Abstract
Key message
Expression of OsHsfB2b was strongly induced by heat, salt, ABA and PEG treatments. Drought and salt tolerances were significantly decreased by OsHsfB2b overexpression, but were enhanced by RNA interference.
Abstract
Plants have more than 20 heat shock factors (Hsfs) that were designated class A, B, and C. Many members of Class A Hsfs were characterized as activators of transcription, but the functional roles of class B and C Hsfs have not been fully recognized. OsHsfB2b is a member of class B Hsfs in rice (Oryza sativa). Expression of OsHsfB2b was strongly induced by heat, salt, abscisic acid (ABA) and polyethylene glycol (PEG) treatments but was almost not affected by cold stress. Drought and salt tolerances were significantly decreased in OsHsfB2b-overexpressing transgenic rice, but were enhanced in the OsHsfB2b-RNAi transgenic rice. Under drought stress, the OsHsfB2b-overexpressing transgenic rice exhibited increased relative electrical conductivity (REC) and content of malondialdehyde (MDA) and decreased proline content compared with the wild type, while the lower REC and MDA content and increased proline content were found in the OsHsfB2b-RNAi transgenic rice. These results suggest that OsHsfB2b functions as a negative regulator in response to drought and salt stresses in rice, with its existing B3 repression domain (BRD) that might be necessary for the repressive activity. The present study revealed the potential value of OsHsfB2b in genetic improvement of rice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Plants cannot escape from unfavorable environmental conditions and thus have developed numerous physiological and biochemical changes to respond and adapt to the adverse conditions during their growth and development. The major modes of plant response to various stresses are perception and transduction of the stress signals through signaling components, which result in expression of a great quantity of stress-related genes and synthesis of diverse functional proteins. Finally, a variety of physiological and metabolic responses are produced in plants to adapt to the stresses (Hirayama and Shinozaki 2010; Lata and Prasad 2011; Tang et al. 2012; Yamaguchi-Shinozaki and Shinozaki 2006; Zhu 2002).
Heat shock proteins (Hsps) in plants function as molecular chaperones which regulate cellular homeostasis and promote plants survival under stressful conditions (Hartl and Hayer-Hartl 2002; Haslbeck and Buchner 2002). The expressions of Hsps are regulated by heat shock factors (Hsfs) through their interaction with heat shock elements (HSE) presented in the Hsp promoter region (Baniwal et al. 2004; Nover et al. 2001). Plants have more than 20 Hsfs, with Arabidopsis and rice having 21 and 25 members, respectively (Guo et al. 2008; Nover et al. 2001). Based on structural characteristics and phylogenetic comparison, plant Hsfs are subdivided into three classes (A, B and C). Class A Hsfs possess the exclusive C-terminal activation domain with AHA motifs, while class B and class C Hsfs are characterized by lacking an activation domain (Baniwal et al. 2004; Nover et al. 2001). In Arabidopsis, the members of the HsfA1 group (HsfA1a, HsfA1b, HsfA1d and HsfA1e) function as the master regulators of heat stress response (HSR) and participate in other abiotic stress responses as important components (Busch et al. 2005; Liu et al. 2011; Lohmann et al. 2004). Overexpression of the Arabidopsis gene AtHsfA1b improved resistance to chilling in transgenic tomato (Li et al. 2003). HsfA1 was characterized as master regulator of thermotolerance in tomato (Mishra et al. 2002). Overexpression of GmHsfA1 significantly enhanced thermotolerance in soybean by the activation of GmHsp70 (Zhu et al. 2006). The HsfA2 was reported as a key regulator in the induction of the defence system under several types of environmental stresses and enhanced tolerance to heat, high light, salt, oxidative, osmotic and anoxia stresses in Arabidopsis (Banti et al. 2010; Charng et al. 2007; Li et al. 2005; Nishizawa-Yokoi et al. 2011; Nishizawa et al. 2006; Ogawa et al. 2007; Schramm et al. 2006). Overexpressing rice gene OsHsfA2e highly expressed certain stress-associated genes under unstressed conditions and improved thermotolerance and salt stress tolerance in transgenic Arabidopsis (Yokotani et al. 2008). Furthermore, HsfA3, HsfA4a and HsfA7 play an important role in abiotic stress tolerance in Arabidopsis, rice and wheat (Larkindale and Vierling 2008; Liu et al. 2013; Schramm et al. 2008; Shim et al. 2009; Yoshida et al. 2008).
In contrast to class A Hsfs, there was no evidence that class B and C Hsfs functioned as transcription activators on their own due to lack of the activation domain (Czarnecka-Verner et al. 2000, 2004; Kotak et al. 2004; Nover et al. 2001). Tomato HsfB proteins were suggested to be coactivators of HsfAs (Bharti et al. 2004; Hahn et al. 2011). OsHsfB4b showed interaction with OsHsfA2c and activated the target genes (Singh et al. 2012). Recent researches show that some members of class B Hsfs are active transcriptional repressors. The repressive activity of HsfB1 and HsfB2b was reported in Arabidopsis and soybean, and a repression domain, designated the B3 repression domain (BRD), was found in HsfB1 and HsfB2b at the C terminus (Czarnecka-Verner et al. 2000, 2004; Ikeda et al. 2011; Ikeda and Ohme-Takagi 2009; Kumar et al. 2009). HsfB1 and HsfB2b were shown to regulate the expression of defensin genes, and the disease resistance were significantly improved in mutant lines compared with the wild type (Kumar et al. 2009), suggesting that HsfB1 and HsfB2b may negatively regulate the disease response. HsfB1 and HsfB2b suppressed the general heat shock response under non-heat-stress conditions and in the attenuating period via direct repression of the expression of the heat stress inducible HsfA2 (Ikeda et al. 2011). On the other hand, HsfB1 and HsfB2b exhibited cell death-inducing activity in Nicotiana benthamiana, and the cell death symptom by HsfB1 and HsfB2b required both their repression activity and their nuclear localization activity (Zhu et al. 2012). The Hsfs interaction showed that HsfB1 and HsfB2b might be involved in complex regulatory networks, which were linked to stress responses and signaling processes (Li et al. 2010).
In this paper, we report the functional analysis of OsHsfB2b, a member of class B Hsfs in rice. Overexpression of OsHsfB2b significantly decreased drought and salt tolerance in transgenic plants, while the OsHsfB2b-RNAi plants enhanced tolerance under these stresses. Our results indicate OsHsfB2b may negatively regulate the abiotic stress tolerance in rice.
Materials and methods
Generation of transgenic rice plants
The full-length cDNA clone of OsHsfB2b (AK101700) was obtained from the National Institute of Agrobiological Sciences (NIAS, Tsukuba, Japan). To create the overexpression construct of OsHsfB2b, the OsHsfB2b coding region was PCR amplified with BamHI-PstI (indicated by underline) linker primers 5′-CGGGATCCTGGCATGGCTGACCAGACCGCT-3′ and 5′-AACTGCAGCGAGACGGGTCTAACACACAAC-3′ based on the template of the cDNA clone AK101700. The PCR fragment of OsHsfB2b was digested by BamHI and PstI and ligated into the BamHI and PstI sites of the binary expression vector pCAMBIA1301-Multi (modified from pCAMBIA1301) under the control of the CaMV 35S promoter. To make double-strand RNA interference (RNAi) construct, the antisense fragment OsHsfB2b-A was PCR amplified with XhoI-KpnI (indcated by underline) linker primers 5′-CCGCTCGAGGTTTACCAGCCCATCCG-3′ and 5′-GGGGTACCAACTCGATCTGATCTTTTATGC-3′, the sense fragment OsHsfB2b-S was PCR amplified with PstI-XbaI (indcated by underline) linker primers 5′-AACTGCAGGTTTACCAGCCCATCCG-3′ and 5′-GCTCTAGAAACTCGATCTGATCTTTTATGC-3′, using the cDNA clone AK101700 as the template. The OsHsfB2b-A fragment was digested by XhoI and KpnI and cloned into pBSK-3Dlinker (containing 128 bp intron sequence of RAmy3D between XhoI and PstI of pBluescript SK, pBS-OsHsfB2b-A-ΔRAmy3D). The OsHsfB2b-S fragment was digested by PstI and XbaI and cloned into pBS-OsHsfB2b-A-ΔRAmy3D (pBS-OsHsfB2b-A-ΔRAmy3D-OsHsfB2b-S). Finally, pBS-OsHsfB2b-A-ΔRAmy3D-OsHsfB2b-S was digested with KpnI and XbaI and cloned into pCAMBIA1301-Multi to yield the 35S::RNAi OsHsfB2b. Both of the constructs were transformed into the japonica rice Nipponbare by the Agrobacterium-mediated transformation method (Toki et al. 2006).
Stress treatment of plant materials
To check the expression level of OsHsfB2b gene under various abiotic stresses or phytohormone treatment, Nipponbare rice plants were grown in sandy soil (for heat and cold stress) or hydroponic culture medium (1/2 MS, for others) for about 3 weeks in a climate chamber at 28 °C, 14 h/10 h (light/dark) and 75 % relative humidity. The seedlings at four-leaf stage were used for the following treatments: osmotic stress (20 % PEG), salt stress (200 mM NaCl), heat stress (exposing plants to 42 °C), cold stress (seedlings were transferred to a growth chamber at 4 °C), and phytohormone treatment (0.1 mM ABA). Samples were taken after treatment for 0, 0.5, 1, 2, 4, 8 and 24 h separately. Ten shoots were pooled together as one biological replicate and each treatment was repeated three times.
To detect the transcript levels of OsHsfB2 in different tissues, Nipponbare rice plants were grown in field or the climate chamber and tissue samples of roots, stems, leaves, sheaths and spikes were collected essentially as described previously (Zou et al. 2009). Seedlings were obtained from the two-week-old rice seedlings grown in a climate chamber. Three replications were performed in this study.
To test various abiotic stress tolerances of transgenic plants at the seedling stage, positive T2 transgenic lines were selected by germinating seeds in 1/2 MS medium containing 50 mg/L hygromycin; the wild type was also germinated in normal 1/2 MS medium. The one-week-old rice seedlings (15 plants each, three repeats) were grown in an equational separation manner (OsHsfB2b-overexpressing:wild type:OsHsfB2b-RNAi = 1:1:1) in plastic pots filled with sandy soil. After 2 weeks growing in the growth chamber, the seedlings were used for abiotic stress treatments. For drought stress treatment, irrigation was withheld for 1 week followed by recovery for 1 week. For salt stress treatment, irrigated with 200 mM NaCl solution for 1 week followed by recovery for 1 week. For heat or cold stress treatment, the seedlings were transferred to a growth chamber at 45 °C for 5 h/day continued for 5 days or 4 °C for 5 days separately, followed by recovery at 28 °C for 1 week.
For testing the osmotic or salt stress tolerance of transgenic seed germination, the homozygous T3 transgenic lines and the wild type (40 seeds each, three repeats) were germinated in 1/2 MS medium containing a gradient concentration of mannitol (0, 50, 100, and 150 mM) or NaCl (0, 50, 100, and 150 mM), and the germination rate of the treated seeds was calculated after 10 days.
For testing the osmotic or salt stress tolerance of transgenic seedling, the homozygous T3 transgenic and wild-type seeds were geminated in normal 1/2 MS medium for 4 days, and the seedlings were then transferred to medium containing 150 mM mannitol or 150 mM NaCl, respectively. The shoot heights were measured at 10 days after being transferred.
Drought testing at the reproductive stage was conducted in plastic pots. One positive transgenic plant and one wild-type plant each were planted per plastic pot (10 plants each). Drought stress was initiated at the booting stage by discontinuing watering of plants. After 2 weeks of water deprivation, a period that can normally cause serious drought stress (almost all leaves became completely rolled and some of the leaves died) in this environment, the plants were recovered with irrigation for 2 weeks.
To measure the water loss rate under dehydration conditions, leaves of positive transgenic plants and wild-type plants (10 leaves each, three repeats) were cut, exposed to air at room temperature (approximately 25 °C). The leaves were weighed at 0, 15, 30, 45 min, 1, 2, 3, 6, 12, and 24 h after being cut down.
RNA isolation, semi-quantitative PCR and quantitative PCR
Total RNA was extracted using the TRIzol reagent (Invitrogen) according to the manufacturer’s instructions. The DNaseI-treated (Fermentas) RNA was reversed transcribed using ReverTra Ace® RT Kit (Toyobo). Semi-quantitative RT-PCR analysis was performed according to Zou et al. (2009), the rice Actin1 gene (Os03g0718100) was used as the endogenous control. The following gene-specific primes were used. OsHsfB2b: 5′-CAGACAACGAGGAGGGAATGA-3′ and 5′-CGGCTCGGTCGTACTCGTAT-3′. Actin1: 5′-CTTCAACACCCCTGCTATG-3′ and 5′-TCCATCAGGAAGCTCGTAG-3′. Real-time PCR was performed on an ABI 7300 real-time PCR system (Applied Biosystems) using SYBR® Premix Ex Taq™ II (TaKaRa) according to the manufacturer’s instructions. The PCR thermal cycles were as follows: 95 °C for 30 s, followed by 40 cycles at 95 °C for 5 s and 60 °C for 31 s. Real-time PCR of OsHsfB2b was performed using primer pair 5′-GCACGGCAAGGACGAACA-3′ and 5′-CGGCTCGGTCGTACTCGTAT-3′. The rice Actin1 gene was used as the endogenous control with primers 5′-ATCGCCCTGGACTATGAC-3′ and 5′-GAAACGCTCAGCACCAAT-3′. Dates represent three biological replicates and three technical replicates. The relative expression levels were quantified using the \(2^{-\Updelta\Updelta C_{\rm t}}\)method (Livak and Schmittgen 2001). The data were expressed as mean ± standard error.
Measurements of relative electrical conductivity (REC), malondialdehyde (MDA) and proline content
The transgenic plants and wild-type plants at the booting stage were used for drought stress treatment. The flag leaves (5 plants each, three repeats) were harvested at the beginning and at third day of discontinuing watering of plants for the measurements of REC, MDA and proline content. The leaf REC was determined as described previously (Yu et al. 2006). MDA content was measured as the method described previously (Kuk et al. 2003). Free proline content was measured in acidic extracts and quantified spectrophotometrically using the acid-ninhydrin reagent with proline as a standard (Bates et al. 1973).
Results
Expression pattern of OsHsfB2b under different stresses
The previous study revealed that OsHsfB2b (Os08g0546800) was strongly induced in heat-tolerant rice cultivar 996 (indica) under heat stress (Zhang et al. 2012). The induction level of this gene was significantly higher than those of the other members of class B Hsfs, class C Hsfs and some members of class A Hsfs. Therefore, we isolated the full-length cDNA of OsHsfB2b (AK101700) from Nipponbare (japonica) for further functional analysis.
The expression pattern of OsHsfB2b under different abiotic stresses and abscisic acid (ABA) treatment as well as at different tissues was measured by quantitative PCR (qPCR). The expression of OsHsfB2b was induced by salt, polyethylene glycol (PEG), and ABA treatments after 2 h, and kept increasing at 24 h (Fig. 1a). The expression of OsHsfB2b was strongly induced by heat stress especially at very early stage and gradually reduced to lower increased level, but the expression was almost not affected by low-temperature treatment (Fig. 1b). The tissue-specific expression analysis indicated that OsHsfB2b expression was high in sheaths and panicles, medium expression in stems and leaves, but very low in roots and seedlings (Fig. 1c). The strong induction of OsHsfB2b gene by abiotic stresses prompted us to check its promoter sequence (2,000 bp upstream the transcription start site) by searching the promoter sequence against the PLACE database (http://www.dna.affrc.go.jp/PLACE/) and PlantCARE (http://bioinformatics.psb.ugent.be/webtools/plantcare/html/). The results revealed that the promoter contains many putative stress response-related cis-elements, such as ABRE (4 hits), DRE (2 hits), MYB recognition site (MYBRS, 5 hits), MYC recognition site (MYCRS, 6 hits), and HSE (1 hit) (Fig. 1d).
Expression analysis of the OsHsfB2b gene by real-time PCR. a Expression level of OsHsfB2b under salt, PEG and ABA treatments (time course 0, 0.5, 1, 2, 4, 8 and 24 h). b Expression level of OsHsfB2b under cold and heat stresses (time course 0, 0.5, 1, 2, 4, 8 and 24 h). c Expression level of OsHsfB2b in different tissues. d Distribution of major stress-related cis-elements in the promoter region of OsHsfB2b
OsHsfB2b negatively regulates drought tolerance in rice
The strong induction of OsHsfB2b expression by abiotic stress and ABA treatment suggested its possible involvement in stress response in rice. To observe the effect of OsHsfB2b on stress tolerance, OsHsfB2b overexpression (Fig. 2a) and RNAi (Fig. 2b) vectors were constructed and were separately transformed into japonica cultivar Nipponbare. The expression level of OsHsfB2b in transgenic plants was analyzed by reverse transcription (RT)-PCR (Fig. 2c, d). Altogether 22 positive OsHsfB2b overexpression lines and 17 positive RNAi lines were obtained. Under normal condition, there was no significant difference in plant morphology (such as root depth, plant height, and numbers of tillers and spikelets) and yield between the transgenic lines and the wild type (data not shown). These results suggested that overexpression or RNA interference of OsHsfB2b had no detrimental effect on the growth and development of rice. Three independent overexpression lines (OE1, OE10 and OE12) and three independent RNAi lines (Ri1, Ri5 and Ri8) were chosen for stress tolerance test.
Expression analysis of OsHsfB2b in transgenic rice lines and wild type under normal condition. a Diagram of the CaMV35S promoter::OsHsfB2b construct. b Diagram of the CaMV35S promoter::RNAi OsHsfB2b construct. c Expression level of OsHsfB2b in overexpressing OsHsfB2b (OsHsfB2b-OE) lines and wild type under normal condition. Rice Actin1 gene was included as a control for constitutive expression in the assays. d Expression level of OsHsfB2b in OsHsfB2b-RNAi lines and wild type under normal condition. Rice Actin1 gene was included as a control for constitutive expression in the assays
Three-week T2 transgenic and non-transgenic control seedlings were tested for drought tolerance. After 5 days of water-withholding, almost all leaves of OsHsfB2b-overexpressing plants became completely rolled, while the leaves of wild-type plants were partially rolled and only a small portion of the leaves of OsHsfB2b-RNAi plants were slightly rolled. After water-withholding for 7 days, almost all leaves of the transgenic plants and the wild-type plants were completely rolled. One week after rewatering, more than 80 % of the OsHsfB2b-RNAi plants recovered, only about 40 % of the wild-type plants recovered, whereas the survival rate of OsHsfB2b-overexpressing plants fell to about 10 % (Fig. 3a, b). The results indicated that overexpressing OsHsfB2b significantly decreased drought tolerance while RNA interference of this gene improved drought tolerance in transgenic rice at the seedling stage.
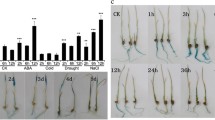
Drought stress tolerance testing of OsHsfB2b-overexpressing and OsHsfB2b-RNAi transgenic rice at the seedling stage. a Phenotype of the OsHsfB2b-overexpressing (OE) plants, the OsHsfB2b-RNAi (Ri) plants and wild-type (WT) plants before drought stress (upper figure) and recovery after drought stress (lower figure) at the seedling stage. b Survival rates of the OsHsfB2b-overexpressing plants (OE1, OE10 and OE12), the OsHsfB2b-RNAi plants (Ri1, Ri5 and Ri8), and wild-type plants recovered for 7 days after drought stress. Error bars indicate SE based on three replicates. c Germination rate of the OsHsfB2b-overexpressing lines, the OsHsfB2b-RNAi lines, and wild-type seeds in 1/2 MS medium with a gradient concentration of mannitol. Error bars indicate SE based on three replicates. d Plant heights of the OsHsfB2b-overexpressing lines, the OsHsfB2b-RNAi lines, and wild type grown in normal or mannitol-containing (150 mM) medium. Plant height was measured after the seedlings growing for 10 days. Values are the mean ± SD (n = 10)
Seed germination experiment of transgenic lines under osmotic stress was performed by adding mannitol in the medium to mimic dehydration stress. After 10 days of seed imbibition, the OsHsfB2b-overexpressing seeds had only about 75 % of the relative germination rate under 100 mM mannitol treatment, while the OsHsfB2b-RNAi seeds and wild type showed more than 90 % of the relative germination rate. Under 150 mM mannitol treatment, the relative germination rates of the OsHsfB2b-RNAi and wild-type seeds were 82 and 67 %, respectively, but that of OsHsfB2b-overexpressing seeds reduced to about 45 % (Fig. 3c). There was no difference between the germination rates of transgenic lines and wild type in normal medium, suggesting that overexpression or RNA interference of OsHsfB2b does not affect seed germination under normal condition. The lower germination rate of overexpression lines and the higher germination rate of RNAi lines than that of wild-type seeds under osmotic stress suggested that OsHsfB2b may have a negative regulatory effect on osmotic stress tolerance in rice. This effect was further confirmed in the postgermination stage rice seedling. In this experiment, transgenic and wild-type rice seedlings geminated in 1/2 MS medium, were transferred to the medium containing 150 mM mannitol for 10 days and the shoot heights were compared. No shoot height difference was observed for the transgenic and wild-type seedlings growing in the medium without mannitol. Under mannitol treatment, the shoot heights of OsHsfB2b-overexpressing seedlings were shorter while that of the OsHsfB2b-RNAi seedlings were longer compared with those of the wild type (Fig. 3d).
Drought stress was also performed at the booting stage by discontinuing watering for 2 weeks and all leaves of transgenic and wild-type plants became completely rolled. After recovering with irrigation for 2 weeks, the overexpression lines failed to be recovered and died, while all the RNAi lines and part of the wild type got recovered (Fig. 4a). The detached leaves of transgenic and wild-type plants were exposed to air to compare the water loss rate in leaves. The results showed that the leaves from the overexpression plants had higher water loss rate, while the leaves from the RNAi plants had lower water loss rate than wild type (Fig. 4b).
Drought stress tolerance testing of OsHsfB2b-overexpressing and OsHsfB2b-RNAi transgenic rice at the booting stage. a Phenotype of the OsHsfB2b-overexpressing (OE) plants, the OsHsfB2b-RNAi (Ri) plants and wild-type (WT) plants before drought stress (upper figure) and recovery after stress (lower figure) at the booting stage. b Water loss rates in leaves detached from the OsHsfB2b-overexpressing plants (OE1, OE10 and OE12), the OsHsfB2b-RNAi plants (Ri1, Ri5 and Ri8), and wild-type plants. Error bars indicate SE based on three replicates. c Relative electrical conductivity (C1/C2) of leaves. C1, conductance of the water solution with leaves in room-temperature water for 30 min; C2, conductance of water solution with leaves in boiled water for 15 min. Values are the mean ± SD (n = 10). d MDA content of the OsHsfB2b-overexpressing plants, the OsHsfB2b-RNAi plants, and wild-type plants under normal and drought stress conditions. Values are the mean ± SD (n = 10). e Proline content of the OsHsfB2b-overexpressing plants, the OsHsfB2b-RNAi plants, and wild-type plants under normal and drought stress conditions. Values are the mean ± SD (n = 10)
Proline enrichment in the stressed plants is a general response to various abiotic stresses and serves as effective adaptation for stress tolerance identification (Akram et al. 2007). In contrast, REC of electrolyte leakage can be used as an indicator of the cell membrane penetrability. MDA, a product of lipid peroxidation, is associated with oxidative degradation of cell membrane lipids and its abundance serves as an indicator of cell membrane damage. The flag leaves of transgenic and wild-type plants at the booting stage were harvested at the beginning and at the third day of discontinuing watering for the measurements of REC, MDA and proline content. There were no significant differences in REC, MDA content and proline content between the transgenic lines and wild type before drought stress. However, after withholding water for 3 days, the REC and the MDA content of the OsHsfB2b-overexpressing lines were significantly higher than that of the wild type, but those of the OsHsfB2b-RNAi lines significantly lower than that of the wild type (Fig. 4c, d). The accumulation of proline in the OsHsfB2b-RNAi lines was higher than that of wild type and the OsHsfB2b-overexpressing lines (Fig. 4e). Lower REC and the MDA content indicate less membrane damage, and proline can improve osmotic adjustment ability of plants. Therefore, the results may partially explain the reason that overexpression of OsHsfB2b resulted in decreased drought tolerance, while RNA interference of this gene improved stress tolerance in rice.
OsHsfB2b negatively regulates salt stress tolerance in rice
To check salt tolerance of the OsHsfB2b transgenic rice, three-week-old plants were irrigated with water containing 200 mM NaCl, and leaf death rate was checked after 7 days of treatment. The results showed that the OsHsfB2b-overexpressing plants had less green leaf area (approximately 63 %) than the wild-type control (approximately 80 %) and the OsHsfB2b-RNAi plants (approximately 88 %; Fig. 5a, b), suggesting that overexpression of OsHsfB2b in rice can also decrease salt tolerance, while RNA interference of this gene can slightly improve salt tolerance of transgenic rice at seedling stage.
Salt stress tolerance testing of OsHsfB2b-overexpressing and OsHsfB2b-RNAi transgenic rice at the seedling stage. a Phenotype of the OsHsfB2b-overexpressing (OE) plants, the OsHsfB2b-RNAi (Ri) plants and wild-type (WT) plants before (upper figure) and recovery after salt stress (lower figure) at the seedling stage. b Percentage of green leaves of the OsHsfB2b-overexpressing plants (OE1, OE10 and OE12), the OsHsfB2b-RNAi plants (Ri1, Ri5 and Ri8), and wild-type plants recovered for 7 days after salt stress. Error bars indicate SE based on three replicates. c Germination rate of the OsHsfB2b-overexpressing lines, the OsHsfB2b-RNAi lines, and wild-type seeds in 1/2 MS medium with a gradient concentration of NaCl. Error bars indicate SE based on three replicates. d Plant heights of the OsHsfB2b-overexpressing lines, the OsHsfB2b-RNAi lines, and wild type grown in normal or NaCl-containing (150 mM) medium. Plant height was measured after the seedlings growing for 10 days. Values are the mean ± SD (n = 10)
The germination ability evaluation of the transgenic lines under salt stress condition revealed that, after 10 days of germination in the medium containing 100 mM NaCl, only about 38 % of the OsHsfB2b-overexpressing seeds were poorly germinated, whereas more than 84 % of wild-type seeds and OsHsfB2b-RNAi seeds were well germinated (Fig. 5c). Under the treatment with 150 mM NaCl, the relative germination rate of the OsHsfB2b-overexpressing seeds fell to 20 %, but that of wild type and OsHsfB2b-RNAi seeds were 66 and 85 %, respectively. In the normal medium, the germination rates of transgenic lines and wild-type had no significant difference. The results indicated that overexpression of OsHsfB2b significantly decreased seed germination, while RNA interference of this gene improved seed germination under salt stress condition.
Transgenic and wild-type rice seedlings geminated in 1/2 MS medium, were transferred to the medium containing 150 mM NaCl for 10 days and the shoot heights were compared. The OsHsfB2b-overexpressing plants had shorter shoots than the OsHsfB2b-RNAi plants and the wild-type plants, and the OsHsfB2b-RNAi plants were taller than the wild-type plants, while no obvious difference was observed for transgenic and wild-type seedlings grown in the medium without NaCl (Fig. 5d). These results indicated that OsHsfB2b negatively regulated salt stress tolerance at postgermination stage.
In addition, three-week-old seedlings of the transgenic rice and wild type with similar vigor were used for high (45 °C) and low (4 °C) temperature treatments. No significant difference was observed between the transgenic lines and the wild type after treatment. After recovery for 1 week, both the transgenic plants and the wild-type plants remained alive and produced new leaves (data not shown). These results showed overexpression or RNA interference of OsHsfB2b gene had no significant effect on temperature tolerance.
Discussion
Plant Hsfs play important roles not only in heat stress, but also in other biotic and abiotic stresses. Class A Hsfs contain the exclusive C-terminal activation domain and are reported as positive regulators in stress tolerance. The recently reported OsHsfC1b played an important role in ABA-mediated salt and osmotic stress tolerance in rice (Schmidt et al. 2012). HsfB1 and HsfB2b had repressive activities and negatively regulated stress tolerance in Arabidopsis and soybean (Czarnecka-Verner et al. 2000, et al. 2004; Ikeda et al. 2011; Ikeda and Ohme-Takagi 2009; Kumar et al. 2009). In this study, we report the characterization of OsHsfB2b, which is a member of class B Hsfs in rice. Our data suggest that OsHsfB2b may be a negative regulator for affecting rice tolerance under drought and salt stresses.
OsHsfB2b expression can be induced by abiotic stress and ABA treatment and have tissue specificity
We investigate the expression pattern of OsHsfB2b under different abiotic stresses and ABA treatment, and the accumulation patterns of OsHsfB2b transcripts in different tissues. OsHsfB2b is strongly up-regulated by heat stress like the other members of Hsfs, which agrees with our previous microarray data very well (Zhang et al. 2012), and is also induced by NaCl, PEG and ABA treatments. The induced expression of OsHsfB2b by heat, high-salinity, drought and oxidative stress and ABA treatment were also reported in previous studies (Chauhan et al. 2011; Mittal et al. 2009). The stress-related cis-acting elements presented in the 2-kb promoter region of OsHsfB2b, including ABRE, DRE, MYBRS, MYCRS, and HSE, may be responsible for the responses under abiotic stress and ABA treatment. ABRE, MYBRS, and MYCRS can be recognized by the AREB/ABF, MYB, and MYC transcription factors, respectively. These cis-elements and the corresponding transcription factors have important roles in ABA signaling and abiotic stress responses (Yamaguchi-Shinozaki and Shinozaki 2005; Yamaguchi-Shinozaki and Shinozaki 2006). Such an enriched presence of various stress responsive cis-acting elements in the OsHsfB2b promoter may suggest an important role of this gene in stress tolerance. We also found the OsHsfB2b exhibited diverse expression in different tissues, although expression could be detected in almost all tissues of rice we checked. The transcript abundances of OsHsfB2b in sheath and spike were significantly higher than those in other tissues. This result is similar to the previous study that higher transcript level of OsHsfB2b was noted at seed developmental stage (Mittal et al. 2009), suggesting OsHsfB2b may be of great importance for seed development.
OsHsfB2b is a negative regulator of drought and salt tolerance
Although OsHsfB2b-overexpressing transgenic plants lead to an abundance of mRNA accumulation and OsHsfB2b-RNAi transgenic plants lead to a reduction of mRNA accumulation, there was no detectable effect on transgenic rice growth and development under normal condition. But overexpression and RNAi transgenic plants showed opposite phenotypes in stress tolerance. The seedlings of the OsHsfB2b-overexpressing lines showed significantly decreased drought and salt tolerance, while the OsHsfB2b-RNAi plants had higher stress tolerance compared with the wild type. This is also true for the osmotic and salt stress tolerance tested at both germination and postgermination stages. Furthermore, drought stress at booting stage revealed that OsHsfB2b-overexpressing lines exhibited significantly poorer drought tolerance than the wild type, but the OsHsfB2b-RNAi lines enhanced drought tolerance.
Leaf REC, MDA and proline content act as three important physiological parameters for evaluating plant stress tolerance. Under drought stress condition, leaf REC and MDA content were higher in the OsHsfB2b-overexpressing lines than those in the wild-type and the OsHsfB2b-RNAi lines, and the accumulation of proline in the OsHsfB2b-overexpressing lines was lower compared with the wild type and the OsHsfB2b-RNAi lines. On the other hand, leaf REC and MDA content were lower in the OsHsfB2b-RNAi lines than those in the wild type, and proline content in the OsHsfB2b-RNAi lines was higher compared with the wild type. Electrolyte leakage is an indirect effect of damage done to plant cell membranes (Bajji et al. 2002), and the higher REC in OsHsfB2b-overexpressing lines indicates that more membrane damage occurred, while the lower REC in OsHsfB2b-RNAi lines means less membrane damage occurred. MDA is one of the end products of lipid peroxidation damage from free radicals (Marnett 1999). The higher level of MDA in the OsHsfB2b-overexpression lines means more lipids damage occured, while the lower level of MDA in the OsHsfB2b-RNAi lines indicated less lipids damage. Proline accumulation in response to environmental stress plays an important role in the osmoregulation and also exhibits many protective effects (Szabados and Savoure 2010). The decreased proline content in OsHsfB2b-overexpressing lines and increased proline content in the OsHsfB2b-RNAi lines indicated negative regulation of OsHsfB2b on proline accumulation under drought stress. The REC, MDA and proline content are related to plants stress tolerance that have also been found in OsHsp17.0, OsHsp23.7 and OsHsfA7 overexpression rice plants under drought and salt stress at seedling stage (Liu et al. 2013; Zou et al. 2012). These findings provided physiological evidences for the negative regulation of OsHsfB2b in stress tolerance.
Recently, HsfB2b has been reported to act as a transcriptional repressor in Arabidopsis. The single mutant hsfb2b and the double mutant hsfb1 hsfb2b showed significantly improved pathogen resistance levels compared with the wild type, and the double mutant hsfb1 hsfb2b enhanced expression of the defensin genes (Kumar et al. 2009). It was also reported that the double mutant hsfb1 hsfb2b exhibited higher thermotolerance and the reason may be that HsfB1 and HsfB2b repress expression of the HS-inducible genes under non-HS conditions, which are important for stress tolerance (Ikeda et al. 2011; Nishizawa et al. 2006). In the similar way, we speculate that OsHsfB2b may act as a transcriptional repressor in rice and repress expression of HS-inducible genes for OsHsfs, some OsHSPs and defensin genes under normal condition. When the transgenic rice plants and the wild-type plants are exposed to drought and salt stresses, the OsHsfB2b-overexpressing plants exhibit lower stress tolerance than the wild type, while OsHsfB2b-RNAi plants exhibit higher stress tolerance, probably due to the different expression level of genes for HSPs and defensin genes repressed by OsHsfB2b. The target genes of OsHsfB2b and the interaction manner of OsHsfB2b and class A Hsfs in rice stress response system will be conducted in a further study.
In our study, there were no detectable changes in temperature stress tolerance for both the OsHsfB2b-overexpressing and the OsHsfB2b-RNAi pants at seedlings stage. This is reminiscent of Arabidopsis. The single mutant hsfb2b showed no obvious effect on the heat stress response (Kumar et al. 2009), but the double mutant hsfb1 hsfb2b enhanced expression of the heat-inducible genes for Hsfs and small HSPs under non-heat-stress conditions and exhibited higher thermotolerance (Ikeda et al. 2011). And in tomato, when the expression of LbHsfB1 was compromised in a transgenic approach, plants showed no obvious effect on the heat stress response (Mishra et al. 2002). These results suggested that HsfB2b may act as a functionally redundant factor to HsfB1 in heat stress response.
BRD may be a critical component of OsHsfB2b acting as transcriptional repressor
In Arabidopsis leaves, HsfB1 and HsfB2b exhibited strong repressive activities in a yeast Gal4 DNA-binding domain (GAL4-DB) fusion transient expression assay (Ikeda and Ohme-Takagi 2009). The core BRD sequence L/VR/KLFGVXM/V/L is conserved in the orthologs of HsfB1 in various plant species and in all five members of Arabidopsis class B Hsfs (Czarnecka-Verner et al. 2004; Ikeda et al. 2011; Ikeda and Ohme-Takagi 2009). When the BRD was mutated, the repressive activity of HsfB1 was abolished (Ikeda et al. 2011). The OsHsfB2b protein contains BRD in the C-terminal amino acid sequence (Fig. 6a). The core BRD sequence P/AR/KLFGVS/CIG, was conserved in HsfB2b orthologs of a variety of plant species (Fig. 6b) and is similar to the core BRD sequence of HsfB1 in Arabidopsis. The core BRD sequence of OsHsfB2b is also conserved in all eight members of rice class B Hsfs (Fig. 6c). We presumed that BRD is necessary for the OsHsfB2b acting as transcriptional repressor just like the HsfB1 in Arabidopsis.
Identification of the repression domain of OsHsfB2b. a Schematic representation of the OsHsfB2b. DNA-binding domain (DBD), oligomerization domain (OD, HR-A/B), nuclear localization signal (NLS), B3 repression domain (BRD) are shown. b An alignment of the core sequence of BRD of HsfB2b orthologs from 12 varieties of plants including rice OsHsfB2b. c An alignment of the core sequence of BRD of rice class B Hsfs
In conclusion, this study showed that OsHsfB2b can be induced by various abiotic stresses and ABA treatment. OsHsfB2b negatively regulates drought and salt stress tolerance in rice, and BRD might be necessary for the repressive activity of OsHsfB2b. Our results also suggest that RNA interference of the OsHsfB2b gene may have a promising utility in improving stress tolerance of crops.
References
Akram NA, Shahbaz M, Ashraf M (2007) Relationship of photosynthetic capacity and proline accumulation with the growth of differently adapted populations of two potential grasses (Cynodon dactylon (L.) Pers. and Cenchrus ciliaris L.) to drought stress. Pak J Bot 39:777–786
Bajji M, Kinet J-M, Lutts S (2002) The use of the electrolyte leakage method for assessing cell membrane stability as a water stress tolerance test in durum wheat. Plant Growth Regul 36:61–70
Baniwal SK, Bharti K, Chan KY, Fauth M, Ganguli A, Kotak S, Mishra SK, Nover L, Port M, Scharf KD, Tripp J, Weber C, Zielinski D, von Koskull-Doring P (2004) Heat stress response in plants: a complex game with chaperones and more than twenty heat stress transcription factors. J Biosci 29:471–487
Banti V, Mafessoni F, Loreti E, Alpi A, Perata P (2010) The heat-inducible transcription factor HsfA2 enhances anoxia tolerance in Arabidopsis. Plant Physiol 152:1471–1483
Bates L, Waldren R, Teare I (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39:205–207
Bharti K, Von Koskull-Doring P, Bharti S, Kumar P, Tintschl-Korbitzer A, Treuter E, Nover L (2004) Tomato heat stress transcription factor HsfB1 represents a novel type of general transcription coactivator with a histone-like motif interacting with the plant CREB binding protein ortholog HAC1. Plant Cell 16:1521–1535
Busch W, Wunderlich M, Schoffl F (2005) Identification of novel heat shock factor-dependent genes and biochemical pathways in Arabidopsis thaliana. Plant J 41:1–14
Charng YY, Liu HC, Liu NY, Chi WT, Wang CN, Chang SH, Wang TT (2007) A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol 143:251–262
Chauhan H, Khurana N, Agarwal P, Khurana P (2011) Heat shock factors in rice (Oryza sativa L.): genome-wide expression analysis during reproductive development and abiotic stress. Mol Genet Genomics 286:171–187
Czarnecka-Verner E, Yuan CX, Scharf KD, Englich G, Gurley WB (2000) Plants contain a novel multi-member class of heat shock factors without transcriptional activator potential. Plant Mol Biol 43:459–471
Czarnecka-Verner E, Pan S, Salem T, Gurley WB (2004) Plant class B HSFs inhibit transcription and exhibit affinity for TFIIB and TBP. Plant Mol Biol 56:57–75
Guo J, Wu J, Ji Q, Wang C, Luo L, Yuan Y, Wang Y, Wang J (2008) Genome-wide analysis of heat shock transcription factor families in rice and Arabidopsis. J Genet Genomics 35:105–118
Hahn A, Bublak D, Schleiff E, Scharf KD (2011) Crosstalk between Hsp90 and Hsp70 chaperones and heat stress transcription factors in tomato. Plant Cell 23:741–755
Hartl FU, Hayer-Hartl M (2002) Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295:1852–1858
Haslbeck M, Buchner J (2002) Chaperone function of sHsps. Prog Mol Subcell Biol 28:37–59
Hirayama T, Shinozaki K (2010) Research on plant abiotic stress responses in the post-genome era: past, present and future. Plant J 61:1041–1052
Ikeda M, Ohme-Takagi M (2009) A novel group of transcriptional repressors in Arabidopsis. Plant Cell Physiol 50:970–975
Ikeda M, Mitsuda N, Ohme-Takagi M (2011) Arabidopsis HsfB1 and HsfB2b act as repressors of the expression of heat-inducible Hsfs but positively regulate the acquired thermotolerance. Plant Physiol 157:1243–1254
Kotak S, Port M, Ganguli A, Bicker F, von Koskull-Doring P (2004) Characterization of C-terminal domains of Arabidopsis heat stress transcription factors (Hsfs) and identification of a new signature combination of plant class A Hsfs with AHA and NES motifs essential for activator function and intracellular localization. Plant J 39:98–112
Kuk YI, Jung HI, Kwon OD, Lee DJ, Burgos NR, Guh JO (2003) Sulfonylurea herbicide-resistant Monochoria vaginalis in Korean rice culture. Pest Manag Sci 59:949–961
Kumar M, Busch W, Birke H, Kemmerling B, Nurnberger T, Schoffl F (2009) Heat shock factors HsfB1 and HsfB2b are involved in the regulation of Pdf1.2 expression and pathogen resistance in Arabidopsis. Mol Plant 2:152–165
Larkindale J, Vierling E (2008) Core genome responses involved in acclimation to high temperature. Plant Physiol 146:748–761
Lata C, Prasad M (2011) Role of DREBs in regulation of abiotic stress responses in plants. J Exp Bot 62:4731–4748
Li H-Y, Chang C-S, Lu L-S, Liu C-A, Chan M-T, Charng Y–Y (2003) Over-expression of Arabidopsis thaliana heat shock factor gene (AtHsfA1b) enhances chilling tolerance in transgenic tomato. Bot Bull Acad Sin 44:129–140
Li C, Chen Q, Gao X, Qi B, Chen N, Xu S, Chen J, Wang X (2005) AtHsfA2 modulates expression of stress responsive genes and enhances tolerance to heat and oxidative stress in Arabidopsis. Sci China C Life Sci 48:540–550
Li M, Doll J, Weckermann K, Oecking C, Berendzen KW, Schoffl F (2010) Detection of in vivo interactions between Arabidopsis class A-HSFs, using a novel BiFC fragment, and identification of novel class B-HSF interacting proteins. Eur J Cell Biol 89:126–132
Liu HC, Liao HT, Charng YY (2011) The role of class A1 heat shock factors (HSFA1s) in response to heat and other stresses in Arabidopsis. Plant Cell Environ 34:738–751
Liu AL, Zou J, Liu CF, Zhou XY, Zhang XW, Luo GY, Chen XB (2013) Over-expression of OsHsfA7 enhanced salt and drought tolerance in transgenic rice. BMB Rep 46:31–36
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408
Lohmann C, Eggers-Schumacher G, Wunderlich M, Schoffl F (2004) Two different heat shock transcription factors regulate immediate early expression of stress genes in Arabidopsis. Mol Genet Genomics 271:11–21
Marnett LJ (1999) Lipid peroxidation-DNA damage by malondialdehyde. Mutat Res 424:83–95
Mishra SK, Tripp J, Winkelhaus S, Tschiersch B, Theres K, Nover L, Scharf KD (2002) In the complex family of heat stress transcription factors, HsfA1 has a unique role as master regulator of thermotolerance in tomato. Genes Dev 16:1555–1567
Mittal D, Chakrabarti S, Sarkar A, Singh A, Grover A (2009) Heat shock factor gene family in rice: genomic organization and transcript expression profiling in response to high temperature, low temperature and oxidative stresses. Plant Physiol Biochem 47:785–795
Nishizawa A, Yabuta Y, Yoshida E, Maruta T, Yoshimura K, Shigeoka S (2006) Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J 48:535–547
Nishizawa-Yokoi A, Nosaka R, Hayashi H, Tainaka H, Maruta T, Tamoi M, Ikeda M, Ohme-Takagi M, Yoshimura K, Yabuta Y (2011) HsfA1d and HsfA1e involved in the transcriptional regulation of HsfA2 function as key regulators for the Hsf signaling network in response to environmental stress. Plant Cell Physiol 52:933–945
Nover L, Bharti K, Doring P, Mishra SK, Ganguli A, Scharf KD (2001) Arabidopsis and the heat stress transcription factor world: how many heat stress transcription factors do we need? Cell Stress Chaperones 6:177–189
Ogawa D, Yamaguchi K, Nishiuchi T (2007) High-level overexpression of the Arabidopsis HsfA2 gene confers not only increased thermotolerance but also salt/osmotic stress tolerance and enhanced callus growth. J Exp Bot 58:3373–3383
Schmidt R, Schippers JH, Welker A, Mieulet D, Guiderdoni E, Mueller-Roeber B (2012) Transcription factor OsHsfC1b regulates salt tolerance and development in Oryza sativa ssp. japonica. AoB Plants 2012:pls011
Schramm F, Ganguli A, Kiehlmann E, Englich G, Walch D, von Koskull-Döring P (2006) The heat stress transcription factor HsfA2 serves as a regulatory amplifier of a subset of genes in the heat stress response in Arabidopsis. Plant Mol Biol 60:759–772
Schramm F, Larkindale J, Kiehlmann E, Ganguli A, Englich G, Vierling E, von Koskull-Doring P (2008) A cascade of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant J 53:264–274
Shim D, Hwang JU, Lee J, Lee S, Choi Y, An G, Martinoia E, Lee Y (2009) Orthologs of the class A4 heat shock transcription factor HsfA4a confer cadmium tolerance in wheat and rice. Plant Cell 21:4031–4043
Singh A, Mittal D, Lavania D, Agarwal M, Mishra RC, Grover A (2012) OsHsfA2c and OsHsfB4b are involved in the transcriptional regulation of cytoplasmic OsClpB (Hsp100) gene in rice (Oryza sativa L.). Cell Stress Chaperones 17:243–254
Szabados L, Savoure A (2010) Proline: a multifunctional amino acid. Trends Plant Sci 15:89–97
Tang N, Zhang H, Li X, Xiao J, Xiong L (2012) Constitutive activation of transcription factor OsbZIP46 improves drought tolerance in rice. Plant Physiol 158:1755–1768
Toki S, Hara N, Ono K, Onodera H, Tagiri A, Oka S, Tanaka H (2006) Early infection of scutellum tissue with Agrobacterium allows high-speed transformation of rice. Plant J 47:969–976
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Yokotani N, Ichikawa T, Kondou Y, Matsui M, Hirochika H, Iwabuchi M, Oda K (2008) Expression of rice heat stress transcription factor OsHsfA2e enhances tolerance to environmental stresses in transgenic Arabidopsis. Planta 227:957–967
Yoshida T, Sakuma Y, Todaka D, Maruyama K, Qin F, Mizoi J, Kidokoro S, Fujita Y, Shinozaki K, Yamaguchi-Shinozaki K (2008) Functional analysis of an Arabidopsis heat-shock transcription factor HsfA3 in the transcriptional cascade downstream of the DREB2A stress-regulatory system. Biochem Biophys Res Commun 368:515–521
Yu X, Peng YH, Zhang MH, Shao YJ, Su WA, Tang ZC (2006) Water relations and an expression analysis of plasma membrane intrinsic proteins in sensitive and tolerant rice during chilling and recovery. Cell Res 16:599–608
Zhang X, Li J, Liu A, Zou J, Zhou X, Xiang J, Rerksiri W, Peng Y, Xiong X, Chen X (2012) Expression profile in rice panicle: insights into heat response mechanism at reproductive stage. PLoS ONE 7:e49652
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Zhu B, Ye C, Lu H, Chen X, Chai G, Chen J, Wang C (2006) Identification and characterization of a novel heat shock transcription factor gene, GmHsfA1, in soybeans (Glycine max). J Plant Res 119:247–256
Zhu X, Thalor SK, Takahashi Y, Berberich T, Kusano T (2012) An inhibitory effect of the sequence-conserved upstream open-reading frame on the translation of the main open-reading frame of HsfB1 transcripts in Arabidopsis. Plant Cell Environ 35:2014–2030
Zou J, Liu A, Chen X, Zhou X, Gao G, Wang W, Zhang X (2009) Expression analysis of nine rice heat shock protein genes under abiotic stresses and ABA treatment. J Plant Physiol 166:851–861
Zou J, Liu C, Liu A, Zou D, Chen X (2012) Overexpression of OsHsp17.0 and OsHsp23.7 enhances drought and salt tolerance in rice. J Plant Physiol 169:628–635
Acknowledgments
This study was supported by Specialized Research Fund for the Doctoral Program of Higher Education of China (20124320110012), Natural Science Foundation of China (30870206), The Education Ministry Program for Innovative Research Team in University (IRT1239) and Construct Program of the Key Discipline in Hunan Province.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by Y. Lu.
Rights and permissions
About this article
Cite this article
Xiang, J., Ran, J., Zou, J. et al. Heat shock factor OsHsfB2b negatively regulates drought and salt tolerance in rice. Plant Cell Rep 32, 1795–1806 (2013). https://doi.org/10.1007/s00299-013-1492-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-013-1492-4