Abstract
Key message
The SlTCP26 negatively regulated auxin signal to relieve the apical dominance and suppressed abscisic acid signal to remove the lateral bud dormancy, promoting lateral branches development.
Abstract
Lateral branches formation from lateral buds is a complex regulatory process in higher plants, and the interaction between transcription factors and hormones is indispensable during this process. TCP transcription factors have been reported to regulate lateral branches development, while the detailed function, especially interacting with auxin and ABA during this process, was still ambiguous in tomato. In this study, a branch regulatory gene, SlTCP26, was identified in tomato, and its role along with its interaction to hormones during branch development, as investigated. The results indicated that overexpression of SlTCP26 would promote lateral branches development, and could suppress the expressing of the genes associated with IAA signaling, presenting similar effects in decapitated plants. Conversely, the exogenous IAA application could inhibit the expression of SlTCP26. Furthermore, the expressing of the ABA signaling-related genes was inhibited in SlTCP26 overexpressed tomato, similar to that in decapitated tomato. Our findings suggested that SlTCP26 may be a crucial adjuster for synergistic action between ABA and IAA signals during the development of lateral branches, and it could promote the lateral buds grow into lateral shoots, via inhibiting IAA signal to relieve the apical dominance and suppressing ABA signal to remove the lateral bud dormancy. Our study provided some insights for the development of tomato lateral branches to understand the apical dominance regulatory network.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Branches development plays an important role in plant morphogenesis, and proper branching can improve the light harvesting efficiency and promote the exchange of plant with environmental substances, subsequently improving the yield (Mar et al. 2011; Ferreira et al. 2018). At present, molecular mechanism of plant lateral branching has been gradually described in Arabidopsis (Finlayson 2007; Aguilar-Martinez et al. 2007), peas (Braun et al. 2012), rice (Hu et al. 2003), Brassica juncea cv. (Shikha et al. 2019), Medicago truncatula (Yin et al. 2020), and other plants. For example, pea brc1 mutant displayed an increased branching phenotype, and the branches development could be regulated coordinately by the strigolactone (SL) and auxin (IAA) (Beveridge 2006; Braun et al. 2012). In rice, OsTB1 gene was expressed with high level in axillary buds and involved in tiller of rice (Hu et al. 2003).
Commonly, main shoot has the apical dominance in higher plants and it could inhibit the outgrowth of lateral branches. When the main shoot was damaged or decapitated, the axillary buds were able to grow into lateral branches (Thimann and Skoog 1934). It was reported that plant hormones were involved in the development of plant lateral buds and the formation of lateral branches (Wang et al. 2019; Zhao et al. 2019; Liu et al. 2020). Among the plant hormones, IAA was found to be synthesized mainly in the shoot apex, and acted as long-distance signal to suppress branching (Leyser 2003). If the auxin content in lateral buds was decreased by decapitating or destroying the main shoot apex, the lateral buds could develop into lateral branches (Thimann and Skoog 1934). Another important phytohormone, abscisic acid, may promote dormancy and inhibit the development of lateral buds (Yiliham-nur et al. 2000). When the ABA signaling was weakened, lateral buds began to grow into lateral branches (Tucker 1977). It was suggested that IAA could induce and maintain high levels of ABA in plant tissues, which may be the reason for growth inhibition of lateral buds (Tucker 1978).
TCP transcription factors, angiosperm-specific transcription factor, functioned in regulating plant growth and development, such as the floral symmetry (Chapman et al. 2012), the differentiation of trichome (Vadde et al. 2018), the photomorphogenesis of shoot apex (López-Juez et al. 2008), bud branching, and the number of lateral branches (Bai et al. 2012; Danisman et al. 2012; Mar et al. 2011). It has been reported that the TCP transcription factors acted importantly in hormones signal pathway (Zhou et al. 2017). In rice, OsTB1 gene, as a TCP family member, was identified as a repressor of lateral organ growth, especially on the formation of branches, and its expression was negatively correlated with tiller level (Finlayson 2007). In Arabidopsis, AtTB1 gene was found to inhibit bud development of plant, while AtTCP12 (BRC2) and AtTCP18 (BRC1) could negatively regulate the morphological formation of lateral branches, and the latter two genes were most closely related to former one (Aguilar-Martinez et al. 2007). The OsTCP19 gene in rice has been reported to regulate the ABA signaling network, through direct interacting with the transcription factor ABI4 (Mukhopadhyay and Tyagi 2015). In tomato, AtTCP1 could actively regulate the biosynthesis of BR by regulating the expression of DWF4 (Guo et al. 2010).
Up to date, about 30 TCP transcription factor genes have been identified in tomato, named as SlTCP (Mar et al. 2011; Parapunova and Busscher 2014). It was reported that SlTCP12, SlTCP15, and SlTCP18 were mainly expressed in the tomato fruit, suggesting their important role in fruit development or ripening (Parapunova and Busscher 2014). In previous report by Mar et al. (Mar et al. 2011), the down-regulation of SlBRC1 could increase number of tomato branches, but the detailed function, especially interacting with plant hormone during this process, was still unclear in tomato. In this study, another TCP transcription factor gene, SlTCP26, was identified with positively promoting effect on tomato branching, so the expression of this gene to exogenous hormones, decapitation, as well as interference and overexpression transgenic plants were analyzed, to investigate its function in tomato branch development, and to understand the relationship of SlTCP26, IAA, ABA, and branch development.
Materials and methods
Plant materials and growth conditions
Tomato plants (Solanum lycopersicum, Micro Tom) were grown in a greenhouse under conditions: 25 ± 2 °C for 16 h in light, 20 ± 2 °C for 8 h in dark, 250 m mol m−2 s−1 light intensity, 80% humidity.
The tomato tissues for experiment contains: root, stem, stem + bud1-2, Stem + bud3-4, Stem + bud5-6, leaf, flower, immature green fruit (IMG), mature green fruit (MG), breaker fruit (Br), and red fruit (Red). All samples collected were immediately frozen in liquid nitrogen and then kept in a refrigerator at – 80 ℃ before use.
Bioinformatics and homology analysis
The sequence of SlTCP26 gene (Solyc03g045030.1) was downloaded from tomato database (https://solgenomics.net/). ORF Finder of NCBI database was used to analyze the sequence of SlTCP26 gene and predict its coding amino acid sequence. 30 TCP protein sequence from tomato was compared with all 24 A. thaliana TCP proteins and 10 potato TCP proteins in MEGA v6.0, and phylogenetic reconstruction was obtained by NJ (neighbor-joining) method.
Cloning of SlTCP26 and plant transformation
Using the cDNA mixture of each tomato tissue as a template, the full length of the SlTCP26 gene was amplified by fast Pfu polymerase and the purification target band was described according to the OMEGA (United States) gel extraction kit.
To construct the RNAi vector and overexpression vector, the positive and negative interference fragments of SlTCP26 gene were amplified using interference fragment primers and cDNA as templates. The positive fragments of SlTCP26 gene were connected to pcambia-1301 vector. The positive transformant of SlTCP26-RNAi was obtained by transforming the positive interference vector into E. coli. Then, the reverse fragment of SlTCP26 gene was linked to the pcambia-1301-SlTCP26 positive ligation vector and transformed into E. coli to obtain transgenic plants. The full-length fragment of SlTCP26 gene with restriction site was linked to pLP100-35S overexpression vector and transformed into E. coli to obtain transgenic plants. Then, GUS staining and direct PCR were performed to verify the transgenic positive plants. The seeds of T0 lines were screened in half-strength Murashige and Skoog medium with kanamycin (100 mg L–1). The expression efficiency of SlTCP26-RNAi and SlTCP26 overexpression were detected by qRT-PCR. Homozygous lines of the T2 generation were used for the following experiments.
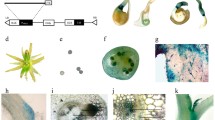
β-Glucosidase (GUS) solution stains transgenic and wild leaves were incubated (0.1% Triton 5-bromo-4-chloro-3-indolyl-β-d-glucuronic acid; PH 7.2, 10 mM EDTA) at 37 ℃ overnight. After staining, samples were decolorized using a series of graded ethanol solution. Direct PCR was performed using pyrolysis liquid from transgenic and WT plant leaves as templates with the Takara (Japan) PCR kit according to the manufacturer's instructions.
Real-time quantitative PCR analysis
The total RNA from root, stem, leaf, node, flower, fruit of transgenic, and WT plant was extracted by a EZNA® Plant RNA Kit(America), and cDNA was synthesized via the TAKARA Reverse Transcription Kit (Japan) according to the manufacturer’s instructions. qRT-PCR was carried out following the Bio-Rad protocol in a Bio-Rad CFX96 real-time PCR detection system. Relative expression levels were calculated based on the 2−ΔΔCT method.
Histological analysis
Stem fragments of wild and SlTCP26 overexpression transgenic tomato plants were fixed in formalin–acetic acid–alcohol solution (50% ethanol, 5% acetic acid and 3.7% methanol) for 24 h. Dehydration was performed in a series of ethanol solutions with increasing concentration (30, 50, 70, 85, 95, and 100%). Xylene was used for transparentizing tissues. Then, samples were embedded in paraffin, and cut into sections with 5 mm thickness using a microtome. The sections were dewaxed in xylene, and rehydrated in ethanol solutions with decreasing concentration (100, 95, 85, 70, 50, and 30%). Dyeing with 1% toluidine blue solution and dehydration was performed in a series of ethanol solutions with increasing concentration (70, 85, 95, and 100%). Resinene was used for sealing the slide, for observation under a Motic-OLYMPUS BX51 microscope.
Hormone treatment
ABA solution (100 μmol L–1), IAA solution (50 μmol L–1), CTK solution (50 μmol L–1), and GA3 solution (50 μmol L–1) were sprayed on several healthy wild-type seedlings with the same size after growth for 35 days, separately (Fujita et al. 2010). Leave materials were collected as sample at 3, 6, 9, 12, 24, 48, and 72 h, respectively, after treatment.
Plant top removal
The top above the fourth node of wild-type plants were cut-off after the first flower bloomed. The partial stem along with petiole base at each node from No.1 to No.4 of the decapitated plants were collected 8, respectively, h after decapitation for qRT-PCR analysis, using the same materials from untreated plants as control.
Results
Identification and sequence analysis of tomato SlTCP26 gene
By mining the annotated tomato genome database (http://solgenomics.net/), totally, 30 TCP family genes currently identified in tomato, and the TCP gene Solyc03g045030.1 was designated as SlTCP26. Using the neighbor connection method on MEGA6, phylogenetic analysis was performed on the amino acid sequences of 30 TCP genes in tomato, along with 24 TCP genes from A. thaliana and 10 from potato, respectively, to investigate their relationship (Fig. 1). The results showed that SlTCP26 was closely related to AtTCP1 gene, the latter mainly regulating the biosynthesis of brassinolide and expressing preferentially in the later stages of axillary bud meristem development (Cubas et al. 2001; Guo et al. 2010).
Expression pattern of SlTCP26 in tomato
To investigate the role of SlTCP26 in tomato, its expression pattern in different tissues of tomato was analyzed. The results showed that SlTCP26 may be transcribed in all tissues, while it presented different levels in various tissues (Fig. 2a). This gene was tested with relative high levels in roots, leaves, developing flowers, and fruits, and the optimal expression was observed in stem and lateral buds, with about 4–7 times than in root; however, it expressed poorly in breaker fruit.
SlTCP26 expression in transgenic plants and wild type. a Expression of SlTCP26 in different tomato tissues. R Roo, S Ste, Sb1 Stem + bud1-, Sb2 Stem + bud3-, Sb3 Stem + bud5-, L Lea, F Flower, IMG immature green frui, MG mature green frui, Br breaker stage frui, Red red fruit. All data representing mean ± SD of three biological replicates. b Relative expression level of SlTCP26 in RNAi lines. c Relative expression level of SlTCP26 in overexpression lines (*indicating significance level by t test, *P < 0.05, **P < 0.01)
Number increase of branches by overexpression of SlTCP26
To identify the function of SlTCP26, overexpression lines and RNAi lines of this gene were constructed, based on the Micro-Tom tomato genetic background, and the phenotype of the transgenic plants was analyzed. Comparatively, SlTCP26 presented lower expressing level in RNAi transgenic plants (Fig. 2b), but much higher in overexpression lines (Fig. 2c), than in wide plants, respectively. As shown in our data, the alteration of the branch number was not observed in the SlTCP26-silencing plants, by comparing with wild type, whereas the branches of the SlTCP26-overexpressing plants were significantly more in number than that of the wild type (Fig. 3a–c). In the overexpression lines, axillary buds occurred in the leaf axil of the transgenic plant at seedling stage, but not in the wild type. After 1 week of the first flower blooming, the lateral branches on the main stem were measured, with an average of above 0.5 cm in length. The results showed that overexpression lines of SlTCP26 possessed increased branching, compared with wide plants. About an average of 6, 5, and 3 lateral branches differentiated from the axillary of the main stem were recorded in Line OE34, Line OE6, and Line OE10 plants, respectively (Fig. 3d). Comparatively, phloem tissue in SlTCP26 overexpressed tomato plants was thicker, and the vascular tissue cells were larger than those of wild type (Supplementary Fig. 6).
Phenotypes comparison between wild-type (WT) and SlTCP26 transgenic plants. a Comparison between wild type, RNAi lines, and overexpression line. b Comparison between wild type and overexpression line OE34, OE6, OE10 at mature stage. c Comparison between wild type and overexpression Line OE34, OE6, OE10 after leaf removal. d Number of branches of wild type and overexpression Lines OE34, OE6, OE10 (*representing significant difference between trangenic group and control group by t test, *P < 0.0, **P < 0.01)
Impact of exogenous hormones on the SlTCP26 expression
Hormones were considered as the crucial factors during plant development. To detect the effects of hormones on SlTCP26 expressing in tomato, the express level of this gene was measured after treating wide plants with CTK, ABA, IAA, and GA3, respectively. The results showed that SlTCP26 expressing significantly increased after 1 h of CTK treatment, and the optimum level was obtained after 2 h of treatment, with approximately 15 times higher than control. Afterward, its expression gradually decreased with time extension (Fig. 4a). For exogenous ABA application, the expression of this gene gradually improved after treatment, till to 30 h (Fig. 4b). However, the application of exogenous IAA significantly inhibited its expressing. After 3 h of IAA treatment, the SlTCP26 gene presented with only the half expression level of the control plants, and even only about 1/58 of control was achieved at 30 h after IAA treatment (Fig. 4c). In addition, GA3 treatment resulted in a reduced expression of SlTCP26 within 2 h of treatment, and then, the gene expressed with fluctuant increasing trend with time prolongation (Fig. 4d). These results indicated that several exogenous hormones could impact the SlTCP26 expression in different degrees. Especially, of interest, the expression of SlTCP26 was promoted by ABA, but inhibited by IAA. Therefore, it was suggested that the development process of tomato regulated by SlTCP26 was associated intensively with the hormone signals, especially with ABA and IAA.
Impact of various stress and hormone treatment on expression of tomato SlTCP26 gene. a Relative expression level of SlTCP26 under CTK treatment. b Relative expression level of SlTCP26 under ABA treatment. c Relative expression level of SlTCP26 under IAA treatment. d Relative expression level of SlTCP26 under GA3 treatment (*representing significant difference between treatment group and control group by t test, *P < 0.05, **P < 0.01)
Overexpression of SlTCP26 affecting transcription of genes associated with auxin IAA signaling
As SlTCP26 was previously detected sensitive to exogenous IAA, the expression of 12 auxin signaling-related core genes was tested in wild type and three lines of SlTCP26 overexpression plants leaves (Leyser 2006). The results showed that the expressions of the tested auxin response genes, including SlARF5, SlARF6, SlARF7, and SlARF8, were all inhibited in the overexpressed transgenic plants, but they showed various inhibition extent. Comparatively, SlARF5, SlARF6, and SlARF7 were inhibited intensively in Line OE6, OE10, and OE34 plants, with silence about 80–90%, compared with wild type, while SlARF8 expressed with only about 50% level of the wild type (Fig. 5a). Concerning to auxin transport-related genes, including SlPIN1, SlPIN3, SlPIN5, and SlPIN6, their expressions were also all significantly inhibited by the SlTCP26 overexpression (Fig. 5b). Furthermore, IAA transcriptional regulator genes, including SlIAA2, SlIAA4, SlIAA7, and SlIAA9, were all detected with significantly decreased expression in overexpression transgenic plants (Fig. 5c). These results implied SlTCP26 gene negatively regulated auxin signal-related genes during the tomato development.
Overexpression of SlTCP26 inhibiting expression of genes related to ABA signal
Based on the above results of the sensitivity of the SlTCP26 gene to exogenous ABA, the effect of SlTCP26 overexpression on the expression of ABA signal transduction and response-related core genes was detected (Hauser et al. 2011). The results showed that a large number of ABA signal transduction-related genes were suppressed in SlTCP26 overexpressed plants. Among them, the core components of ABA signal transduction SnRK2 genes, including SAPK2, SAPK3, SnRK2C, and SnRK2I, were strongly suppressed in SlTCP26 overexpressed transgenic tomato plants. Meanwhile, SlABF4, a key gene of ABA signal response, was also strongly inhibited, while the ABA signal transduction gene SnRK2B and ABA signal response gene ABI5-LIKE2 (1) did not present significant difference from the wild type (Fig. 6). These results indicated that the SlTCP26 gene may be involved in the inhibition or negative regulation of ABA signaling during the development of tomato plant lateral branches.
Expression level of SlTCP26 and lateral branch length in decapitated plants
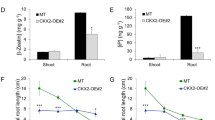
In general, the apical dominance may inhibit axillary bud origin and lateral branch growth. In this experiment, tomato plants were decapitated to investigate the effect of apical dominance on the lateral branch growth, as well as SlTCP26 expression. It was found that the branches from 1 to 4 nodes were recorded as 1.24, 2.78, 3.38, and 4.96 cm in length, respectively, on the decapitated plants, whereas, the axillary bud length in the control plants was not observed (Fig. 7d). Meanwhile, SlTCP26 gene was found expressing with un-regulation after decapitation, especially more intense in node 1 and node 4 of the decapitated plants (Fig. 7c). These results indicated that apical dominance not only inhibited the development of lateral buds but also suppressed the expression of SlTCP26, suggesting a potential connection between the gene expression of SlTCP26 and the lateral bud development.
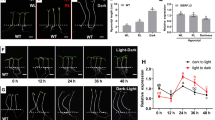
SlTCP26 response to decapitation. a Schematic representation of a tomato plant with node numbers as mentioned in text and figures. b Detail of nodes 3 and 4 of intact (left) and decapitated (right) tomato plants. One week after decapitation, lateral branches grown out in decapitated plants (right), while axillary buds of intact plants remained arrested (left). Red arrow and numbers indicate node positions; yellow straight line indicates decapitation site. c SlTCP26 expression levels in axillary buds of decapitated plants 8 h after treatment, relative to levels in buds of intact plants, analyzed by qRT-PCR. Data representing mean ± SD of three biological replicates. d Lateral branch length in intact and decapitated plants 1 week after the treatment (*representing significant difference between treatment group and control group by t test, *P < 0.05, **P < 0.01) (color figure online)
The transcription of genes associated with IAA pathway in decapitated plants
Previous results in this experiment showed that in SlTCP26 overexpressed tomatoes, the expression of genes related to the IAA signal pathway was down-regulated. To further verify the relationship between tomato lateral bud development and IAA signal pathway, the expression changes of auxin response factors, auxin transport-related genes, and Aux/IAA transcriptional regulators were analyzed during the lateral bud development of tomato after decapitation. The results showed that numerous auxin-related genes were down-regulated, including auxin response gene SlARF5, SlARF6, SlARF7, SlARF8, auxin transport gene SlPIN1, SlPIN3, SlPIN5, SlPIN6 and IAA family gene SlIAA2, SlIAA4, SlIAA7, and SlIAA9 (Fig. 8). This is consistent with the achieved results in transgenic tomatoes, which further proved that the regulation of tomato lateral branches development is associated with IAA signaling, and also indicated that the SlTCP26 gene is associated with IAA signaling in regulating tomato lateral branch development.
The transcription of genes associated with ABA pathway in decapitated plants
As the lateral branches in decapitated tomatoes presented more in number than the non-decapitated plants, to further investigate the relation between the occurrence of tomato lateral buds and ABA signal, the expression changes of ABA signal transduction and response genes were tested during the development of lateral buds after removing apical dominance. The results showed that numerous ABA signal transduction and response-related genes were expressed with down-regulation during the process of lateral bud generation after apical removal, including the SAPK2, SAPK3, SnRK2B, SnRK2C, and SnRK2I in the core original SnRK2 family of ABA signal transduction pathway and the ABI5-LIKE2 (1) and SlABF4 in the ABA signal response (Fig. 9). This is also in consistence with the obtained results in transgenic tomatoes. These results proved that the ABA signaling would be involved in regulating the development of tomato lateral branch, and SlTCP26 gene may be associated with the inhibition or negative regulation of ABA signal in regulating the development of lateral branches of tomato plants.
Overexpression of SlTCP26 light reducing tomato fruit yield
To explore the effect of increased lateral branches due to SlTCP26 overexpression on tomato fruit yield, the fruit set per plant, weight of single fruit, and yield per plant were measured on the overexpression transgenic and wild-type plants, respectively. Line OE6 and Line OE34 presented only few fruit sets per plant, while that of Line OE10 was significantly reduced compared with wild type (Fig. 10a). There was no significant change in the single fruit weight of all overexpression transgenic plant lines compared with wild type (Fig. 10b). By comparing with wild type, the fruit yield per plant of Line OE6 did not change significantly, while the yield per plant of Line OE10 and Line OE34 decreased significantly (Fig. 10c). These interesting results indicated that the increase of lateral branches did not lead the yield raise in transgenic tomato.
Yield of tomato fruit of overexpression of SlTCP26. a Fruit set of per plant between wild type and line OE34, OE6, OE10. b Weight of single fruit between wild type and line OE34, OE6, OE10. c Fruit yield per plant between wild type and line OE34, OE6, OE10 (*representing significant difference between treatment group and control group by t test, *P < 0.05, **P < 0.01)
Discussion
Plant branching is an important developmental process, and the genes, hormones, and environment could affect this process from axillary meristems to lateral branches. When the leaf primordia form on the meristem, a small and slowly dividing cell boundary area was formed, and the axillary meristem formed in this area. When axillary meristems were formed, they could grow or enter in a dormant state, usually forming axillary buds (Janssen et al. 2014). Once the main shoot tip was damaged or decapitated by herbivorous or pruning, the axillary buds should relief their dormancy, and then develop into lateral branches (Thimann and Skoog 1934). During this process, the regulation of the gene regulatory networks was indispensible for the development for branches. It was reported that the pea TCP transcription factor, PsBRC1, acted as a downstream element of strigolactones to control branching (Braun et al. 2012). In Arabidopsis, TCP family gene AtTCP12 and AtTCP18 functioned crucially in the development of lateral branches (Aguilar-Martinez et al. 2007). SlBRC1a and SlBRC1b were expressed in arrested axillary buds and both are down-regulated upon bud activation in tomato (Mar et al. 2011).
In this study, MIC-TOM tomato was used to investigate the function of the TCP family gene SlTCP26 during the tomato lateral branch development (Fig. 1). As a result SlTCP26 gene was found mainly expressing in the axillary bud, with significantly higher level than that in other tissues of tomato (Fig. 2a). The overexpression of SlTCP26 could induce the emergence of multiple lateral branches in tomato plants (Fig. 3d). These results indicated that the SlTCP26 gene is involved in the outgrowth of tomato axillary buds, opposite to AtTCP12 and AtTCP18 in Arabidopsis, playing a positive regulatory role.
As known, in the apical dominance, the main shoot could produce high concentrations of inhibitory hormone, auxin, moving downwards within the stem, and subsequently inhibiting the development of lateral branches from axillary buds below (Barbier et al. 2017). In this study, the expression of SlTCP26 was strongly inhibited by exogenous IAA, and the inhibitory action was more intense with time prolongation after treatment (Fig. 4c). The results implied that high concentrations of IAA could not only inhibit the lateral branches development, but also affect the expression of SlTCP26 in tomato. Furthermore, the expression of auxin signal-related genes were found significantly down-regulated in SlTCP26 overexpression plants compared with wild type (Fig. 5). These results proved that SlTCP26 gene overexpression could reduce the inhibitory effect of auxin on apical dominance, similar to the effect of apical dominance removal after mechanical damage to the apical tip of plant. Furthermore, the expression of genes related to the IAA signal pathway was down-regulated in the four nodes of the decapitated tomato plant (Fig. 8). These results verified an inhibitory interaction between the SlTCP26 gene and auxin in tomato apical dominance. Previous reports showed that the expressions of PsIPT1 and PsIPT2 genes were upregulated, and the auxin content was reduced, suggesting that their interaction in the apical dominance may jointly regulate the development of lateral branches (Tanaka et al. 2010). Similar results obtained in this study demonstrated that the SlTCP26 gene could interact with IAA signal in the apical dominance of tomato, and this gene may regulate IAA signal process, as well as the dormancy of lateral buds and development of lateral branches.
Commonly, the formation of lateral branches was inhibited due to the dormancy of lateral buds in plant (Shimizu-Sato and Mori 2001; Horvath et al. 2003). Concerning to plant hormones affecting lateral bud development, ABA presented mainly inhibition on the lateral buds by maintaining their dormancy. If the ABA signal was weakened, the dormancy of the lateral buds could be relieved, and the buds should develop into lateral branches promptly (Tucker 1977; Cline and Oh 2006). In this study, the overexpression of SlTCP26 significantly inhibited the expression of numerous ABA signal transduction and response-related genes (Fig. 6), indicating that the SlTCP26 gene may require suppressing the ABA signal to achieve the relief of lateral bud dormancy during the tomato lateral branch development. In the decapitated tomatoes, the lateral branches increased in number significantly, and most ABA signal transduction and response-related genes were significantly down-regulated (Fig. 9). A similar dormancy-releasing effect was presented when SlTCP26 gene overexpression. Therefore, the ABA signal would be involved in the regulation of lateral bud dormancy in apical dominance formation, and SlTCP26 gene could promote lateral buds develop into lateral branches, via suppressing ABA signal and removing lateral bud dormancy, after plant apex was mechanical damaged.
The development of plant lateral branches is a complex regulatory process. The synergistic effect of ABA and IAA signals plays a crucial role in this process (Galoch et al. 1998; Cline and Oh 2006). Perhaps, the SlTCP26 may play a crucial role, as an adjuster of synergistic action between ABA and IAA signals, in regulating the development of lateral branches. On one hand, the SlTCP26 gene inhibited the expression of IAA-related genes, to relieve the apical dominance. On the other hand, it suppressed the ABA signal to accelerate the removal of dormancy from lateral buds, subsequently facilitate the differentiation of lateral buds into lateral branches. Maybe, as a more developed phloem and vascular tissue was also conducive to the supply of nutrients and energy (Supplementary. Fig. S6), beneficial for development of lateral branches (Wang et al. 2005; Golan et al. 2013), SlTCP26 can promote the differentiation of vascular tissue, correspondingly enhancing the ability of the nutrient transport to facilitate the axillary buds convert into lateral branches in tomato. However, the increase of lateral branches did not lead to the increase of the tomato yields (Fig. 10), which needs further research.
Conclusion
In this study, the function of SlTCP26 was analyzed in lateral branches development in tomato, using exogenous hormones, decapitation, as well as interference and overexpression transgenic transformation technology. Overexpression of SlTCP26 would promote lateral branches development. Exogenous IAA application may inhibit SlTCP26 expressing, and this gene could, in turn, down-regulate the gene expression associated with IAA signaling, similar to decapitated plants. Furthermore, the expression of ABA signal-related genes could be down-regulated in SlTCP26 overexpressed tomatoes, and ABA signal showed similar inhibiting effects in decapitated tomato. Our findings suggested that SlTCP26 may be a crucial adjuster for synergistic action between ABA and IAA signals during the development of lateral branches and it could promote the lateral buds grow into lateral branches via inhibiting IAA signal to relieve the apical dominance and suppressing ABA signal to remove the lateral bud dormancy. Our finds would provide some insights into the regulation of lateral branches formation in tomato.
Change history
04 May 2021
A Correction to this paper has been published: https://doi.org/10.1007/s00299-021-02694-5
References
Aguilar-Martinez JA, Poza-Carrion C, Cubas P (2007) Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell 19:458–472. https://doi.org/10.1105/tpc.106.048934
Bai F, Reinheimer R, Durantini D, Kellogg EA, Schmidt RJ (2012) TCP transcription factor, BRANCH ANGLE DEFECTIVE 1 (BAD1), is required for normal tassel branch angle formation in maize. P Natl Acad Sci USA 109:12225–12230. https://doi.org/10.1073/pnas.1202439109
Barbier FF, Dun EA, Beveridge CA (2017) Apical dominance. Curr Biol 27:864–865. https://doi.org/10.1016/j.cub.2017.05.024
Beveridge CA (2006) Axillary bud outgrowth: sending a message. Curr Opin Plant Biol 9:35–40. https://doi.org/10.1016/j.pbi.2005.11.006
Braun N, Saint GA, Pillot JP, Boutet-Mercey S, Dalmais M, Antoniadi I, Li X, Maia-Grondard A, Le Signor C, Bouteiller N, Luo D, Bendahmane A, Turnbull C, Rameau C (2012) The pea TCP transcription factor PsBRC1 acts downstream of strigolactones to control shoot branching1. Plant Physiol 158:225–238. https://doi.org/10.1104/pp.111.182725
Chapman MA, Tang S, Draeger D, Nambeesan S, Shaffer H, Barb JG, Knapp SJ, Burke JM (2012) Genetic analysis of floral symmetry in van Gogh’s sunflowers reveals independent recruitment of CYCLOIDEA genes in the asteraceae. PLoS Genet 8:1–10. https://doi.org/10.1371/journal.pgen.1002628
Cline MG, OH C, (2006) A reappraisal of the role of abscisic acid and its interaction with auxin in apical dominance. Ann Bot Lond 98:891–897. https://doi.org/10.1093/aob/mcl173
Cubas P, Coen E, Zapater JMM (2001) Ancient asymmetries in the evolution of flowers. Curr Biol 11:1050–1052. https://doi.org/10.1016/S0960-9822(01)00295-0
Danisman S, Van DWF, Dhondt S, Waites R, De FS, Bimbo A, Dijk ADJV, Muino JM, Cutri L, Dornelas MC, Angenent GC, Immink RGH (2012) Arabidopsis class I and class II TCP transcription factors regulate jasmonic acid metabolism and leaf development antagonistically. Plant Physiol 159:1511–1523. https://doi.org/10.1104/pp.112.200303
Ferreira AS, Zucareli C, Werner F, Balbinot J (2018) Plant spatial arrangement affects grain production from branches and stem of soybean cultivars. Bragantia 77:567–576. https://doi.org/10.1590/1678-4499.2017285
Finlayson SA (2007) Arabidopsis TEOSINTE BRANCHED1-LIKE 1 regulates axillary bud outgrowth and is homologous to monocot TEOSINTE BRANCHED1. Plant Cell Physiol 48:667–677. https://doi.org/10.1093/pcp/pcm044
Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LSP, Yamaguchi-Shinozaki K, Shinozaki K (2010) A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signaling pathway. Plant J 39:863–876. https://doi.org/10.1111/j.1365-313x.2004.02171.x
Galoch E, Marlena Z, Burkacka-Aukajtys E (1998) The effect of decapitation on the levels of IAA and ABA in the lateral buds of Betula pendularoth. Acta Physiol Plant 20:399–403. https://doi.org/10.1007/s11738-998-0026-0
Golan G, Betzer R, Wolf S (2013) Phloem-specific expression of a melon Aux/IAA in tomato plants alters auxin sensitivity and plant development. Front Plant Sci 4:329. https://doi.org/10.3389/fpls.2013.00329
Guo Z, Fujioka S, Blancaflor EB, Miao S, Gou X, Li J (2010) TCP1 modulates brassinosteroid biosynthesis by begulating the expression of the key biosynthetic gene DWARF4 in Arabidopsis thaliana. Plant Cell 22:1161–1173. https://doi.org/10.1105/tpc.109.069203
Hauser F, Waadt R, Schroeder JI (2011) Evolution of abscisic acid synthesis and signaling mechanisms. Curr Biol 21:346–355. https://doi.org/10.1016/j.cub.2011.03.015
Horvath DP, Anderson JV, Chao WS, Foley ME (2003) Knowing when to grow: signals regulating bud dormancy. Trends Plant Sci 8:534–540. https://doi.org/10.1016/j.tplants.2003.09.013
Hu W, Zhang S, Zhao Z, Sun C, Zhao Y, Luo D (2003) The analysis of the structure and expression of OsTBl gene in rice. Acta Photophysiologica Sinica 29:507–514. https://doi.org/10.3321/j.issn:1671-3877.2003.06.005
Janssen BJ, Drummond RS, Snowden KC (2014) Regulation of axillary shoot development. Curr Opin Plant Biol 17:28–35. https://doi.org/10.1016/j.pbi.2013.11.004
Leyser O (2003) Regulation of shoot branching by auxin. Trends Plant Sci 8:541–545. https://doi.org/10.1016/j.tplants.2003.09.008
Leyser O (2006) Dynamic integration of auxin transport and signalling. Curr Biol 16:424–433. https://doi.org/10.1016/j.cub.2006.05.014
Liu L, Wu Y, Zhao D, Tao J (2020) Integrated mRNA and microRNA transcriptome analyses provide insights into paclobutrazol inhibition of lateral branching in herbaceous peony. Biotech 10:1–9. https://doi.org/10.1007/s13205-020-02489-7
López-Juez E, Dillon E, Magyar Z, Khan S, Hazeldine S, De Jager SM, Murray JAH, Beemster GTS, Bogre L, Shanahan H (2008) Distinct light-initiated gene expression and cell cycle programs in the shoot apex and cotyledons of Arabidopsis. Plant Cell 20:947–968. https://doi.org/10.1105/tpc.107.057075
Mar MT, Grandío EG, Serra F, Marcel F, Rodríguez-Buey ML, Schmitz G, Theres K, Bendahmane A, Dopazo H, Cubas P (2011) Role of tomato BRANCHED1-like genes in the control of shoot branching. Plant J 67:701–714. https://doi.org/10.1111/j.1365-313X.2011.04629.x
Mukhopadhyay P, Tyagi AK (2015) OsTCP19 influences developmental and abiotic stress signaling by modulating ABI4-mediated pathways. Sci Rep 5:9998. https://doi.org/10.1038/srep09998
Parapunova V, Busscher M, Busscher-Lange J, Lammers M, Karlova R, Bovy AG, Angenent GC, Maagd RA (2014) Identification, cloning and characterization of the tomato TCP transcription factor family. BMC Plant Biol 14:157. https://doi.org/10.1186/1471-2229-14-157
Shikha T, Tanu S, Anupama S, Pratiksha M, Shivaraj SM, Prateek S, Anandita S (2019) SUPPRESSOR OF OVEREXPRESSION OF CONSTANS1 influences flowering time, lateral branching, oil quality, and seed yield in Brassica juncea cv. Varuna. Funct Integr Genomic 19:43–60. https://doi.org/10.1007/s10142-018-0626-8
Shimizu-Sato S, Mori H (2001) Control in outgrowth and dormancy in axillary buds. Plant Physiol 127:1405–1413. https://doi.org/10.2307/4280206
Tanaka M, Takei K, Kojima M, Sakakibara H, Mori H (2010) Auxin controls local cytokinin biosynthesis in the nodal stem in apical dominance. Plant J 45:1028–1036. https://doi.org/10.1111/j.1365-313x.2006.02656.x
Thimann KV, Skoog F (1934) On the inhibition of bud development and other functions of growth substance in Vicia faba. P R Soc B Biol Sci 114:317–339. https://doi.org/10.1098/rspb.1934.0010
Tucker DJ (1977) Hormonal regulation of lateral bud outgrowth in the tomato. Plant Sci Lett 8:105–111. https://doi.org/10.1016/0304-4211(77)90019-0
Tucker DJ (1978) Apical dominance in the tomato: the possible roles of auxin and abscisic acid. Plant Sci Lett 12:273–278. https://doi.org/10.1016/0304-4211(78)90078-0
Vadde BVL, Challa KR, Nath U (2018) The TCP4 transcription factor regulates trichome cell differentiation by directly activating GLABROUS INFLORESCENCE STEMS in Arabidopsis thaliana. Plant J 93:259–269. https://doi.org/10.1111/tpj.13772
Wang H, Jones B, Li Z, Frasse P, Delalande C, Regad F, Chaabouni S, Latche A, Pech JC, Bouzayena M (2005) The tomato aux/IAA transcription factor IAA9 is involved in fruit development and leaf morphogenesis. Plant Cell 17:2676–2692. https://doi.org/10.1105/tpc.105.033415
Wang P, Zhang S, Qiao J, Sun Q, Shi Q, Cai C, Mo J, Chu Z, Yuan Y, Du X, Miao Y, Zhang X, Cai Y (2019) Functional analysis of the GbDWARF14 gene associated with branching development in cotton. Peer J 7:e6901. https://doi.org/10.7717/peerj.6901
Yiliham-nur MH, Omori H, Hirano T (2000) Effect of four plant hormones on growth, flowering and seed-setting in buckwheat (fagopyrum esculentum moench). Rep Tokai Branch Crop Sci Soc Jpn 6:19–26
Yin P, Ma Q, Wang H, Feng D, Wang X, Pei Y, Wen J, Tadege M, Niu L, Lin H (2020) SMALL LEAF AND BUSHY1 controls organ size and lateral branching by modulating the stability of BIG SEEDS1 in Medicago truncatula. New Phytolo 226:1399–1412. https://doi.org/10.1111/nph.16449
Zhao J, Gao P, Li C, Lin X, Guo X, Liu S (2019) PhePEBP family genes regulated by plant hormones and drought are associated with the activation of lateral buds and seedling growth in Phyllostachys edulis. Tree Physiol 39:1387–1404. https://doi.org/10.1093/treephys/tpz056
Zhou Y, Zhang D, An J, Yin H, Fang S, Chu J, Zhao Y, Li J (2017) TCP transcription factors regulate shade avoidance via directly mediating the expression of both phytochrome interacting factors and auxin biosynthetic genes. Plant Physiol. https://doi.org/10.1104/pp.17.01566
Acknowledgements
This work is financially supported by National Natural Science Fund of China (31171587), Innovation Team Program of Sichuan Education Department (16TD0020), and Talent Fund of China West Normal University (17YC354).
Author information
Authors and Affiliations
Contributions
DL and XYW designed and completed the experiment, collected and analyzed the data, and XYW wrote and revised the manuscript. JY involved in experimental design and manuscript editing and JZ involved in manuscript editing. HF and GQW contributed new reagents. ZYH analyzed data. WJZ and ZAY involved in experimental operation process. All authors approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Communicated by Xian Sheng Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wei, X., Yang, J., Lei, D. et al. The SlTCP26 promoting lateral branches development in tomato. Plant Cell Rep 40, 1115–1126 (2021). https://doi.org/10.1007/s00299-021-02680-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-021-02680-x