Abstract
Although resveratrol-forming stilbene synthase (STS) genes have been well characterized in many plant species, there are only a few descriptions about STS genes from Polygonum cuspidatum Sieb. et Zucc, an important medicinal crop in Asian countries. To evaluate the biological functions of a Polygonum cuspidatum resveratrol synthase gene (PcRS), the PcRS gene was expressed in Arabidopsis under the control of Cauliflower mosaic virus (CaMV) 35S promoter. Integration and expression of transgene in the plant genome of Arabidopsis was confirmed by Southern blot and Northern blot analyses. Transgenic plants accumulated a new compound in both the leaves and seeds, which was identified as trans-piceid by high-pressure liquid chromatography (HPLC) and electrospray mass spectrometry (HPLC–ESI–MS). Overexpression of PcRS in transgenic Arabidopsis caused restriction of Colletotrichum higginsianum colonization by inhibition of spore production, resulting in enhanced resistance against C. higginsianum. So, the PcRS gene could be deployed in other crop plants to significantly enhance resistance to fungal pathogens and improve the nutritional quality. In addition, altered seed coat pigmentation and significant reduction in anthocyanin levels were observed in transgenic Arabidopsis, while the expression of endogenous chalcone synthase (CHS) gene was not down-regulated. These results suggest that additional STS activities cause a lack of precursors for CHS which leads to the disturbance of the subsequent flavonoid biosynthesis steps in Arabidopsis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Resveratrol (3, 5, 4′-trihydroxystilbene) is known since the 1940s when it was first isolated from the roots of white hellebore and later from Polygonum cuspidatum Sieb. et Zucc (Aggarwal et al. 2004). Polygonum cuspidatum is a medicinal plant used historically in Asia, known for its medicinal properties and traditionally used in the treatment of neuropsychiatric disorders, such as Parkinson’s disease (Chen et al. 2007; Wang et al. 2011). To date, it is known that resveratrol targets include a wide range of enzymes, such as cyclo- and lipooxygenases, protein and lipid kinases, ribonucleotide reductase and DNA polymerases (Pirola and Frojdo 2008). The biological activity of resveratrol has been well documented in recent years through a number of physiological and pharmacological studies which indicate that resveratrol plays an important role in prevention of cancers, heart diseases and neurodegenerative diseases (Anekonda 2006; Athar et al. 2007; Saiko et al. 2008). Moreover, resveratrol has been shown to increase longevity in eukaryotes through activation of sirtuins (Yang et al. 2009). Recently, Xu et al. (2010) reported that the antidepressant-like effect of trans-resveratrol derived from P. cuspidatum might be related to serotonergic and noradrenergic activation.
Biosynthesis of resveratrol is catalyzed by resveratrol synthase (RS; EC 2.3.1.95), also known as stilbene synthase (STS), which utilizes the same substrates as chalcone synthase (CHS), but a different cyclization mechanism is involved. Resveratrol-forming stilbene synthase genes have been well characterized in many plant species, including peanut (Chung et al. 2001, 2003) and grapevine (Melchior and Kindl 1991; Wiese et al. 1994). Subsequently, a root-specific STS cDNA was identified in the medicinal plant Rehmannia glutinosa L., which accumulates resveratrol in roots (Samappito et al. 2003). The first example of a monocot STS gene, SbSTS1, was isolated from sorghum (Yu et al. 2005). More recent analyses of the draft assembly of the grapevine genome has revealed a large array of STS genes, with 43 genes identified and 20 of these being shown to be expressed (Jaillon et al. 2007).
Most of the plant species transformed with various sources of STS genes showed increased tolerance against microbial pathogens (Hain et al. 1993; Coutos-Thévenot et al. 2001; Leckband and Lörz 1998; Stark-Lorenzen et al. 1997; Thomzik et al. 1997; Serazetdinova et al. 2005; Hipskind and Paiva 2000; Lim et al. 2005; Zhu et al. 2004). But no increased resistance against fungal pathogen was observed in transgenic poplar (Giorcelli et al. 2004) and kiwi (Kobayashi et al. 2000). On the other hand, engineering resveratrol might help increasing the nutritional quality of crops (Liu et al. 2006; Schwekendiek et al. 2007; D’Introno et al. 2009). Generally, stilbene accumulation does not trigger any detrimental effect on plant development, growth, fertility or morphology, and no influence of newly synthesized stilbenes has been reported on other secondary metabolite pools (Delaunois et al. 2009). Nevertheless, transformation events may have a negative impact on the general fitness of a plant. Fischer et al. (1997) reported altered flower morphology as well as male sterility in transgenic tobacco. The expression of the gene encoding the STS enzyme in tomato tissues resulted in an altered flavonoid profile (Nicoletti et al. 2007). The transgenic strawberry engineered with the NS-vitis3 gene showed an altered phenylpropanoid metabolism, leading to increased sensitivity to grey mould infection (Hanhineva et al. 2009).
Despite the fact that STS has been well characterized in many plant species and widely introduced in transgenic plants, there are only a few descriptions about resveratrol biosynthesis-related genes in P. cuspidatum. PcPKS2, a three-intron type III polyketide synthase (PKS) gene was isolated from P. cuspidatum and functional analyses revealed recombinant PcPKS2 overexpressed in Escherichia coli to be a benzalacetone synthase (Ma et al. 2009a). Subsequently, another three-intron type III PKS gene (PcPKS1) was isolated from P. cuspidatum, and sequence analyses demonstrated that PcPKS1 is a chalcone synthase (Ma et al. 2009b). Recently, a P. cuspidatum resveratrol synthase gene (PcRS) was isolated (NCBI Accession No. DQ900615). However, the effectiveness of PcRS in improving stilbene accumulation and/or increasing tolerance to pathogenic microorganisms in foreign species has not been studied. In this paper, to assess the biological functions of PcRS, the PcRS gene was expressed in Arabidopsis under the control of CaMV 35S promoter. We showed that overexpression of PcRS in transgenic Arabidopsis resulted in accumulation of trans-piceid and enhanced resistance against Colletotrichum higginsianum. These observations indicate that the PcRS gene can be deployed in other crop plants to significantly enhance resistance to fungal pathogens and improve the nutritional quality.
Materials and methods
Plant material and growth conditions
Wild-type Arabidopsis thaliana (ecotype Columbia) was used for raising transgenics. Arabidopsis plants were grown in a growth chamber (22°C, 16-h light/8-h dark).
Overexpression construct and floral dip transformation of Arabidopsis
The full-length cDNA of PcRS (NCBI Accession No. DQ900615) was PCR amplified with gene-specific primers 5′-CCGGATCCATGGCAGCTTCAACTGAAG-3′ (forward, BamHI site underlined) and 5′-GAAGATCTTTAAATGATGGGCACACTTC-3′ (reverse, Bgl II site underlined). The PCR product was cloned into the BamHI and Bgl II sites of pCAMBIA 1380 vector under the CaMV35S promoter. The 35S:PcRS recombinant plasmid was introduced into Agrobacterium tumefaciens LBA 4404 by electroporation and transformed into Arabidopsis by the floral dip method (Clough and Bent 1998). The dipped plants were grown to maturity, and the seeds harvested. The seeds of transformed plants were selected after being sown on agar plates supplemented with hygromycin (25 μg/ml). Resistant seedlings were transferred to soil for further growth. Transformed plants were identified by PCR using gene-specific primers and their seeds were harvested separately. Independent transgenic lines were identified by Southern blot analysis, and PcRS expression was analyzed by Northern blot analysis. Homozygous T3 progenies were selected by hygromycin resistance and used for further experiments.
Southern blot analysis
About 10 μg of genomic DNA was isolated from leaves of wild-type and transgenic Arabidopsis and digested with EcoRI overnight. The completely digested DNA was size-fractioned in 0.8% agarose gel and transferred to a Hybond N+ membrane (Amersham). Southern blotting analysis was performed according to the DIG DNA Labeling and Detection Kit instruction (Roche). The full-length cDNA of PcRS labelled with the DIG-High Prime kit (Roche Diagnostics) was used as a probe.
Northern blot analysis
Total RNA was isolated from leaves of 4-week-old wild-type and transgenic Arabidopsis. About 20 μg of RNA was size-fractioned in 1% formaldehyde agarose gel and transferred to Hybond N+ membrane (Amersham). The hybridization conditions and labelling of probes were prepared as described above. For individual experiments, probes included the PcRS cDNA and cDNA of the Arabidopsis CHS gene (At5g13939).
HPLC and HPLC–ESI–MS analysis of transgenic Arabidopsis
Transgenic lines with strong expression of PcRS were analysed for the presence of stilbene-related metabolites. T3 plants were grown to maturity, and the seeds harvested. Triplicate samples (0.5 g each) were collected from leaves of 4-week-old plants and ground to a fine powder in liquid nitrogen. Ground tissues were extracted with 5 ml of 80% methanol and supernatants were collected after centrifugation. The methanol fraction was evaporated to dryness under nitrogen. The residue was immediately redissolved in 0.5 ml 80% methanol. Extracts from seeds (20 mg) were prepared as described above. The sample was filtered through a 0.45 μl PVDF filter (Millipore) into HPLC vial. HPLC experiments were performed at room temperature on a HP 1100 series HPLC system (Agilent Technologies, Palo Alto, CA, USA) using a Nucleosil C18 column (5 mm, 4.6 × 250 mm) and H2O-acetonitrile as eluent (acetonitrile: H2O = 25:75, flow rate 0.2 ml/min), and 306 nm of detection wavelength was used. The LC effluent was then introduced into a turbo ion-spray source on a Q/STAR-XL quadrupole/time-of-flight (TOF) hybrid mass spectrometer (Applied Biosystems, Foster City, CA, USA). Negative ESI mass spectra were acquired over the range from m/z 100 to 400. The electrospray voltage was set at −4.5 kV and the source temperature was maintained at 350°C.
Disease resistance evaluation of wild-type and transgenic Arabidopsis
The Colletotrichum higginsianum strain IMI 349063 was kindly provided by Dr. Minhui Li (South China Agriculture University, China). Cultures of the isolate were maintained on potato dextrose agar (PDA) at 25°C in the dark. Four-week-old wild-type and transgenic Arabidopsis were used for inoculation test. For spot inoculations, 10 μl of spore suspension at concentrations of 104–106 spores/ml was used to inoculate detached leaves. Detached leaves were inoculated in petri dishes (with wet filter paper inside) and maintained with a 12-h photoperiod at a light intensity of 30 μE m−2 s−1. The disease symptoms were evaluated by lesion size at 6 days after inoculation. Number of spore on inoculated leaves of Arabidopsis was evaluated by using a previous method (Kim et al. 1999). Each experiment was repeated at three times and each included 15 individual plants. Data are the mean ± standard errors from three independent experiments.
Quantitation of anthocyanin
To determine anthocyanin levels in transgenic Arabidopsis, seeds were sown on Murashige and Skoog (MS) basal medium without a nitrogen source and grown for 5 days under continuous white light (2 mW cm−2). Accumulation of anthocyanin on cotyledons was observed in 4–5 days. The quantitation was done as described before (Shirley et al. 1995). Briefly, 50 seedlings were picked and soaked in 0.5 ml of the extraction solution (100% methanol + 0.5% HCl) overnight at 4°C. The next day, samples were centrifuged briefly and the supernatant was used for spectrophotometric assay. Anthocyanin was quantitated by absorbance at OD530. Each experiment was run in triplicate.
Results
Molecular analysis of transgenic Arabidopsis overexpressing PcRS
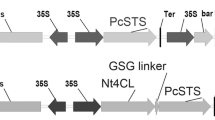
The full-length cDNA of PcRS was expressed in Arabidopsis under the control of CaMV 35S promoter (Fig. 1a). The presence of PcRS in hygromycin-resistant transgenic Arabidopsis was confirmed by PCR using gene-specific primers (data not shown). Seven transgenic lines were further evaluated for the integration of the transgene by Southern blot analysis (Fig. 1b). Between one and seven insertions of PcRS were detected in transgenic lines. The hybridization pattern indicated that they carried independent integration events for the transgenes and some of them (L4, L6 and L15) are single-copy-carrying transgenic lines. The expression of the transgene in transgenic plants was analysed by Northern blot (Fig. 1c). No hybridization signal was observed in the wild-type plants. The single-copy-carrying transgenic lines (L4, L6 and L15) showed relatively high-level expression of PcRS. The expression level of the transgene was very low or absent in the high-copy intergrated transgenic lines (L3,L5 and L8). The homozygous T3 progenies of three transgenic lines (L4, L6 and L15) were selected for further studies.
Identification of transgenic Arabidopsis overexpressing PcRS. a Schematic representation of the construct used to overexpress PcRS in Arabidopsis. b Southern blot analysis to confirm the presence and copy number of transgene. DNA samples were digested by EcoRI and blots were probed by a PCR amplified fragment of PcRS. Lines L4, L6 and L15 represent single copy integration. c Northern analysis of transgenic Arabidopsis using the PCR amplified PcRS as a probe. Lines L4, L6, L7 and L15 represent relatively high expression of PcRS. rRNA was used as a loading control
High levels of piceid were detected in transgenic Arabidopsis overexpressing PcRS
For analysis of plant metabolites, extracts were prepared from leaves of 4-week-old plants. HPLC chromatograms of the extracts from transgenic plants had an obvious peak (Fig. 2a) which was not detected in the extracts of the wild-type plants (Fig. 2b). The retention time (7.5 min) of the peak was the same as the trans-piceid standard (Fig. 2c), indicating the production of trans-piceid in transgenic plants. HPLC–ESI–MS was performed to further confirm the new compound. We confirmed its identity as trans-piceid from the spectrum (Fig. 3) of the [M–H]− ion (m/z 389) with a prominent [M–H–C6H10O5]− product ion at m/z 227 (deprotonated resveratrol). As shown in Table 1, trans-piceid accumulated in the leaves of T3 transgenic lines ranged from 93.31 to 183.73 μg/g fresh weight (FW). In addition, trans-piceid was also detected in the extracts of transgenic seeds, although to a much lesser extent. The amount of trans-piceid reached an average of 26.77 μg/g FW in seeds (mean value of lines L4, L6 and L15).
Enhanced resistance of transgenic Arabidopsis overexpressing PcRS against Colletotrichum higginsianum
We investigated whether the accumulation of trans-piceid in transgenic Arabidopsis would improve fungal resistance. Three independent homozygous T3 lines (L4, L6 and L15) that consistently accumulated the high levels of trans-piceid and the wild-type plants were tested for resistance to C. higginsianum. The detached leaves of 4-week-old Arabidopsis plants were used for spot inoculations. Fungal colonization was arrested around the inoculation sites in the transgenic plants; by contrast, wild-type plants showed the typical susceptible symptoms in which spreading lesions were formed (Fig. 4a). At 6 days after inoculation, the lesion diameter of the transgenic plants was 2.5- to threefold smaller than that of the wild-type plants (Fig. 4b). To verify if the restricted symptom of transgenic plants is caused by inhibited fungal colonization, spore production on infected plants was measured. At 6 days after inoculation, spore production of the transgenic plants was over fivefold less than the wild-type plants (Fig. 4c). This observation was consistent with the smaller lesion size of the resistance response.
Disease evaluation of the PcRS transgenic T3 Arabidopsis plants. a Comparison of lesion formation. Four-week-old wild-type plants (WT) and PcRS transgenic T3 plants were inoculated with Colletotrichum higginsianum. Pictures were made 6 days post infection. b Comparison of lesion diameter. Data were recorded 6 days post infection and are the mean ± SE of three replications, with each replication comprising three subreplications. c Comparison of spore production. Data were taken 6 days post infection and are the mean ± SE of three replications, with each replication comprising three subreplications
Reduced pigmentation in transgenic Arabidopsis overexpressing PcRS
To study the effect of overexpression of PcRS on growth and development of transgenic Arabidopsis, the homozygous T3 progenies of three transgenic lines (L4, L6 and L15) were selected for phenotypic studies. The mature seed coats of wild-type Arabidopsis were brown, resulting from the presence of tannins in the endothelium, whereas the seeds of transgenic Arabidopsis appeared pale fawn (Fig. 5a). To test if the overexpression of PcRS influences accumulation of anthocyanin, both wild-type and transgenic Arabidopsis plants were grown on medium devoid of nitrogen sources and anthocyanin content was quantitated using a spectrophotometer. Transgenic seedlings showed reduced anthocyanin pigmentation in contrast to the purple cotyledons of the wild-type plants (Fig. 5a). The transgenic lines had a remarkable decrease in anthocyanin accumulation, resulting in OD530 values approximately 50% of those in the wild-type plants (Fig. 5b). Northern analysis showed that expression of endogenous Arabidopsis chalcone synthase gene (AtCHS) in the transgenic lines was not decreased (Fig. 5c). No significant phenotypic difference was observed in adult plants overexpressing PcRS.
Reduced pigmentation in transgenic Arabidopsis. a Pigment accumulation of 5-day-old Arabidopsis seedlings grown on MS plates without nitrogen sources and of seed coats. b Anthocyanin content in 5-day-old seedlings was quantitated by a spectrophotometer. Data are the mean ± SE of three replications, with each replication comprising three subreplications. c Endogenous AtCHS and ectopic PcRS expression in wild-type and transgenic plants. Blots were probed by the PCR amplified fragment of PcRS and AtCHS, respectively
Discussion
Our results demonstrated that PcRS was integrated into the Arabidopsis genome in varying copy numbers and expressed to varying degrees. There are conflicting reports about the relationship between transgene copy number and expression. They have been shown to be negatively correlated (Tang et al. 2007), not correlated (Kobayashi et al. 2000; Zhu et al. 2004) or positively correlated (Hobbs et al. 1993). In white poplar, a high transgene copy number correlating with low levels of transcripts suggest that homology-dependent gene silencing should have occurred in these transgenic lines (Giorcelli et al. 2004). This is in agreement with the findings of the present study that the expression level of PcRS was very low or absent in the high-copy intergrated transgenic lines (L3, L5 and L8).
In a number of cases, transformation of plants with STS has led to the production of resveratrol or piceid. But the anticipated resveratrol or its derivatives were not detected in transgenic strawberry (Hanhineva et al. 2009). The products of PcRS found in the transgenic Arabidopsis were identified piceid instead of resveratrol. This is consistent with the reports from poplar (Giorcelli et al. 2004), kiwi (Kobayashi et al. 2000), and Brassica napus seeds (Hüsken et al. 2005). Furthermore, we found that only trans-piceid was detected in 35S:PcRS transgenic Arabidopsis. By contrast, Yu et al. (2006) reported that cis-piceid was accumulated as the major stilbene in the 35S:SbSTS1 Arabidopsis plants and inferred that the cis-isoform were likely to result from endogenous isomerase activities in Arabidopsis. The quantitative HPLC analysis of leaves from selected transgenic lines allowed us to demonstrate a high PcRS expression, with levels of trans-piceid in the range 93.31–183.73 μg/g FW. The stilbene contents were higher than those in transgenic alfalfa (Hipskind and Paiva 2000), pea (Richter et al. 2006) and lettuce (Liu et al. 2006), but lower than those in transgenic poplar (Giorcelli et al. 2004) and hop (Schwekendiek et al. 2007). In addition, trans-piceid was also detected in the seeds of transgenic Arabidopsis. The amount of trans-piceid produced in transgenic Arabidopsis seeds, 36.30 μg/g FW, was much lower than that produced in transgenic Brassica napus (Hüsken et al. 2005), the maximum value of piceid being around 616 μg/g FW in seeds of a homozygous T3-plant. Though piceid contents in our STS-overexpressing Arabidopsis seeds were much lower than those in STS-overexpressing Brassica napus seeds, altered seed coats pigmentation was observed in our report, which is not present in the transgenic Brassica napus (Hüsken et al. 2005).
The accumulation of stilbene derivatives has been reported to enhance protection in plants against various fungal pathogens; however, no significant increase in resistance was reported to Colletotrichum fungi. The hemibiotrophic fungus Colletotrichum higginsianum is pathogenic to Arabidopsis (Narusaka et al. 2004; O’Connell et al. 2004). Colletotrichum higginsianum was reported to cause typical anthracnose lesions on the leaves, petioles and stems. The anthracnose disease caused on the model plant Arabidopsis by C. higginsianum provides a valuable new system for analysis of host responses in a hemibiotrophic disease interaction. Using the new system, Narusaka et al. (2004) provided evidence that Glycerol-3-phosphate (G3P) levels in plants were associated with defence to C. higginsianum. In our experiments, 35S:PcRS transgenic plants accumulating piceid showed a significant reduction in the size of lesions following infection with the pathogen C. higginsianum. Transgenic plants over-expressing PcRS had restriction of C. higginsianum colonization by inhibition of spore production. Taken together, these results suggest that PcRS products in the transgenic plants may inhibit fungal mycelia formation of C. higginsianum, resulting in enhanced resistance against C. higginsianum.
A clear relationship between stilbenes and disease resistance has been demonstrated in most of transgenic plants except kiwifruit (Kobayashi et al. 2000) and poplar (Giorcelli et al. 2004). Based on the fact that piceid was not synthesized in wild-type Arabidopsis, we refer the reduction of susceptibility in transgenic Arabidopsis lines to the presence of stilbenes. Disease symptoms in transgenic Arabidopsis were reduced but not completely inhibited as described for a horizontal type of resistance. This type of disease resistance was characteristic for other transgenic plant species expressing stilbene synthase genes (Hain et al. 1993; Thomzik et al. 1997; Hipskind and Paiva 2000; Zhu et al. 2004; Serazetdinova et al. 2005). Transgenic wheat plants showed a different level of resistance against two different pathogens (Serazetdinova et al. 2005). A pathogen-dependent resistance response was also reported for transgenic tomato (Thomzik et al. 1997). Therefore, further experiments in planta, and especially with other fungal and bacterial pathogens, are necessary in larger-scale experiments to determine whether the PcRS-overexpressing lines have broad-spectrum resistance to pathogen.
Induced defence responses are regulated through a network of signal transduction pathways in which the small molecules, salicylic acid (SA), jasmonic acid (JA) and ethylene (ET), act as secondary messengers. Although signalling pathways regulating stilbene biosynthesis in plant cells are not fully understood. The results from previous study clearly demonstrated that the stress hormones (such as ET, SA and JA) were involved in the resveratrol synthase gene expression in grapevine (Melchior and Kindl 1991; Wiese et al. 1994) and peanut (Chung et al. 2001, 2003). In Arabidopsis, SA controls the expression of PR1, PR2 (β-1.3-glucanase) and PR5 (thaumatin-like) genes and is required for the resistance to certain biotrophic pathogens as well as for mediating systemic acquired resistance (Thomma et al. 1998). In contrast, induction of distinct defense genes, including the defensin PDF1.2, the PR3 (basic chitinase) and PR4 (chitinase type I and II) genes together with the resistance to necrotrophic pathogens are abolished in mutant plants compromised in JA- or ET-dependent pathways (Glazebrook 2005). It is possible that enhanced resistance to pathogen attack is a result of the constitutive expression of PcRS in transgenic Arabidopsis plants. But the possibility cannot be completely excluded that overexpression of PcRS in Arabidopsis may result in activation of a signal transduction pathway that may be directly or indirectly related to pathogen disease resistance. To ascertain whether PcRS overexpression affects the expression levels of a series of defense genes (such as PR1, PR2 and PDF1.2), expression analysis of these defense genes by RT–PCR or Northern blot are planned.
Generally, stilbene accumulation does not affect plant development, growth, fertility, morphology and other secondary metabolite pools (Giovinazzo et al. 2005; Schwekendiek et al. 2007; Delaunois et al. 2009). However, very high constitutive STS expression also has a dramatic influence on flower colour and pollen development. These undesirable side effects may be due to a competition between CHS and STS for their common substrates, leading to reduced levels of flavonoids, which may give rise to male sterility in some species (Van der Meer et al. 1992; Ylstra et al. 1992). Other reports suggest that the reason for diminished flower pigmentation and male sterility is competition for the substrates rather than suppression of endogenous CHS expression by STS sequences (Fettig and Hess 1999; Fischer et al. 1997; Jeandet et al. 2002). More evidence for the hypothesis that the observed pleiotropic effects in STS-overexpressing plants are due to reduced flavonoid levels comes from partial chemical complementation of the male sterility phenotype by exogenous addition of flavonols as well as flavonoid precursors from the phenylpropanoid pathway (Fischer et al. 1997). In this paper, we observed that transgenic plants had seeds with a pale fawn colour and accumulated less anthocyanin in cotyledons in contrast to the wild-type plants. These results indicate that both tannin and anthocyanin accumulation are decreased due to ectopic expression of PcRS, though this reduction was not as dramatic as in the Arabidopsis tt4 mutants which has yellow seeds and cotyledons. Moreover, we found that the expression of endogenous AtCHS in PcRS-overexpressing Arabidopsis was not down-regulated. These results suggest that the significant reduction in flavonoid levels in transgenic plants may be due to competition for the substrates, instead of suppression of endogenous AtCHS expression. None of the previous studies on transgenic plants modified by introducing STS demonstrates down-regulation of endogenous genes. Recently, one paper described a significant decrease in flavonols in transgenic strawberry modified with STS gene (NS-Vitis3) from Vitis riparia (Hanhineva et al. 2009). But the endogenous CHS mRNA level was dramatically diminished in the fully expanded leaves of the 35S:NS-Vitis3 line. One explanation for the changes in the CHS expression levels may lay in the regulatory control of the phenylpropanoid pathway; the introduction of the transgene may have resulted in changes in the control of gene expression, shown as diminished CHS mRNA levels (Hanhineva et al. 2009).
Remarkably, significant reduction in anthocyanin levels observed in PcRS-overexpressing Arabidopsis was similar to that previously observed in transgenic Arabidopsis expressing the mutated CHS genes (Hanumappa et al. 2007). Likewise, altered flower colour observed in STS-overexpressing tobacco (Fischer et al. 1997) is in line with the observation of (Nakatsuka et al. 2007) that the pigmentation of CHS RNAi tobacco flowers was suppressed in comparison with that of the wild type. Similarly, the occurrence of parthenocarpic fruits observed in STS-overexpressing tomato (Giovinazzo et al. 2005; Schijlen et al. 2006) has also been reported in CHS RNAi tomato (Schijlen et al. 2007), indicating that overexpressing of STS and silencing of CHS may cause a similar phenotype.
The present investigation shows that the expression of PcRS in transgenic Arabidopsis leads to accumulation of trans-piceid and also confers enhanced resistance against C. higginsianum. Moreover, additional PcRS activities cause a lack of precursors for CHS which leads to the disturbance of the subsequent flavonoid biosynthesis steps in Arabidopsis.
References
Aggarwal BB, Bhardwaj A, Aggarwal RS, Seeram NP, Shishodia S, Takada Y (2004) Role of resveratrol in prevention and therapy of cancer: preclinical and clinical studies. Anticancer Res 24:2783–2840
Anekonda TS (2006) Resveratrol—a boon for treating Alzheimer’s disease? Brain Res Rev 52:316–326
Athar M, Back JH, Tang X, Kim KH, Kopelovich L, Bickers DR, Kim AL (2007) Resveratrol: a review of preclinical studies for human cancer protection. Toxicol Appl Pharmacol 224:274–283
Chen LW, Wang YQ, Wei LC, Shi M, Chan YS (2007) Chinese herbs and herbal extracts for neuroprotection of dopaminergic neurons and potential therapeutic treatment of Parkinson’s disease. CNS Neurol Disord Drug Targets 6:273–281
Chung IM, Park MR, Rehman S, Yun SJ (2001) Tissue specific and inducible expression of resveratrol synthase gene in peanut plants. Mol Cells 12:353–359
Chung IM, Park MR, Chun JC, Yun SJ (2003) Resveratrol accumulation and resveratrol synthase gene expression in response to abiotic stresses and hormones in peanut plants. Plant Sci 164:103–109
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Coutos-Thévenot P, Poinssot B, Bonomelli A, Yean H, Breda C, Buffard D, Esnault R, Hain R, Boulay M (2001) In vitro tolerance to Botrytis cinerea of grapevine 41B rootstock in transgenic plants expressing the stilbene synthase Vst 1 gene under the control of a pathogen-inducible PR 10 promoter. J Exp Bot 52:901–910
D’Introno A, Paradiso A, Scoditti E, D’Amico L, De Paolis A, Carluccio MA, Nicoletti I, DeGara L, Santino A, Giovinazzo G (2009) Antioxidant and anti-inflammatory properties of tomato fruits synthesizing different amounts of stilbenes. Plant Biotech J 7:422–429
Delaunois B, Cordelier S, Conreux A, Clement C, Jeandet P (2009) Molecular engineering of resveratrol in plants. Plant Biotechnol J 7:2–12
Fettig S, Hess D (1999) Expression of a chimeric stilbene synthase gene in transgenic wheat lines. Transgenic Res 8:179–189
Fischer R, Budde I, Hain R (1997) Stilbene synthase gene expression causes changes in flower colour and male sterility in tobacco. Plant J 11:489–498
Giorcelli A, Sparvoli F, Mattivi F, Tava A, Balestrazzi A, Vrhovsek U, Calligari P, Bollini R, Confalonieri M (2004) Expression of the stilbene synthase (StSy) gene from grapevine in transgenic white poplar results in high accumulation of the antioxidant resveratrol glucosides. Transgenic Res 13:203–214
Giovinazzo G, D’Amico L, Paradiso A, Bollino R, Sparvoli F, DeGara L (2005) Antioxidant metabolite profiles in tomato fruit constitutively expressing the grapevine stilbene synthase gene. Plant Biotech J 3:57–69
Glazebrook J (2005) Contrasting mechanisms of defense against biotrophic and necrotrophic pathogens. Annu Rev Phytopathol 43:205–227
Hain R, Reif HJ, Krause E, Langebartels R, Kindl H, Vornam B, Wiese W, Schmelzer E, Schreier P, Stöcker R, Stenzel K (1993) Disease resistance results from foreign phytoalexin expression in a novel plant. Nature 361:153–156
Hanhineva K, Kokko H, Siljanen H, Rogachev I, Aharoni A, Karenlampi SO (2009) Stilbene synthase gene transfer caused alterations in the phenylpropanoid metabolism of transgenic strawberry (Fragaria × ananassa). J Exp Bot 60:2093–2106
Hanumappa M, Choi G, Ryu S, Choi G (2007) Modulation of flower colour by rationally designed dominant-negative chalcone synthase. J Exp Bot 58:2471–2478
Hipskind JD, Paiva NL (2000) Constitutive accumulation of a resveratrol glucoside in transgenic alfalfa increases resistance to Phoma medicaginis. Mol Plant Microbe Interact 13:551–562
Hobbs SL, Warkentin TD, DeLong CM (1993) Transgene copy number can be positively or negatively associated with transgene expression. Plant Mol Biol 21:17–26
Hüsken A, Baumert A, Milkowski C, Becker HC, Strack D, Mollers C (2005) Resveratrol glucoside (Piceid) synthesis in seeds of transgenic oilseed rape (Brassica napus L.). Theor Appl Genet 111:1553–1562
Jaillon O, Aury JM, Noel B, Policriti A, Clepet C, Casagrande A, Choisne N, Aubourg S, Vitulo N, Jubin C, Vezzi A, Legeai F, Hugueney P, Dasilva C, Horner D, Mica E, Jublot D, Poulain J, Bruyere C, Billault A, Segurens B, Gouyvenoux M, Ugarte E, Cattonaro F, Anthouard V, Vico V, Del FC, Alaux M, Di Gaspero G, Dumas V, Felice N, Paillard S, Juman I, Moroldo M, Scalabrin S, Canaguier A, Le Clainche I, Malacrida G, Durand E, Pesole G, Laucou V, Chatelet P, Merdinoglu D, Delledonne M, Pezzotti M, Lecharny A, Scarpelli C, Artiguenave F, Pe ME, Valle G, Morgante M, Caboche M, Adam-Blondon AF, Weissenbach J, Quetier F, Wincker P (2007) The grapevine genome sequence suggests ancestral hexaploidization in major angiosperm phyla. Nature 449:463–467
Jeandet P, Douillet-Breuil AC, Bessis R, Debord S, Sbaghi M, Adrian M (2002) Phytoalexins from the Vitaceae: biosynthesis, phytoalexins gene expression in transgenic plants, antifungal activity and metabolism. J Agric Food Chem 50:2731–2741
Kim KD, Oh BJ, Yang JM (1999) Differential interactions of a Colletotrichum gloeosporioides isolate with green and red pepper fruits. Phytoparasitica 27:1–10
Kobayashi S, Ding CK, Nakamura Y, Nakajima I, Matsumoto R (2000) Kiwifruits (Actinidia deliciosa) transformed with a Vitis stilbene synthase gene produce piceid (resveratrol-glucoside). Plant Cell Rep 19:904–910
Leckband G, Lörz H (1998) Transformation and expression of a stilbene synthase gene of Vitis vinifera L. in barley and wheat for increased fungal resistance. Theor Appl Genet 96:1004–1012
Lim JD, Yun SJ, Chung IM, Yu CY (2005) Resveratrol synthase transgene expression and accumulation of resveratrol glycoside in Rehmannia glutinosa. Mol Breed 16:219–233
Liu S, Hu Y, Wang X, Zhong J, Lin Z (2006) High content of resveratrol in lettuce transformed with a stilbene synthase gene of Parthenocissus henryana. J Agric Food Chem 54:8082–8085
Ma LQ, Guo YW, Gao DY, Ma DM, Wang YN, Li GF, Liu BY, Wang H, Ye HC (2009a) Identification of a Polygonum cuspidatum three-intron gene encoding a type III polyketide synthase producing both naringenin and p-hydroxybenzalacetone. Planta 229:1077–1086
Ma LQ, Pang XB, Shen HY, Pu GB, Wang HH, Lei CY, Wang H, Li GF, Liu BY, Ye HC (2009b) A novel type III polyketide synthase encoded by a three-intron gene from Polygonum cuspidatum. Planta 229:457–469
Melchior F, Kindl H (1991) Coordinate- and elicitor-dependent expression of stilbene synthase and phenylalanine ammonialyase genes in Vitis cv. Optima. Arch Biochem Biophys 288:552–557
Nakatsuka T, Pitaksutheepong C, Yamamura S, Nishihara M (2007) Induction of differential flower pigmentation patterns by RNAi using promoters with distinct tissue-specific activity. Plant Biotechnol Rep 1:251–257
Narusaka Y, Narusaka M, Park P, Kubo Y, Hirayama T, Seki M, Shiraishi T, Ishida J, Nakashima M, Enju A, Sakurai T, Satou M, Kobayashi M, Shinozaki K (2004) RCH1, a locus in Arabidopsis that confers resistance to the hemibiotrophic fungal pathogen Colletotrichum higginsianum. MPMI 17:749–762
Nicoletti I, De Rossi A, Giovinazzo G, Corradini D (2007) Identification and quantification of stilbenes in fruits of transgenic tomato plants (Lycopersicon esculentum Mill.) by reversed phase HPLC with photodiode array and mass spectrometry detection. J Agric Food Chem 55:3304–3311
O’Connell R, Herbert C, Sreenivasaprasad S, Khatib M, Esquerré-Tugayé MT, Dumas B (2004) A novel Arabidopsis-Colletotrichum pathosystem for the molecular dissection of plant-fungal interactions. MPMI 17:272–282
Pirola L, Frojdo S (2008) Resveratrol: one molecule, many targets. IUBMB Life 60:323–332
Richter A, de Kathen A, de Lorenzo G, Briviba K, Hain R, Ramsay G, Jacobsen HJ, Kiesecker H (2006) Transgenic peas (Pisum sativum) expressing polygalacturonase inhibiting protein from raspberry (Rubus idaeus) and stilbene synthase from grape (Vitis vinifera). Plant Cell Rep 25:1166–1173
Saiko P, Szakmary A, Jaeger W, Szekeres T (2008) Resveratrol and its analogs: defense against cancer, coronary disease and neurodegenerative maladies or just a fad? Mutat Res 658:68–94
Samappito S, Page JE, Schmidt J, De-Eknamkul W, Kutchan TM (2003) Aromatic and pyrone polyketides synthesized by a stilbene synthase from Rheum tataricum. Phytochemistry 62:313–323
Schijlen EGWM, de Vos CHR, Jonker H, van den Broeck H, Molthoff J, van Tunen AJ, Martens S, Bovy A (2006) Pathway engineering for healthy phytochemicals leading to the production of novel flavonoids in tomato fruit. Plant Biotech J 4:433–444
Schijlen EGWM, de Vos CHR, Martens S, Jonker HH, Rosin FM, Molthoff JW, Tikunov YM, Angenent GC, van Tunen AJ, Bovy AG (2007) RNAi silencing of chalcone synthase, the first step in the flavonoid biosynthesis pathway, leads to parthenocarpic tomato fruits. Plant Physiol 144:1520–1530
Schwekendiek A, Spring O, Heyerick A, Pickel B, Pitsch NT, Peschke F, de Keukeleire D, Weber G (2007) Constitutive expression of a grapevine stilbene synthase gene in transgenic hop (Humulus lupulus L.) yields resveratrol and its derivatives in substantial quantities. J Agric Food Chem 55:7002–7009
Serazetdinova L, Oldach K, Lörz H (2005) Expression of transgenic stilbene synthases in wheat causes the accumulation of unknown stilbene derivatives with antifungal activity. J Plant Physiol 162:985–1002
Shirley BW, Kubasek WL, Storz G, Bruggemann E, Koornneef M, Ausubel FM, Goodman HM (1995) Analysis of Arabidopsis mutants deficient in flavonoid biosynthesis. Plant J 8:659–671
Stark-Lorenzen P, Nelke B, Hänbler G, Mühlbach HP, Thomzik JE (1997) Transfer of a grapevine stilbene synthase gene to rice (Oryza sativa L.). Plant Cell Rep 16:668–673
Tang W, Newton RJ, Weidner DA (2007) Genetic transformation and gene silencing mediated by multiple copies of a transgene in eastern white pine. J Exp Bot 58:545–554
Thomma BPHJ, Eggermont K, Penninckx IAMA, Mauch-Mani B, Vogelsang R, Cammue BPA, Broekaert WF (1998) Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc Natl Acad Sci USA 95:15107–15111
Thomzik JE, Stenzel K, Stöcker R, Schreier PH, Hain R, Stahl DJ (1997) Synthesis of a grapevine phytoalexin in transgenic tomatoes (Lycopersicon esculentum Mill.) conditions resistance against Phytophthora infestans. Physiol Mol Plant Pathol 51:265–278
Van der Meer IM, Stam ME, Van Tunen AJ, Mol JNM, Stuitje AR (1992) Antisense inhibition of flavonoid biosynthesis in petunia anthers results in male sterility. Plant Cell 4:253–262
Wang YC, Xu HL, Fu Q, Ma R, Xiang JZ (2011) Protective effect of resveratrol derived from Polygonum cuspidatum and its liposomal form on nigral cells in Parkinsonian rats. J Neurol Sci 304:29–34
Wiese W, Vornam B, Krause E, Kindl H (1994) Structural organization and differential expression of three stilbene synthase genes located on a 13 kb grapevine DNA fragment. Plant Mol Biol 26:667–677
Xu Y, Wang ZC, You WT, Zhang XH, Li S, Barish PA, Vernon MM, Du X, Li GW, Pan JC, Ogle WO (2010) Antidepressant-like effect of trans-resveratrol: involvement of serotonin and noradrenaline system. Euro Neuropsychopharm 20:405–413
Yang J, Kong X, Martins-Santos ME, Aleman G, Chaco E, Liu GE, Wu SY, Samols D, Hakaimi P, Chiang CM, Hanson RW (2009) The activation of Sirt1 by resveratrol represses transcription of the gene for the cytosolic form of phosphoenolpyruvate carboxykinase (GTP) by deacetylating HNF4α. J Biol Chem 284:27042–27053
Ylstra B, Touraev A, Benito Moreno RM, Stöger E, van Tunen AJ, Vicente O, Mol JNM, Heberle-Bors E (1992) Flavonols stimulate development, germination and tube growth of tobacco pollen. Plant Physiol 100:902–907
Yu CK, Springob K, Schmidt J, Nicholson RL, Chu IK, Yip WK, Lo C (2005) A stilbene synthase gene (SbSTS1) is involved in host and nonhost defense responses in Sorghum. Plant Physiol 138:393–401
Yu CK, Lam CN, Springob K, Schmidt J, Chu IK, Lo C (2006) Constitutive accumulation of cis-piceid in transgenic Arabidopsis overexpressing a Sorghum stilbene synthase gene. Plant Cell Physiol 47:1017–1021
Zhu YJ, Agbayani R, Jackson MC, Tang CS, Moore PH (2004) Expression of the grapevine stilbene synthase gene VST1 in papaya provides increased resistance against diseases caused by Phytophthora palmivora. Planta 220:241–250
Acknowledgments
We are grateful to Dr. Minhui Li (South China Agriculture University, China) for providing Colletotrichum higginsianum strain. We thank Dr. Yuchun Luo (University of Colorado, USA) for critical reading of the manuscript. This work was funded by grants from Guangdong Provincial Natural Science Foundation (10151001002000012) and Technologies Program of Guangdong Province Foundation (2010B060200009).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. R. Liu.
Rights and permissions
About this article
Cite this article
Liu, Z., Zhuang, C., Sheng, S. et al. Overexpression of a resveratrol synthase gene (PcRS) from Polygonum cuspidatum in transgenic Arabidopsis causes the accumulation of trans-piceid with antifungal activity. Plant Cell Rep 30, 2027–2036 (2011). https://doi.org/10.1007/s00299-011-1110-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-011-1110-2