Abstract
Key message
VaPAT1 functions as a stress-inducible GRAS gene and enhanced cold, drought and salt tolerance in transgenic Arabidopsis via modulation of the expression of a series of stress-related genes.
Abstract
The plant-specific GRAS transcription factor family regulates diverse processes involved in plant growth, development and stress responses. In this study, VaPAT1, a GRAS gene from Vitis amurensis was isolated and functionally characterized. Sequence alignment and phylogenetic analysis showed that VaPAT1 has a high sequence identity to CmsGRAS and OsCIGR1, which belong to PAT1 branch of GRAS family and function in stress resistance. The transcription of VaPAT1 was markedly induced by stress-related phytohormone abscisic acid (ABA) and various abiotic stress treatments such as cold, drought and high salinity, however, it was repressed by exogenous gibberellic acid (GA) application. Overexpression of VaPAT1 increased the cold, drought and high salinity tolerance in transgenic Arabidopsis. When compared with wild type (WT) seedlings, the VaPAT1-overexpression lines accumulated higher levels of proline and soluble sugar under these stress treatments. Moreover, stress-related genes such as AtSIZ1, AtCBF1, AtATR1/MYB34, AtMYC2, AtCOR15A, AtRD29A and AtRD29B showed higher expression levels in VaPAT1 transgenic lines than in WT Arabidopsis under normal growth conditions. Together, our results indicated that VaPAT1 functions as a positive transcriptional regulator involved in grapevine abiotic stress responses.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Abiotic stresses, such as cold, drought and high salinity adversely affect plant growth and crop production (Shinozaki and Yamaguchi-Shinozaki 2000; Qiu and Yu 2009). In response to these environmental adversities, plants have developed a number of strategies to get better tolerance or resistance. One primary model of these strategies is to modulate the expression of genes involved in stress response pathways (Yang et al. 2015). Many transcription factors, such as DREB/CBF (dehydration responsive element/C-repeat binding factor), ERF (ethylene responsive factor), WRKY, bHLH (basic helix-loop-helix), MYB (myeloblastosis), MYC (myelocytomatosis) and NAC (NAM, ATAF1/2 and CUC2) families have been found to play critical roles in controlling the transcription of stress-responsive genes and therefore regulating plants adaptation to stresses separately or cooperatively (Singh et al. 2002). Overexpression of these transcription factors mentioned above could increase stress tolerance in corresponding plants (Yamaguchi-Shinozaki and Shinozaki 2006; Lindemose et al. 2013; Li et al. 2013).
GRAS transcription factor family comprises plant-specific proteins (Benfey et al. 1993). The acronym GRAS was named after the first three identified family members: GIBBERELLIN ACID INSENSITIVE (GAI), REPRESSOR of GA1 (RGA), and SCARECROW (SCR) genes (Pysh et al. 1999). Based on the 33 and 55 GRAS members from Arabidopsis and rice, respectively, the GRAS protein family can be divided into eight branches which include SCL9, SCR, DELLA, SHR, Ls, SCL4/7, PAT1 and HAM (Bolle 2004; Lee et al. 2008). GRAS proteins have been shown to be involved in various biological processes in plants such as root development (Benfey et al. 1993), axillary meristem initiation (Schumacher et al. 1999), shoot meristem maintenance (Helariutta et al. 2000), GA and phytochrome A signal transduction (Bolle et al. 2000; Boss and Thomas 2002; Fu et al. 2001; Heo et al. 2011). Moreover, recent studies have reported the participation of GRAS proteins in stress response in several species. For instance, NtGRAS1 from tobacco was strongly induced by various stimulants that raise the intracellular reactive oxygen (ROS) levels (Czikkel and Maxwell 2007). AtSCL14 is also involved in the activation of stress-inducible promoters and could increase plants tolerance to noxious chemicals (Fode et al. 2008). In addition, overexpressing of PeSCL7 confers improved drought and salt tolerance in transgenic Arabidopsis plants (Ma et al. 2010), while OsGRAS23 could positively modulate drought tolerance in rice (Xu et al. 2015). However, the underlying mechanisms on how GRAS proteins are involved in stress response in plants are still largely unknown.
Vitis amurensis is a wild grapevine species with remarkable cold and drought tolerance and therefore widely used during cross breeding to increase the cold and drought tolerance for grapevine (Su et al. 2015; Li 2015). Previous study on whole genome transcriptome analysis of V. amurensis found that the expression of two GRAS genes was significantly increased under cold stress (Xin et al. 2013). The coding region of one of the GRAS genes, GSVIVT01014570001, was cloned from V. amurensis. Sequence alignment and phylogenetic analysis showed that the gene belongs to PAT1 branch of GRAS protein family, and thus we named it as VaPAT1.
In the present study, the expression pattern of VaPAT1 induced by phytohormones ABA and GA, and subjected to various abiotic stresses was analyzed. Moreover, the VaPAT1 overexpressed lines were constructed in Arabidopsis by the floral dip method to examine the effects of VaPAT1 gene in response to cold, drought and high salinity stresses. To further understand the molecular mechanisms of enhanced stresses tolerance by VaPAT1, the expression levels of stress-related genes in VaPAT1 transgenic Arabidopsis were tested. Together, the investigation increased the comprehensive understanding toward the function of VaPAT1 in stress responses. These results suggest that VaPAT1 play a positive role in plant stress responses and thus could be a promising candidate for commercial crop species stresses-tolerant trait improvement.
Materials and Methods
Plant materials and growth conditions
Tissue culture plantlets of V. amurensis (collected from Changbai Mountain in Jilin province in China) were grown on half-strength (1/2) Murashige and Skoog (MS) medium containing 1 % sucrose and 0.7 % agar in a growth chamber at constant 26 °C under a 16-h light/8-h dark photoperiod and 100 μmol m−2 s−1 light intensity.
Arabidopsis thaliana (ecotype Columbia Col-0) was used for gene transformation in this study. Seeds of Arabidopsis were vernalized at 4 °C in the dark for two days and then sown in same pots with equiponderate soil in a growth chamber at 22 °C under a 16-h light/8-h dark cycle and 60 % relative humidity.
Exogenous phytohormone and stress treatments
Forty-day-old grapevine plantlets with 5–6 totally expanded leaves were used for the expression analysis under exogenous phytohormones and stress treatments. Phytohormones ABA and GA treatments were induced by carefully transferring plantlets to 1/2 MS liquid medium (without 0.7 % agar) containing 100 μM ABA or 200 μM GA. Moreover, leaves of the plantlets were separately sprayed with 100 μM ABA or 200 μM GA solutions. The control (no stress treatment) were transferred to liquid 1/2 MS medium with leaves sprayed with same amount of deionized water. For cold treatment, plantlets were transferred to a low-temperature chamber at 4 °C with a 16-h light/8-h darkness cycle. For drought and salt stress, roots of plantlets were immersed into liquid 1/2 MS medium containing 2.5 % PEG (polyethylene glycol) 6000 or 200 mM NaCl, respectively.
The shoot apex with the first fully expanded leaf was harvested at specific time after initiating the treatments. The collected samples at each time point has three independent replicates. Harvested samples were immediately frozen in liquid nitrogen and stored at -80 °C for later RNA extraction.
RNA extraction and quantitative real-time PCR (qRT-PCR) analysis
Total RNA was extracted from collected samples using TIANDZ Column Plant RNAout 2.0 Kit (Tiandz, Beijing, China) following the manufacturer’s procedure. A maximum of 1 μg total RNA was used for synthesizing cDNA by TransScript One-Step gDNA Removal and cDNA Synthesis SuperMix Kit (TransGen, Beijing, China). The qRT-PCR experiment was performed on a StepOnePlus™ realtime PCR Instrument (Applied Biosystems) using FastStart Universal SYBR Green Master (Roche Diagnostics, Mannheim, Germany). The β-actin gene (VvACT, GenBank accession: EC969944) and malate dehydrogenase gene (VvMDH, GenBank accession: EC921711) were used as reference genes to normalize the expression of VaPAT1 in V. amurensis, While AtACT2 (GenBank accession: BAH20120.1) was used to normalize the expression level of genes in Arabidopsis. Gene specific primer pairs for qRT-PCR (listed in Table 1) were designed by Primer 5.0 and their specificities were tested by NCBI Primer BlAST. Three biological replicates were performed to ensure the accuracy of results. The relative expression was estimated via the relative quantization method (2−ΔΔCt).
The cDNA cloning and sequence analysis
Full coding sequence of VaPAT1 was obtained by homologous cloning using a primer pair (VaPAT1CDS-F/R, Table 1) designed according to the putative cDNA sequence of GSVIVT01014570001 from the published 12 × Vitis vinifera cv. Pinot Noir genome sequences (http://www.genoscope.cns.fr/externe/GenomeBrowser/Vitis/). The fragment was cloned into pGEMT-easy vector (Promega, WI, USA) and sequenced using ABI3730.
ExPASy server (http://www.expasy.org/) was applied to translate the confirmed CDS sequence of VaPAT1 into amino acids and calculate the theoretical isoelectric point (pI) and molecular weight of VaPAT1 protein. Alignment of amino acid sequences was performed with the software DNAMAN 6.0. The phylogenetic tree was generated by the MEGA program (v.5.0) using the neighbor-joining (NJ) method (Hall 2013).
Vector construction and Arabidopsis transformation
The full coding sequence of VaPAT1 was amplified with primers containing restriction sites BamHI and EcoRI (VaPAT1-BamH and VaPAT1-EcoR, Table 1). The amplified fragment was digested with BamHI and EcoRI and inserted into the binary vector pCAMBIA 1301, which contains 35S CaMV (cauliflower mosaic virus) promoter. The expression vectors of VaPAT1 were introduced into Agrobacterium tumefaciens GV3101, and then transformed into Arabidopsis by the floral dip method (Clough and Bent 1998). Seeds of WT and transgenic Arabidopsis were harvested, surface-sterilized with 2 % sodium hypochlorite for 5 min and then washed with sterile water for 3 times. Transgenic seedlings were selected on 1/2 MS medium containing hygromycin (50 mg/L). Homozygous T3 transgenic lines were further tested by hygromycin resistance screening and used for subsequent stress tolerance assay experiments.
Abiotic stress tolerance assays of transgenic Arabidopsis
WT and T3 generation transgenic Arabidopsis seedlings at 2-week-old stage were employed for detection of the cold, drought and salt stress tolerance. Cold stress was induced by transferring Arabidopsis seedlings inside a cold treatment incubator at −1 °C for 8 h to initiate freezing, followed by the temperature decreased at a rate of 2 °C/h from −1 to −9 °C, each temperature point (−3, −5, −7, −9 °C) was kept for 3 h, respectively. After that, seedlings thawed overnight at 4 °C and then incubated at 22 °C for recovery. Drought stress was imposed to previously adequately watered plants by withholding water for 12 days. Finally, seedlings were rewatered to analyze the recovery from drought stress. For salt treatment, 2-week-old seedlings were watered with 200 mM NaCl twice at 5-days intervals.
About 20 plants per line (WT, 35S::VaPAT1-1, -2 or -4) were used in each stress treatment experiment. The seedlings exhibiting >50 % green on their above-ground tissues were judged as surviving plants. Survival rates of WT and transgenic Arabidopsis were calculated based on three independent repeats for each stress test (about 60 plants for per line).
Measurement of proline and soluble sugar contents
Leaves sampled from normal and stresses-treated WT and transgenic Arabidopsis were used for determination of proline and soluble sugar contents. Proline was assayed as described by Bates et al. (1973) with some modifications. Briefly, fresh leaf material (200 mg) of Arabidopsis was extracted with 3 ml of 3 % sulfosalicylic acid at 100 °C for 10 min. Thereafter, 2 ml of the extract was heated at 100 °C for 30 min mixing with same volume of glacial acetic acid and acid ninhydrin reagent. After cooling, 3 ml toluene was added to partition the mix, and the absorbance of the supernatant extract containing proline was measured at 520 nm. The soluble sugar content was determined by an anthrone-sulfuric acid method (Dubois et al. 1956). Absorbance of proline and soluble sugar hereby were all measured using a Tecan Infinite M1000 PRO Multiskan Spectrum Microplate Spectrophotometer (TECAN, Austria).
Results
VaPAT1 encodes a GRAS protein that belongs to the PAT1 branch
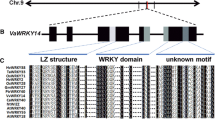
Full coding sequence (CDS) of the VaPAT1 was cloned from V. amurensis leaf cDNA using gene-specific primers VaPAT1CDS-F and VaPAT1CDS-R (Table 1). The length of the nucleotide sequence of VaPAT1 was 1752 bp, encoding a protein of 583 amino acid residues with a calculated molecular mass of 65.0 kDa and a pI (predicted theoretical isoelectric point) of 5.75. Alignment of VaPAT1 with their homologues derived from other plant species showed that it contained a variable N-terminal and a conserved C-terminal domain similar to other members of the GRAS family. The conserved C-terminal part contains the typical motifs defined for GRAS proteins, namely two leucine-rich (LR) domains flanking a conserved V/I HIID domain (Fig. 1a). Furthermore, the motifs LXXLL, PFYRE, RVER and SAW were highly conserved. A phylogenetic tree based on the conserved domains was constructed. The result confirmed that VaPAT1 protein is a member of the PAT1-branch of the GRAS protein family (Fig. 1b). In the PAT1-branch, VaPAT1 presented close evolutionally relationship with CmsGRAS from Citrus medica and OsCIGR1 from Oryza sativa (74 and 52 % identities, respectively) (Shi et al. 2011; Day et al. 2003). Both CmsGRAS and OsCIGR1 were previously identified as positive components that functioned in stress resistance, which implies that VaPAT1 may also have a similar role in stress-responsive processes.
Alignment and phylogenetic analysis of VaPAT1 with other GRAS proteins. a Alignment of amino acid sequences of VaPAT1 with other GRAS proteins. Sequences were from Vitis vinifera (VvPAT1, XP_002282942.1), Arabidopsis thaliana (AtPAT1, At5g48150), Oryza sativa (OsCIGR1, AAL61820.1 and OsCIGR2, Q8GVE1.1), Citrus medica var. sarcodactylis (CmsGRAS, JF440647.1), Nicotiana tabacum (NtGRAS1, ABE02823.1) and Populus euphratica (PeSCL7, AHZ13509.1). Conserved sequences are coloured. Gaps introduced to facilitate alignment are indicated as dashes. The five main conserved domains in the C-terminal are indicated with uppercase letters above the alignment. Asterisks mark the conserved Leucine residues in the leucine-rich domains flanking the V/I HIID motif. W(X)7G, W(X)10W, the two pairs of conserved residues in SAW domain is double-underlined. b Phylogenetic tree of the GRAS protein family. The red solid diamond represents VaPAT1. All Arabidopsis PAT1 branch proteins (AtPAT1; AtSCL5, At1g50600; AtSCL13, At4g17230; AtSCL21, At2g04890) as well as several stress-related proteins belong to PAT1-branch from other specises (CmsGRAS, OsCIGR1, OsCIGR2) are presented. Representatives of other seven branches of the GRAS family are shown (AtSCL9, At2g37650; AtSHR, At4g37650; AtRGA, At2g01570; AtGAI, At1g14920; AtSCR, At3g54220; SlLs, AAD05242.1; AtSCL4, At5g66770; AtSCL7, At3g50650; and PhHAM, AAM90848.1). Sl Solanum lycopersicum; Ph Petunia × hybrid
VaPAT1 responded to exogenous ABA and GA applications and could be induced by cold, drought and high salinity treatments
ABA is a representative stress-related phytohormone, with a role in the regulation of stomatal aperture and genes expression when plants were subjected to environmental stress factors (Hubbard et al. 2010; Lindemose et al. 2013). GRASs were previously reported to be involved in GA signal transduction pathway. Upon GA application, DELLA-branch GRAS proteins bind to GID1 (GA receptor) to form a GA-GID1-DELLA complex, thus induces the degradation of DELLAs via the ubiquitin proteasome pathway (Silverstone et al. 1998, 2001; Fu et al. 2001; Heo et al. 2011). The inducible expression profiles of VaPAT1 in V. amurensis by exogenous ABA and GA applications were hereby investigated. The expression of VaPAT1 was quickly induced by ABA application with an early peak at 2 h and the second peak at 24 h after the plantlets were subjected to ABA application, which suggests that VaPAT1 might function via an ABA-dependent pathway (Fig. 2). On the contrary, the expression of VaPAT1 was reduced gradually within the first 3 h (with a minimum level at 3 h, about one-third compared to the control), and then raised back to a relatively higher level (about two-thirds compared to the control) in the following time course after GA application (Fig. 2).
Expression patterns of VaPAT1 in response to cold (4 °C), drought (2.5 % PEG6000), salt (200 mM NaCl), 100 μM ABA and 200 μM GA treatments. Error bars refer to three biological replicates. The expression level of VaPAT1 in the controls was set at 1.0. The relative expression level of VaPAT1 in the control plant without treatment (0 h) was defined as 1 to evaluate the dynamic changes
To confirm whether VaPAT1 was involved in abiotic stress response in V. amurensis, its expression patterns under various stresses were investigated by real-time PCR. As shown in Fig. 2, the expression levels of VaPAT1 were increased in all the stress conditions used. Under cold stress at 4 °C and drought stress simulated by PEG, VaPAT1 expression was induced quickly and the level of the products peaked at 2 h or 4 h (about 5.5-fold or 3.4-fold that of the untreated control plants), respectively. As regard to high salinity treatment, the VaPAT1 transcripts raised constantly and reached to a maximum at 48 h (up to 22-fold higher than that before salt treatment).
Overexpression of VaPAT1 in Arabidopsis enhances cold, drought and salt tolerance
To examine the biological function of VaPAT1, transgenic Arabidopsis plants that overexpresses VaPAT1 were constructed. Three independent homozygous transgenic lines of T3 transgenic plants (35S::VaPAT1-1, -2, and -4) with higher VaPAT1 expression levels were selected to assess the cold, drought, or salt tolerance (Fig. S1). Moreover, there were no significant differences in morphological traits between the WT and transgenic plants during their life cycles.
After the cold treatment, all the leaves of WT and transgenic plants were flooded. Upon returning them back to normal growth conditions for five days, the leaves of most WT plants became dry and withered. However, most transgenic plants recovered and had green leaves (Fig. 3a1). The survival rates of transgenic lines were 78 % compared with 23 % of WT plants under cold stress. To determine their drought resistance, 2 weeks old WT and transgenic Arabidopsis plants were watered once deeply and then subjected to water withholding for an additional 12 days. The WT plants wilted more severely and only about 13 % plants could survive after rewatering. As regards VaPAT1-overexpressing lines, 72 % of plants recovered after rewatering (Fig. 3b). Moreover, the development of surviving WT plants was retarded by drought stress compared with VaPAT1-overexpressing lines (Fig. 3a2). VaPAT1-overexpressing lines bolted even during the drought stress treatment (Fig. 3a2. Similarly, after 200 μM NaCl treatment for 10 days, the WT plants showed a much more severe etiolating phenotype than transgenic seedlings (Fig. 3a3). On average, 22 % of total WT plants were able to survive, whereas 71 % of transgenic lines continued growing and exhibited much bigger leaves than in normal growth conditions (Fig. 3b). These results indicated that the overexpression of VaPAT1 could improve the tolerance to cold, drought and high salinity in Arabidopsis.
Stress tolerance assays of WT and VaPAT1-overexpressing Arabidopsis plants: a Phenotypic differences between WT and transgenic Arabidopsis seedlings under cold, drought and salt stresses. 1 Cold treatment. Two weeks old seedlings grown at 22 °C under long days (16-h light/8-h dark) were incubated at −1 °C for 8 h, then followed by a gradient freezing process decreasing at a rate of −2 °C/h to −9 °C, each temperature point kept for 3 h. Seedlings were photographed after 5 days recovery at normal growth conditions (22 °C). 2 Drought treatment. Two weeks old WT and transgenic Arabidopsis were subjected to water withholding for 12 days then seedlings were rewatered for recovery. Photographs were taken 5 days after rewatering. 3 Salt treatment. Two weeks old seedlings were watered with 200 mM NaCl twice at an interval of 5 days. Photographs were taken 10 days after salt stress treatment. b Survival rate of WT and VaPAT1-overexpressing lines after stresses treatments. Control plants without stresses have a survival rate of nearly 100 %. Error bars represent standard deviation of three biological replicates, 20 plants per line (WT, 35S::VaPAT1-1, -2 or -4) in each experiment. ** and *** indicate significant differences between WT and VaPAT1-overexpressing lines at P < 0.01 and P < 0.001 level (Student’s t test), respectively
Overexpression of VaPAT1 led to significantly higher proline and soluble sugar contents in transgenic lines
To deeply uncover the physiological changes in transgenic Arabidopsis, several physiological index related to abiotic stress were measured in WT and transgenic plants. A total of six indexes including malondialdehyde (MDA), superoxide dismutase (SOD), peroxidase (POD), catalase (CAT), proline and soluble sugar contents were measured. The content of MDA, activity of SOD, POD and CAT did not show significant differences between WT and VaPAT1 transgenic plants either in normal growth conditions or exposed to stresses (Fig. S2). The proline concentration in overexpression lines was higher than that in WT plants under normal growth conditions (Fig. 4a), whereas soluble sugar contents in transgenic lines were similar to that in WT plants (Fig. 4b). When the seedlings were subjected to stresses including cold, drought and salt, the proline and soluble sugar contents increased distinctly in both VaPAT1-overexpression and WT plants. Meanwhile, VaPAT1-overexpression lines accumulated more proline and soluble sugar than in WT plants. Overall, these findings indicated that overexpression of VaPAT1 led to the increase of the proline and soluble sugar contents, which were two important factors for the enhanced cold, drought and salt stress tolerance in transgenic Arabidopsis.
Proline and soluble sugar contents of WT and VaPAT1-overexpression Arabidopsis with or without stress treatments. a Proline content. b Soluble sugar content. Normal: three-week-old plants grown under normal conditions; Cold treatment: incubating seedlings at 4 °C for 3 days; Drought treatment: withholding water for 8 days; Salt treatment: irrigated with 200 mM NaCl for a week. The bars represent standard deviation of three biological replicates. *, ** and *** indicate significant differences between WT and VaPAT1-overexpressing lines at P < 0.05, P < 0.01 and P < 0.001 level based on Student’s t test, respectively
Altered expression of stress-related genes in VaPAT1-overexpression lines
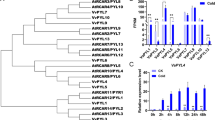
The expression profiles of a set of stress-related genes (Table S1) were analyzed to further elucidate the possible molecular mechanisms underlying enhanced stress tolerance by VaPAT1. Under normal conditions, the test genes including AtSIZ1, AtCBF1, AtATR1/MYB34, AtMYC2, AtCOR15A, AtRD29A and AtRD29B showed significantly higher expression levels in VaPAT1 transgenic lines compared to that of WT plants as shown in Fig. 5. These results suggest that VaPAT1 could affect the expression of those genes. Moreover, AtProDH, a gene encoding a proline dehydrogenase was significantly down-regulated in VaPAT1-overexpression lines, which may partly confer to the higher proline contents in VaPAT1-overexpression lines even under normal growth conditions (Fig. 4a).
Expression of stress-related genes in WT and VaPAT1-overexpression Arabidopsis plants under normal growth conditions. Total RNA was obtained from two-week-old WT and T3 generation VaPAT1 transgenic seedlings (35S::VaPAT1-1, -2 and -4, which were also marked as OE1, OE2 and OE4 in the figure) without stress treatment. Expression values were calculated by real-time qRT-PCR using the gene-specific primers listed in Table 1. The putative downstream stress-related genes are AtSIZ1 (AT5G60410), AtCBF1 (AT4G25490), AtATR1/MYB34 (AT5G60890), AtMYC2 (AT1G32640), AtCOR15A (AT2G42540), AtRD29A (AT5G52310), AtRD29B (AT5G52300) and AtProDH1(AT3G30775). Real-time RT-PCR quantifications were normalized to the expression of AtACTIN2 (AT3G18780). Error bars indicate SD from three independent experiments. The asterisk *, ** and *** indicate significant differences between WT and VaPAT1-overexpressing lines at P < 0.05, P < 0.01 and P < 0.001 level based on Student’s t test, respectively
Discussion
In the present study, VaPAT1, a GRAS gene that participates in the cold, drought and high salinity stresses responses in V. amurensis, has been characterized. Further phylogenetic analysis demonstrated that VaPAT1 has high sequences similarity with CmsGRAS and OsCIGR1 and belongs to PAT1 branch of GRAS proteins. CmsGRAS from Citrus medica was significantly up-regulated under low-temperature stress (Shi et al. 2011). OsCIGR1 was rapidly induced upon co-cultivation with rice blast fungus and N-acetylchitooligosaccharide perception, functioning in the early defense response in rice (Day et al. 2003). Other members such as OsCIGR2 and AtSCL13, which also belongs to the PAT1 branch, were reported to have a role in stress responses (Day et al. 2003; Torres-Galea et al. 2006). In addition to PAT1 branch, genes from other branches of the GRAS family such as PeSCL7 (SCL4/7-branch) in poplar (Populus euphratica Oliv.) and OsGRAS23 (Ls-branch) in rice, were also shown to improve stress tolerance in corresponding overexpressed lines (Ma et al. 2010; Xu et al. 2015). Moreover, AtSCL14, a gene in SCL9 branch, were reported to interact with Class II TGA (TGACG MOTIF-BINDING FACTOR) transcription factors to activate the expression of genes involved in the detoxification of harmful chemicals (Fode et al. 2008). These results therefore indicate that GRAS gene family members widely participate in abiotic stress related signal transduction pathways in plants.
A promoter prediction of VaPAT1 using PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/) showed it contains many stress and phytohormone responsive cis-acting regulatory elements, including the defense and stress responsiveness element, heat stress responsive element, anaerobic inducible element, GA and SA (salicylic acid) responsive element (data not shown). These motifs may be responsible for the dynamic transcription of VaPAT1 during stress conditions and exogenous hormones applications. The expression of VaPAT1 decreased under GA application, similar to a typical feature of DELLA-branch GRAS proteins (Silverstone et al. 1998; Wen and Chang 2002). Previous studies demonstrated that high level expression of DELLAs confer reduced GA responses associated with dwarfism phenotypes (Fu et al. 2001; Boss and Thomas 2002; Achard et al. 2008). In this study however, VaPAT1-overexpression Arabidopsis exhibited no phenotype differences compared with WT under normal growth conditions. In accordance with our findings is that, the expression of PeSCL7 (a member of the SCL4/7-branch GRASs) decreased under GA application, but PeSCL7-overexpression Arabidopsis exhibited normal phenotypes (Ma et al. 2010). These results implied that VaPAT1 and PeSCL7 may be involved in GA response pathway, but in a way different from that of DELLAs.
Proline and soluble sugar are common compatible osmolytes to protect cell membrane system from detrimental effects of abiotic stresses, which are widely used as physiological indicators estimating cellular damage upon various stresses (Ashraf and Foolad 2007; Karan and Subudhi 2012; Yu et al. 2013). The contents of proline and soluble sugar increased in both VaPAT1-overexpression and WT Arabidopsis plants under various stresses, but accumulated significantly higher in VaPAT1-overexpression lines than in WT. Moreover, it was found that the effect of VaPAT1 overexpression on the accumulation of proline was different from that on soluble sugar. Prior to subjection to stresses, transgenic Arabidopsis contained higher proline than WT plants, which suggests that overexpression of VaPAT1 could increase the proline content. With regard to soluble sugar, the increased content in overexpression lines was only found under stresses. This phenomenon indicates that soluble sugar maybe regulated by VaPAT1 with the help of other factors under stresses. Furthermore, AtProDH was found down-regulated in VaPAT1 transgenic Arabidopsis. This gene encodes proline dehydrogenase 1, an enzyme functioning in proline metabolism process (Cecchini et al. 2011). Down-regulation of AtProDH may partly contribute to higher proline content in VaPAT1-overexpression lines in normal growth conditions, which, in turn, resulted in increased stress tolerance in transgenic Arabidopsis.
In order to further study the molecular mechanisms underlying enhanced abiotic stress tolerance in VaPAT1 transgenic Arabidopsis, the expression levels of many stress-related genes were examined in WT and VaPAT1 transgenic Arabidopsis under normal growth conditions. From the real time RT-PCR analysis, among the examined genes, transcripts of seven genes, including AtSIZ1, AtCBF1, AtATR1/MYB34, AtMYC2, AtCOR15A, AtRD29A and AtRD29B were significantly higher in VaPAT1-overexpression lines than in WT (Fig. 5). AtSIZ1 and AtCBF1 are upstream transcriptional regulation factors in the ICE-CBF-COR pathway under cold stress (Yamaguchi-Shinozaki and Shinozaki 2006; Novillo et al. 2007; Miura and Nozawa 2014). AtSIZ1 encodes a small ubiquitin-like modifier (SUMO) E3 ligase, which mediates sumoylation of ICE1, thereby enhancing the stability of ICE1, leading to increased induction of CBFs and its target COR genes (COR15A and RD29A) (Miura et al. 2007). In addition, a study carried out by Catala et al. (2007) showed that AtSIZ1 plays a role in Arabidopsis drought tolerance through the regulation of a set of drought-responsive genes, including AtATR1/MYB34, AtMYC2, AtCOR15A and AtRD29B. These genes were probably up-regulated by AtSIZ1 in VaPAT1-overexpression Arabidopsis plants in our investigation. AtATR1/MYB34 and AtMYC2 also function as transcription factors in stress response processes (Abe et al. 1997). AtATR1/MYB34 is a R2R3 MYB gene which can respond to various hormone applications and salt stress treatment (Chen et al. 2006). AtMYC2, which encodes a bHLH transcription factor, is involved in activating ABA-inducible gene expression under drought stress (Abe et al. 2003; Nakata et al. 2013). It has been confirmed that AtCOR15A, AtRD29A and AtRD29B are downstream genes implicated in multiple stress responses (Thalhammer et al. 2014; Miyazaki et al. 2015; Kim et al. 2014) and regulated by various upstream transcription factors (Yamaguchi-Shinozaki and Shinozaki 2006). Taken together, VaPAT1 triggered the constitutive expression of many stress-related genes and enhanced abiotic stresses in transgenic plants. These genes could be considered as direct or indirect downstream genes of VaPAT1 and further works still need to uncover the VaPAT1 related signal transduction during abiotic stress.
In summary, we obtained a GRAS transcription factor VaPAT1 from V. amurensis. VaPAT1 responds to exogenous ABA and GA, as well as cold, drought, and salt stress treatments. Ectopic expression of VaPAT1 in Arabidopsis modulated the expressions of a series of stress-related genes, increased proline and soluble sugar contents and enhanced cold, drought and high salinity tolerance in VaPAT1-overexpression lines. Hence, it is concluded that VaPAT1 is a novel stress-responsive GRAS gene playing positive roles in plants stress responses. This new gene identified can be used as a candidate gene in molecular breeding of commercial crop species to get better stress tolerance.
Author contribution statement
YY and LF designed and conducted experiments. YY wrote the manuscript. LZ and YG contributed by helping some experiments presented in the manuscript. SKK helped to edit the manuscript. SL and HX supervised the studies and revised the manuscript. All authors read and approved the manuscript.
Abbreviations
- GRAS:
-
GAI (gibberellin acid insensitive), RGA (repressor of GAI), and SCR (scarecrow)
- PAT:
-
Phytochrome A signal transduction
- ABA:
-
Abscisic acid
- GA:
-
Gibberellic acid
- WT:
-
Wild type
- WRKY:
-
DELLA name derived from the most prominent amino acid residues in the protein
- SCL:
-
Scarecrow-like
- SHR:
-
Short-root
- Ls:
-
Lateral suppressor
- HAM:
-
Hairy meristem
- ROS:
-
Reactive oxygen
- CDS:
-
Coding sequence
- CIGR:
-
Chitin-inducible gibberellin-responsive
- ICE:
-
Inducer of CBF expression
References
Abe H, YamaguchiShinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868. doi:10.1105/tpc.9.10.1859
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15:63–78. doi:10.1105/tpc.006130
Achard P, Renou JP, Berthome R, Harberd NP, Genschik P (2008) Plant DELLAs restrain growth and promote survival of adversity by reducing the levels of reactive oxygen species. Curr Biol 18:656–660. doi:10.1016/j.cub.2008.04.034
Ashraf M, Foolad MR (2007) Roles of glycine betaine and proline in improving plant abiotic stress resistance. Environ Exp Bot 59:206–216. doi:10.1016/j.envexpbot.2005.12.006
Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207. doi:10.1007/bf00018060
Benfey PN, Linstead PJ, Roberts K, Schiefelbein JW, Hauser MT, Aeschbacher RA (1993) Root development in Arabidopsis-four mutants with dramatically altered root morphogenesis. Development 119:57–70
Bolle C (2004) The role of GRAS proteins in plant signal transduction and development. Planta 218:683–692. doi:10.1007/s00425-004-1203-z
Bolle C, Koncz C, Chua NH (2000) PAT1, a new member of the GRAS family, is involved in phytochrome A signal transduction. Gene Dev 14:1269–1278. doi:10.1101/gad.14.10.1269Genes&Dev.2000
Boss PK, Thomas MR (2002) Association of dwarfism and floral induction with a grape ‘green revolution’ mutation. Nature 416:847–850. doi:10.1038/416847a
Catala R, Ouyang J, Abreu IA, Hu Y, Seo H, Zhang X, Chua NH (2007) The Arabidopsis E3 SUMO ligase SIZ1 regulates plant growth and drought responses. Plant Cell 19:2952–2966. doi:10.1105/tpc.106.049981
Cecchini NM, Monteoliva MI, Alvarez ME (2011) Proline dehydrogenase contributes to pathogen defense in Arabidopsis. Plant Physiol 155:1947–1959. doi:10.1104/pp.110.167163
Chen YH, Yang XY, He K, Liu MH, Li JG, Gao ZF, Lin ZQ, Zhang YF, Wang XX, Qiu XM, Shen YP, Zhang L, Deng XH, Luo JC, Deng XW, Chen ZL, Gu HY, Qu LJ (2006) The MYB transcription factor superfamily of arabidopsis: expression analysis and phylogenetic comparison with the rice MYB family. Plant Mol Biol 60:107–124. doi:10.1007/s11103-005-2910-y
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743. doi:10.1046/j.1365-313x.1998.00343.x
Czikkel BE, Maxwell DP (2007) NtGRAS1, a novel stress-induced member of the GRAS family in tobacco, localizes to the nucleus. J Plant Physio 164:1220–1230. doi:10.1016/j.jplph.2006.07.010
Day RB, Shibuya N, Minami E (2003) Identification and characterization of two new members of the GRAS gene family in rice responsive to N-acetylchitooligosaccharide elicitor. BBA Gene Struct Expr 1625:261–268. doi:10.1016/s0167-4781(02)00626-7
Dubois M, Gilles KA, Hamilton JK, Rebers PA, Smith F (1956) Colorimetric method for determination of sugars and related substances. Anal Chem 28:350–356. doi:10.1021/ac60111a017
Fode B, Siemsen T, Thurow C, Weigel R, Gatz C (2008) The Arabidopsis GRAS protein SCL14 interacts with class II TGA transcription factors and is essential for the activation of stress-inducible promoters. Plant Cell 20:3122–3135. doi:10.1105/tpc.108.058974
Fu XD, Sudhakar D, Peng JR, Richards DE, Christou P, Harberd NP (2001) Expression of arabidopsis GAI in transgenic rice represses multiple gibberellin responses. Plant Cell 13:1791–1802. doi:10.1105/tpc.13.8.1791
Hall BG (2013) Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol 30:1229–1235. doi:10.1093/molbev/mst012
Helariutta Y, Fukaki H, Wysocka-Diller J, Nakajima K, Jung J, Sena G, Hauser MT, Benfey PN (2000) The SHORT-ROOT gene controls radial patterning of the Arabidopsis root through radial signaling. Cell 101:555–567. doi:10.1016/s0092-8674(00)80865-x
Heo JO, Chang KS, Kim IA, Lee MH, Lee SA, Song SK, Lee MM, Lim J (2011) Funneling of gibberellin signaling by the GRAS transcription regulator SCARECROW-LIKE 3 in the Arabidopsis root. P Natl Acad Sci USA 108:2166–2171. doi:10.1073/pnas.1012215108
Hubbard KE, Nishimura N, Hitomi K, Getzoff ED, Schroeder JI (2010) Early abscisic acid signal transduction mechanisms: newly discovered components and newly emerging questions. Gene Dev 24:1695–1708. doi:10.1101/gad.1953910
Karan R, Subudhi PK (2012) A stress inducible SUMO conjugating enzyme gene (SaSce9) from a grass halophyte Spartina alterniflora enhances salinity and drought stress tolerance in Arabidopsis. BMC Plant Biol 12:187. doi:10.1186/1471-2229-12-187
Kim EY, Seo YS, Park KY, Kim SJ, Kim WT (2014) Overexpression of CaDSR6 increases tolerance to drought and salt stresses in transgenic Arabidopsis plants. Gene 552:146–154. doi:10.1016/j.gene.2014.09.028
Lee H, Kim B, Song S-K, Heo J-O, Yu N-I, Lee SA, Kim M, Kim DG, Sohn SO, Lim CE, Chang KS, Lee MM, Lim J (2008) Large-scale analysis of the GRAS gene family in Arabidopsis thaliana. Plant Mol Biol 67:659–670. doi:10.1007/s11103-008-9345-1
Li SH (2015) Grape breeding and genetics in China: history, current status and the future. Acta Hort 1082(165–176):2015. doi:10.17660/ActaHortic.1082.22
Li JT, Wang N, Xin HP, Li SH (2013) Overexpression of VaCBF4, a transcription factor from Vitis amurensis, improves cold tolerance accompanying increased resistance to drought and salinity in Arabidopsis. Plant Mol Biol Rep 31:1518–1528. doi:10.1007/s11105-013-0627-7
Lindemose S, O’Shea C, Jensen MK, Skriver K (2013) Structure, function and networks of transcription factors involved in abiotic stress responses. Int J Mol Sci 14:5842–5878. doi:10.3390/ijms14035842
Ma HS, Liang D, Shuai P, Xia XL, Yin WL (2010) The salt- and drought-inducible poplar GRAS protein SCL7 confers salt and drought tolerance in Arabidopsis thaliana. J Exp Bot 61:4011–4019. doi:10.1093/jxb/erq217
Miura K, Nozawa R (2014) Overexpression of SIZ1 enhances tolerance to cold and salt stresses and attenuates response to abscisic acid in Arabidopsis thaliana. Plant Biotechnol 31:167–172. doi:10.5511/plantbiotechnology.14.0109a
Miura K, Jin JB, Lee J, Yoo CY, Stirm V, Miura T, Ashworth EN, Bressan RA, Yun DJ, Hasegawa PM (2007) SIZ1-mediated sumoylation of ICE1 controls CBF3/DREB1A expression and freezing tolerance in Arabidopsis. Plant Cell 19:1403–1414. doi:10.1105/tpc.106.048397
Miyazaki Y, Abe H, Takase T, Kobayashi M, Kiyosue T (2015) Overexpression of LOV KELCH PROTEIN 2 confers dehydration tolerance and is associated with enhanced expression of dehydration-inducible genes in Arabidopsis thaliana. Plant Cell Rep 34:843–852. doi:10.1007/s00299-015-1746-4
Nakata M, Mitsuda N, Herde M, Koo AJ, Moreno JE, Suzuki K, Howe GA, Ohme-Takagi M (2013) A bHLH-Type transcription factor, ABA-INDUCIBLE BHLH-TYPE TRANSCRIPTION FACTOR/JA-ASSOCIATED MYC2-LIKE1, acts as a repressor to negatively regulate jasmonate signaling in Arabidopsis. Plant Cell 25:1641–1656. doi:10.1105/tpc.113.111112
Novillo F, Medina J, Salinas J (2007) Arabidopsis CBF1 and CBF3 have a different function than CBF2 in cold acclimation and define different gene classes in the CBF regulon. P Natl Acad Sci USA 104:21002–21007. doi:10.1073/pnas.0705639105
Pysh LD, Wysocka-Diller JW, Camilleri C, Bouchez D, Benfey PN (1999) The GRAS gene family in Arabidopsis: sequence characterization and basic expression analysis of the SCARECROW-LIKE genes. Plant J 18:111–119. doi:10.1046/j.1365-313X.1999.00431.x
Qiu YP, Yu DQ (2009) Over-expression of the stress-induced OsWRKY45 enhances disease resistance and drought tolerance in Arabidopsis. Environ Exp Bot 65:35–47. doi:10.1016/j.envexpbot.2008.07.002
Schumacher K, Schmitt T, Rossberg M, Schmitz C, Theres K (1999) The Lateral suppressor (Ls) gene of tomato encodes a new member of the VHIID protein family. P Natl Acad Sci USA 96:290–295. doi:10.1073/pnas.96.1.290
Shi R, Cao YB, Chen WR, Guo WD (2011) On cDNA cloning and expression analysis of GRAS gene in fingered citron. J Zhejiang Norm Univ (Nat Sci) 34:446–451 (in chinese)
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223. doi:10.1016/s1369-5266(00)80068-0
Silverstone AL, Ciampaglio CN, Sun TP (1998) The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10:155–169. doi:10.1105/tpc.10.2.155
Silverstone AL, Jung HS, Dill A, Kawaide H, Kamiya Y, Sun TP (2001) Repressing a repressor: gibberellin-induced rapid reduction of the RGA protein in Arabidopsis. Plant Cell 13:1555–1565. doi:10.1105/tpc.13.7.1555
Singh KB, Foley RC, Onate-Sanchez L (2002) Transcription factors in plant defense and stress responses. Curr Opin Plant Biol 5:430–436. doi:10.1016/s1369-5266(02)00289-3
Su LY, Dai ZW, Li SH, Xin HP (2015) A novel system for evaluating drought-cold tolerance of grapevines using chlorophyll fluorescence. BMC Plant Biol 15:82. doi:10.1186/s12870-015-0459-8
Thalhammer A, Bryant G, Sulpice R, Hincha DK (2014) Disordered cold regulated 15 proteins protect chloroplast membranes during freezing through binding and folding, but do not stabilize chloroplast enzymes in vivo. Plant Physiol 166:190–201. doi:10.1104/pp.114.245399
Torres-Galea P, Huang LF, Chua NH, Bolle C (2006) The GRAS protein SCL13 is a positive regulator of phytochrome-dependent red light signaling, but can also modulate phytochrome A responses. Mol Genet Genomics 276:13–30. doi:10.1007/s00438-006-0123-y
Wen CK, Chang C (2002) Arabidopsis RGL1 encodes a negative regulator of gibberellin responses. Plant Cell 14:87–100. doi:10.1105/tpc.010325
Xin HP, Zhu W, Wang LN, Xiang Y, Fang LC, Li JT, Sun XM, Wang N, Londo JP, Li SH (2013) Genome wide transcriptional profile analysis of Vitis amurensis and Vitis vinifera in response to cold stress. PLoS One 8:e58740. doi:10.1371/journal.pone.0058740
Xu K, Chen SJ, Li TF, Ma XS, Liang XH, Ding XF, Liu HY, Luo LJ (2015) OsGRAS23, a rice GRAS transcription factor gene, is involved in drought stress response through regulating expression of stress-responsive genes. BMC Plant Biol 15:141. doi:10.1186/s12870-015-0532-3
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803. doi:10.1146/annurev.arplant.57.032905.105444
Yang XW, Wang XY, Ji L, Yi ZL, Fu CX, Ran JC, Hu RB, Zhou GK (2015) Overexpression of a Miscanthus lutarioriparius NAC gene MlNAC5 confers enhanced drought and cold tolerance in Arabidopsis. Plant Cell Rep 34:943–958. doi:10.1007/s00299-015-1756-2
Yu LH, Chen X, Wang Z, Wang SM, Wang YP, Zhu QS, Li SG, Xiang CB (2013) Arabidopsis enhanced drought tolerance1/HOMEODOMAIN GLABROUS11 confers drought tolerance in transgenic rice without yield penalty. Plant Physiol 162:1378–1391. doi:10.1104/pp.113.217596
Acknowledgments
Financial support for this work was provided by the National Science Foundation of China (NSFC Accession No.:31130047, 31471857), National Key Technology R&D Program of the Ministry of Science and Technology during the Twelfth Five-year Plan Period (2013BAD02B04-1) and Youth Innovation Promotion Association of CAS (2015281).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interests.
Additional information
Communicated by E. Benvenuto.
Y. Yuan and L. Fang are equal contributors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yuan, Y., Fang, L., Karungo, S.K. et al. Overexpression of VaPAT1, a GRAS transcription factor from Vitis amurensis, confers abiotic stress tolerance in Arabidopsis. Plant Cell Rep 35, 655–666 (2016). https://doi.org/10.1007/s00299-015-1910-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-015-1910-x