Abstract
Key message
Overexpression of the iron transporter NtPIC1 increases iron concentration in shoots and reduces Cd uptake/accumulation in plants, mediating tolerance to cadmium.
Abstract
Cadmium (Cd) is toxic to plant cells and causes plants to display a typical iron (Fe) deficiency phenotype. NtPIC1 (Permease In Chloroplast1) is an Fe transporter protein in tobacco, required for Fe homeostasis. Based on preliminary results in transformed Saccharomyces cerevisiae BY4741 cells, which showed that NtPIC1 expression increased Cd tolerance, this study evaluated Cd tolerance in tobacco plants overexpressing NtPIC1 (NtPIC1-OE). We show that these plants have longer roots and higher fresh weights than wild-type (WT) plants after Cd exposure. Under Cd stress, WT plants display more chlorosis, stronger growth inhibition, and lower chlorophyll concentrations than NtPIC1-OE plants. Importantly, NtPIC1-OE plants had higher Fe concentrations in shoots and lower Fe concentrations in roots, and Cd concentrations in NtPIC1-OE plants were significantly lower compared to those in WT plants. Moreover, Fe transport-related genes (NtPIC1, NtNRAMP1, and NtFER1) were upregulated in NtPIC1-OE plants, while Fe deficiency-related genes (NtFRO1, NtIRT1, and NtZIP1) that mediate Cd uptake were downregulated. We also found that the activities of several antioxidative enzymes were significantly higher in NtPIC1-OE plants than in WT plants under Cd stress. Overall, our results demonstrate that overexpression of NtPIC1 is an efficient way to increase shoot Fe concentrations and reduce Cd uptake/accumulation in plants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Heavy metal pollution is a significant environmental problem that affects many physiological and biochemical processes in plants (Williams and Salt 2009; Clemens et al. 2013). Cadmium (Cd), a non-essential element in plants, is a major, widespread industrial and agricultural pollutant (Sanità di Toppi and Gabbrielli 1999). Environmental Cd is toxic to plants and animals, including humans (Raskin et al. 1997; Sanità di Toppi and Gabbrielli 1999).
In plants, Cd toxicity disrupts photosynthesis and slows thylakoid membrane development, as well as the synthesis of chloroplast–protein complexes (Das et al. 1997; Fagioni et al. 2008). It also disturbs plant water status (Perfus-Barbeoch et al. 2002) and injures cell components by producing reactive oxygen species (ROS) (Schützendübel and Polle 2002; Lee et al. 2003). As a result, visible symptoms occur in affected plants, including chlorosis, the browning of root tips, and growth retardation. The toxicity of Cd is due to binding competition with essential metals [e.g., iron (Fe), magnesium (Mn), and zinc (Zn)] for low specificity metal transporters such as iron-regulated transporter 1 (IRT1: Eide et al. 1996; Cohen et al. 1998; Vert et al. 2002; Yoshihara et al. 2006) and natural resistance-associated macrophage protein 1 (NRAMP1: Curie et al. 2000; Thomine et al. 2000; Williams et al. 2000; Thomine et al. 2003). This competition reduces the uptake of essential metals, especially Fe (Garty et al. 1992; Cohen et al. 1998; Clemens 2001), an essential micronutrient for all organisms.
Previous research has demonstrated the tight link between Fe homeostasis and Cd. For example, Cd induces Fe deficiency in tobacco plants by inhibiting the expression of Fe-uptake-related genes (Yoshihara et al. 2006). In addition, phytosiderophore-mediated Fe acquisition can protect plants from Cd toxicity (Meda et al. 2007). Furthermore, mutant OPT3 (essential to shoot–root Fe signaling) in Arabidopsis causes increased Cd accumulation in the seed and root, as well as redistribution of Fe and Cd (Mendoza-Cózatl et al. 2014; Zhai et al. 2014). These findings clearly emphasize the important relationship between Fe homeostasis and Cd partitioning.
Thus, a clear understanding of the components in Fe transport is necessary to combat the effects of Cd toxicity in plants. This study investigates NtPIC1, a chloroplast iron transporter in tobacco. When overexpressed, NtPIC1 was found to increase iron concentration in chloroplasts and overall chlorophyll concentrations in the plant (Gong et al. 2014). This study was initiated to investigate the mechanisms underlying decreased Cd uptake and translocation to shoots following overexpression of NtPIC1 in tobacco. Elucidation of these mechanisms will contribute to enhancing the efficiency of micronutrients and reducing the levels of non-essential toxic metals in plants.
Materials and methods
Cd tolerance assays in yeast expressing NtPIC1
Cells from the Saccharomyces cerevisiae BY4741 strain (MATa his3 lue2 met15 ura3), transformed either with a pYES2 empty vector (EV) or with a pYES2-NtPIC1 (Gong et al. 2014), were grown on synthetic-defined (SD) medium (0.67 % bacto-yeast nitrogen base, pH 5.8; Difco, Detroit, MI), supplemented with 2 % d-glucose. An amino acid mixture without uracil was used for transformant selection. Transformed cells were then grown on SD-ura medium. SD-ura liquid cultures, inoculated with the respective transformants, were adjusted to an optical density at 600 nm (OD600) of 1.0 and diluted to an OD600 of 0.1–0.001. Subsequently, 10 μL each of the dilutions was spotted onto SD-ura agar medium (control) or onto plates containing different concentrations of NiSO4, CuSO4, CoCl2, or CdCl2. All cultures were incubated at 30 °C for 3 days. Growth of the yeast transformants in SD-ura liquid medium supplemented with 50 μM CdCl2 was monitored at OD600.
Plant growth conditions and heavy metal treatments
Experiments were performed using three transgenic tobacco lines (Nicotiana tabacum cv. ‘NtPIC1-OE1’, ‘NtPIC1-OE13’, ‘NtPIC1-OE33’) that overexpressed NtPIC1 (Gong et al. 2014), and wild-type (WT) tobacco plants (N. tabacum cv. SR1). NtPIC1-OE and WT seedlings were germinated on half-strength Murashige and Skoog (1/2 MS) basal agar medium (pH 5.8) (Murashige and Skoog 1962) in a growth chamber under a 16 h/8 h light/dark cycle at 25 °C. After 4 days, the seedlings were transferred to 1/2 MS basal medium supplemented with 2 % sucrose, 0.8 % agar, and 0, 75, or 100 μM CdCl2. Seedlings were weighed and root lengths were measured; changes in root length were determined after 7 days of further vertical growth. Ten seedlings of each line were measured and three replicate experiments were performed.
Fe and Cd concentrations were compared across NtPIC1-OE and WT plants grown on either MS medium (control) or medium plus 100 μM Cd (Cd). Two-week-old NtPIC1-OE and WT seedlings were transferred to MS agar plates with 100 μM CdCl2. After 10 days in a growth chamber under a 16-h light period, the shoots and roots of the plants were separately harvested. Part of each was used for total RNA extraction to determine gene expression profiles, as well as for measuring Fe and Cd concentrations.
Measurement of Fe and Cd concentrations
The concentrations of Fe and Cd were assessed using roots and shoots of NtPIC1-OE and WT plants grown as described above. At harvest, intact roots of NtPIC1-OE and WT plants were soaked in 20 mM EDTA and washed three times with deionized water (Liu et al. 2011a, b). The tissues were then oven-dried at 65 °C for 3 days. Dried samples were placed into a solution of 65 % HNO3 and 30 % H2O2 (9:1) at 140 °C for 10 h in a digestion stove (DongYe, China). Concentrations were determined using an ICP-OES (Perkin Elmer) as described by Yuan et al. (2005).
Measurement of chlorophyll concentration
Chlorophyll was extracted from NtPIC1-OE and WT leaves following the methods of Johnston et al. (1984). Weighed tobacco leaves were ground in a mortar and pestle with 20 mL of 80 % acetone and 0.4 g CaCO3 per gram of fresh leaves. Homogenates were centrifuged (10,000×g for 10 min), and the pellets were again extracted with 10 mL of 80 % acetone. Extractions were performed 4–5 times until the leaf samples were colorless. All extraction procedures were carried out in the dark at 0–5 °C. Absorbance of the combined supernatants was measured at 663 and 645 nm using a spectrophotometer (UV-5100, Shanghai, China). The reference sample for these measurements was 80 % acetone. Chlorophyll extractions were performed for at least three plants grown under each condition, and the experiment was repeated three times. Results are expressed as milligrams of chlorophyll per gram fresh weight.
Quantitative real-time PCR analysis of gene expression
Total RNA was extracted from root and shoot tissues of NtPIC1-OE and WT plants, using an RNAprep Pure Plant kit (TIANGEN, Beijing, China). Each RNA sample was isolated from three seedlings, grown and harvested at the same time. Single-strand cDNA was synthesized from total RNA using a ReverTra Ace kit (Toyobo Life Science, Shanghai, China). Quantitative RT-PCR (qRT-PCR) was performed using an Applied Biosystems 7300 real-time cycler. The thermocycling protocol was as follows: 5 min at 95 °C, followed by 40 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 20 s. PCR products were then subjected to a dissociation protocol of 95 °C for 15 s, 60 °C for 20 s, and 97 °C for 15 s. Primers used for these reactions are listed in Supplemental Table 1. NtACT served as the internal control and all data were normalized against NtACT levels (Hodoshima et al. 2007). Each 20 μL reaction contained 10 μL of SYBR Premix Ex Taq II (TaKaRa), 2 μL of sample, 1.6 μL of 10 μM primer pairs, 6.0 μL of water, and 0.4 μL of ROX reference dye (50×). Gene expression levels were determined using the \(2^{{ - \varDelta \varDelta C_{\text{T}} }}\) method (Livak and Schmittgen 2001). Three replicate PCR amplifications were performed for each sample.
Measurement of antioxidant enzyme activities and MDA content
NtPIC1-OE and WT plants were grown in MS medium (control) for 4 weeks and transferred to MS with 100 μM CdCl2 (Cd) for 10 days. We determined superoxide dismutase (SOD) activity by monitoring the inhibition of nitroblue tetrazolium (NBT) reduction (Beauchamp and Fridovich 1971). Peroxidase (POD) activity was measured via a standard method (Moerschbacher et al. 1988). Catalase (CAT) activity was monitored using the decomposition of H2O2 at 240 nm (Aebi 1984). Malondialdehyde (MDA) was determined by measuring the concentration of thiobarbituric acid-reacting substances (Buege and Aust 1978). Three plants were analyzed from each line. All of the above experiments were carried out three times.
Statistical analysis
Statistical analyses were performed using the Statistical Product and Service Solutions software package, version 16.0 (SPSS Inc.). Data were evaluated for normality and homogeneity of variance. A one-way analysis of variance (ANOVA) was used to test for significant differences between treatments. The ANOVA was followed by a Duncan’s Multiple Range test to compare individual means. Differences were considered significant at p < 0.05. Data represent mean values of three replicates.
Results
Overexpression of NtPIC1 results in increased tolerance to Cd
The BY4741-pYES2-NtPIC1-transformed S. cerevisiae yeast fet3fet4 cells were more sensitive to Ni, Cu, and Co than EV cells. However, the NtPIC1-transformed cells showed greater growth than EV cells in the Cd-supplemented SD-ura medium (Supplemental Fig. 1). We hypothesized that NtPIC1 expression was related to increased Cd tolerance in yeast cells.
To test this hypothesis, we compared the growth of NtPIC1-OE1, -OE13, and -OE33 with WT plants, under a series of Cd treatments. We used fresh weight and primary root length as measures of growth.
Four-day-old OE1, OE13, OE33, and WT seedlings were grown vertically on medium containing 0, 75, or 100 μM CdCl2 for 7 days. Figure 1a shows representative OE1 plants compared with WT plants; the latter exhibited significant growth inhibition under all Cd concentrations, but the effect was especially pronounced under 100 μM Cd. Specifically, compared with OE1 plants, WT plants were much smaller and showed signs of chlorosis. Next, the fresh weights of the three OE plants were significantly higher (Fig. 1b) and their roots significantly longer (Fig. 1c) than those of WT plants. Even in the absence of Cd, OE plants were likely able to accumulate fresh weight faster than WT plants because higher NtPIC1 levels result in the transport of more Fe to the chloroplast. Finally, the relative growth inhibition rates for roots of OE1, OE13, and OE33 plants were lower than WT plants under 100 μM Cd stress (Fig. 1d).
Fresh weights and root lengths of NtPIC1-OE and WT plants exposed to different Cd concentrations. a Phenotypes of representative plant OE1 grown on half-strength MS agar plates supplemented with 0, 75, or 100 μM CdCl2 were photographed 7 days after transfer to the Cd treatments. Scale bars 1 cm. b Fresh weights were measured after 7 days. c Root lengths were measured after 7 days. d The relative inhibitions of root growth in NtPIC1-OE and WT plants. WT seedlings show severe root growth inhibition at 100 μM CdCl2. Fresh weight and root length values correspond to mean ± SE (n = 10). Letters above the error bars (SD) indicate significant differences (p < 0.05)
NtPIC1-OE and chlorophyll, Fe and Cd concentrations
To examine the basis for the increased Cd tolerance of NtPIC1-OE plants, we measured chlorophyll levels in leaves and Cd and Fe concentrations in roots and shoots of OE1, OE13, OE33, and WT plants. The phenotypes of OE1, OE13, OE33, and WT plants treated with 100 μM Cd are shown in Fig. 2a; the WT plants were more chlorotic than the OE1, OE13, OE33 plants. The chlorophyll concentrations in the leaves of OE1, OE13, OE33 plants were significantly higher (p < 0.05) than in WT leaves (Fig. 2b–d). Compared with WT plants under both control and Cd stress, the three OE plants had lower Fe concentrations in the roots and higher Fe concentrations in the shoots (Fig. 2e, f). In addition, when under Cd stress, compared with WT plants, the shoots and roots of OE plants had significantly lower Cd concentrations (p < 0.05; Fig. 2g). These results support the interpretation that higher Fe levels in plants overexpressing NtPIC1 contribute to a reduction in both Cd uptake and Cd transport from the roots to the shoots.
The concentrations of Fe, Cd, and chlorophyll in NtPIC1-OE and WT plants. a Phenotypes of NtPIC1-OE and WT plants grown on MS medium containing 100 μM CdCl2 for 10 days. b–d Chlorophyll concentrations in leaves of NtPIC1-OE and WT plants. e–g Fe and Cd concentrations in root and shoot tissues of NtPIC1-OE and WT plants with or without Cd treatment. Letters above the error bars (SD) indicate significant differences (p < 0.05). Scale bars 1 cm
Expression of genes associated with iron transport
The above analysis indicated that root-to-shoot transport activity was reduced in NtPIC1-OE plants exposed to Cd. To determine whether this effect involves genes responsible for Fe uptake and transport, we analyzed the expression of the Fe transporter-related genes NtPIC1, NtFER1 (ferritin), NtNRAMP1, NtFRO1 (ferric-chelate reductase oxidase), NtIRT1, and NtZIP1 (zinc-regulated transporter/Fe-regulated transporter) in NtPIC1-OE and WT plants, under both control and Cd stress conditions. Representative results concerning the OE1 line are shown in Figs. 3 and 4. We used both NtACT and NtGAPDH as reference genes for qRT-PCR and performed parallel experiments with the two standards. We obtained equivalent results from both standards, and so we have chosen to show the data obtained from NtACT here and included the data from NtGAPDH as online supplemental material (Supplemental Fig. 2, 3). Our analysis showed that Cd stress had little effect on NtPIC1 expression in WT roots (Fig. 3a). In contrast, the Fe storage protein NtFER1 and the Fe transporter NtNRAMP1 were both significantly downregulated (p < 0.05; Fig. 3b, c), while the Fe deficiency-response genes NtFRO1, NtIRT1, and NtZIP1 were all significantly upregulated (p < 0.05; Fig. 3d–f). In NtPIC1-OE plants, NtPIC1 expression was higher than in WT plants regardless of the experimental conditions, because the OE plants were ectopically overexpressing NtPIC1 (Figs. 3g, 4g). However, in comparison to WT plants, NtFER1 and NtNRAMP1 expression was significantly upregulated in OE1 roots (p < 0.05; Fig. 3h, i), while NtFRO1 and NtIRT1 expression was significantly downregulated (p < 0.05; Fig. 3j, k). NtZIP1 expression was significantly upregulated in control conditions, but significantly downregulated under Cd treatment in the roots of OE1 plants compared with WT plants (p < 0.05; Fig. 3l).
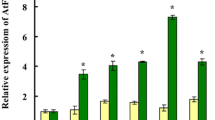
Quantitative RT-PCR analysis of iron metabolism-related genes in NtPIC1-OE1 and WT plant roots. Levels of NtPIC1, NtFER1, NtNRAMP1, NtFRO1, NtIRT1, and NtZIP1 transcripts were normalized against NtACT. Data represent the means of three technical replicates. a–f WT plants in control or Cd treatments. g–l NtPIC1-OE1 plants and WT plants in control and Cd treatments. a–f Asterisks significant differences from control (p < 0.05; Student’s t test). g–l Asterisks significant differences from WT (p < 0.05; Student’s t test)
Quantitative RT-PCR analysis of iron transporter-related genes in NtPIC1-OE1 and WT plant shoots. Levels of NtPIC1, NtFER1, NtNRAMP1, NtFRO1, NtIRT1, and NtZIP1 transcripts were normalized against NtACT. Data represent the mean of three technical replicates. a–f WT plants in control and Cd treatments. g–l NtPIC1-OE1 plants and WT plants in control or Cd treatments. a–f Asterisks significant differences from control (p < 0.05; Student’s t test). g–l Asterisks significant differences from WT (p < 0.05; Student’s t test)
A similar analysis of gene expression was performed using shoot tissues. In WT plants under Cd treatment, NtPIC1 and NtFER1 expressions were significantly downregulated (p < 0.05) (Fig. 4a, b), while NtNRAMP1, NtFRO1, and NtIRT1 expressions were significantly higher than under control conditions (p < 0.05; Fig. 4d–f). Unlike that in WT plants, NtFER1 and NtNRAMP1 expression levels were significantly upregulated in the shoots of OE1 plant under control conditions (p < 0.05; Fig. 4h, i). Again, unlike that in WT, NtFRO1 and NtIRT1 expression levels were significantly downregulated in the shoots of OE1 plants under both control and Cd treatment conditions (Fig. 4j, k); NtZIP1 expression (Siemianowski et al. 2014) was also downregulated in control conditions (Fig. 4l). Based on these results, we suggest that reduction of Cd uptake in NtPIC1-OE plants might be related to the decreased expression of the NtIRT1, NtFRO1, and NtZIP1 genes.
Antioxidant index analysis
Finally, we investigated antioxidant enzymes in NtPIC1-OE plants by screening the activities of SOD, POD, and CAT (Fig. 5a–c) and assessing MDA contents (Fig. 5d). Compared to WT plants, OE1, OE13, and OE33 plants showed significantly higher SOD, POD, and CAT activities under both control and 100 μM CdCl2 stress conditions (p < 0.05). The MDA contents of WT plants did not differ significantly from those of OE1, OE13, OE33 plants in control conditions, but after Cd exposure, MDA levels in WT plants were significantly higher than in the three OE plants (p < 0.05; Fig. 5d). This result indicates that Cd-induced membrane damage was more severe in WT plants than in NtPIC1-OE plants.
Effect of NtPIC1 overexpression on antioxidant enzymes and MDA amounts in NtPIC1-OE and WT plants treated with Cd. a Superoxide dismutase (SOD) activity. b Peroxidase (POD) activity. c Catalase (CAT) activity. d Malondialdehyde (MDA) content in leaves of transgenic and WT plants grown in 100 μM CdCl2 for 10 days. Values shown are mean ± SE. Three plants were measured in each line. Letters above the error bars (SD) indicate significant differences (p < 0.05)
Discussion
Cd toxicity is mainly due to the disruption of essential metal uptake and homeostasis (Clemens et al. 2013). This disruption is caused by binding competition between Cd and other essential metals, especially Fe (Schützendübel and Polle 2002). Thus, we hypothesized that NtPIC1 overexpression should lead to higher cell Fe levels, reducing Cd ion accumulation, and thereby alleviating the impact of Cd toxicity.
In higher plants, Cd phytotoxicity inhibits root elongation (Schützendübel and Polle 2002). Our results demonstrated that under Cd stress, NtPIC1-OE plants had significantly longer roots, higher fresh weights, and lower growth suppression than WT plants (Fig. 1). Moreover, NtPIC1-OE plants had higher chlorophyll levels and remained green, while WT plants exhibited severe chlorosis (Fig. 2a–d). These results indicate that NtPIC1-OE plants were more Cd tolerant than WT plants. In contrast to Fe, the Cd concentrations in the roots and shoots of NtPIC1-OE plants were significantly lower than in WT plants (Fig. 2e–g). This pattern suggests that NtPIC1 overexpression increased Fe transport from roots to shoots and decreased Cd accumulation in shoots and roots. At the same time, Cd interferes with Fe homeostasis, causing Fe deficiency in the upper plant parts. Low Fe concentrations then induce two processes in the roots: the upregulation of Fe deficiency-response genes NtFRO1 and NtIRT1 (Fig. 3d, e) (Yoshihara et al. 2006) and the downregulation of the Fe storage protein NtFER1 (Figs. 3b, 4b). A recent study found that upregulation of IRT1 resulted in Cd accumulation because of a wide range of substrate transport properties, leading to Cd accumulation (Korshunova et al. 1999; Mendoza-Cózatl et al. 2014). In this study, changes to transport components were also observed: the potential metal uptake-related genes NtFRO1, NtIRT1, and NtZIP1 were expressed at lower levels in Cd-exposed NtPIC1-OE plants (Figs. 3j–l, 4j–l), leading to lower Cd accumulation. Additionally, the iron transporter NtNRAMP1 was upregulated in NtPIC1-OE roots, possibly facilitating Fe accumulation under Cd stress. These results indicate that Cd exposure interferes with the Fe homeostasis mechanism (Bovet et al. 2006; Yoshihara et al. 2006; Hodoshima et al. 2007), but the resultant downregulation of NtFRO1, NtIRT1, and NtZIP1 may prevent toxic levels of Cd accumulation.
Plant tolerance to Cd involves numerous physiological and molecular mechanisms, such as cell wall binding, chelation with phytochelatins, and compartmentalization in vacuoles (Wu et al. 2012). According to the results presented here, higher Cd tolerance in NtPIC1-OE plants is a consequence of increasing Fe entry into plants, particularly root-to-shoot partitioning, which results in the restriction of Cd uptake (Wu et al. 2012). Although Cd does not participate in the Fenton and Haber–Weiss reactions, exposure to the metal can still cause oxidative stress (Clemens 2006). Cd increases ROS in plants (Cuypers et al. 2011; Liu et al. 2011a, b), injuring cell components (Romero-Puertas et al. 2004; Heyno et al. 2008). Following previous research (Halliwell 2006; Møller et al. 2007; Gill and Tuteja 2010), we used MDA levels and antioxidant enzyme activity as diagnostics for oxidative stress (ROS presence). We found the NtPIC1-OE plants exhibited lower levels of MDA content and higher levels of antioxidant enzyme activity (Fig. 5), which is in accord with the decrease in Cd accumulation. Our results thus indicate that NtPIC1-OE plants experience less oxidative stress than WT plants when exposed to Cd (Fig. 5).
In summary, this work has shown clearly that under Cd stress, NtPIC1-OE plants were more efficient than WT in Fe translocation from roots to shoots. Further, NtPIC1-mediated Fe acquisition effectively mitigated Cd toxicity by reducing the risk of Cd oxidative damage at the cellular level. Higher Fe concentrations in NtPIC1-OE plants also appeared to suppress the expression of iron deficiency genes (NtFRO1, NtIRT1, and NtZIP1) (Zhao and Eide 1996; Connolly et al. 2002; Enomoto et al. 2007), which are putatively involved in Cd uptake. Thus, their downregulation likely resulted in the reduction of root and shoot Cd concentrations. These findings indicate that Fe plays a considerable role in the alleviation of Cd toxicity in plants, which should contribute greatly to biofortification efforts aimed at reducing heavy metal contamination of crops.
Author contribution statement
Changhong Guo conceived and designed the experiments. Xun Gong and LinWei Yin performed the experiments. Xun Gong wrote the manuscript. Jiaqi Chen analyzed data.
References
Aebi H (1984) Catalase in vitro. Methods Enzymol 105:121–126
Beauchamp C, Fridovich I (1971) Superoxide dismutase: improved assays and an assay applicable to acrylamide gels. Anal Biochem 44:276–287
Bovet L, Rossi L, Lugon-Moulin N (2006) Cadmium partitioning and gene expression studies in Nicotiana tabacum and Nicotiana rustica. Physiol Plant 128:466–475
Buege JA, Aust SD (1978) Microsomal lipid peroxidation. Methods Enzymol 53:302–310
Clemens S (2001) Developing tools for phytoremediation: towards a molecular understanding of plant metal tolerance and accumulation. Int J Occup Med Environ Health 14:235–239
Clemens S (2006) Toxic metal accumulation, responses to exposure and mechanisms of tolerance in plants. Biochimie 88:1707–1719
Clemens S, Aarts MGM, Thomine S, Verbruggen N (2013) Plant science: the key to preventing slow cadmium poisoning. Trends Plant Sci 18:92–99
Cohen CK, Fox TC, Garvin DF, Kochian LV (1998) The role of iron-deficiency stress responses in stimulating heavy-metal transport in plants. Plant Physiol 116:1063–1072
Connolly EL, Fett JP, Guerinot ML (2002) Expression of the IRT1 metal transporter is controlled by metals at the levels of transcript and protein accumulation. Plant Cell 14:1347–1357
Curie C, Alonso JM, Jean ML, Ecker JR, Briat JF (2000) Involvement of NRAMP1 from Arabidopsis thaliana in iron transport. Biochem J 347:749–755
Cuypers A, Karen S, Jos R, Kelly O, Els K, Tony R, Nele H, Nathalie V, Suzy VS, Frank VB, Yves G, Jan C, Jaco V (2011) The cellular redox state as a modulator in cadmium and copper responses in Arabidopsis thaliana seedlings. J Plant Physiol 168:309–316
Das P, Samantaray S, Rout GR (1997) Studies on cadmium toxicity in plants: a review. Environ Pol 98:29–36
Eide D, Broderius M, Fett J, Guerinot ML (1996) A novel iron-regulated metal transporter from plants identified by functional expression in yeast. Proc Natl Acad Sci 93:5624–5628
Enomoto Y, Hodoshima H, Shimada H, Shoji K, Yoshihara T, Goto F (2007) Long-distance signals positively regulate the expression of iron uptake genes in tobacco roots. Planta 227:81–89
Fagioni M, D’Amici GM, Timperio AM, Zolla L (2008) Proteomic analysis of multiprotein complexes in the thylakoid membrane upon cadmium treatment. J Proteome Res 8:310–326
Garty J, Karary Y, Harel J (1992) Effect of low pH, heavy metals and anions on chlorophyll degradation in the lichen Ramalina duriaei (de not.) bagl. Environ Exp Bot 32:229–241
Gill SS, Tuteja N (2010) Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol Biochem 48:909–930
Gong X, Guo CH, Terachi T, Cai HS, Yu DS (2014) Tobacco PIC1 mediates iron transport and regulates chloroplast development. Plant Mol Biol Rep 33:401–413
Halliwell B (2006) Reactive species and antioxidants. Redox biology is a fundamental theme of aerobic life. Plant Physiol 141:312–322
Heyno E, Klose C, Krieger-Liszkay A (2008) Origin of cadmium-induced reactive oxygen species production: mitochondrial electron transfer versus plasma membrane NADPH oxidase. New Phytol 179:687–699
Hodoshima H, Enomoto Y, Shoji K, Shimada H, Goto F, Yoshihara T (2007) Differential regulation of cadmium-inducible expression of iron-deficiency-responsive genes in tobacco and barley. Physiol Plant 129:622–634
Johnston M, Grof CPL, Brownell PF (1984) Effect of sodium nutrition on chlorophyll a/b ratios in C4 plants. Funct Plant Biol 11:325–332
Korshunova Y, Eide D, Gregg Clark W, Lou Guerinot M, Pakrasi H (1999) The IRT1 protein from Arabidopsis thaliana is a metal transporter with a broad substrate range. Plant Mol Biol 40:37–44
Lee S, Moon JS, Ko TS, Petros D, Goldsbrough PB, Korban SS (2003) Overexpression of Arabidopsis phytochelatin synthase paradoxically leads to hypersensitivity to cadmium stress. Plant Physiol 131:656–663
Liu GY, Zhang YX, Chai TY (2011a) Phytochelatin synthase of Thlaspi caerulescens enhanced tolerance and accumulation of heavy metals when expressed in yeast and tobacco. Plant Cell Rep 30:1067–1076
Liu YT, Chen ZS, Hong CY (2011b) Cadmium-induced physiological response and antioxidant enzyme changes in the novel cadmium accumulator, Tagetes patula. J Hazard Mater 189:724–731
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the \(2^{{ - \varDelta \varDelta C_{\text{T}} }}\) method. Methods 25:402–408
Meda AR, Scheuermann EB, Prechsl UE, Erenoglu B, Schaaf G, Hayen H, Weber G, von Wiren N (2007) Iron acquisition by phytosiderophores contributes to cadmium tolerance. Plant Physiol 143:1761–1773
Mendoza-Cózatl DG, Xie Q, Akmakjian GZ, Jobe TO, Patel A, Stacey MG, Song L, Demoin DW, Jurisson SS, Stacey G, Schroeder JI (2014) OPT3 is a component of the iron-signaling network between leaves and roots and misregulation of OPT3 leads to an over-accumulation of cadmium in seeds. Mol Plant 7:1455–1469
Moerschbacher BM, Noll UM, Flott BE, Reisener HJ (1988) Lignin biosynthetic enzymes in stem rust infected, resistant and susceptible near-isogenic wheat lines. Physiol Mol Plant Pathol 33:33–46
Møller IM, Jensen PE, Hansson A (2007) Oxidative modifications to cellular components in plants. Annu Rev Plant Biol 58:459–481
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Perfus-Barbeoch L, Leonhardt N, Vavasseur A, Forestier C (2002) Heavy metal toxicity: cadmium permeates through calcium channels and disturbs the plant water status. Plant J 32:539–548
Raskin I, Smith RD, Salt DE (1997) Phytoremediation of metals: using plants to remove pollutants from the environment. Curr Opin Biotechnol 8:221–226
Romero-Puertas MC, RodríGuez-Serrano M, Corpas FJ, Gómez M, Del Río LA, Sandalio LM (2004) Cadmium-induced subcellular accumulation of O2− and H2O2 in pea leaves. Plant, Cell Environ 27:1122–1134
Sanità di Toppi L, Gabbrielli R (1999) Response to cadmium in higher plants. Environ Exp Bot 41:105–130
Schützendübel A, Polle A (2002) Plant responses to abiotic stresses: heavy metal-induced oxidative stress and protection by mycorrhization. J Exp Bot 53:1351–1365
Siemianowski O, Barabasz A, Kendziorek M, Ruszczyńska A, Bulska E, Williams LE, Antosiewicz DM (2014) AtHMA4 expression in tobacco reduces Cd accumulation due to the induction of the apoplastic barrier. J Exp Bot. doi:10.1093/jxb/ert471
Thomine S, Wang R, Ward JM, Crawford NM, Schroeder JI (2000) Cadmium and iron transport by members of a plant metal transporter family in Arabidopsis with homology to Nramp genes. Proc Natl Acad Sci 97:4991–4996
Thomine S, Lelièvre F, Debarbieux E, Schroeder JI, Barbier-Brygoo H (2003) AtNRAMP3, a multispecific vacuolar metal transporter involved in plant responses to iron deficiency. Plant J 34:685–695
Vert G, Grotz N, Dédaldéchamp F, Gaymard F, Guerinot ML, Briat JF, Curie C (2002) IRT1, an Arabidopsis transporter essential for iron uptake from the soil and for plant growth. Plant Cell 14:1223–1233
Williams L, Salt DE (2009) The plant ionome coming into focus. Curr Opin Plant Biol 12:247–249
Williams LE, Pittman JK, Hall JL (2000) Emerging mechanisms for heavy metal transport in plants. BBA Biomembr 1465:104–126
Wu HL, Chen CL, Du J, Liu HF, Cui Y, Zhang Y, He YJ, Wang YQ, Chu CC, Feng ZY, Liu JM, Ling HQ (2012) Co-overexpression FIT with AtbHLH38 or AtbHLH39 in Arabidopsis-enhanced cadmium tolerance via increased cadmium sequestration in roots and improved iron homeostasis of shoots. Plant Physiol 158:790–800
Yoshihara T, Hodoshima H, Miyano Y, Shoji K, Shimada H, Goto F (2006) Cadmium inducible Fe deficiency responses observed from macro and molecular views in tobacco plants. Plant Cell Rep 25:365–373
Yuan YX, Zhang J, Wang DW, Ling HQ (2005) AtbHLH29 of Arabidopsis thaliana is a functional ortholog of tomato FER involved in controlling iron acquisition in strategy I plants. Cell Res 15:613–621
Zhai Z, Gayomba SR, Hi Jung, Vimalakumari NK, Piñeros M, Craft E, Rutzke MA, Danku J, Lahner B, Punshon T, Guerinot ML, Salt DE, Kochian LV, Vatamaniuk OK (2014) OPT3 is a phloem-specific iron transporter that is essential for systemic iron signaling and redistribution of iron and cadmium in Arabidopsis. Plant Cell 26:2249–2264
Zhao H, Eide D (1996) The yeast ZRT1 gene encodes the zinc transporter protein of a high-affinity uptake system induced by zinc limitation. Proc Natl Acad Sci USA 93(6):2454–2458
Acknowledgments
We thank Prof. Shaojian Zheng (Zhejiang University) for providing the yeast strain BY4741. This work was supported by the MOST 863 Project (2013AA102607-5), and the Graduate Innovation Fund of Harbin Normal University (HSDBSCX2012-05) and Graduate Innovation Fund of Harbin Normal University (HSDSSCX2014-07).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by J. Register.
X. Gong and L. Yin are co-first authors of this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2015_1843_MOESM1_ESM.tif
Supplemental Fig. 1 Expression of NtPIC1 enhances Cd tolerance in Saccharomyces cerevisiae. A. Cells of S. cerevisiae strain BY4741 (WT) carrying pYES2 (Empty vector) or pYES2-NtPIC1 were grown in SD-ura medium (control) and SD-ura medium containing different concentrations of NiSO4, CuSO4, CoCl2, or CdCl2, respectively, for 3 d. B, C. Growth curves of yeast strains in SD-ura liquid control medium or medium supplemented with 50 μM CdCl2. The growth of the yeast cells was monitored at OD600. The data shows means ± standard deviation (SD) from three independent experiments. Asterisks indicate a significant difference from BY4741-pYES2 (p < 0.05; Student’s t test) (TIFF 1298 kb)
299_2015_1843_MOESM2_ESM.tif
Supplemental Fig. 2 Quantitative RT-PCR analysis of iron transporter-related genes in NtPIC1-OE1 and WT plant roots. Levels of NtPIC1, NtFER1, NtNRAMP1, NtFRO1, NtIRT1, and NtZIP1 transcripts were normalized against NtGAPDH. Data represent the mean of three technical replicates. A-F. WT plants in control and Cd treatments. Asterisks indicate significant differences from control (p < 0.05; Student’s t-test). G-L. NtPIC1-OE1 plants and WT plants in control or Cd treatments. Asterisks indicate significant differences from WT (p < 0.05; Student’s t-test) (TIFF 2677 kb)
299_2015_1843_MOESM3_ESM.tif
Supplemental Fig. 3 Quantitative RT-PCR analysis of iron transporter-related genes in NtPIC1-OE1 and WT plant shoots. Levels of NtPIC1, NtFER1, NtNRAMP1, NtFRO1, NtIRT1, and NtZIP1 transcripts were normalized against NtGAPDH. Data represent the mean of three technical replicates. A-F. WT plants in control and Cd treatments. Asterisks indicate significant differences from control (p < 0.05; Student’s t-test). G-L. NtPIC1-OE1 plants and WT plants in control or Cd treatments. Asterisks indicate significant differences from WT (p < 0.05; Student’s t-test) (TIFF 2731 kb)
Rights and permissions
About this article
Cite this article
Gong, X., Yin, L., Chen, J. et al. Overexpression of the iron transporter NtPIC1 in tobacco mediates tolerance to cadmium. Plant Cell Rep 34, 1963–1973 (2015). https://doi.org/10.1007/s00299-015-1843-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-015-1843-4