Abstract
Key message
Our findings indicated that the SlERF.J2-IAA23 module integrates hormonal signals to regulate hypocotyl elongation and plant height in tomato.
Abstract
Light and phytohormones can synergistically regulate photomorphogenesis-related hypocotyl elongation and plant height in tomato. AP2/ERF family genes have been extensively demonstrated to play a role in light signaling and various hormones. In this study, we identified a novel AP2/ERF family gene in tomato, SlERF.J2. Overexpression of SlERF.J2 inhibits hypocotyl elongation and plant height. However, the plant height in the slerf.j2ko knockout mutant was not significantly changed compared with the WT. we found that hypocotyl cell elongation and plant height were regulated by a network involving light, auxin and gibberellin signaling, which is mediated by regulatory relationship between SlERF.J2 and IAA23. SlERF.J2 protein could bind to IAA23 promoter and inhibit its expression. In addition, light–dark alternation can activate the transcription of SlERF.J2 and promote the function of SlERF.J2 in photomorphogenesis. Our findings indicated that the SlERF.J2-IAA23 module integrates hormonal signals to regulate hypocotyl elongation and plant height in tomato.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Light and darkness have different effects on seedlings after germination. Photomorphogenesis is characterized by suppressed hypocotyl elongation, cotyledons open and green, without apical hooks; skotomorphogenesis is characterized by hypocotyl elongation, cotyledons closed and yellowish, apical hooks (Von Arnim and Deng 1996). Cells in the apical meristem divide and elongate resulting in the growth of the hypocotyl (de Wit et al. 2016). Light is the most important energy and signal source for plant development (Li and He 2016). Light signals can be perceived by families of photoreceptors, including phytochromes, cryptochromes, phototropins, and UV resistance locus 8 (UVR8) (Casal 2013). Downstream of these photoreceptors are several transcription factors, including the bHLH protein phytochrome-interacting factor (PIFs) (Leivar and Quail 2011) and the bZIP protein elongating hypocotyl 5 (HY5) (Osterlund et al. 2000). PIFs (phytochrome-interacting factors) are a family of basic helix–loop–helix (bHLH) transcription factors that promote hypocotyl elongation in the dark, and phytochromes regulate light responses by promoting the degradation of PIFs (Lorrain et al. 2008). The basic leucine zipper (bZIP) transcription factor (HY5) is required to inhibit hypocotyl growth under light conditions (Shi et al. 2011). Many targets of HY5 are modulators of hormone signaling, including modulators of auxin, gibberellin (GA), abscisic acid (ABA), ethylene, brassinolide (BR), and jasmonic acid (Wang et al. 2012; Lau and Deng 2010).
Plant hormones are the main regulators of plant growth and development. The signaling pathway of the plant hormone auxin and gibberellin (GA) has been studied extensively (Sun 2010; Weijers and Wagner 2016). Auxin has long been regarded as a major regulator of plant growth and development. AUXIN/INDOLE-3-ACETIC ACID (Aux/IAA) and auxin response factors (ARF) were two transcriptional regulators of auxin signaling (Liu et al. 2018). The Aux/IAA protein plays a pivotal role in the perception and signaling of the plant hormone auxin (Audran-Delalande et al. 2012), and Aux/IAA gain-of-function mutants display multiple auxin-related phenotypes such as apical dominance, root formation, and hypocotyl elongation (Chaabouni et al. 2009; Bassa et al. 2012; Deng et al. 2012; Su et al. 2014). Silencing of Sl-IAA27 resulted in impaired auxin sensitivity and reduced leaf chlorophyll content in tomato (Bassa et al. 2012). Silencing of Sl-IAA3 exhibits altered apical dominance, decreased auxin sensitivity, and exaggerated curvature of the apical hook in the dark (Chaabouni et al. 2009). ARFs bind to auxin response elements to activate or repress transcription of target genes (Zouine et al. 2014). The double mutant arf6 arf8 displayed a short hypocotyl phenotype in the dark (Nagpal et al. 2005).
It has been well studied that the synergistic regulation of GA, auxin, and light signaling of cell elongation during seedling morphogenesis (Bai et al. 2012; de Lucas et al. 2008; Chaiwanon et al. 2016). GA and auxin are inextricably regulated in plant growth and development, and auxin appears to alter GA responses by interacting with DELLA proteins that act as repressors of GA signaling (Fleet and Sun 2005). DELLA interacts with PIF and inhibits its DNA-binding activity (Feng et al. 2008). GA is sensed by GA-INSENSITIVE DWARF1 (GID1) (Shimada et al. 2008). Upregulation of AtGA20ox1, AtGA20ox2 and AtGA20ox3 increased hypocotyl length by increasing GA production in Arabidopsis (Huang et al. 1998). Overexpression of GA2oxs results in a dwarf phenotype in Arabidopsis (Schomburg et al. 2003). GID2 is a positive regulator of gibberellin signaling, and inhibition of this gene can lead to a dwarf phenotype in tomato (Liu et al. 2016).
The AP2/ERF TFs (APETALA2/ETHYLENE RESPONSIVE FACTOR) family plays an important role in plant development processes and stress response (Licausi et al. 2013; Muller and Munne-Bosch 2015). There have been many studies on the AP2/ERF family of genes involved in the regulation of light and hormone signals. For example, it has been reported that Sl-ERF.B3 integrates ethylene and auxin signaling by regulating the expression of the auxin signaling component Sl-IAA27, and overexpression of Sl-ERF.B3 inhibits the expression of Sl-IAA27 to suppress hypocotyl length and plant height (Liu et al. 2018). ERF109 binds directly to the GCC-box in the promoters of ASA1 and YUC2, two key enzymes in auxin biosynthesis. Overexpression of ERF109 resulted in a root phenotype similar to auxin overproduction mutants (Cai et al. 2014). SlERF.F12 negatively regulates fruit ripening by regulating the transcription of ripening-related genes 1-amino-cyclopropane-1-carboxylic acid synthase 2 (ACS2), ACS4, POLYGALACTURONASE 2a and PECTATE LYASE (Deng et al. 2022). SlERF.D6 regulates the expression of multiple genes in the SGA synthesis pathway, affects the SGA (Steroidal glycoalkaloids) content of the fruit, and promotes fruit ripening (Guo et al. 2022). SlERF.E4 can transcriptionally regulate tomato fruit ripening specific expression of the β-D-N-acetylhexosaminidase (β-Hex) gene (Irfan et al. 2022). The genes of the AP2/ERF family play an important role in plant growth and development. Recently, more and more gene functions of this family have been revealed, and it has become increasingly important to study the function of this family of genes.
In this study, we found that transgenic tomato seedlings overexpressing of SlERF.J2 (Solyc02g090790) exhibited the constitutive photomorphogenesis phenotype of a short hypocotyl, and open cotyledons in darkness compared with wild type (WT). In addition, our data revealed that SlERF.J2 is involved in the regulation of auxin and GA homeostasis, and plays a role in the dwarf phenotype affecting plant growth and development. We further demonstrated that SlERF.J2 could repress the transcription of IAA23. Our results suggested that the SlERF.J2-IAA23 module is involved in integrating hormone signaling pathways to control hypocotyl cell elongation and plant height in tomato.
Materials and methods
Plant materials and growth conditions
The WT tomato (Solanum lycopersicum Mill. cv. Ailsa Craig) and transgenic seedlings (SlERF.J2-OE, slerf.j2ko mutant) were grown in a greenhouse under standard conditions (16-h day (28 °C)/8-h night (18 °C) cycle, 80% relative humidity). All seeds were surface sterilized and sown on 1/2 MS medium after incubation and germination. and incubated for 7 days under white light (60 µmol m−2 s−1, 16-h day (28 °C)/8-h night (18 °C) cycle, 80% relative humidity), red light (60 µmol m−2 s−1, 16-h day (28 °C)/8-h night (18 °C) cycle, 80% relative humidity) and dark (24-h night, 16-h day (28 °C)/8-h night (18 °C) cycle, 80% relative humidity) conditions. For the light-to-dark transition experiments, seedlings were grown under continuous light or dark conditions for 6 days and then transferred to the opposite condition. Hypocotyl length was measured using the ImageJ software. All these samples were collected and immediately frozen in liquid nitrogen and then stored at − 80 °C freezer.
Construction of overexpression and CRISPR/Cas9 knockout vectors and plant transformation
For construction of the SlERF.J2 overexpression vector, the ORF sequence of the SlERF.J2 gene was amplified using SlERF.J2-all-F/R primers. Then, the open reading frame (ORF) sequence was cloned into the plant binary vector pBI121 with CaMV 35S as its promoter. The knockout target of SlERF.J2 was designed on the first exon sequence using targetdesign (http://skl.scau.edu.cn/targetdesign). Then the CRISPR/Cas9-SlERF.J2 vector was constructed. The constructed vector was transformed into Agrobacterium tumefaciens strain LBA4404. Finally, the constructed vector was transformed into WT tomato cotyledons using the method previously described (Chen et al. 2004). Plants with transgenes were selected using kanamycin and positive transgenic plants were identified by PCR using NPTII-F/R primers. All primers used in this study are listed in Table S1.
Total RNA extraction and qRT-PCR analysis
Trizol reagent (Invitrogen, Shanghai, China) was used to extract total RNA. The RNA extraction method was based on previous research (Xie et al. 2014). RNA was reverse transcribed into cDNA using a kit (Promega, Beijing, China).
Quantitative reverse-transcription PCR (qRT-PCR) was performed by using a CFX96™ RealTime System (Bio-Rad, USA) (Zhang et al. 2020). Relative gene expression levels were analyzed using the 2−ΔΔCT method (Nicot et al. 2005) and normalized with the SlCAC (Solyc08g006960) gene (Exposito-Rodriguez et al. 2008). Three independent biological replicates were performed for each sample. All primer sequences used in this experiment are listed in Table S1.
Determination of chlorophyll content
Chlorophyll was extracted from frozen tissue in 80% acetone. The extraction method was determined according to previous studies (Arnon 1949). The content of chlorophyll was extracted from cotyledons and hypocotyls (0.1 g) of WT and SlERF.J2-OE lines. For the determination of chlorophyll content, total Chl (mg mL−1) = 20.29 × A646 + 8.02 × A663.
Gibberellic acid and paclobutrazol treatment
To test the response of SlERF.J2 to gibberellins (GA3), 4-week-old WT and SlERF.J2-overexpressing tomato seedlings were sprayed with GA3 (50 μM) every 2 days for 8 days. These GA3-sprayed leaves were collected for RNA extraction. The plant height was measured. All samples were subjected to three biological replicates.
Transient expression assay in tobacco leaves
The ORF sequence of SlERF.J2 gene was amplified and cloned into the pGreen II 62-SK vector and used as an effector. The promoter fragment of IAA23 and IAA27 were amplified and cloned into the pGreen II 0800-LUC vector and used as a reporter (Hellens et al. 2005; Xu et al. 2018). The effector and reporter were co-transformed into N. benthamiana leaves. Firefly luciferase and Renilla luciferase were measured using a dual-luciferase reporter assay (Promega) according to the manufacturer’s instructions. The binding activity was calculated by detecting the LUC–REN ratio. All primer sequences used in this experiment are listed in Table S1.
Yeast one-hybrid assay
Yeast one-hybrid (Y1H) assay was performed according to the instructions for the Matchmaker Gold Yeast One Hybrid System (TaKaRa). The ORF sequence of SlERF.J2 gene was amplified and transferred into the pGADT7 vector as a prey vector. The IAA23 promoter sequence was amplified and transferred into pAbAi as a bait vector. According to the TaKaRa’s instructions, the pAbAi-proIAA23 plasmid was linearized and transformed into the Y1H Gold yeast cells. Aureobasidin A (AbA) screened the minimum concentration that inhibits the bait strain. The prey vector was transformed into the bait yeast strain and screened on SD/-Leu (with or without AbA) medium. The pAbAi-p53 ( +) and pGADT7-p53 ( +) plasmids were used as positive controls. Incubate in the dark (30 °C) for 2–3 days. All primer sequences used in this experiment are listed in Table S1.
Statistical analysis
Data were subjected to analysis of variance with SPSS 26.0. Student’s t test (*P < 0.05, **P < 0.01) was performed to analyze the significant difference. ANOVA statistical analyses were performed using SPSS 26.0. Significant differences (P < 0.05) between treatments, as determined by Tukey’s tests, are indicated with different letters. All data are taken from the average of at least three independent biological replicates.
Results
Light/dark regulates transcription of SlERF.J2
Light and darkness are the primary regulators of photomorphogenesis and skotomorphogenesis in plant. To explore the effects of specific light and darkness on tomato seedling growth, we evaluated seedling growth of WT under continuous white light (WL), red light (RL) and dark conditions. Under WL and RL conditions, WT seedlings exhibited short hypocotyls, open cotyledons, and no apical hooks. Under dark conditions, WT seedlings exhibited long hypocotyls, yellowed cotyledons, and apical hooks (Fig. 1A–C). Under light conditions, the seedlings maintained photomorphogenesis and their hypocotyl length was shorter than that of seedlings under dark conditions (Fig. 1D). To obtain some clues of whether SlERF.J2 was involved in the morphogenesis of seedlings under different light conditions, we investigated the expression patterns of SlERF.J2 under different light treatments by quantitative RT-PCR technique (qRT-PCR). The results showed that the expression level of SlERF.J2 in hypocotyls under WL and RL treatments was higher than that of under dark treatments (Fig. 1E).
Expression profile of SlERF.J2 in WT. A–C Growth of WT seedlings for 6 days under continuous white light (WL), red light (RL) and dark. D Hypocotyl length of WT seedlings under the conditions of (A–C). E Expression levels of SlERF.J2 in the hypocotyls of WT seedlings under the conditions of (A–C). F, G Seedlings were grown under continuous light or dark conditions for 6 d and then transferred to the opposite conditions for the indicated times. H The expression level of SlERF.J2 was determined by qRT-PCR. Seedlings were grown under continuous light or dark conditions for 6 d and then transferred to the opposite conditions. Data are means (SD) of at least 20 seedlings. Scale bars are 1 μm. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
To further investigate the expression of SlERF.J2 under different light conditions, we transferred light-grown seedlings to the dark and dark-grown seedlings to light (Fig. 1F, G), and qRT-PCR was performed to detect the expression of SlERF.J2. As shown in Fig. 1H, the expression level of SlERF.J2 in light-grown seedlings was lower than that in dark-grown seedlings. After the seedlings were transferred to dark conditions, SlERF.J2 expression increased for 24 h and then decreased up to 48 h. In contrast, after seedlings were transferred from dark to light conditions, SlERF.J2 expression decreased within 12 h and a low level was maintained up to 48 h. These results suggested that the increased expression level of SlERF.J2 was mainly caused by the light-to-dark transition. To verify whether SlERF.J2 has functional redundancy with other tomato ERFs genes, a phylogenetic tree of tomato ERF family proteins was constructed (Fig. S1). The results showed that SlERF.J2 did not have high homology to other ERF family proteins. We speculate that the function of the SlERF.J2 gene may not be affected by other homologous genes in tomatoes.
Overexpression of SlERF.J2 triggers a constitutive photomorphogenetic responses under dark and light conditions
To investigate the role of the SlERF.J2 gene, we generated transgenic plants overexpressing of SlERF.J2 (SlERF.J2-OE) (Fig. S2). Under light conditions, the hypocotyl length of SlERF.J2-OE plants was significantly shorter compared with WT (Fig. 2A). We further evaluated the growth of transgenic and WT seedlings under dark conditions. Consistent with the results under light conditions, the hypocotyl length of SlERF.J2-OE plants was significantly shortened and the apical hook were inconspicuous compared to WT (Fig. 2E). To further investigate the effect of light–dark alternation on the growth of SlERF.J2-OE and WT seedlings, we treated the seedlings with light–dark alternation. More interestingly, the hypocotyl length of SlERF.J2-OE seedlings did not change during transition from light to dark, while the hypocotyl length of WT seedlings became longer (Fig. 2B, I). During the transition from dark to light, the hypocotyl length of SlERF.J2-OE and WT seedlings did not change (Fig. 2F, I). In addition, during the transition from light to dark, the hypocotyls of WT seedlings were elongated and the chlorophyll content decreased compared with that of SlERF.J2-OE seedlings (Fig. 2C, D, J). During the transition from dark to light, the cotyledons of the SlERF.J2-OE seedlings opened and accumulated more chlorophyll than that of WT (Fig. 2G, H, J). These results indicated that overexpression of SlERF.J2 in WT plants triggered a constitutive photomorphogenic-like response under both dark and light conditions, and the resulting in shorter hypocotyls of SlERF.J2-OE lines seedlings.
Overexpression of SlERF.J2 in tomato triggers a constitutive photomorphogenetic response. A WT and SlERF.J2-OE seedlings were grown under continuous light for 6 days. B Seedlings grown in A were transferred to dark conditions for 2 days. C, D The stem of the WT and SlERF.J2-OE seedling in B. E WT and SlERF.J2-OE seedlings were grown under continuous dark for 6 days. F Seedlings grown in (E) were transferred to light conditions for 2 days. G, H The cotyledon of the WT and SlERF.J2-OE seedling in (F). I Hypocotyl length of WT and SlERF.J2-OE seedlings in A, B, E, F. J Chlorophyll content in cotyledons and hypocotyls of WT and SlERF.J2-OE seedlings in A, B, E, F. Data are means (SD) of at least 20 seedlings. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
Gene expression profiles in SlERF.J2-OE and WT tomato seedlings
To evaluate the role of SlERF.J2 in controlling hypocotyl growth, we performed qRT-PCR analysis on WT and SlERF.J2-OE seedlings grown under light conditions. Aux/IAA mutants exhibit multiple auxin-related developmental phenotypes, including apical dominance, hypocotyl elongation, and leaf expansion (Tatematsu et al. 2004). In this study, expression of IAA2, IAA3, IAA13, IAA19, and IAA23 were detected in seedlings of WT and SlERF.J2-OE lines (Fig. 3A). The expression levels of all genes were down-regulated in the SlERF.J2-OE lines compared with WT. The basic leucine zipper (bZIP) transcription factor (HY5) is required to inhibit hypocotyl growth under light conditions. PIF3 is a transcription factor that inhibits photomorphogenesis. In addition, the expression of light response-related genes HY5 was significantly higher and PIF3 was decreased in overexpression lines compared with WT.
Expression levels of genes related to auxin/gibberellin in wild-type and SlERF.J2-overexpressing seedlings. A Expression levels of IAA2, IAA3, IAA13, IAA19 and IAA23 in WT and SlERF.J2-OE seedlings. B Expression levels of HY5, PIF1 and PIF3 in WT and SlERF.J2-OE seedlings. C Expression levels of GA20ox2, KAO, CPS, GAI, GID1 and GID2 in WT and SlERF.J2-OE seedlings. D Expression levels of XTH2 and XTH5 in WT and SlERF.J2-OE seedlings. Expression values are relative to the SlCAC gene. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
Gibberellin (GA) and cell elongation-related genes were detected in seedlings of WT and SlERF.J2-OE lines. The expression of all tested GA important biosynthesis enzymes GA20ox2; two genes in the early steps of the GA biosynthetic pathway, KAO and CPS; and GID1 and GID2 encoding GA receptors were down-regulated in SlERF.J2-OE seedlings compared to wild type (Fig. 3C). The expression of GAI, encoding the DELLA protein as a repressor of GA signaling (Murase et al. 2008), was up-regulated in the SlERF.J2-OE lines. XTH2 and XTH5 play a role in cell expansion process (Saladie et al. 2006; Catala et al. 2001) and were significantly down-regulated in overexpressing lines.
SlERF.J2 affects tomato plant height
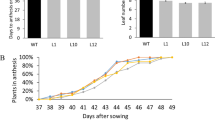
To gain further understanding of the function of SlERF.J2 in tomato, we cultivated tomato seedlings of WT and SlERF.J2-OE lines under the same growth conditions. Four weeks after sowing, the plant height of the SlERF.J2-OE lines was significantly reduced compared with WT (Fig. 4A). From the top view of tomato seedlings, we observed that the leaves of SlERF.J2-OE plants were more compact than those of WT (Fig. 4B–E). This phenotype continued until the 12th week flowering and fruiting stage. Compared with WT, the SlERF.J2-OE lines had significantly lower plant height and also smaller internode lengths than WT plants (Fig. 4F). As shown in Fig. 4F, when the WT plants grew to 110 cm, the plant height of the SlERF.J2-OE lines was 42–60 cm, which was obviously about half lower than that of the WT plants. This demonstrates that the dwarf transgenic lines depicted the shortening of internodal length. To further confirm the role of SlERF.J2 in plant development, We generated slerf.j2 knockout mutants in tomatoes using the CRISPR/Cas9 system. The type of knockout is shown in Fig. S3. During tomato growth and development, we found that overexpression of SlERF.J2 significantly suppressed plant height, while the slerf.j2ko mutants had no obvious change compared with WT (Fig. 4J). This phenotype was maintained when the tomato was in the flowering stage (Fig. S4A–G). Statistical analysis of the plant height of WT, SlERF.J2-OE and slerf.j2ko lines showed that the SlERF.J2-OE lines was significantly lower than WT, while the slerf.j2ko lines had no significant change compared with WT (Fig. S4H). These results indicated that SlERF.J2 affected the internodal length and height of tomato plants.
SlERF.J2 affects plant growth and the expression levels of growth-related genes. A Plant height phenotypes of WT and SlERF.J2-OE lines. B–E Top view of plants in (A). F Height of WT and SlERF.J2-OE tomato plants at 12 weeks. G, H qRT-PCR analysis of auxin-related genes IAA1, IAA2, IAA3, IAA4, IAA7, IAA8, IAA9, IAA11, IAA12, IAA13, IAA14, IAA15, IAA16, IAA17, IAA19, IAA22, IAA23, IAA27, PIN1, PIN3, PIN4, PIN5, PIN6, PIN7, PIN8, PIN9, LAX1, LAX2, LAX3, LAX4, ARF5, ARF6a, ARF18, ARF19, and ARF24 expression level. I Expression levels of leaves cell development-related genes XTH2, XTH5, XTH7, and PRE2. Expression values are relative to the SlCAC gene. The relative expression of each gene in WT leaves was normalized to 1. J Plant height phenotypes of SlERF.J2-OE lines, WT and slerf.j2ko lines. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
Overexpression of SlERF.J2 affects auxin signaling
Auxin has long been considered a major regulator of plant growth and development (Zouine et al. 2014). In this study, we analyzed the transcript accumulation of auxin-related genes in leaves of WT and SlERF.J2-OE lines, including eighteen Aux/IAA transcription factor genes (IAA1, IAA2, IAA3, IAA4, IAA7, IAA8, IAA9, IAA11, IAA12, IAA13, IAA14, IAA15, IAA16, IAA17, IAA19, IAA22, IAA23, IAA27) (Audran-Delalande et al. 2012), eight encoding PIN auxin efflux transport protein (PIN1, PIN3, PIN4, PIN5, PIN6, PIN7, PIN8, PIN9) (Pattison and Catala 2012), four auxin transporter genes (LAX1, LAX2, LAX3, LAX4) (Pattison and Catala 2012) and five auxin response genes (ARF5, ARF6a, ARF18, ARF19, ARF24) (Fig. 4G, H) (Zouine et al. 2014). Compared with WT, the expression levels of all other genes were down-regulated except for IAA7, IAA13, IAA19, IAA27, PIN8 and ARF24 which were up-regulated in SlERF.J2-OE lines. These results suggested that auxin homeostasis may be disrupted in the SlERF.J2-overexpressing plants.
In addition, the expression levels of genes related to plant cell expansion and cell division, XTH2, XTH5, XTH7, and PRE2, showed significant changes in overexpression lines compared with WT (Fig. 4I). According to previous reports, XTH2, XTH5 and XTH7 play roles in cell wall reorganization and cell expansion process (Catala et al. 2001; Saladie et al. 2006). PRE2 plays a role in the regulation of cell size (Zhu et al. 2019). These genes were down-regulated in the SlERF.J2-OE lines, suggesting that SlERF.J2 may affect cell development to affect plant height.
The role of SlERF.J2 in gibberellin-dependent growth and development
Gibberellin (GA) is an important hormone that regulates plant growth (Depuydt and Hardtke 2011). Given that overexpression of SlERF.J2 caused a dwarf phenotype, we sought to further explore the role of GA in the SlERF.J2-OE lines. Here, we tested the response of WT and SlERF.J2-OE lines to GA treatment. We first sprayed 50 μM GA3 or aqueous solution as the control to 4-week-old plant leaves every 2 days for 8 days. Thereby, these plants were divided into two groups: CK-0 d and CK-8 d (Fig. 5A, C); GA3-0 d and GA3-8 d (Fig. 5B, D). The WT plants were significantly taller than SlERF.J2-OE plants before GA3 treatment (Fig. 5A, B). In the GA3 treatment experiment, WT plants in the control group were still significantly higher than SlERF.J2-OE plants (Fig. 5C); after exogenous spraying of GA3, both WT and SlERF.J2-OE plants grew significantly taller, but WT plants were a slightly taller than the SlERF.J2-OE plants (Fig. 5D). Plant heights of transgenic and WT plants were measured as shown in Fig. 5E, F, and showed that after GA3 treatment, WT plants were about 7 cm longer than the control group, while SlERF.J2-OE plants were 11–15 cm longer than the control group. These results suggested that the dwarf phenotype of SlERF.J2-OE lines could be partly rescued by GA3 application, and that SlERF.J2-OE plants were more sensitive to exogenous GA3 stimulation.
The phenotype of WT and SlERF.J2-OE transgenic tomato plants under control and GA3 treatments. A, B Growth characteristics of wild-type and transgenic tomato plants before treatment. C Growth characteristics of control wild-type and transgenic tomato plants. D Growth characteristics of wild-type and transgenic tomato plants after 8 days of GA3 treatment. E, F Height of WT and SlERF.J2-OE tomato plants after control and GA3 treatment. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
We further speculated that overexpression of SlERF.J2 might change the sensitivity of the transgenic lines to GA3, so the sensitivity of tomato seedlings to GA3 was performed (Fig. S5A). To examine the response of seedlings to GA3, we transferred the germinated seeds to MS medium without or with GA3. After GA3 treatment, the hypocotyl lengths of WT and SlERF.J2-OE lines were obviously promoted, but the hypocotyls of the seedlings of SlERF.J2-OE lines were promoted by 66–70%, and the hypocotyls of WT seedlings were promoted by 61% (Fig. S5B), suggesting that the SlERF.J2-OE seedlings were more sensitive to GA3.
Overexpression of SlERF.J2 regulates the transcripts accumulation of gibberellins-related genes
To further explore the possibility of SlERF.J2-mediated GA pathway change, we examined the expression levels of GA-related genes in tomato leaves of WT and SlERF.J2-OE lines before and after GA treatment. Except for CPS, the expression of all tested GA biosynthetic genes KS, and KAO was down-regulated, while the GA catabolism gene GA2ox2 was up-regulated (Fig. 6E) in SlERF.J2-OE tomato leaves. In addition, the GA receptor genes GID1 and GID2 were down-regulated (Voegele et al. 2011) (Fig. 6C), while the expression of GID1b was up-regulated (Fig. 6C). Meanwhile, the expression of the gibberellin-inducible gene GAST1 (Shi and Olszewski 1998) was significantly down-regulated (Fig. 6E). These results indicated that the overexpression of SlERF.J2 could affect gibberellin metabolism and signal strength in leaf development.
Expression levels of GA-related genes in WT and SlERF.J2-OE transgenic plants under control and GA3 treatments. A, C, E The expression of CPS, KS, KAO, GID1, GID1b, GID2, GA2ox2 and GAST1 genes was analyzed by qRT-PCR under control conditions. B, D, F The expression of CPS, KS, KAO, GID1, GID1b, GID2, GA2ox2 and GAST1 genes was analyzed by qRT-PCR under GA3 treatments conditions. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01)
We identified the responsiveness of SlERF.J2-OE and WT plants to GA3. With the 8 days application of GA3, these plants showed the response in plant height increment, and GA-related genes were further detected using qRT-PCR. After GA3 treatment, the expression levels of the GA biosynthetic genes CPS and KS genes did not change compared with WT, while the KAO gene was significantly up-regulated (Fig. 6B) in the SlERF.J2-OE lines. GA receptor genes (GID1, GID1b, GID2) were also up-regulated (Fig. 6D). The GA catabolism gene GA2ox2 was unchanged, while the gibberellin-inducible gene GAST1 was significantly up-regulated (Fig. 6F). These results suggested that GA3 treatment increases the expression of GA-related genes in SlERF.J2-OE lines in tomato.
SlERF.J2 directly targets the auxin transcription factor IAA23 gene
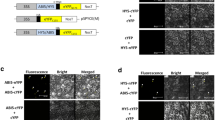
Given that overexpression of SlERF.J2 suppressed tomato seedling hypocotyl length and plant height, among the differentially expressed genes, the sequence analysis of IAA-related genes showed that IAA23 and IAA27 promoter sequences contained DRE (dehydration response element) elements (Fig. S6). It was previously reported that Sl-ERF.B3 regulates the expression of Sl-IAA27 by directly binding to its promoter (Liu et al. 2018). In addition, qRT-PCR analysis results revealed that the gene expression levels of IAA23 and IAA27 were down-regulated in the SlERF.J2-OE lines compared with WT (Fig. 4G). To investigate whether SlERF.J2 can regulate the activity of the IAA23 and IAA27 promoters, a transient transactivation assays was performed in N. benthamiana leaves. The promoters of IAA23 and IAA27 genes were, respectively, cloned into pGreen II 0800-LUC vector, and then co-transformed into tobacco leaves with the effector 35S::SlERF.J2 or empty vector (Fig. 7A). As shown in Fig. 7B, the LUC/REN ratio of IAA23pro was inhibited by approximately 40% in the presence of SlERF.J2 compared to the control (empty vector). As shown in Fig. 7C, the LUC/REN ratio of IAA27pro did not change significantly in the presence of SlERF.J2 compared to the control (empty vector). To further determine whether IAA23 was the direct target of SlERF.J2, we examined the ability of SlERF.J2 to directly bind to the IAA23 promoter using a yeast one-hybrid (Y1H) assay (Fig. 7D, E). The results showed that the promoter of IAA23 could be recognized by SlERF.J2. Taken together, these results indicated that SlERF.J2 can directly target the auxin transcription factor IAA23 and that the activity of the IAA23 promoter was negatively regulated by SlERF.J2 in vivo.
SlERF.J2 interacts with XI and PLA8 and regulates the activity of the IAA23 promoter. A Schematic map of the transient expression vectors pGreenII-0800-LUC and pGreenII-62-SK. REN renilla luciferase, LUC firefly luciferase. B, C Transcriptional regulation of IAA23 and IAA27 promoters by SlERF.J2. All data are means (± SE) of three independent biological replicates (*P < 0.05, **P < 0.01). D The IAA23 promoter and the positive control in yeast grown on SD/-Ura medium containing aureobasidin A (− Ura + AbA) did not detect the auto-activation ability. E The interaction between SlERF.J2 and the IAA23 promoter was determined by the yeast one-hybrid assay. The interactions were determined on SD/-Leu medium in the presence of AbA (− Leu + AbA)
Discussion
The AP2/ERF transcription factor family is involved in many growth and developmental processes in plants (Xie et al. 2019). It has been reported that CBF1, an AP2/ERF family transcription factor, increases the protein abundance of PIF4 and PIF5 and promotes hypocotyl growth under ambient temperatures in Arabidopsis (Dong et al. 2020). Two independent Arabidopsis mutations of AtERF71 and GmERF75 showed shorter hypocotyls, and overexpression of GmERF75 rescued the short hypocotyl phenotypes in these mutants (Zhao et al. 2019). Members of the plant-specific transcriptional regulator group-VII ERF (ERF-VII) family were involved in the regulation of shoot elongation and photomorphogenesis (Gibbs et al. 2014). Here, we report that SlERF.J2 was a negative regulator of hypocotyl elongation and plant height in tomato. In this study, expression of SlERF.J2 was up-regulated during light-to-dark transition (Fig. 1H), and the hypocotyl length of the seedlings of the SlERF.J2-OE lines was shorter than that of the WT under both light and dark conditions (Fig. 2A, E). When the light-treated seedlings were transferred to dark conditions, the hypocotyl length of the WT seedlings increased significantly, whereas the hypocotyl length of the SlERF.J2-OE lines was unchanged (Fig. 2B, I). In addition, the expression of SlERF.J2 was induced under dark conditions, further indicating that overexpression of SlERF.J2 could inhibit the length of hypocotyl.
The phytochrome-interacting factor pif3 mutants have been reported to exhibit a short hypocotyls under red light (Job and Datta 2021). In addition, PIF3 is a positive regulator of chloroplast development, and the pif1pif3 double mutant showed elevated protochlorophyllide levels, decreased hypocotyl elongation and increased cotyledon opening in the dark (Stephenson et al. 2009). Here, we examined the expression levels of PIF1 and PIF3 in WT and SlERF.J2-OE plant seedlings. Compared with WT tomato plants, the expression level of PIF1 did not change significantly, while the expression level of PIF3 was down-regulated in SlERF.J2-OE plants. After dark-to-light culture, the cotyledons of SlERF.J2-OE seedlings were opened compared with WT seedlings (Fig. 2G, H), and the chlorophyll content of SlERF.J2-OE seedlings was found to be higher in cotyledons (Fig. 2J), indicating that SlERF.J2-OE seedlings accumulated more protochlorophyllide in the dark, which is similar to the phenotype of pif3 mutants. This result indicated that overexpression of SlERF.J2 affected the expression of PIF3, and the transgenic seedlings maintained photomorphogenic properties. We speculate that SlERF.J2 may be involved in light signaling and plastid development, thereby affecting hypocotyl elongation.
Plant growth and development was coordinated by a complex network of interacting hormones. Sl-ERF.B3 inhibits the expression of Sl-IAA27 by directly binding to its promoter. Ectopic expression of Sl-ERF.B3 leads to impaired sensitivity to auxin, resulting in shortened hypocotyls and dwarfing plants (Liu et al. 2018). AtERF109 mediates crosstalk between JA signaling and auxin biosynthesis, thereby regulating lateral root formation (Cai et al. 2014). In this study, we further cultivated tomato seedlings and found that overexpression of SlERF.J2 resulted in a dwarf phenotype (Fig. 4A). The slerf.j2 knockout mutant did not affect plant height (Fig. 4J). In this study, we found that knocking out the mutant slerf.j2 had no effect on plant height, possibly due to the low background expression level of the SlERF.J2 gene in tomato tissues, and the expression of SlERF.J2 gene may not be needed during the growth and development of tomato, on the contrary, overexpression of SlERF.J2 gene will inhibit the elongation of tomato hypocotyl and plant height. This phenotype was further verified by detecting the expression of auxin and GA-related genes (Figs. 4G, H; 6A, C, E). First, we demonstrated that overexpression of SlERF.J2 altered the mRNA accumulation of some auxin-related genes by qRT-PCR analysis. The results showed that the expression of most auxin-related genes was down-regulated in the SlERF.J2-OE lines compared to WT. Subsequently, it was demonstrated that SlERF.J2 could bind to the promoter of IAA23 and inhibit its expression by a dual-luciferase reporter system (Fig. 7). These results suggested that overexpression of SlERF.J2 may lead to impaired sensitivity to auxin, resulting in shortened hypocotyl and dwarfing plants. In addition, GA also plays an important role in regulating cell expansion and plant height (Schomburg et al. 2003). We found that the dwarf phenotype of SlERF.J2-OE lines could be partly rescued by exogenous application of GA3, and SlERF.J2-OE plants were more sensitive to exogenous GA3 stimulation. By detecting GA-related genes, it was found that the expression of gibberellin-inducible gene (GAST1) genes was down-regulated in SlERF.J2-OE lines compared with WT. These results suggested that overexpression of SlERF.J2 may inhibit gibberellin synthesis by inhibiting gibberellin-inducible genes, resulting in a dwarf phenotype. However, exogenous application of GA3 partially rescued the dwarf phenotype.
In short, we report an ethylene transcription factor, SlERF.J2. We demonstrate that overexpression of SlERF.J2 affects hypocotyl elongation and plant height from phenotypic analysis and related gene expression levels. In addition, SlERF.J2 directly binds to the promoter of IAA23 to inhibit its activity, thereby suppressing the plant height of the SlERF.J2-OE lines. We propose a model to elucidate the potential function of SlERF.J2 in tomato hypocotyl and plant height (Fig. 8), and provide a molecular mechanism for studies on how to decipher crosstalk between different hormones to control plant growth and development in the future.
A proposed model illustrates the regulatory role of SlERF.J2 in the hypocotyl and plant high. Under the alternation of darkness and light condition, the expression of SlERF.J2 can be induced; SlERF.J2 can regulate the promoter activity of IAA23 and inhibit its expression; overexpression of SlERF.J2 may inhibits the expression of GA-related genes. In conclusion, overexpression of SlERF.J2 inhibited hypocotyl length and plant height, and played an important role in the regulation of light, auxin, and gibberellin signaling
Data availability
All data supporting the findings of this study are available within the paper and within its supplementary materials published online.
References
Arnon DI (1949) Copper enzymes in isolated chloroplasts. polyphenoloxidase in beta vulgaris. Plant Physiol 24(1):1–15
Audran-Delalande C, Bassa C, Mila I, Regad F, Zouine M, Bouzayen M (2012) Genome-wide identification, functional analysis and expression profiling of the Aux/IAA gene family in tomato. Plant Cell Physiol 53(4):659–672
Bai MY, Fan M, Oh E, Wang ZY (2012) A triple helix-loop-helix/basic helix-loop-helix cascade controls cell elongation downstream of multiple hormonal and environmental signaling pathways in Arabidopsis. Plant Cell 24(12):4917–4929
Bassa C, Mila I, Bouzayen M, Audran-Delalande C (2012) Phenotypes associated with down-regulation of Sl-IAA27 support functional diversity among Aux/IAA family members in tomato. Plant Cell Physiol 53(9):1583–1595
Cai XT, Xu P, Zhao PX, Liu R, Yu LH, Xiang CB (2014) Arabidopsis ERF109 mediates cross-talk between jasmonic acid and auxin biosynthesis during lateral root formation. Nat Commun 5:5833
Casal JJ (2013) Photoreceptor signaling networks in plant responses to shade. Annu Rev Plant Biol 64:403–427
Catala C, Rose JK, York WS, Albersheim P, Darvill AG, Bennett AB (2001) Characterization of a tomato xyloglucan endotransglycosylase gene that is down-regulated by auxin in etiolated hypocotyls. Plant Physiol 127(3):1180–1192
Chaabouni S, Jones B, Delalande C, Wang H, Li Z, Mila I, Frasse P, Latche A, Pech JC, Bouzayen M (2009) Sl-IAA3, a tomato Aux/IAA at the crossroads of auxin and ethylene signalling involved in differential growth. J Exp Bot 60(4):1349–1362
Chaiwanon J, Wang W, Zhu JY, Oh E, Wang ZY (2016) Information integration and communication in plant growth regulation. Cell 164(6):1257–1268
Chen G, Hackett R, Walker D, Taylor A, Lin Z, Grierson D (2004) Identification of a specific isoform of tomato lipoxygenase (TomloxC) involved in the generation of fatty acid-derived flavor compounds. Plant Physiol 136(1):2641–2651
de Lucas M, Daviere JM, Rodriguez-Falcon M, Pontin M, Iglesias-Pedraz JM, Lorrain S, Fankhauser C, Blazquez MA, Titarenko E, Prat S (2008) A molecular framework for light and gibberellin control of cell elongation. Nature 451(7177):480–484
de Wit M, Galvao VC, Fankhauser C (2016) Light-mediated hormonal regulation of plant growth and development. Annu Rev Plant Biol 67:513–537
Deng W, Yang Y, Ren Z, Audran-Delalande C, Mila I, Wang X, Song H, Hu Y, Bouzayen M, Li Z (2012) The tomato SlIAA15 is involved in trichome formation and axillary shoot development. New Phytol 194(2):379–390
Deng H, Chen Y, Liu Z, Liu Z, Shu P, Wang R, Hao Y, Su D, Pirrello J, Liu Y, Li Z, Grierson D, Giovannoni JJ, Bouzayen M, Liu M (2022) SlERF.F12 modulates the transition to ripening in tomato fruit by recruiting the co-repressor TOPLESS and histone deacetylases to repress key ripening genes. Plant Cell 34(4):1250–1272
Depuydt S, Hardtke CS (2011) Hormone signalling crosstalk in plant growth regulation. Curr Biol 21(9):R365-373
Dong X, Yan Y, Jiang B, Shi Y, Jia Y, Cheng J, Shi Y, Kang J, Li H, Zhang D, Qi L, Han R, Zhang S, Zhou Y, Wang X, Terzaghi W, Gu H, Kang D, Yang S, Li J (2020) The cold response regulator CBF1 promotes Arabidopsis hypocotyl growth at ambient temperatures. EMBO J 39(13):e103630
Exposito-Rodriguez M, Borges AA, Borges-Perez A, Perez JA (2008) Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. Bmc Plant Biol 8:1–12
Feng S, Martinez C, Gusmaroli G, Wang Y, Zhou J, Wang F, Chen L, Yu L, Iglesias-Pedraz JM, Kircher S, Schafer E, Fu X, Fan LM, Deng XW (2008) Coordinated regulation of Arabidopsis thaliana development by light and gibberellins. Nature 451(7177):475–479
Fleet CM, Sun TP (2005) A DELLAcate balance: the role of gibberellin in plant morphogenesis. Curr Opin Plant Biol 8(1):77–85
Gibbs DJ, Md Isa N, Movahedi M, Lozano-Juste J, Mendiondo GM, Berckhan S, Marin-de la Rosa N, Vicente Conde J, Sousa Correia C, Pearce SP, Bassel GW, Hamali B, Talloji P, Tome DF, Coego A, Beynon J, Alabadi D, Bachmair A, Leon J, Gray JE, Theodoulou FL, Holdsworth MJ (2014) Nitric oxide sensing in plants is mediated by proteolytic control of group VII ERF transcription factors. Mol Cell 53(3):369–379
Guo H, Mao M, Deng Y, Sun L, Chen R, Cao P, Lai J, Zhang Y, Wang C, Li C, Li Y, Bai Q, Tan T, Yang J, Wang S (2022) Multi-omics analysis reveals that SlERF.D6 synergistically regulates SGAs and fruit development. Front Plant Sci 13:860577
Hellens RP, Allan AC, Friel EN, Bolitho K, Grafton K, Templeton MD, Karunairetnam S, Gleave AP, Laing WA (2005) Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods 1:13
Huang S, Raman AS, Ream JE, Fujiwara H, Cerny RE, Brown SM (1998) Overexpression of 20-oxidase confers a gibberellin-overproduction phenotype in Arabidopsis. Plant Physiol 118(3):773–781
Irfan M, Kumar P, Kumar V, Datta A (2022) Fruit ripening specific expression of beta-D-N-acetylhexosaminidase (beta-Hex) gene in tomato is transcriptionally regulated by ethylene response factor SlERF.E4. Plant Sci 323:111380
Job N, Datta S (2021) PIF3/HY5 module regulates BBX11 to suppress protochlorophyllide levels in dark and promote photomorphogenesis in light. New Phytol 230(1):190–204
Lau OS, Deng XW (2010) Plant hormone signaling lightens up: integrators of light and hormones. Curr Opin Plant Biol 13(5):571–577
Leivar P, Quail PH (2011) PIFs: pivotal components in a cellular signaling hub. Trends Plant Sci 16(1):19–28
Li QF, He JX (2016) BZR1 interacts with HY5 to mediate brassinosteroid- and light-regulated cotyledon opening in Arabidopsis in darkness. Mol Plant 9(1):113–125
Licausi F, Ohme-Takagi M, Perata P (2013) APETALA2/Ethylene Responsive Factor (AP2/ERF) transcription factors: mediators of stress responses and developmental programs. New Phytol 199(3):639–649
Liu Q, Guo X, Chen G, Zhu Z, Yin W, Hu Z (2016) Silencing SlGID2, a putative F-box protein gene, generates a dwarf plant and dark-green leaves in tomato. Plant Physiol Biochem 109:491–501
Liu M, Chen Y, Chen Y, Shin JH, Mila I, Audran C, Zouine M, Pirrello J, Bouzayen M (2018) The tomato Ethylene Response Factor Sl-ERF.B3 integrates ethylene and auxin signaling via direct regulation of Sl-Aux/IAA27. New Phytol 219(2):631–640
Lorrain S, Allen T, Duek PD, Whitelam GC, Fankhauser C (2008) Phytochrome-mediated inhibition of shade avoidance involves degradation of growth-promoting bHLH transcription factors. Plant J 53(2):312–323
Muller M, Munne-Bosch S (2015) Ethylene response factors: a key regulatory hub in hormone and stress signaling. Plant Physiol 169(1):32–41
Murase K, Hirano Y, Sun TP, Hakoshima T (2008) Gibberellin-induced DELLA recognition by the gibberellin receptor GID1. Nature 456(7221):459–463
Nagpal P, Ellis CM, Weber H, Ploense SE, Barkawi LS, Guilfoyle TJ, Hagen G, Alonso JM, Cohen JD, Farmer EE, Ecker JR, Reed JW (2005) Auxin response factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 132(18):4107–4118
Nicot N, Hausman JF, Hoffmann L, Evers D (2005) Housekeeping gene selection for real-time RT-PCR normalization in potato during biotic and abiotic stress. J Exp Bot 56(421):2907–2914
Osterlund MT, Hardtke CS, Wei N, Deng XW (2000) Targeted destabilization of HY5 during light-regulated development of Arabidopsis. Nature 405(6785):462–466
Pattison RJ, Catala C (2012) Evaluating auxin distribution in tomato (Solanum lycopersicum) through an analysis of the PIN and AUX/LAX gene families. Plant J 70(4):585–598
Saladie M, Rose JK, Cosgrove DJ, Catala C (2006) Characterization of a new xyloglucan endotransglucosylase/hydrolase (XTH) from ripening tomato fruit and implications for the diverse modes of enzymic action. Plant J 47(2):282–295
Schomburg FM, Bizzell CM, Lee DJ, Zeevaart JA, Amasino RM (2003) Overexpression of a novel class of gibberellin 2-oxidases decreases gibberellin levels and creates dwarf plants. Plant Cell 15(1):151–163
Shi L, Olszewski NE (1998) Gibberellin and abscisic acid regulate GAST1 expression at the level of transcription. Plant Mol Biol 38(6):1053–1060
Shi QM, Yang X, Song L, Xue HW (2011) Arabidopsis MSBP1 is activated by HY5 and HYH and is involved in photomorphogenesis and brassinosteroid sensitivity regulation. Mol Plant 4(6):1092–1104
Shimada A, Ueguchi-Tanaka M, Nakatsu T, Nakajima M, Naoe Y, Ohmiya H, Kato H, Matsuoka M (2008) Structural basis for gibberellin recognition by its receptor GID1. Nature 456(7221):520–523
Stephenson PG, Fankhauser C, Terry MJ (2009) PIF3 is a repressor of chloroplast development. Proc Natl Acad Sci USA 106(18):7654–7659
Su L, Bassa C, Audran C, Mila I, Cheniclet C, Chevalier C, Bouzayen M, Roustan JP, Chervin C (2014) The auxin Sl-IAA17 transcriptional repressor controls fruit size via the regulation of endoreduplication-related cell expansion. Plant Cell Physiol 55(11):1969–1976
Sun TP (2010) Gibberellin-GID1-DELLA: a pivotal regulatory module for plant growth and development. Plant Physiol 154(2):567–570
Tatematsu K, Kumagai S, Muto H, Sato A, Watahiki MK, Harper RM, Liscum E, Yamamoto KT (2004) MASSUGU2 encodes Aux/IAA19, an auxin-regulated protein that functions together with the transcriptional activator NPH4/ARF7 to regulate differential growth responses of hypocotyl and formation of lateral roots in Arabidopsis thaliana. Plant Cell 16(2):379–393
Voegele A, Linkies A, Muller K, Leubner-Metzger G (2011) Members of the gibberellin receptor gene family GID1 (GIBBERELLIN INSENSITIVE DWARF1) play distinct roles during Lepidium sativum and Arabidopsis thaliana seed germination. J Exp Bot 62(14):5131–5147
Von Arnim A, Deng XW (1996) Light control of seedling development. Annu Rev Plant Physiol Plant Mol Biol 47:215–243
Wang ZY, Bai MY, Oh E, Zhu JY (2012) Brassinosteroid signaling network and regulation of photomorphogenesis. Annu Rev Genet 46:701–724
Weijers D, Wagner D (2016) Transcriptional responses to the auxin hormone. Annu Rev Plant Biol 67:539–574
Xie Q, Hu Z, Zhu Z, Dong T, Zhao Z, Cui B, Chen G (2014) Overexpression of a novel MADS-box gene SlFYFL delays senescence, fruit ripening and abscission in tomato. Sci Rep 4:4367
Xie Z, Nolan TM, Jiang H, Yin Y (2019) AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front Plant Sci 10:228
Xu N, Chu Y, Chen H, Li X, Wu Q, Jin L, Wang G, Huang J (2018) Rice transcription factor OsMADS25 modulates root growth and confers salinity tolerance via the ABA-mediated regulatory pathway and ROS scavenging. PLoS Genet 14(10):e1007662
Zhang L, Kang J, Xie Q, Gong J, Shen H, Chen Y, Chen G, Hu Z (2020) The basic helix-loop-helix transcription factor bHLH95 affects fruit ripening and multiple metabolisms in tomato. J Exp Bot 71(20):6311–6327
Zhao MJ, Yin LJ, Liu Y, Ma J, Zheng JC, Lan JH, Fu JD, Chen M, Xu ZS, Ma YZ (2019) The ABA-induced soybean ERF transcription factor gene GmERF75 plays a role in enhancing osmotic stress tolerance in Arabidopsis and soybean. Bmc Plant Biol 19(1):506
Zhu Z, Liang H, Chen G, Li F, Wang Y, Liao C, Hu Z (2019) The bHLH transcription factor SlPRE2 regulates tomato fruit development and modulates plant response to gibberellin. Plant Cell Rep 38(9):1053–1064
Zouine M, Fu Y, Chateigner-Boutin AL, Mila I, Frasse P, Wang H, Audran C, Roustan JP, Bouzayen M (2014) Characterization of the tomato ARF gene family uncovers a multi-levels post-transcriptional regulation including alternative splicing. PLoS ONE 9(1):e84203
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31872121), and the Natural Science Foundation of Chongqing of China (csts2019jcyj-msxmX0094), and the Innovation project of people returned from studying abroad of Chongqing (cx2019158).
Author information
Authors and Affiliations
Contributions
ZH and QX designed and managed the research work and improved the manuscript. YC, HY designed the experiments and analyzed the data. YC, BT, FL, GC performed the experiments. YC wrote the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors have read and approved this version of the article, and due care has been taken to ensure the integrity of this work. All the authors have declared no conflict of interest.
Additional information
Communicated by Sukhpreet Sandhu.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chen, Y., Yang, H., Tang, B. et al. The AP2/ERF transcription factor SlERF.J2 functions in hypocotyl elongation and plant height in tomato. Plant Cell Rep 42, 371–383 (2023). https://doi.org/10.1007/s00299-022-02963-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-022-02963-x