Abstract
Key message
We report that GhWRKY21, a WRKY transcription factor, plays essential roles in regulating the intensity of the drought-induced ABA signalling pathway by facilitating the expression of GhHAB in cotton (Gossypium hirsutum).
Abstract
Abscisic acid (ABA) is one of the most important plant hormones in response to abiotic stress. However, activation of the ABA signalling pathway often leads to growth inhibition. The mechanisms that regulate the intensity of ABA signals are poorly understood. Here, we isolated and analysed the cotton group IId WRKY transcription factor (TF) gene GhWRKY21. Functional analysis indicated that GhWRKY21 plays a negative role in the drought response of cotton. Silencing of GhWRKY21 in cotton dramatically increased drought tolerance, whereas ectopic GhWRKY21 overexpression in Nicotiana benthamiana decreased drought tolerance. Furthermore, the GhWRKY21-mediated drought tolerance was ABA dependent. To clarify the mechanism underlying the GhWRKY21-mediated regulation of drought tolerance, 17 clade-A-type type 2C protein phosphatase (PP2C) genes, which are negative regulators of ABA signalling, were identified in cotton. Notably, GhWRKY21 interacted specifically with the W-box element within the promoter of GhHAB and regulated its expression. Silencing of GhHAB in cotton yielded a phenotype similar to that of GhWRKY21-silenced cotton. These results suggest that GhWRKY21 regulates the intensity of ABA signals by facilitating the expression of GhHAB. In summary, these findings dramatically improve our understanding of the function of WRKY TFs and provide insights into the mechanism of ABA-mediated drought tolerance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
To accommodate dynamic growth environments, plants employ complex regulatory mechanisms to quickly optimize their response to environmental stress (Li et al. 2013). Phytohormones are key signals that regulate a series of plant physiological processes in response to various environmental stresses (Fujita et al. 2006; Yang et al. 2017). Abscisic acid (ABA) is a crucial phytohormone that is largely involved in response to abiotic stress, such as high salinity, extreme temperature and drought, in plants (Finkelstein et al. 2002; Fujita et al. 2006). ABA is often referred to as a “stress hormone” and plays essential roles in integrating a wide range of stress signals and in regulating multiple downstream stress responses (Assmann and Jegla 2016; Zhu 2016; Rodrigues et al. 2017). ABA is suppressed by uridine diphosphate glucosyltransferase (SlUGT75C1), which impacts ABA‐mediated drought responses and fruit ripening in tomato (Sun et al. 2017). TabHLH1 participates in the ABA-dependent signalling pathway and mediates plant adaptations to osmotic stress (Yang et al. 2016). ABA is generated or accumulated in guard cells to give rise to stomatal closure, thus leading to the retainment of plant moisture; this has been confirmed in Arabidopsis ABA-deficient mutant aba3-1 (Lee and Luan 2012; Bauer et al. 2013). However, mounting evidence has indicated that, although ABA plays essential roles in increasing plant tolerance to abiotic stress, activation of the ABA signalling pathway often leads to growth inhibition (Julkowska and Testerink 2015; Wang et al. 2018c; Ma et al. 2018). Under stress, the ABA-mediated signalling pathway phosphorylates TOR kinase, which then contributes to ABA- and stress-induced growth inhibition (Wang et al. 2018c). The ABA signalling pathway interacts with the auxin and gibberellin signalling pathways to control root development (Gou et al. 2010; Zhao et al. 2014). ABA also causes dephosphorylation of the plasma membrane H+-ATPase, which inhibits hypocotyl elongation in Arabidopsis (Hayashi et al. 2014). Overexpression of MPK3 increased ABA sensitivity in ABA-induced growth retardation after germination (Lu et al. 2002). Therefore, the intensity of ABA signals must be strictly controlled in plants. Although many studies have explored this topic, the mechanism underlying the regulation of the ABA signalling pathway remains unclear.
In Arabidopsis, five core components drive signal perception and conduction in the classic ABA signalling pathway: ABA receptors (pyrabactin resistance 1 [PYR]/PYR1-like [PYL]/regulatory component of ABA receptor [RCAR]), clade-A-type type 2C protein phosphatases (PP2Cs), snf1-related protein kinase 2 (SnRK2), ABA-responsive transcription factors (TFs) and ABA-responsive genes (Ma et al. 2009; Fujii et al. 2009; Cutler et al. 2010; Hauser et al. 2011). When receptors perceive ABA, clade-A-type PP2Cs bind to the receptor complex, which in turn inhibits clade-A-type PP2Cs (Park et al. 2009; Umezawa et al. 2009). Inhibition of clade-A-type PP2C activity increases the activation of SnRK2 and then targets downstream ABA-responsive TFs and ABA-responsive genes (Fujii et al. 2009; Park et al. 2009). In the absence of ABA, clade-A-type PP2Cs, through physical interaction and dephosphorylation, maintain SnRK2 kinases in an inactive state (Soon et al. 2012). Clade-A-type PP2Cs are key regulators of ABA signal transduction, negatively regulating ABA responses (Umezawa et al. 2009; Ma et al. 2009; Park et al. 2009). In Arabidopsis, six clade-A-type PP2Cs (ABI1, ABI2, HAB1, HAB2, AHG1, AHG3) have been demonstrated to be associated with ABA signalling (Schweighofer et al. 2004; Fujii et al. 2009; Wang et al. 2018b). EAR1 affects ABA signalling by interacting with the N-termini of clade-A-type PP2Cs and increasing their activity (Wang et al. 2018b). PUB12/13 specifically targets ABI1 for degradation to alter the ABA response (Kong et al. 2015). Proteasomal degradation of clade-A-type PP2Cs was shown to be promoted by BPM3 and BPM5, which counteracted the clade-A-type PP2C accumulation induced in response to ABA to desensitize ABA signalling (Julian et al. 2019).
WRKY TFs compose a plant-specific family of TFs and are kinds of key regulators of hormone-mediated signalling pathways (Ülker and Somssich 2004; Rushton et al. 2010; Jiang et al. 2017). WRKY TFs share the highly conserved motif (WRKYGQK) and a specific zinc finger-like motif, CX4-5CX22-23HXH (Yamasaki et al. 2005). According to their specific structural domains, WRKY TFs are classified into three groups, of which the second group can be divided into five subgroups (IIa-e) (Eulgem et al. 2000; Rushton et al. 2010). WRKY TFs bind to W-box (C/TTGACT/C) elements to regulate the expression of downstream genes and as such play a crucial role in response to various environmental stresses (Eulgem et al. 2000; Jiang et al. 2017). WRKY17 and WRKY11 are negative regulators of biological stress resistance and are involved in the regulation of jasmonic acid (JA)-dependent Pst-induced responses (Journot-Catalino et al. 2006). Overexpression of AtWRKY57 increases the drought tolerance of plants. AtWRKY57 can directly bind to the W-box element within the promoters of NCED3 and RD29A and positively regulate their expression (Jiang et al. 2012). When wheat (Triticum aestivum) is exposed to high temperature, the transcription level of TaWRKY70 increases, which affects the resistance of the species to stripe rust (Wang et al. 2017b). Overexpression of GhWRKY17 markedly decreases drought resistance and salt tolerance of Nicotiana benthamiana (Yan et al. 2014). Although a large number of studies have demonstrated the function of WRKY proteins, the regulatory mechanisms underlying the response of WRKY TFs to stress still need further study.
As the main raw material of the cotton textile industry, cotton (Gossypium hirsutum) is involved in many industrial supply chains and is an economically important crop species (Qaim and Zilberman 2003; Fang et al. 2017). However, both the production and quality of cotton are often affected by drought stress (Ullah et al. 2017). Drought can disrupt photosynthesis, water metabolism, and membrane stability and can increase photorespiration, severely threatening crop production (Krasensky and Jonak 2012; Zhang et al. 2018). Therefore, it is important to understand the molecular mechanism of drought resistance of cotton. In our study, a group IId WRKY gene, GhWRKY21, was isolated from cotton, and the mechanism by which GhWRKY21 regulates drought tolerance was verified. Moreover, the results from biochemical and a series of genetic analyses indicated that GhWRKY21 negatively regulates ABA-mediated drought tolerance by directly interacting with the W-box element within the promoter of GhHAB. Our study not only broadens the knowledge of the role of group IId WRKY TFs but also provides new insight for future studies of the ABA-mediated abiotic stress mechanism.
Materials and methods
Plants and growth conditions
Cotton (G. hirsutum L. cv. Lumian 22) and N. benthamiana were the main plant materials used in this study. Cotton seeds were covered with wet cheesecloth until they germinated, and the budlets were transplanted into pots or in water under greenhouse conditions consisting of a temperature of 26 ± 3 °C, a 16 h light (illumination intensity of 200 mmol m–2 s−1)/8 h dark photoperiod and a relative humidity of 65–75%. N. benthamiana seeds were sown and grown in soil-based mixture [peat moss/perlite/vermiculite/general soil (1/1/1/1, v/v/v/v)] and were irrigated with tap water. The greenhouse conditions were as follows: temperature, 24 °C; photoperiod, 16 h light (illumination intensity of 100 μmol m−2 s−1)/8 h darkness; and relative humidity, 50 ± 10%.
GhWRKY21 isolation, bioinformatic analysis, and plant transformation
The GhWRKY21 open reading frame (ORF) was cloned and isolated via PCR (the relevant primer sequences are listed in Table S1). The amino acid sequence was acquired from the NCBI database (https://blast.ncbi.nlm.nih.gov/Blast.cgi). PlantCARE was subsequently used to analyse the promoter sequence, and cluster analysis was performed by MEGA 5.0 software (via the neighbour-joining method).
The full-length GhWRKY21 cDNA sequence connected to the cauliflower mosaic virus (CaMV) 35S promoter was inserted into a pBI121 plant expression vector. The Agrobacterium-mediated leaf disc method (Zhang et al. 2011; Lu et al. 2013) was then used to obtain transgenic N. benthamiana.
Subcellular localization analysis of GhWRKY21
35S::GhWRKY21-GFP recombinant plasmids were constructed (the relevant primer sequences are listed in Table S1). A control 35S::GFP plasmid and the 35S::GhWRKY21-GFP recombinant plasmid were transferred into the Agrobacterium tumefaciens strain GV3101. To achieve transient expression, two A. tumefaciens strains were prepared according to a previously described method (Wang et al. 2017a), and 8-week-old leaves of N. benthamiana were used. After routine culture for 68–72 h, the fluorescent signals were observed with confocal microscopy.
Agrobacterium-mediated VIGS and transient overexpression in protoplasts
On the basis of the method of virus-induced gene silencing (VIGS) described by Gu et al. (2014), the specific sequence of GhWRKY21 or GhHAB was inserted into a pCLCrV-A vector (the relevant primer sequences are listed in Table S1). The recombinant plasmid was subsequently transformed into A. tumefaciens strain EHA105. Seven-day-old cotton cotyledons that had fully expanded were inoculated with a mixed Agrobacterium suspension containing a pCLCrV-A recombinant vector (pCLCrV-A-GhWRKY21 or pCLCrV-A-GhHAB) and pCLCrV-B in equal amounts. A mixed Agrobacterium suspension containing pCLCrV-B and pCLCrV-A was used for a control inoculation. Three weeks after inoculation, the silenced cotton plants were subjected to functional analysis. Protoplasts were isolated from 7-day-old cotton cotyledons. The cotyledons were sliced and digested based on the method described by Wang et al. (2018a) to obtain protoplasts. Transformation of protoplasts was performed as described by Wang et al. (2017a). Each assay was independently repeated at least three times.
Drought stress treatment
For the germination and root growth assay, the GhWRKY21 overexpression (OE) plants and Vec seeds of tobacco were used. After their surface was disinfected with 75% ethanol and 26% sodium hypochlorite, the seeds were spread on Murashige and Skoog (MS) media that consisted of different concentrations of mannitol and sodium tungstate (Tu) for daily measurements of germination rates. Additionally, when the radicle developed after approximately 3 days, the buds were transplanted to different MS media (supplemented with mannitol or mannitol + Tu) and were oriented perpendicularly in an incubator for measurements of taproot lengths.
For the drought response assay, 5-week-old cotton, 4-week-old tobacco and 8-week-old tobacco were used. The cotton grown in soil was completely deprived of water for 7 days (5 days for tobacco), and the leaves were collected for RNA extraction or chemical staining analysis. Leaves of 5-week-old cotton plants were selected and weighed immediately to determine their fresh weight. The detached leaves were incubated at room temperature in darkness to record the weight changes at designated times. Eight-week-old tobacco exposed to drought stress was also subjected to the same water loss rate assay. In addition, 4-week-old transgenic tobacco was subjected to progressive drought for 5 days to collect images of their phenotype. The survival percentages were subsequently documented after the plants had been re-watered for 2 days.
All the experiments were independently repeated, each involving at least three replicates.
RNA isolation and qRT-PCR-based analysis
A Plant RNA EasySpin Plus Kit (Aidlab, Beijing, China) and a TRIzol reagent (Invitrogen, Carlsbad, CA, USA) were used to extract the total RNA from cotton and N. benthamiana, respectively. A cDNA synthesis kit (Vazyme, Nanjing, China) was used to synthesize single-strand cDNA from the total RNA. qRT-PCR-based analysis was performed with SYBR Premix Ex Taq (Takara) via a CFX96™ Real-time Detection System (Bio-Rad). The standard controls included UBI from G. hirsutum and the β-actin gene from N. benthamiana (the sequences of the primers are listed in Table S1).
Y1H assays
Yeast one-hybrid (Y1H) assays were performed with the Y1H strain as described by the manufacturer of the MATCHMAKER One-Hybrid System (Clontech). The aureobasidin A (AbA) antibiotic reporter gene with high screening intensity was used in the system. The ORF of GhWRKY21 was inserted into the GAL4 activation domain in a pGADT7 prey vector. ProPP2Cs-pAbAi was generated with the pAbAi reporter vector and the clade-A-type PP2C promoter, which contains 2 bait sequences of repeated sequences of target DNA (W-box elements). The linearized bait plasmids of proPP2C-pAbAi were integrated into the genome of yeast to generate a proPP2C-pAbAi-Y1H strain. The minimum AbA concentration that completely inhibited colony growth was 600 ng ml−1. The GhWRKY21-AD and pGADT7 plasmids were transformed into the proPP2C-pAbAi-Y1H strain, which was subsequently cultured on media lacking Ura (-Ura) or Leu (-Leu). The transformed strains were then screened for 3 days on -Leu media supplemented with 600 ng ml−1 AbA (-Leu/600 ng ml−1 AbA).
Recombinant protein expression, purification and EMSAs
To express protein in Escherichia coli Transetta (DE3), the ORF of GhWRKY21 was inserted into a pET30a vector, after which the pET-30a ( +)-GhWRKY21 plasmid was transformed into E. coli Transetta. A final concentration of 2 mM isopropyl-1-thio-β-galactopyranoside (IPTG) was selected for the liquid coliform bacillus at OD600 values of 0.4–0.6. After being incubated for an additional 10 h at 16 °C, the cells were collected and centrifuged by 5000 × g for 15 min at 4 °C. The sediment was resuspended in lysates to perform ultrasonic crushing and centrifuged again. A MagneHis™ Protein Purification System (Promega, Madison, WI, USA) was used to purify the recombinant protein. Finally, 12% SDS–polyacrylamide gel electrophoresis (SDS–PAGE) was used to determine the expression levels of the target recombinant protein (Fig. S1). Electrophoretic mobility shift assay (EMSA) experiments based on the manufacturer’s protocol were then performed (LightShift™ Chemiluminescent EMSA Kit, Thermo, USA).
Results
Characterization of GhWRKY21 in cotton
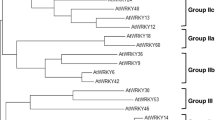
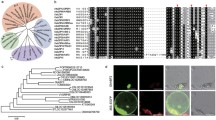
Our previous studies have indicated that group IId WRKY TFs play significant roles in regulating resistance to environmental stress in cotton (Yu et al. 2012; Yan et al. 2014; Shi et al. 2014). To clarify the function of group IId WRKY TFs more comprehensively, GhWRKY21 (GenBank Number: KT983420) was isolated from cotton. The full-length cDNA sequence of GhWRKY21 is 1494-bp in length, comprising a 173-bp 5′-untranslated region (UTR), a 324-bp 3′-UTR and a 996-bp ORF. Multiple-sequence analysis revealed that GhWRKY21 shared high identities (57.59–81.30%) with its homologous sequences (Fig. 1a). GhWRKY21 contains the single classic WRKY DNA-binding domain (WRKYGQK) and a conserved C-terminal motif, which showed that GhWRKY21 belongs to the group IId WRKY subfamily (Fig. 1a). Analysis of the phylogenetic tree also confirmed that GhWRKY21 belongs to the group IId WRKY subfamily (Fig. 1b). In addition, a nuclear localization signal (NLS) KKRK was identified at amino acids 289–292 (Fig. 1a). To verify the intracellular localization of GhWRKY21, a 35S::GhWRKY21-GFP fusion plasmid was constructed (Fig. S2a). The control 35S::GFP plasmid and the 35S::GhWRKY21-GFP plasmid were transiently expressed separately via agroinfiltration in N. benthamiana to detect the fluorescent signals. As shown in Fig. S2b, the fluorescent signals from 35S::GFP were visible in the cytoplasm and nucleus in multiple cells, and the fluorescent signals from 35S::GhWRKY21-GFP were visible specifically in the nucleus.
Sequence characterization and phylogenetic analysis of GhWRKY21. a Amino acid sequence alignment of WRKY21 from different plant species. The black shadows indicate the same amino acids. The WRKY domain is marked by a double-headed arrow, and the highly conserved amino acid sequence WRKYGQK is boxed. The conserved C-terminal motif is indicated by the frame. The putative nuclear localization signal, KKRK, is marked by asterisks. b Phylogenetic tree of the WRKY protein. GhWRKY21 is boxed. The tree was constructed with MEGA 5.0
GhWRKY21 negatively regulates the drought response of cotton
To explore the potential function of GhWRKY21 in response to environmental stress, GhWRKY21-silenced cotton lines were generated according to the Agrobacterium-mediated VIGS technique. After infiltration for 21 days, the GhWRKY21 expression level was typically reduced (Fig. 2c). The CRV::00 (empty vector control) and GhWRKY21-silenced (CRV::GhWRKY21-01/02) cotton lines were exposed to drought (water was withheld). Seven days later, the CRV::00 cotton plants displayed severe wilting, while the CRV::GhWRKY21 cotton plants were not as strongly affected (Fig. 2a). 3,3′-Diaminobenzidine (DAB) staining also revealed that the degree of cell damage in the leaves of CRV::00 cotton plants was more severe than that in the leaves of CRV::GhWRKY21 cotton plants after drought treatment (Fig. 2b). Further experiments showed that, compared with those of the CRV::00 cotton plants, detached leaves of the CRV::GhWRKY21 cotton plants presented a lower water loss rate, which matched the drought-tolerant phenotype (Fig. 2d). These loss-of-function experimental data indicated that GhWRKY21 is a negative regulator of cotton tolerance to drought stress.
Silencing of GhWRKY21 reduced the sensitivity to drought stress in cotton. a Phenotypes of GhWRKY21-silenced plants after drought stress. b Phenotypes of leaves of GhWRKY21-silenced plants after drought treatment. c Expression level of GhWRKY21 in GhWRKY21-silenced plants. d Relative water loss rates of the leaves of CRV::00 and CRV::GhWRKY21 plants. The data are presented as the means ± SEs of three independent experiments. The different letters in c indicate significant differences (P < 0.05) according to Tukey’s HSD test
Overexpression of GhWRKY21 enhances susceptibility to drought stress in transgenic N. benthamiana
To further explore the function of GhWRKY21 under drought stress, GhWRKY21 OE transgenic N. benthamiana were generated, and two independent overexpression lines (#2 and #4, named OE2 and OE4, respectively) were identified by kanamycin selection and qRT-PCR-based analysis (Fig. S3). T2 transgenic and Vec plants were planted for further experiments.
Seeds of the two lines of transgenic plants (OE2, OE4) and the Vec plants were sterilized and sown on MS agar media consisting of different concentrations of mannitol. As shown in Fig. 3a, the approximate germination percentage and growth status were recorded for the Vec and OE plants when no mannitol treatment was applied. Compared with those of the Vec plants, the seeds of the OE plants on the MS media with the increased mannitol concentration germinated later, and the seedlings grew lower (Figs. 3a and S4). Notably, the germination percentage of the OE plants was less than 60% of that of the Vec plants under 200 mM mannitol (Fig. 3c). In addition, the roots of the OE plants were shorter than those of the Vec plants (Fig. 3b). These results showed that GhWRKY21 overexpression increases susceptibility to drought stress during seed germination.
Overexpression of GhWRKY21 increased the sensitivity to drought stress in transgenic N. benthamiana. a, c Seed germination phenotype and germination percentages between the Vec and OE plants exposed to 0 and 200 mM mannitol. b Taproot phenotype of Vec and OE plants exposed to 0 and 200 mM mannitol. d Phenotypes of 4-week-old Vec and OE plants under drought stress. e Survival rates of 4-week-old plants after 2 days of re-watering following drought treatment. f Relative water loss rates of the leaves of Vec and OE plants. g Representative phenotypes of 8-week-old Vec and OE plants before and after drought treatment. h Images showing DAB and NBT staining of leaves of 8-week-old plants after drought. The data are presented as the means ± SEs of three independent experiments. The different letters in e indicate significant differences (P < 0.05) according to Tukey’s HSD test
Wilting is a typical phenotype of plants experiencing drought stress. To determine the drought response of the transgenic plants, 4- and 8-week-old tobacco plants were exposed to drought conditions (water was withheld). After drought treatment for 5 days, the 4-week-old OE plants began to wilt, whereas the Vec plants displayed mild symptoms (Fig. 3d). When the plants were re-watered, the survival of the OE plants (69.7%) was lower than that of the Vec plants (92%) (Fig. 3e). Additionally, after irrigation was withheld for 5 days, the 8-week-old OE plants exhibited more leaf wilting than did the Vec plants, and histochemical staining with DAB and nitro blue tetrazolium (NBT) revealed that the cell damage of the OE plants was more severe than that in Vec plants (Fig. 3g, h). Moreover, compared with the Vec plants, the OE plants had a higher water loss rate (Fig. 3f). These results collectively showed that GhWRKY21 is a negative regulator that affected the drought tolerance of the transgenic plants.
GhWRKY21-mediated drought tolerance in cotton is ABA dependent
To explore the mechanism of action of GhWRKY21 in response to drought stress, an effective ABA biosynthesis inhibitor (Tu) was used. On MS media supplemented with 5 mM Tu, the germination of seeds from OE and Vec plants did not distinctly differ. However, on MS media supplemented with 200 mM mannitol, compared with that of the Vec plants, the germination of seeds from the OE plants was lower (Fig. 4a and c). When 5 mM Tu was added to the media that were supplemented with 200 mM mannitol, the drought sensitivity of the OE plants decreased (Fig. 4a and c). Moreover, the difference in root length between the Vec and OE plants on MS media supplemented with both 200 mM mannitol and 5 mM Tu diminished (Fig. 4b and d). Furthermore, the expression level of GhWRKY21 in 7-day-old cotton plants was upregulated in response to ABA and peaked (3.5-fold induction) at 10 h after ABA treatment (Fig. 4e). Changes in stomatal aperture of the Vec and OE plants in response to ABA were then measured. In the absence of ABA, the degree of stomatal closure did not significantly differ between the OE and Vec plants (Fig. 4f, g). ABA was added to the solution, and the plants were exposed to light for 3 h; the degree of stomatal closure of the OE plants was lower than that of the Vec plants (Fig. 4f, g), the latter of which was highly similar to the stomatal aperture observed under drought-stress conditions (Fig. S5). The above results indicate that the sensitivity of GhWRKY21 to drought stress is related to ABA in the transgenic plants.
GhWRKY21 was involved in ABA-mediated drought tolerance in cotton. a, c Seed germination and germination percentages of Vec and OE plants grown on MS media consisting of 200 mM mannitol or 200 mM mannitol with 5 mM Tu. b, d Root elongation of Vec and OE plants after treatment with 200 mM mannitol or 200 mM mannitol with 5 mM Tu. e Expression pattern of GhWRKY21 after ABA treatment. f, g Stomatal aperture of Vec and OE plants in response to ABA treatment. The data represent the means ± SEs of 40 stomata (length/width) of three independent experiments. The different letters in d, e and g indicate significant differences (P < 0.05) according to Tukey’s HSD test
Identification of clade-A-type PP2Cs in cotton
Clade-A-type PP2Cs are core negative regulators of ABA signalling (Ma et al. 2009; Umezawa et al. 2009; Park et al. 2009). We speculated that GhWRKY21 affects ABA-mediated drought responses by regulating the expression level of clade-A-type PP2C genes. To verify this supposition, we identified a total of 17 apparent clade-A-type PP2C genes throughout the whole cotton genome and cloned their promoters. A phylogenetic tree was subsequently constructed that indicated that all the clade-A-type PP2C proteins in cotton were highly homologous to clade-A-type PP2Cs in Arabidopsis, which have been widely suggested to be associated with ABA signal transduction (Fig. 5a). W-box is typical cis-element that binds WRKY TFs. Analysis of the promoters of the clade-A-type PP2C genes revealed that eight clade-A-type PP2Cs (CotAD_00293, CotAD_39701, CotAD_09702, CotAD_53998, CotAD_22992, CotAD_60815, CotAD_06730, CotAD_21748) contain W-boxes, and these were classified into seven types according to their surrounding bases (Fig. 5b).
Phylogenetic relationships, conserved motifs, and W-box within the promoters of clade-A-type PP2C genes. a Phylogenetic tree and conserved motifs of clade-A-type PP2Cs in G. hirsutum and Arabidopsis. The tree was constructed with MEGA 5.0, and conserved motifs are displayed by 20 different coloured boxes. b Sequence of the W-box element within the promoters of clade-A-type PP2Cs. The W-boxes are indicated by small red bars, and the surrounding bases follow
GhWRKY21 regulates the expression of GhHAB by directly binding to the W-box element within its promoter
To prove that GhWRKY21 can directly interact with the promoter of clade-A-type PP2Cs, a Y1H analysis was conducted. The results indicated that only the pro00293-pAbAi-Y1H strain transformed with the GhWRKY21-AD combination plasmid grew well in -Leu media supplemented with 600 ng ml−1 AbA, while the rest did not survive (Fig. 6a). Database queries and bioinformatic analyses revealed that pro00293 was the W-box element within the promoter of both CotAD_00293 and CotAD_39701 (designated GhHAB), which are homologous genes in the cotton genome. An EMSA was performed to further confirm the interaction. The results indicated that GhWRKY21 bound to the W-box element within the promoter of GhHAB, and the bands diminished when a part of the biotin probe was replaced with a cold probe, showing that the binding was specific (Fig. 6b). The expression pattern of GhHAB was then measured and found to be induced after ABA treatment (Fig. S6). Moreover, overexpression of GhWRKY21 in protoplasts of cotton increased the expression level of GhHAB (Fig. 6c), whereas silencing of GhWRKY21 in cotton reduced the expression level of GhHAB (Fig. 6d). According to these data, GhWRKY21 directly binds to the W-box element within the promoter of GhHAB to regulate its expression.
GhWRKY21 interacted with the W-box element within the promoter of GhHAB. a GhWRKY21 interaction with the W-box element within the promoter of GhHAB according to Y1H analysis. b Interaction between the GhWRKY21 protein and the promoter fragments of GhHAB according to an EMSA. c Expression level of GhHAB in GhWRKY21 OE protoplasts. d Expression level of GhHAB in GhWRKY21-silenced cotton. The data are presented as the means ± SEs of three independent experiments. The different letters in c and d indicate significant differences (P < 0.05) according to Tukey’s HSD test
GhHAB negatively regulates the drought response of cotton
To verify the function of GhHAB in response to drought tolerance in cotton, GhHAB-silenced cotton plants were generated according to the VIGS technique (Fig. 7c). The GhHAB-silenced (CRV::GhHAB-01/02) and CRV::00 (empty vector control) cotton plants were exposed to drought conditions (water was withheld) for 7 days, and the CRV::00 cotton plants were more shrivelled than were the CRV::GhHAB cotton plants (Fig. 7a). There was no obvious difference between the leaves of the CRV::00 and CRV::GhHAB plants under the control conditions, while the leaves of the CRV::GhHAB plants exhibited less damage than the leaves of the CRV::00 cotton did after drought treatment (Fig. 7b). In addition, the water loss rate was measured, and the results indicated that, compared with those of the those from the CRV::00 plants, the detached leaves of the CRV::GhHAB plants presented a lower water loss rate (Fig. 7d). These loss-of-function experimental data indicated that GhHAB is a negative regulator of drought resistance in cotton.
Silencing of GhHAB increased the tolerance to drought in cotton. a Phenotypes of GhHAB-silenced plants after drought stress. b Phenotypes of leaves of GhHAB-silenced plants after drought treatment. c Expression level of GhHAB in GhHAB-silenced cotton. d Relative water loss rates were calculated by the use of detached leaves of CRV::00 and CRV::GhHAB plants. The data are presented as the means ± SEs of three independent experiments. The different letters in c indicate significant differences (P < 0.05) according to Tukey’s HSD test
Discussion
WRKY TFs are among the most important TFs in plants and have been reported in various model plant species. Group IId WRKY TFs play a crucial role in regulating resistance to environmental stress in plants. In Arabidopsis, wrky11 and wrky17 mutants exhibit enhanced basal resistance in the regulation of Pst induction, which indicated that WRKY11 and WRKY17 play a negative regulatory role in response to biotic stress (Journot-Catalino et al. 2006). Salt and drought stress can induce the expression of ZmWRKY58 in Zea mays, and overexpression of ZmWRKY58 was shown to reduce salt and drought sensitivity in transgenic rice by enhancing the relative water content and survival (Cai et al. 2014). To clarify the function of group IId WRKY TFs in cotton, many genes have been isolated. Our previous studies indicated that overexpression of GhWRKY17 in tobacco increases salt and drought susceptibility through ABA signalling and regulates the production of ROS (Yan et al. 2014). Additionally, increased salt tolerance and resistance to pathogenic bacteria of GhWRKY39 OE tobacco indicated that GhWRKY39 is a positive regulator of the plant stress response (Shi et al. 2014). Yu et al. (2012) reported that overexpression of GhWRKY15 not only increases the resistance of tobacco to fungal or viral pathogens by increasing the transcript levels of pathogen-related genes but also promotes growth by elongating the stems at the early shoot stages of transgenic tobacco (Yu et al. 2012). The above results indicate that the functions of group IId WRKY TFs are complex and are redundant or divergent. Therefore, exploring the function of IId WRKY subfamily genes in detail in cotton is necessary. In this study, we isolated a group IId WRKY TF gene, GhWRKY21 (Fig. 1). Silencing of GhWRKY21 increased the cotton tolerance to drought (Fig. 2). Further experiments revealed that, compared with that of the Vec plants, the drought tolerance of the GhWRKY21 OE plants was weaker, as reflected by their low germination percentage, increased DAB and NBT stain accumulation, and poor survival under drought stress (Fig. 3). These results imply that GhWRKY21 plays a negative role in regulating the drought response in cotton.
ABA is a crucial phytohormone that is involved in the plant response to drought stress. Extensive efforts have been made to discover the mechanisms of ABA signalling, and substantial progress has been made. In the absence of ABA, SnRK2 kinases are maintained in an inactive state by PP2Cs through physical interaction and dephosphorylation. In contrast, ABA gives rise to receptors (PYR/PYL/RCAR) that undergo a conformational change that allows clade-A-type PP2C, core negative regulators of ABA signalling, to bind to the receptor complex (Boudsocq et al. 2007; Cutler et al. 2010; Li et al. 2013; Wang et al. 2018c). Drought stress induces ABA synthesis in plants. Activation of the ABA signalling pathway can affect the germination of seedlings, the growth of roots and interactions with auxin, all of which can inhibit plant growth and development (Lu et al. 2002; Gou et al. 2010; Hayashi et al. 2014; Zhao et al. 2014; Wang et al. 2018a). Analysis of the molecular mechanism underlying the regulation of the ABA signalling pathway is highly important for further elucidation of ABA-mediated drought tolerance of plants. Previous studies have suggested that WRKY TFs acts as key nodes in ABA-responsive signalling pathway (Chen et al. 2010; Ren et al. 2010; Shang et al. 2010; Rushton et al. 2012). Shang et al. (2010) indicated that WRKY40 binds to the promoter of ABI5 and represses its expression. WRKY63 has been shown to regulate the expression of AREB1/ABF2 through binding to the W-box within its promoter (Ren et al. 2010). Moreover, WRYK18 and WRKY40 are induced by ABA signalling, which induces WRKY60 through binding to its W-box (Chen et al. 2010). In the present study, germination and root growth were obviously inhibited by mannitol in the GhWRKY21 OE transgenic plants, but with the addition of Tu, a widely used inhibitor of ABA biosynthesis, the inhibition diminished (Fig. 4). In addition, a small extent of stomatal closure was observed in the OE plants after ABA treatment (Fig. 4). Taken together, these results implied that, as a negative regulatory factor, GhWRKY21 functions in ABA-mediated drought tolerance and may be one of the important mechanisms involved in mitigating the intensity of the ABA signalling pathway.
Clade-A-type PP2Cs have been annotated as negative regulators of the ABA response (Umezawa et al. 2009). Inhibition of clade-A-type PP2C activity facilitates activation of SnRK2 protein kinases and then activates downstream ABA-responsive TFs and increases the expression of ABA-responsive genes (Fujii et al. 2009; Park et al. 2009). Kong et al. (2015) indicated that ABI1 was degraded specifically by PUB12/13 to desensitize ABA signalling. Increased accumulation of ABI1 resulted in increased loss of water in pub12pub13 mutants, which were more sensitive to drought stress than were wild-type plants. BPM3 and BPM5 promote proteasomal degradation of clade-A-type PP2Cs, and the sensitivity to ABA-mediated inhibition of seedling establishment and root growth decreased in bpm3bpm5 mutants compared with wild-type plants (Julian et al. 2019). In addition to components that degrade clade-A-type PP2Cs, a component that increases an ABA co-receptor (EAR1) has been identified (Wang et al. 2018b). The N-termini of clade-A-type PP2Cs interact with EAR1 to increase the activity of the PP2Cs during ABA signalling, which plays a part in seed germination, primary root growth, and drought response (Wang et al. 2018b). To illustrate the molecular mechanism underlying the negative regulation of ABA signalling by GhWRKY21, we identified all clade-A-type PP2CA genes throughout the whole cotton genome. The promoters of these clade-A-type PP2Cs were subsequently cloned. On the basis of the Y1H assay and EMSA results, GhWRKY21 binds specifically to the W-box element within the promoter of GhHAB to regulate its expression (Fig. 6). Further experiments revealed that GhWRKY21 negatively regulates ABA-mediated drought tolerance by fine-tuning the expression of GhHAB.
To promote plant growth appropriately under abiotic stress, ABA stress signalling must be suppressed. EL1-like (AEL) casein kinases phosphorylate PYLs at partially conserved sites and promote both the ubiquitination and degradation of PYLs, the process of which suppresses ABA signalling (Chen et al. 2018). PR5 receptor-like kinase 2 (PR5K2) phosphorylates ABI1/2 and increases its protein phosphatase activity to repress ABA and stress signalling (Baek et al. 2019). In our study, a new mechanism for fine-tuning ABA signalling was proposed. The results of our study indicated that GhWRKY21 regulates ABA-mediated drought tolerance of cotton by facilitating the expression of GhHAB. Future exploration of the role of WRKY TFs in ABA response will help reveal unknown regulatory mechanisms involved in ABA signalling, and our study provides new insight into the mechanism of ABA-mediated drought tolerance.
References
Assmann SM, Jegla T (2016) Guard cell sensory systems: recent insights on stomatal responses to light, abscisic acid, and CO2. Curr Opin Plant Biol 33:157–167. https://doi.org/10.1016/j.pbi.2016.07.003
Baek D, Kim MC, Kumar D et al (2019) AtPR5K2, a PR5-Like receptor kinase, modulates plant responses to drought stress by phosphorylating protein phosphatase 2Cs. Front Plant Sci 10:1146. https://doi.org/10.3389/fpls.2019.01146
Bauer H, Ache P, Lautner S et al (2013) The stomatal response to reduced relative humidity requires guard cell-autonomous ABA synthesis. Curr Biol 23:53–57. https://doi.org/10.1016/j.cub.2012.11.022
Boudsocq M, Droillard MJ, Barbier-Brygoo H, Lauriere C (2007) Different phosphorylation mechanisms are involved in the activation of sucrose non-fermenting 1 related protein kinases 2 by osmotic stresses and abscisic acid. Plant Mol Biol 63:491–503. https://doi.org/10.1007/s11103-006-9103-1
Cai R, Zhao Y, Wang Y, Lin Y, Peng X, Li Q, Chang Y, Jiang H, Xiang Y, Cheng B (2014) Overexpression of a maize WRKY58 gene enhances drought and salt tolerance in transgenic rice. Plant cell. Tissue Org Cult (PCTOC) 119:565–577. https://doi.org/10.1007/s11240-014-0556-7
Chen H, Lai Z, Shi J, Xiao Y, Chen Z, Xu X (2010) Roles of arabidopsis WRKY18, WRKY40 and WRKY60 transcription factors in plant responses to abscisic acid and abiotic stress. BMC Plant Biol 10:281. https://doi.org/10.1186/1471-2229-10-281
Chen HH, Qu L, Xu ZH, Zhu JK, Xue HW (2018) EL1-like Casein kinases suppress ABA signaling and responses by phosphorylating and destabilizing the ABA receptors PYR/PYLs in Arabidopsis. Mol Plant 11:706–719. https://doi.org/10.1016/j.molp.2018.02.012
Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR (2010) Abscisic acid: emergence of a core signaling network. Annu Rev Plant Biol 61:651–679. https://doi.org/10.1146/annurev-arplant-042809-112122
Eulgem T, Rushton PJ, Robatzek S, Somssich IE (2000) The WRKY superfamily of plant transcription factors. Trends Plant Sci 5:199–206. https://doi.org/10.1016/s1360-1385(00)01600-9
Fang L, Wang Q, Hu Y et al (2017) Genomic analyses in cotton identify signatures of selection and loci associated with fiber quality and yield traits. Nat Genet 49:1089–1098. https://doi.org/10.1038/ng.3887
Finkelstein RR, Gampala SS, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14(Suppl):S15–45. https://doi.org/10.1105/tpc.010441
Fujii H, Chinnusamy V, Rodrigues A et al (2009) In vitro reconstitution of an abscisic acid signalling pathway. Nature 462:660–664. https://doi.org/10.1038/nature08599
Fujita M, Fujita Y, Noutoshi Y, Takahashi F, Narusaka Y, Yamaguchi-Shinozaki K, Shinozaki K (2006) Crosstalk between abiotic and biotic stress responses: a current view from the points of convergence in the stress signaling networks. Curr Opin Plant Biol 9:436–442. https://doi.org/10.1016/j.pbi.2006.05.014
Gou J, Strauss SH, Tsai CJ, Fang K, Chen Y, Jiang X, Busov VB (2010) Gibberellins regulate lateral root formation in populus through interactions with auxin and other hormones. Plant Cell 22:623–639. https://doi.org/10.1105/tpc.109.073239
Gu Z, Huang C, Li F, Zhou X (2014) A versatile system for functional analysis of genes and microRNAs in cotton. Plant Biotechnol J 12:638–649. https://doi.org/10.1111/pbi.12169
Hauser F, Waadt R, Schroeder JI (2011) Evolution of abscisic acid synthesis and signaling mechanisms. Curr Biol 21:R346–355. https://doi.org/10.1016/j.cub.2011.03.015
Hayashi Y, Takahashi K, Inoue S, Kinoshita T (2014) Abscisic acid suppresses hypocotyl elongation by dephosphorylating plasma membrane H(+)-ATPase in Arabidopsis thaliana. Plant Cell Physiol 55:845–853. https://doi.org/10.1093/pcp/pcu028
Jiang Y, Liang G, Yu D (2012) Activated expression of WRKY57 confers drought tolerance in Arabidopsis. Mol Plant 5:1375–1388. https://doi.org/10.1093/mp/sss080
Jiang J, Ma S, Ye N, Jiang M, Cao J, Zhang J (2017) WRKY transcription factors in plant responses to stresses. J Integr Plant Biol 59:86–101. https://doi.org/10.1111/jipb.12513
Journot-Catalino N, Somssich IE, Roby D, Kroj T (2006) The transcription factors WRKY11 and WRKY17 act as negative regulators of basal resistance in Arabidopsis thaliana. Plant Cell 18:3289–3302. https://doi.org/10.1105/tpc.106.044149
Julian J, Coego A, Lozano-Juste J et al (2019) The MATH-BTB BPM3 and BPM5 subunits of Cullin3-RING E3 ubiquitin ligases target PP2CA and other clade A PP2Cs for degradation. Proc Natl Acad Sci USA 116:15725–15734. https://doi.org/10.1073/pnas.1908677116
Julkowska MM, Testerink C (2015) Tuning plant signaling and growth to survive salt. Trends Plant Sci 20:586–594. https://doi.org/10.1016/j.tplants.2015.06.008
Kong L, Cheng J, Zhu Y et al (2015) Degradation of the ABA co-receptor ABI1 by PUB12/13 U-box E3 ligases. Nat Commun 6:8630. https://doi.org/10.1038/ncomms9630
Krasensky J, Jonak C (2012) Drought, salt, and temperature stress-induced metabolic rearrangements and regulatory networks. J Exp Bot 63:1593–1608. https://doi.org/10.1093/jxb/err460
Lee SC, Luan S (2012) ABA signal transduction at the crossroad of biotic and abiotic stress responses. Plant Cell Environ 35:53–60. https://doi.org/10.1111/j.1365-3040.2011.02426.x
Li J, Besseau S, Toronen P, Sipari N, Kollist H, Holm L, Palva ET (2013) Defense-related transcription factors WRKY70 and WRKY54 modulate osmotic stress tolerance by regulating stomatal aperture in Arabidopsis. New Phytol 200:457–472. https://doi.org/10.1111/nph.12378
Lu C, Han MH, Guevara-Garcia A, Fedoroff NV (2002) Mitogen-activated protein kinase signaling in postgermination arrest of development by abscisic acid. Proc Natl Acad Sci USA 99:15812–15817. https://doi.org/10.1073/pnas.242607499
Lu W, Chu X, Li Y, Wang C, Guo X (2013) Cotton GhMKK1 induces the tolerance of salt and drought stress, and mediates defence responses to pathogen infection in transgenic Nicotiana benthamiana. PLoS ONE 8:e68503. https://doi.org/10.1371/journal.pone.0068503
Ma Y, Szostkiewicz I, Korte A, Moes D, Yang Y, Christmann A, Grill E (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Sci (New York, NY) 324:1064–1068. https://doi.org/10.1126/science.1172408
Ma Y, Cao J, He J, Chen Q, Li X, Yang Y (2018) Molecular mechanism for the regulation of ABA homeostasis during plant development and stress responses. Int J Mol Sci. https://doi.org/10.3390/ijms19113643
Park SY, Fung P, Nishimura N et al (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Sci (New York, NY) 324:1068–1071. https://doi.org/10.1126/science.1173041
Qaim M, Zilberman D (2003) Yield effects of genetically modified crops in developing countries. Sci (New York, NY) 299:900–902. https://doi.org/10.1126/science.1080609
Ren X, Chen Z, Liu Y et al (2010) ABO3, a WRKY transcription factor, mediates plant responses to abscisic acid and drought tolerance in Arabidopsis. Plant J 63:417–429. https://doi.org/10.1111/j.1365-313X.2010.04248.x
Rodrigues O, Reshetnyak G, Grondin A, Saijo Y, Leonhardt N, Maurel C, Verdoucq L (2017) Aquaporins facilitate hydrogen peroxide entry into guard cells to mediate ABA- and pathogen-triggered stomatal closure. Proc Natl Acad Sci USA 114:9200–9205. https://doi.org/10.1073/pnas.1704754114
Rushton PJ, Somssich IE, Ringler P, Shen QJ (2010) WRKY transcription factors. Trends Plant Sci 15:247–258. https://doi.org/10.1016/j.tplants.2010.02.006
Rushton DL, Tripathi P, Rabara RC et al (2012) WRKY transcription factors: key components in abscisic acid signalling. Plant Biotechnol J 10:2–11. https://doi.org/10.1111/j.1467-7652.2011.00634.x
Schweighofer A, Hirt H, Meskiene I (2004) Plant PP2C phosphatases: emerging functions in stress signaling. Trends Plant Sci 9:236–243. https://doi.org/10.1016/j.tplants.2004.03.007
Shang Y, Yan L, Liu ZQ et al (2010) The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 22:1909–1935. https://doi.org/10.1105/tpc.110.073874
Shi W, Liu D, Hao L, Wu C-A, Guo X, Li H (2014) GhWRKY39, a member of the WRKY transcription factor family in cotton, has a positive role in disease resistance and salt stress tolerance. Plant cell. Tissue Org Cult (PCTOC) 118:17–32. https://doi.org/10.1007/s11240-014-0458-8
Soon FF, Ng LM, Zhou XE et al (2012) Molecular mimicry regulates ABA signaling by SnRK2 kinases and PP2C phosphatases. Sci (New York, NY) 335:85–88. https://doi.org/10.1126/science.1215106
Sun Y, Ji K, Liang B et al (2017) Suppressing ABA uridine diphosphate glucosyltransferase (SlUGT75C1) alters fruit ripening and the stress response in tomato. Plant J 91:574–589. https://doi.org/10.1111/tpj.13588
Ülker B, Somssich IE (2004) WRKY transcription factors: from DNA binding towards biological function. Curr Opin Plant Biol 7:491–498. https://doi.org/10.1016/j.pbi.2004.07.012
Ullah A, Sun H, Yang X, Zhang X (2017) Drought coping strategies in cotton: increased crop per drop. Plant Biotechnol J 15:271–284. https://doi.org/10.1111/pbi.12688
Umezawa T, Sugiyama N, Mizoguchi M et al (2009) Type 2C protein phosphatases directly regulate abscisic acid-activated protein kinases in Arabidopsis. Proc Natl Acad Sci USA 106:17588–17593. https://doi.org/10.1073/pnas.0907095106
Wang C, He X, Wang X, Zhang S, Guo X (2017a) ghr-miR5272a-mediated regulation of GhMKK6 gene transcription contributes to the immune response in cotton. J Exp Bot 68:5895–5906. https://doi.org/10.1093/jxb/erx373
Wang J, Tao F, An F et al (2017b) Wheat transcription factor TaWRKY70 is positively involved in high-temperature seedling plant resistance to Puccinia striiformis f. sp. Tritici Mol Plant Pathol 18:649–661. https://doi.org/10.1111/mpp.12425
Wang C, He X, Li Y, Wang L, Guo X, Guo X (2018a) The cotton MAPK kinase GhMPK20 negatively regulates resistance to fusarium oxysporum by mediating the MKK4-MPK20-WRKY40 cascade. Mol Plant Pathol 19:1624–1638. https://doi.org/10.1111/mpp.12635
Wang K, He J, Zhao Y et al (2018b) EAR1 negatively regulates ABA signaling by enhancing 2C protein phosphatase activity. Plant Cell 30:815–834. https://doi.org/10.1105/tpc.17.00875
Wang P, Zhao Y, Li Z et al (2018c) Reciprocal regulation of the TOR kinase and ABA receptor balances plant growth and stress response. Mol Cell 69(100–112):e106. https://doi.org/10.1016/j.molcel.2017.12.002
Yamasaki K, Kigawa T, Inoue M et al (2005) Solution structure of an Arabidopsis WRKY DNA binding domain. Plant Cell 17:944–956. https://doi.org/10.1105/tpc.104.026435
Yan H, Jia H, Chen X, Hao L, An H, Guo X (2014) The cotton WRKY transcription factor GhWRKY17 functions in drought and salt stress in transgenic Nicotiana benthamiana through ABA signaling and the modulation of reactive oxygen species production. Plant Cell Physiol 55:2060–2076. https://doi.org/10.1093/pcp/pcu133
Yang T, Yao S, Hao L, Zhao Y, Lu W, Xiao K (2016) Wheat bHLH-type transcription factor gene TabHLH1 is crucial in mediating osmotic stresses tolerance through modulating largely the ABA-associated pathway. Plant Cell Rep 35:2309–2323. https://doi.org/10.1007/s00299-016-2036-5
Yang W, Zhang W, Wang X (2017) Post-translational control of ABA signalling: the roles of protein phosphorylation and ubiquitination. Plant Biotechnol J 15:4–14. https://doi.org/10.1111/pbi.12652
Yu F, Huaxia Y, Lu W, Wu C, Cao X, Guo X (2012) GhWRKY15, a member of the WRKY transcription factor family identified from cotton (Gossypium hirsutum L.), is involved in disease resistance and plant development. BMC Plant Biol 12:144. https://doi.org/10.1186/1471-2229-12-144
Zhang L, Xi D, Luo L et al (2011) Cotton GhMPK2 is involved in multiple signaling pathways and mediates defense responses to pathogen infection and oxidative stress. FEBS J 278(8):1367–1378. https://doi.org/10.1111/j.1742-4658.2011.08056.x
Zhang H, Li Y, Zhu J-K (2018) Developing naturally stress-resistant crops for a sustainable agriculture. Nat Plants 4(12):989–996. https://doi.org/10.1038/s41477-018-0309-4
Zhao Y, Xing L, Wang X et al (2014) The ABA receptor PYL8 promotes lateral root growth by enhancing MYB77-dependent transcription of auxin-responsive genes. Sci Signal 7:53. https://doi.org/10.1126/scisignal.2005051
Zhu JK (2016) Abiotic stress signaling and responses in plants. Cell 167:313–324. https://doi.org/10.1016/j.cell.2016.08.029
Acknowledgements
This work was financially supported by grants from the National Natural Science Foundation of China (Grant Nos. 31901431 and 31971823) and the Natural Science Foundation of Shandong Province (Grant No. ZR2019BC015).
Funding
This work was financially supported by grants from the National Natural Science Foundation of China (31901431 to C.W. and 31971823 to X.G.) and the Natural Science Foundation of Shandong Province (ZR2019BC015 to C.W.).
Author information
Authors and Affiliations
Contributions
C.W. and X.G. conceived the project and designed the experiments. X.G. was responsible for the material distribution. J.W., L.W. and Y.Y. performed most of the experiments with the assistance of H.L., Z.G. and S.Z.. J.W. and C.W. wrote the article with the assistance of S.Z.
Corresponding authors
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethics approval
This article does not contain any studies with human participants or animals.
Additional information
Communicated by Neal Stewart.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2020_2590_MOESM2_ESM.pdf
Additional file2 (PDF 1322 kb) Fig. S1 Expression of the GhWRKY21-His protein analysed by SDS-PAGE. lane M: protein marker; lane U: recombinant GhWRKY21-His protein without induction; lanes I: recombinant GhWRKY21-His protein with IPTG-induction. Fig. S2 Subcellular localization of GhWRKY21. a Schematic diagram of the control 35S::GFP construct and the 35S::GhWRKY21-GFP fusion protein construct. b Fluorescent signals of the 35S::GhWRKY21-GFP fusion construct and the 35S::GFP construct in N. benthamiana leaves at 2 days after transformation. Green fluorescence was observed with an LSM 880 META confocal microscope (Carl Zeiss). Fig. S3 Expression of GhWRKY21 in T2 Vec and OE plants as measured by qRT-PCR-based analysis. The different letters indicate significant differences (P<0.05) according to Tukey’s honestly significant difference (HSD) test. Fig. S4 Seed germination rates and phenotypes of Vec and OE plants on MS media consisting of different concentrations of mannitol. Fig. S5 Stomatal aperture of Vec and OE plants under drought stress. The data represent the means ± SEs of 40 stomata (length/width) from three independent experiments. The different letters indicate significant differences (P<0.05) according to Tukey’s HSD test. Fig. S6 Expression pattern of GhHAB under ABA treatment. Different letters indicate significant differences (P<0.05) based on Tukey’s HSD test.
Rights and permissions
About this article
Cite this article
Wang, J., Wang, L., Yan, Y. et al. GhWRKY21 regulates ABA-mediated drought tolerance by fine-tuning the expression of GhHAB in cotton. Plant Cell Rep 40, 2135–2150 (2021). https://doi.org/10.1007/s00299-020-02590-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-020-02590-4