mice lacking tartrate-resistant acid phosphatase (Acp5) have disrupted endochondral ossification and mild osteopetrosis. Development 122:3151–3162
Hegeman CE, Grabau EA (2001) A novel phytase with sequence similarity to purple acid phosphatases is expressed in cotyledons of germinating soybean seedlings. Plant Physiol 126:1598–1608
Hur YJ, Jin BR, Nam J, Chung YS, Lee JH, Choi HK, Yun DJ, Yi G, Kim YH, Kim DH (2010) Molecular characterization of OsPAP2: transgenic expression of a purple acid phosphatase up-regulated in phosphate-deprived rice suspension cells. Biotechnol Lett 32:163–170
Hurley BA, Tran HT, Marty NJ, Park J, Snedden WA, Mullen RT, Plaxton WC (2010) The dual-targeted purple acid phosphatase isozyme AtPAP26 is essential for efficient acclimation of Arabidopsis to nutritional phosphate deprivation. Plant Physiol 153:1112–1122
Jeong K, Baten A, Waters DL, Pantoja O, Julia CC, Wissuwa M, Heuer S, Kretzschmar T, Rose TJ (2017) Phosphorus remobilization from rice flag leaves during grain filling: an RNA-seq study. Plant Biotechnol J 15:15–26
Kaida R, Hayashi T, Kaneko TS (2008) Purple acid phosphatase in the walls of tobacco cells. Phytochemistry 69:2546–2551
Kaida R, Satoh Y, Bulone V, Yamada Y, Kaku T, Hayashi T, Kaneko TS (2009) Activation of β-glucan synthases by wall-bound purple acid phosphatase in tobacco cells. Plant Physiol 150:1822–1830
Kaida R, Serada S, Norioka N, Norioka S, Neumetzler L, Pauly M, Sampedro J, Zarra I, Hayashi T, Kaneko TS (2010) Potential role for purple acid phosphatase in the dephosphorylation of wall proteins in tobacco cells. Plant Physiol 153:603–610
Ketcham CM, Baumbach GA, Bazer FW, Roberts RM (1985) The type 5, acid phosphatase from spleen of humans with hairy cell leukemia. Purification, properties, immunological characterization, and comparison with porcine uteroferrin. J Biol Chem 260:5768–5776
Ketcham CM, Roberts RM, Simmen RCM, Nick HS (1989) Molecular cloning of the type 5, iron-containing, tartrate-resistant acid phosphatase from human placenta. J Biol Chem 264:557–563
Klabunde T, Sträter N, Krebs B, Witzel H (1995) Structural relationship between the mammalian Fe(III)/Fe(II) and the Fe(III)/Zn(II) plant purple acid phosphatases. FEBS Lett 367:56–60
Klabunde T, Sträter N, Fröhlich R, Witzel H, Krebs B (1996) Mechanism of Fe (III)–Zn (II) purple acid phosphatase based on crystal structures. J Mol Biol 259:737–748
Kong Y, Li X, Ma J, Li W, Yan G, Zhang C (2014) GmPAP4, a novel purple acid phosphatase gene isolated from soybean (Glycine max), enhanced extracellular phytate utilization in Arabidopsis thaliana. Plant Cell Rep 33:655–667
Kong Y, Li X, Wang B, Li W, Du H, Zhang C (2018) The soybean purple acid phosphatase GmPAP14 predominantly enhances external phytate utilization in plants. Front Plant Sci 9:292
Koonin EV (1994) Conserved sequence pattern in a wide variety of phosphoesterases. Protein Sci 3:356–358
Kuang R, Chan KH, Yeung E, Lim BL (2009) Molecular and biochemical characterization of AtPAP15, a purple acid phosphatase with phytase activity, in Arabidopsis. Plant Physiol 151:199–209
Lambers H, Martinoia E, Renton M (2015) Plant adaptations to severely phosphorus-impoverished soils. Curr Opin Plant Biol 25:23–31
Li D, Zhu H, Liu K, Liu X, Leggewie G, Udvardi M, Wang D (2002) Purple acid phosphatases of Arabidopsis thaliana. Comparative analysis and differential regulation by phosphate deprivation. J Biol Chem 277:27772–27781
Li WYF, Shao G, Lam HM (2008) Ectopic expression of GmPAP3 alleviates oxidative damage caused by salinity and osmotic stresses. New Phytol 178:80–91
Li C, Gui S, Yang T, Walk T, Wang X, Liao H (2012a) Identification of soybean purple acid phosphatase genes and their expression responses to phosphorus availability and symbiosis. Ann Bot 109:275–285
Li RJ, Lu WJ, Guo CJ, Li XJ, Gu JT, Kai XI (2012b) Characterization and functional analysis of OsPHY1, a purple acid phosphatase (pap)–type phytase gene in rice (Oryza sativa L.). J Integr Agric 11:1217–1226
Li C, Li C, Zhang H, Liao H, Wang X (2017) The purple acid phosphatase GmPAP21 enhances internal phosphorus utilization and possibly plays a role in symbiosis with rhizobia in soybean. Physiol Plant 159:215–227
Li S, Tian Y, Wu K, Ye Y, Yu J, Zhang J, Liu Q, Hu M, Li H, Tong Y, Harberd NP (2018) Modulating plant growth–metabolism coordination for sustainable agriculture. Nature 560:595–600
Li C, Zhou J, Wang X, Liao H (2019) A purple acid phosphatase, GmPAP33, participates in arbuscule degeneration during arbuscular mycorrhizal symbiosis in soybean. Plant Cell Environ 42:2015–2027
Liang C, Tian J, Lam HM, Lim BL, Yan X, Liao H (2010) Biochemical and molecular characterization of PvPAP3, a novel purple acid phosphatase isolated from common bean enhancing extracellular ATP utilization. Plant Physiol 152:854–865
Liang C, Sun L, Yao Z, Liao H, Tian J (2012) Comparative analysis of PvPAP gene family and their functions in response to phosphorus deficiency in Common Bean. PLoS ONE 7:16–20
Liao H, Wong FL, Phang TH, Cheung MY, Li WY, Shao G, Yan X, Lam HM (2003) GmPAP3, a novel purple acid phosphatase-like gene in soybean induced by NaCl stress but not phosphorus deficiency. Gene 318:103–111
Lindqvist Y, Johansson E, Kaija H, Vihko P, Schneider G (1999) Three-dimensional structure of a mammalian purple acid phosphatase at 2.2 Å resolution with a μ-(hydr) oxo bridged di-iron center. J Mol Biol 291:135–147
Liu F, Wang Z, Ren H, Shen C, Li Y, Ling HQ, Wu C, Lian X, Wu P (2010a) OsSPX1 suppresses the function of OsPHR2 in the regulation of expression of OsPT2 and phosphate homeostasis in shoots of rice. Plant J 62:508–517
Liu JQ, Allan DL, Vance CP (2010b) Systemic signaling and local sensing of phosphate in common bean: cross-talk between photosynthate and microRNA399. Mol Plant 3:428–437
Liu P, Cai Z, Chen Z, Mo X, Ding X, Liang C, Liu G, Tian J (2018) A root-associated purple acid phosphatase, SgPAP23, mediates extracellular phytate-P utilization in Stylosanthes guianensis. Plant Cell Environ 41:2821–2834
Liu J, Li L, Foroud NA, Gong X, Li C, Li T (2019) Proteomics of bulked rachides combined with documented QTL uncovers genotype nonspecific players of the Fusarium head blight responses in wheat. Phytopathology 109:111–119
López-Arredondo DL, Leyva-González MA, González-Morales SI, López-Bucio J, Herrera-Estrella L (2014) Phosphate Nutrition: improving low-phosphate tolerance in crops. Annu Rev Plant Biol 65:95–123
Lu L, Qiu W, Gao W, Tyerman SD, Shou H, Wang C (2016) OsPAP10c, a novel secreted acid phosphatase in rice, plays an important role in the utilization of external organic phosphorus. Plant Cell Environ 39:2247–2259
Lung SC, Leung A, Kuang R, Wang Y, Leung P, Lim BL (2008) Phytase activity in tobacco (Nicotiana tabacum) root exudates is exhibited by a purple acid phosphatase. Phytochemistry 69:365–373
Lv Q, Zhong Y, Wang Y, Wang Z, Zhang L, Shi J, Wu Z, Liu Y, Mao C, Yi K, Wu P (2014) SPX4 negatively regulates phosphate signaling and homeostasis through its interaction with PHR2 in rice. Plant Cell 26:1586–1597
Ma XF, Wright E, Ge Y, Bell J, Xi Y, Bouton JH, Wang ZY (2009) Improving phosphorus acquisition of white clover (Trifolium repens L.) by transgenic expression of plant-derived phytase and acid phosphatase genes. Plant Sci 176:479–488
Ma XF, Tudor S, Butler T, Ge Y, Xi Y, Bouton J, Harrison M, Wang ZY (2012) Transgenic expression of phytase and acid phosphatase genes in alfalfa (Medicago sativa) leads to improved phosphate uptake in natural soils. Mol Breeding 1:377–391
Mehra P, Pandey BK, Giri J (2015) Genome-wide DNA polymorphisms in low phosphate tolerant and sensitive rice genotypes. Sci Rep 5:13090
Mehra P, Pandey BK, Giri J (2016) Comparative morphophysiological analyses and molecular profiling reveal Pi-efficient strategies of a traditional rice genotype. Front Plant Sci 6:1184
Mehra P, Pandey BK, Giri J (2017) Improvement in phosphate acquisition and utilization by a secretory purple acid phosphatase (OsPAP21b) in rice. Plant Biotechnol J 15:1054–1067
Miller SS, Liu J, Allan DL, Menzhuber CJ, Fedorova M, Vance CP (2001) Molecular control of acid phosphatase secretion into the rhizosphere of proteoid roots from phosphorus-stressed white lupin. Plant Physiol 127:594–606
Mitić N, Smith SJ, Neves A, Guddat LW, Gahan LR, Schenk G (2006) The catalytic mechanisms of binuclear metallohydrolases. Chem Rev 106:3338–3363
Mitić N, Noble CJ, Gahan LR, Hanson GR, Schenk G (2009) Metal-ion mutagenesis: conversion of a purple acid phosphatase from sweet potato to a neutral phosphatase with the formation of an unprecedented catalytically competent MnII/MnII active site. J Am Chem Soc 131:8173–8179
Mukatira UT, Liu C, Varadarajan DK, Raghothama KG (2001) Negative regulation of phosphate starvation-induced genes. Plant Physiol 127:1854–1862
Nakazato H, Okamoto T, Nishikoori M, Washio K, Morita N, Haraguchi K, Thompson GA, Okuyama H (1998) The glycosylphosphatidylinositol-anchored phosphatase from Spirodela oligorrhiza is a purple acid phosphatase. Plant Physiol 118:1015–1020
Nishikoori M, Washio K, Hase A, Morita N, Okuyama H (2001) Cloning and characterization of cDNA of the GPI-anchored purple acid phosphatase and its root tissue distribution in Spirodela oligorrhiza. Physiol Plant 113:241–248
Oelkers EH, Valsami-Jones E (2008) Phosphate mineral reactivity and global sustainability. Elements 4:83–87
Olczak M, Olczak T (2002) Diphosphonucleotide phosphatase/phosphodiesterase from yellow lupin (Lupinus luteus L.) belongs to a novel group of specific metallophosphatases. FEBS Lett 519:159–163
Olczak M, Olczak T (2007) N-Glycosylation sites of plant purple acid phosphatases important for protein expression and secretion in insect cells. Arch Biochem Biophys 461:247–254
Olczak M, Kobialka M, Wa̧torek WX (2000) Characterization of diphosphonucleotide phosphatase/phosphodiesterase from yellow lupin (Lupinus luteus) seeds. Biochim Biophys Acta 1478:239–247
Olczak M, Morawiecka B, Wa̧torek W (2003) Plant purple acid phosphatases—genes, structures and biological function. Acta Biochim Pol 50:1245–1256
Olczak M, Ciuraszkiewicz J, Wójtowicz H, Maszczak D, Olczak T (2009) Diphosphonucleotide phosphatase/phosphodiesterase (PPD1) from yellow lupin (Lupinus luteus L.) contains an iron-manganese center. FEBS Lett 583:3280–3284
Pandey BK, Mehra P, Giri J (2013) Phosphorus starvation response in plants and opportunities for crop improvement. In: Tuteja N, Gill SS (eds) Climate change and plant abiotic stress tolerance. Wiley, New York, pp 991–1012
Plaxton WC, Shane MW (2015) The role of post-translational enzyme modifications in the metabolic adaptations of phosphorus-deprived plants. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants. Wiley, Hoboken, pp 99–124
Poirier Y, Jung JY (2015) Phosphate transporters. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants. Wiley, Hoboken, pp 125–158
Puga MI, Mateos I, Charukesi R, Wang Z, Franco-Zorrilla JM, de Lorenzo L, Irigoyen ML, Masiero S, Bustos R, Rodríguez J, Leyva A (2014) SPX1 is a phosphate-dependent inhibitor of phosphate starvation response 1 in Arabidopsis. Proc Natl Acad Sci USA 111:14947–14952
Qi W, Iain WM, Stephen PM, Alison B (2017) AtSPX1 affects the AtPHR1–DNA-binding equilibrium by binding monomeric AtPHR1 in solution. Biochem J 474:3675–3687
Raghothama KG (1999) Phosphate acquisition. Annu Rev Plant 50:665–693
Raghothama KG (2005) Phosphorus and plant nutrition: an overview. In: Sims JT, Sharpley AN, Cabrera ML (eds) Phosphorus: agriculture and the environment. Agronomy Monograph Madison, Madison, pp 353–378
Ravichandran S, Stone SL, Benkel B, Prithiviraj B (2013) Purple acid phosphatase 5 is required for maintaining basal resistance against Pseudomonas syringae in Arabidopsis. BMC Plant Biol 13:1–2
Ravichandran S, Stone SL, Benkel B, Zhang J, Berrue F, Prithiviraj B (2015) Optimal level of purple acid phosphatase5 is required for maintaining complete resistance to Pseudomonas syringae. Front Plant Sci 6:568
Reddy CS, Kim KM, James D, Varakumar P, Reddy MK (2017) PgPAP18, a heat-inducible novel purple acid phosphatase 18-like gene (PgPAP18-like) from Pennisetum glaucum, may play a crucial role in environmental stress adaptation. Acta Physiol Plant 39:54
Rivera-Solís RA, Peraza-Echeverria S, Echevarría-Machado I, Herrera-Valencia VA (2014) Chlamydomonas reinhardtii has a small family of purple acid phosphatase homologue genes that are differentially expressed in response to phytate. Ann Microbiol 64:551–559
Robinson WD, Park J, Tran HT, Del Vecchio HA, Ying S, Zins JL, Patel K, McKnight TD, Plaxton WC (2012) Eliminating the purple acid phosphatase AtPAP26 in Arabidopsis thaliana delays leaf senescence and impairs phosphorus remobilization. New Phytol 196:1024–1029
Rubio V, Linhares F, Solano R, Martín AC, Iglesias J, Leyva A, Paz-Ares J (2001) A conserved MYB transcription factor involved in phosphate starvation signaling both in vascular plants and in unicellular algae. Genes Dev 15:2122–2133
Scheible WR, Rojas-Triana M (2015) Sensing, signalling, and control of phosphate starvation in plants: molecular players and applications. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants. Wiley, Hoboken, pp 25–64
Schenk G, Ge Y, Carrington LE, Wynne CJ, Searle IR, Carroll BJ, Hamilton S, de Jersey J (1999) Binuclear metal centers in plant purple acid phosphatases: Fe–Mn in sweet potato and Fe–Zn in soybean. Arch Biochem Biophys 370:183–189
Schenk G, Guddat LW, Ge Y, Carrington LE, Hume DA, Hamilton S, De Jersey J (2000a) Identification of mammalian-like purple acid phosphatases in a wide range of plants. Gene 250:117–125
Schenk G, Korsinczky ML, Hume DA, Hamilton S, DeJersey J (2000b) Purple acid phosphatases from bacteria: similarities to mammalian and plant enzymes. Gene 255:419–424
Schenk G, Elliott TW, Leung E, Carrington LE, Mitić N, Gahan LR, Guddat LW (2008) Crystal structures of a purple acid phosphatase, representing different steps of this enzyme’s catalytic cycle. BMC Struct Biol 8:6
Schenk G, Mitić NŠ, Hanson GR, Comba P (2013) Purple acid phosphatase: a journey into the function and mechanism of a colorful enzyme. Coordin Chem Rev 257:473–482
Secco D, Jabnoune M, Walker H et al (2013) Spatio-temporal transcript profiling of rice roots and shoots in response to phosphate starvation and recovery. Plant Cell 25:4285–4304
Shane MW, Stigter K, Fedosejevs ET, Plaxton WC (2014) Senescence-inducible cell wall and intracellular purple acid phosphatases: implications for phosphorus remobilization in Hakea prostrata (Proteaceae) and Arabidopsis thaliana (Brassicaceae). J Exp Bot 65:6097–6106
Shen J, Yuan L, Zhang J, Li H, Bai Z, Chen X, Zhang W, Zhang F (2011) Phosphorus dynamics: from soil to plant. Plant Physiol 156:997–1005
Shi J, Hu H, Zhang K, Zhang W, Yu Y, Wu Z, Wu P (2014) The paralogous SPX3 and SPX5 genes redundantly modulate Pi homeostasis in rice. J Exp Bot 65:859–870
Spiro RG (2002) Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12:43R-56R
Srivastava S, Upadhyay MK, Srivastava AK, Abdelrahman M, Suprasanna P, Tran LP (2018) Cellular and subcellular phosphate transport machinery in plants. Int J Mol Sci 19:1914
Srivastava R, Akash PAP, Chauhan PK, Kumar R (2020) Identification, structure analysis, and transcript profiling of purple acid phosphatases under Pi deficiency in tomato (Solanum lycopersicum L.) and its wild relatives. Int J Biol Macromol 165:2253–2266
Sträter N, Klabunde T, Tucker P, Witzel H, Krebs B (1995) Crystal structure of a purple acid phosphatase containing a dinuclear Fe(III)-Zn(II) active site. Science 268:1489–1492
Suen PK, Zhang S, Sun SSM (2015) Molecular characterization of a tomato purple acid phosphatase during seed germination and seedling growth under phosphate stress. Plant Cell Rep 34:981–992
Sun F, Carrie C, Law S, Murcha MW, Zhang R, Law YS, Suen PK, Whelan J, Lim BL (2012a) AtPAP2 is a tail-anchored protein in the outer membrane of chloroplasts and mitochondria. Plant Signal Behav 7:927–932
Sun F, Suen PK, Zhang Y, Liang C, Carrie C, Whelan J, Ward JL, Hawkins ND, Jiang L, Lim BL (2012b) A dual-targeted purple acid phosphatase in Arabidopsis thaliana moderates carbon metabolism and its overexpression leads to faster plant growth and higher seed yield. New Phytol 194:206–219
Sun F, Liang C, Whelan J, Yang J, Zhang P, Lim BL (2013) Global transcriptome analysis of AtPAP2-overexpressing Arabidopsis thaliana with elevated ATP. BMC Genomics 14:1–3
Sun L, Song L, Zhang Y, Zheng Z, Liu D (2016) Arabidopsis PHL2 and PHR1 act redundantly as the key components of the central regulatory system controlling transcriptional responses to phosphate starvation. Plant Physiol 170:499–514
Sun Q, Li J, Cheng W, Guo H, Liu X, Gao H (2018) AtPAP2, a unique member of the PAP family, functions in the plasma membrane. Genes 9:257
Sundararajan TA, Sarma PS (1954) Purification and properties of phosphoprotein phosphatase from ox spleen. Biochem J 56:125–130
Tao S, Zhang Y, Wang X, Xu L, Fang X, Lu ZJ, Liu D (2016) The THO/TREX complex active in miRNA biogenesis negatively regulates root-associated acid phosphatase activity induced by phosphate starvation 1. Plant Physiol 171:2841–2853
Tian J, Liao H (2015) The role of intracellular and secreted purple acid phosphatases in plant phosphorus scavenging and recycling. In: Plaxton WC, Lambers H (eds) Phosphorus metabolism in plants. Wiley, Hoboken, pp 265–288
Tian J, Wang C, Zhang Q, He X, Whelan J, Shou H (2012) Overexpression of OsPAP10a, A root-associated acid phosphatase, increased extracellular organic phosphorus utilization in rice. J Integr Plant Biol 54:631–639
Tran HT, Hurley BA, Plaxton WC (2010a) Feeding hungry plants: the role of purple acid phosphatases in phosphate nutrition. Plant Sci 179:14–27
Tran HT, Qian W, Hurley BA, She YM, Wang D, Plaxton WC (2010b) Biochemical and molecular characterization of AtPAP12 and AtPAP26: the predominant purple acid phosphatase isozymes secreted by phosphate-starved Arabidopsis thaliana. Plant Cell Environ 33:1789–1803
Tsyguelnaia I, Doolittle RF (1998) Presence of a fibronectin type III domain in a plant protein. J Mol Evol 46:612–614
Ullah AHJ, Cummins BJ (1988) Aspergillus ficuum extracellular pH 6.0 optimum acid phosphatase: purification, n-terminal amino acid sequence, and biochemical characterization. Prep Biochem 18:37–65
Uppenberg J, Lindqvist F, Svensson C, Ek-Rylander B, Andersson G (1999) Crystal structure of a mammalian purple acid phosphatase. J Mol Biol 290:201–211
Veljanovski V, Vanderbeld B, Knowles VL, Snedden WA, Plaxton WC (2006) Biochemical and molecular characterization of AtPAP26, a vacuolar purple acid phosphatase up-regulated in phosphate-deprived Arabidopsis suspension cells and seedlings. Plant Physiol 142:1282–1293
Venkidasamy B, Selvaraj D, Ramalingam S (2019) Genome-wide analysis of purple acid phosphatase (PAP) family proteins in Jatropha curcas L. Int J Biol Macromol 123:648–656
Wang X, Wang Y, Jiang T, Lim BL, Yan X, Liao H (2009) Overexpressing AtPAP15 enhances phosphorus efficiency in soybean. Plant Physiol 151:233–240
Wang L, Li Z, Qian W, Guo W, Gao X, Huang L, Wang H, Zhu H, Wu JW, Wang D, Liu D (2011) The Arabidopsis purple acid phosphatase AtPAP10 is predominantly associated with the root surface and plays an important role in plant tolerance to phosphate limitation. Plant Physiol 157:1283–1299
Wang C, Huang W, Ying Y, Li S, Secco D, Tyerman S, Whelan J, Shou H (2012) Functional characterization of the rice SPX-MFS family reveals a key role of OsSPX-MFS1 in controlling phosphate homeostasis in leaves. New Phytol 196:139–148
Wang L, Lu S, Zhang Y, Li Z, Du X, Liu D (2014a) Comparative genetic analysis of Arabidopsis purple acid phosphatases AtPAP10, AtPAP12, and AtPAP26 provides new insights into their roles in plant adaptation to phosphate deprivation. J Integr Plant Biol 56:299–314
Wang S, Zhang S, Sun C, Xu Y, Chen Y, Yu C, Qian Q, Jiang DA, Qi Y (2014b) Auxin response factor (OsARF12), a novel regulator for phosphate homeostasis in rice (Oryza sativa). New Phytol 201:91–103
Wang Z, Ruan W, Shi J, Zhang L, Xiang D, Yang C, Li C, Wu Z, Liu Y, Yu Y, Shou H (2014c) Rice SPX1 and SPX2 inhibit phosphate starvation responses through interacting with PHR2 in a phosphate-dependent manner. Proc Natl Acad Sci USA 111:14953–14958
Wang J, Si Z, Li F, Xiong X, Lei L, Xie F, Chen D, Li Y, Li Y (2015) A purple acid phosphatase plays a role in nodule formation and nitrogen fixation in Astragalus sinicus. Plant Mol Biol 88:515–529
Wang F, Rose T, Jeong K, Kretzschmar T, Wissuwa M (2016) The knowns and unknowns of phosphorus loading into grains, and implications for phosphorus efficiency in cropping systems. J Exp Bot 67:1221–1229
Wang X, Balamurugan S, Liu S-F, Ji C-Y, Liu Y-H, Yang W-D, Jiang L, Li H-Y (2021) Organophosphorus hydrolysis by diatom purple acid phosphatase and sequential regulation of cell metabolism. J Exp Bot 72:2918–2932
Wu P, Shou H, Xu G, Lian X (2013) Improvement of phosphorus efficiency in rice on the basis of understanding phosphate signaling and homeostasis. Curr Opin Plant Biol 16:205–212
Wu W, Lin Y, Liu P, Chen Q, Tian J, Liang C (2018) Association of extracellular dNTP utilization with a GmPAP1-like protein identified in cell wall proteomic analysis of soybean roots. J Exp Bot 69:603–617
Xiao K, Harrison MJ, Wang ZY (2005) Transgenic expression of a novel M. truncatula phytase gene results in improved acquisition of organic phosphorus by Arabidopsis. Planta 222:27–36
Xiao K, Katagi H, Harrison M, Wang ZY (2006) Improved phosphorus acquisition and biomass production in Arabidopsis by transgenic expression of a purple acid phosphatase gene from M. truncatula. Plant Sci 170:191–202
Xie L, Shang Q (2018) Genome-wide analysis of purple acid phosphatase structure and expression in ten vegetable species. BMC Genomics 19:1–2
Xue Y, Zhuang Q, Zhu S, Xiao B, Liang C, Liao H, Tian J (2018) Genome wide transcriptome analysis reveals complex regulatory mechanisms underlying phosphate homeostasis in soybean nodules. Int J Mol Sci 19:2924
Yao MZ, Zhang YH, Lu WL, Hu MQ, Wang W, Liang AH (2012) Phytases: crystal structures, protein engineering and potential biotechnological applications. J Appl Microbiol 112:1–4
Yao Z, Tian J, Liao H (2014a) Comparative characterization of GmSPX members reveals that GmSPX3 is involved in phosphate homeostasis in soybean. Ann Bot 114:477–488
Yao ZF, Liang CY, Zhang Q, Chen ZJ, Xiao BX, Tian J, Liao H (2014b) SPX1 is an important component in the phosphorus signalling network of common bean regulating root growth and phosphorus homeostasis. J Exp Bot 65:3299–3310
Yeung SL, Cheng C, Lui TK, Tsang JS, Chan WT, Lim BL (2009) Purple acid phosphatase-like sequences in prokaryotic genomes and the characterization of an atypical purple alkaline phosphatase from Burkholderia cenocepacia J2315. Gene 440:1–8
Yin C, Wang F, Fan H, Fang Y, Li W (2019) Identification of tea plant purple acid phosphatase genes and their expression responses to excess iron. Int J Mol Sci 20:1954
Younessi-Hamzekhanlu M, Izadi-Darbandi A, Malboobi MA, Ebrahimi M, Abdipour M, Sparvoli F, Paolo D (2018) Agrobacterium rhizogenes transformed soybeans with AtPAP18 gene show enhanced phosphorus uptake and biomass production. Biotechnol Biotechnol Equip 32:865–873
Zamani K, Sabet M, Lohrasebi T, Mousavi A, Malboobi M (2012) Improved phosphate metabolism and biomass production by overexpression of AtPAP18 in tobacco. Biologia 67:713–720
Zamani K, Lohrasebi T, Sabet MS, Malboobi MA, Mousavi A (2014) Expression pattern and subcellular localization of Arabidopsis purple acid phosphatase AtPAP9. Gene Expr Patterns 14:9–18
Zambonelli C, Roberts MF (2003) An iron-dependent bacterial phospholipase D reminiscent of purple acid phosphatases. J Biol Chem 278:13706–13711
Zhang W, Gruszewski HA, Chevone BI, Nessler CL (2008) An Arabidopsis purple acid phosphatase with phytase activity increases foliar ascorbate. Plant Physiol 146:431–440
Zhang Q, Wang C, Tian J, Li K, Shou H (2011) Identification of rice purple acid phosphatases related to phosphate starvation signalling. Plant Biol 13:7–15
Zhang Y, Yu L, Yung KF, Leung DY, Sun F, Lim BL (2012) Over-expression of AtPAP2 in Camelina sativa leads to faster plant growth and higher seed yield. Biotechnol Biofuels 5:19
Zhang H, Uddin MS, Zou C, Xie C, Xu Y, Li WX (2014a) Meta-analysis and candidate gene mining of low-phosphorus tolerance in maize. J Integr Plant Biol 56:262–270
Zhang Y, Wang X, Lu S, Liu D (2014b) A major root-associated acid phosphatase in Arabidopsis, AtPAP10, is regulated by both local and systemic signals under phosphate starvation. J Exp Bot 65:6577–6588
Zhang R, Guan X, Law YS, Sun F, Chen S, Wong KB, Lim BL (2016) AtPAP2 modulates the import of the small subunit of Rubisco into chloroplasts. Plant Signal Behav 11:e1239687
Zhou X, Zha M, Huang J, Li L, Imran M, Zhang C (2017) StMYB44 negatively regulates phosphate transport by suppressing expression of PHOSPHATE1 in potato. J Exp Bot 68:1265–1281
Zhu H, Qian W, Lu X, Li D, Liu X, Liu K, Wang D (2005) Expression patterns of purple acid phosphatase genes in Arabidopsis organs and functional analysis of AtPAP23 predominantly transcribed in flower. Plant Mol Biol 59:581–594
Zhu X, Lee SY, Yang WT, Lee SW, Baek D, Li M, Kim DH (2019) The Burholderia pyrrocinia purple acid phosphatase pap9 mediates phosphate acquisition in plants. J Plant Biol 62:342–350
Zhu S, Chen M, Liang C, Xue Y, Lin S, Tian J (2020) Characterization of purple acid phosphatase family and functional analysis of GmPAP7a/7b involved in extracellular ATP utilization in soybean. Front Plant Sci 11:661
Zhuo S, Clemenson JC, Stone RL, Dixon JE (1994) Mutational analysis of a Ser/Thr phosphatase. Identification of residues important in phosphoesterase substrate binding and catalysis. J Biol Chem 269:26234–26238
Zimmermann P, Regierer B, Kossmann J, Frossard E, Amrhein N, Bucher M (2004) Differential expression of three purple acid phosphatases from potato. Plant Biol 6:519–528
Acknowledgements
JB acknowledges research fellowship from DBT, India and NIPGR. JB acknowledges the help from Shweta Singh in computational work. JG acknowledges the grant from DBT, India, under the Innovative Young Biotechnologist Award (BT/010/IYBA/2016/4) and Swarnajayanti fellowship award (SB/SJF/2019-20/07), DST, India.
Author information
Authors and Affiliations
Contributions
JG conceived the idea. JB collected the literature and wrote the manuscript. JG edited and finalized the manuscript.
Corresponding author
Additional information
Communicated by Neal Stewart.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bhadouria, J., Giri, J. Purple acid phosphatases: roles in phosphate utilization and new emerging functions. Plant Cell Rep 41, 33–51 (2022). https://doi.org/10.1007/s00299-021-02773-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02773-7