Abstract
Objectives
Familial Mediterranean Fever (FMF) is the most common hereditary autoinflammatory disorder characterized by recurrent fever and serositis episodes. Identification of low penetrant or heterozygous MEFV mutations in clinically diagnosed FMF patients did raise a concern on whether epigenetic or environmental factors play an additional role in FMF pathogenesis. We aimed to investigate the expression profile of apoptosis-related miRNAs in FMF and their influence on clinical manifestations in the present study.
Method
191 pediatric FMF patients and 31 healthy children included in the study. Expressions of 33 apoptosis-related, circulating cell-free miRNAs were evaluated by a quantitative polymerase chain reaction, statistically calculated within ΔΔCt values and fold changes were evaluated by Welch T test, in which p < 0.05 were considered to be significant.
Results
Nineteen miRNAs, including let-7a-5p, let-7c, let-7 g-5p, miR-15b-5p, miR-16-5p, miR-17-5p, miR-23a-3p, miR-24-3p, miR-25-3p, miR-26a-5p, miR-26b-5p, miR-27a-3p, miR-29c-3p, miR-30a-5p, miR-30d-5p, miR-30e-5p, miR-106b-5p, miR-146a-5p, and miR-195-5p, were found down-regulated; miR-15a-5p, miR-29b-3p, miR-181a-5p, miR-181b-5p, miR-181c-5p, miR-214-3p, and miR-365a-3p were up-regulated in FMF patients. In detail, these miRNAs were similar among FMF patients in terms of genotype, colchicine response, and having an inflammatory attack during analysis.
Conclusion
We found that 26 apoptosis-related circulating miRNAs were deregulated in children with FMF. Thus, we speculate that these miRNAs have a role in FMF pathogenesis via apoptotic mechanisms.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Familial Mediterranean Fever (FMF) is a monogenic autoinflammatory disorder worldwide, which usually presents with recurrent fever and serositis episodes since childhood. Fever episodes usually last 12–72 h, resolve spontaneously, and are often accompanied by sterile peritonitis, synovitis, pleuritis, and seldom pericarditis and erysipelas-like erythema [1, 2]. The severity and frequency of FMF attacks can be alleviated by colchicine, which is the standard treatment option in FMF [3, 4]. The most life-threatening complication of FMF is secondary amyloidosis, particularly involving kidneys due to the accumulation of amyloid resultant from excessive inflammation [2, 5].
In addition to the diversity of symptoms, duration, frequency, and intensity of attacks are highly variable between FMF patients. Based on genotype–phenotype studies, it was generally accepted that M694V and M694I mutations are related to severe disease and amyloidosis in distinct ethnicities [2, 3, 6]. Nevertheless, genotype alone cannot be held responsible for this phenotypic variability.
Following the discovery of Mediterranean FeVer (MEFV) gene on the 16th chromosome in 1997, the confirmation of FMF diagnosis became more challenging [7, 8]. Besides the identification of low penetrant mutations, patients with heterozygous and even no MEFV mutations have been diagnosed as having FMF [9].
Epigenetics can be roughly described as heritable changes affecting gene expression without any changes in the genome. The major epigenetic mechanisms include DNA methylation, histone modifications, and chromatin remodeling [10]. miRNAs are a main group of small non-coding RNAs and an important component of epigenetic mechanisms. In 1993, Lee et al. first determined lin-4 in Caenorhabditis elegans and suggested that it encodes small RNA products [11]. It is followed by the discovery of let-7, the first human miRNA in 2000, and there are more than 2000 human miRNA annotated in databases currently [12, 13]. The most noteworthy postulated functions of miRNAs are the regulation of epigenetic modifications and maintaining tissue homeostasis. miRNAs are also suggested to be transported by directly gap junctions, extracellular vesicles, exosomes, apoptotic bodies, lipoproteins, and ribonucleoproteins, and play a particular role in cell–cell communication [13, 14]. The role of miRNAs in FMF pathogenesis remains doubtful, because there are only few studies investigating inflammation- and autoimmunity-related miRNAs in FMF [15,16,17,18,19,20,21].
The proposed mechanisms of FMF include overactivation of caspase-1, thus excessive IL-1 production due to loss of function of pyrin, a negative regulator of NLRP3 inflammasome assembly [9, 22]. However, recent studies have postulated pyrin as a sensor protein which may be triggered by small GTPases of the Rho family and cytoskeleton changes by interacting with microtubules and actin filaments. Therefore, a gain-of-function mutation in MEFV which causes an overactive pyrin was proposed to cause FMF [23, 24]. Another cellular process in which pyrin was suggested to take part is apoptosis [9, 25,26,27]. Furthermore, it is well accepted that numerous miRNAs, which were defined as having oncogenic and tumor suppressor roles, take part in apoptotic pathways [28]. Therefore, this study was conducted to investigate apoptosis-related miRNAs in FMF and their influence on clinical manifestations.
Materials and methods
Participants
This is a cross-sectional observational study conducted to compare the expressions of certain miRNAs between FMF patients and healthy controls. Patients with FMF, who admitted to our outpatient clinic between March 2017 and July 2017, were included in this study. All patients had been diagnosed as having FMF according to Tel Hashomer diagnostic criteria, in the same department, and were under colchicine treatment at the time of the study [7]. Patients with a follow-up duration of less than 6 months were excluded from the study. Demographic parameters including age, age at disease onset, age at diagnosis, clinical manifestations, duration and dosage of colchicine treatment, treatment responses, and MEFV gene-sequencing results were retrospectively collected from medical files of the patients. Patients who lack homozygosity for exon 10 mutations in MEFV gene underwent additional genetic analysis for autoinflammatory diseases with recurrent fever, including Cryopyrin-Associated Autoinflammatory Syndromes (CAPS), Tumor Necrosis Factor Receptor Associated Periodic Syndrome, mevalonate kinase deficiency, and Deficiency of Adenosine Deaminase 2 by next-generation sequencing system. Patients with other identifiable mutations in genes related to the aforementioned hereditary autoinflammatory disorders and the presence of recurrent urticarial rash, hearing loss, and skeletal abnormalities suggestive for CAPS were excluded from the study [29]. Colchicine resistance was defined as having one or more attacks each month despite receiving the maximally tolerated dose for at least 6 months [30]. We also grouped the responder patients if the attack frequency decreased at least 50% of baseline and had normal acute-phase reactants (APRs) during attack-free periods as having a favorable response, and if they could not meet these criteria but better than before, the patients were grouped as having a partial response.
APRs, including erythrocyte sedimentation rate, C-reactive protein, and Serum Amyloid-A obtained at study enrollment. The control group consisted of 31 healthy, age and sex-matched participants, admitted to our hospital for well-child preventive care visits. Children with an active infection sign, such as fever, cough, vomiting, diarrhea, and APR elevation, were excluded from control group. Written informed consent was obtained from all participants and their parents prior to the study. The study was approved by the local ethics committee of Cukurova University Medical Faculty (number: 53/2, date: 13.05.2016).
MEFV gene analysis
Leukocyte DNA was isolated from all cases by standard methods. We performed MEFV gene analysis by a molecular diagnostics tool, next-generation sequencing platform (MiSeq System, Illumina). The test covers all exons for MEFV gene, at least 50 nucleotides upstream and downstream of each exon and 1 kb of both the 5′ promoter regions and the 3′ UTRs.
miRNA analysis
Metanalysis of miRNAs in relation to both disease phenotype and target interactions had been performed by miRbase (searching for known miRNA informations) and miRWalk 2.0 (searching for predicted and validated miRNA interactions); afterward, the apoptosis-related miRNAs were selected. Venous blood samples were obtained from the participants and further centrifuged in 3500 rpm to the supernatant. Total RNA was extracted by utilizing Qiagen RNeasy Mini Kit (Hilden, Germany) according to the manufacturer’s instructions. Qiagen miScript II RT kit was used for reverse transcription of RNA to cRNA. MiRNAs were isolated from cRNAs by preamplification with Qiagen miScript Microfluidics PreAmp kit. Assay plate containing miRNAs was diluted with miScrip Microfluidics Universal Primer, Assay Loading Reagent (Fluidigm), and Rnase-free water subsequently. This primer was confronted by Qiagen miScript MicroFluidics PCR kit, which were further analyzed in BioMark (Fluidgm, Germany) for the following 33 miRNAs; let-7a-5p, let-7c, let-7 g-5p, miR-15a-5p, miR-15b-5p, miR-16-5p, miR-17-5p, miR-23a-3p, miR-24-3p, miR-25-3p, miR-26b-5p, miR-26a-5p, miR-27a-3p, miR-29a-3p, miR-29b-3p, miR-29c-3p, miR-30a-5p, miR-30b-5p, miR-30c-5p, miR-30d-5p, miR-30e-5p, miR-98-5p. miR-101-3p, miR-106b-5p, miR-145-5p, miR-146a-5p, miR-181a-5p, miR-181b-5p, miR-181c-5p, miR-195-5p, miR-214-3p, miR-222-3p, and miR-365a-3p.
Statistical analysis
The minimum sample size was calculated as 24, in accordance to the formula including type 1 error rate (α = 5%) and type II fault (β = 0.20), power (1−β = 80%), and predicted means for Group A (FMF patients) and B (healthy volunteer) as 1.5 and 1, respectively. In addition, the standard deviation was considered as 0.8 and sampling ratio (Group A/B) as 6. Categorical variables were presented as numbers and percentages. The distribution of continuous variables was tested by Kolmogorov–Smirnov test for normality, and continuous variables including demographic data were given as median and minimum–maximum (range). miRNA expressions were statistically calculated by Flexsix GE Chipi Fludigm (Biomark) system and obtained Ct values were analyzed according to related software (https://www.qiagen.com/us/shop/genes-and-pathways/data-analysis-center-overview-age/web). The best housekeeping gene was chosen as SNORD68 among the nominate genes by normfinder analysis (https://moma.dk/normfinder-software). ΔCt values of both patients and control group were calculated by subtracting Ct values in the control SNORD68 gene from Ct values in each assay. By subtraction of control Ct values from ΔCt values resulted as ΔΔCt values. Fold regulation results were compared with Welch T test. Negative fold regulation value showed decreased expression lower than the control group, whereas positive fold regulations meant increased expression higher than controls. Statistical significance was considered to be p < 0.005 in each test.
Results
Demographic and clinical features
This study included 191 patients with FMF, of whom 86 (45%) were female and 105 (55%) were male. Twenty-five patients (13.1%) were during an FMF attack and 166 (86.9%) were on an attack-free period at the time of the study. In FMF study group, the median ages at symptom onset, diagnosis, and study enrollment were 4.42 (range, 1–16.02), 7.13 (1.52–17.02), and 11.89 (range, 3.38–17.8) years, respectively. All of the patients were under colchicine treatment, of whom seven (3.7%) did not respond to colchicine and treated simultaneously with canakinumab. Clinical characteristics and genotypes of the patients are summarized in Table 1. Control group included 31 healthy children, 15 (48.4%) females and 16 (51.2%) males, and the median age of healthy controls was 9.62 (range, 1.2–17.5) years.
miRNA analysis results
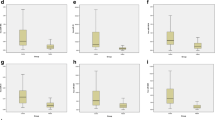
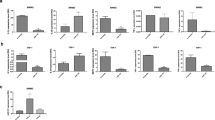
The mean plasma miRNA expression levels of both patients with FMF and the control group were elaborately given in Table 2 as ΔCt values. According to the results of the Welch t test, which was performed to compare the ΔΔCt values between groups including unequal sample sizes, we found that 26 of 33 miRNAs differed between FMF patients and controls. Nineteen miRNAs, including let-7a-5p, let-7c, let-7 g-5p, miR-15b-5p, miR-16-5p, miR-17-5p, miR-23a-3p, miR-24-3p, miR-25-3p, miR-26a-5p, miR-26b-5p, miR-27a-3p, miR-29c-3p miR-30a-5p, miR-30d-5p, miR-30e-5p, miR-106b-5p, miR-146a-5p, and miR-195-5p, were found as down-regulated in FMF patients than healthy controls, whereas expression levels of miR-15a-5p, miR-29b-3p, miR-181a-5p, miR-181b-5p, miR-181c-5p, miR-214-3p, and miR-365a-3p were elevated in FMF. The test results were summarized in Table 3. In detail, when we compared the miRNA expression profiles among FMF patients in terms of genotype, including the presence of M694V positivity, exon 10 mutations, and heterozygosity, we did not find any statistically significant difference. Furthermore, miRNA expression levels were found similar between patients with favorable colchicine response and without. Expression of 33 apoptosis-related miRNAs was also similar between patients in an inflammatory attack and 166 patients during attack-free period.
Discussion
Our study revealed that 26 of 33 apoptosis-related miRNAs’ expressions were altered in serum of FMF patients with respect to healthy controls. In detail, nineteen miRNAs, including let-7a-5p, let-7c, let-7 g-5p, miR-15b-5p, miR-16-5p, miR-17-5p, miR-23a-3p, miR-24-3p, miR-25-3p, miR-26a-5p, miR-26b-5p, miR-27a-3p, miR-29c-3p, miR-30a-5p, miR-30d-5p, miR-30e-5p, miR-106b-5p, miR-146a-5p and miR-195-5p were found down-regulated; miR-15a-5p, miR-29b-3p, miR-181a-5p, miR-181b-5p, miR-181c-5p, miR-214-3p, and miR-365a-3p, were up-regulated. These miRNAs were suggested to target various genes and upregulation of these non-coding miRNAs were linked to oncogenesis and cancer progression; however, except for miR-146a-5p, miR-16-5p, miR-26a-5p, let-7a-5p, and miR-181, they have not been investigated on monogenic autoinflammatory diseases, including FMF.
To the best of our knowledge, the first preliminary miRNA expression study in FMF was conducted in 2016 by Latsoidis et al. Among nine adult FMF patients, they found differently expressed 29 miRNAs, of which miR-4520 was the most promising candidate implicated in biological processes, by further bioinformatic analysis [15]. Subsequently, Koga et al. studied expression profiles of only four miRNAs in nine FMF patients and found extremely low miR-204-3p expression in FMF patients during an attack and even suggested miR-204-3p as a useful biomarker. In the same study, miR-204-3p was also shown to inhibit the secretion of inflammatory cytokines [16].
The most intriguing results came from another study, including 24 FMF patients, grouped by localization of MEFV mutations. The expression of circulating miRNAs differed between the groups according to having a typical phenotype with or without an exon 10 mutation [17]. In the same study, the authors proposed that elevated miR-320 expression may be a compensatory mechanism for controlling excessive inflammation caused by exon 10 mutations in FMF [17]. A recent study found 14 miRNAs to be differentially expressed in 12 FMF patients, of which miR-20a-5p, miR-197-3p, let-7d-3p, and miR-574-3p were associated with inflammatory pathways. Patients with homozygote M694V mutations were shown to have up-regulated miR-20a-5p and down-regulated miR-197-3p, for which the authors commented as they may have a role on severe disease phenotype [18].
Hortu et al. performed a more comprehensive workup about miRNAs in 51 pediatric FMF patients, of whom 27.5% were at an FMF attack and 39.2% were colchicine-naïve. Only 15 miRNAs (miR-15a, miR-146a, miR-155, miR-26, miR-21, miR-223, miR-16, miR-181, miR-125a, miR-34a, miR-124a, miR-203, miR-346, miR-132, and miR-23b) were evaluated. Eleven of them, which were previously linked to inflammatory pathways in other studies including rheumatic and autoimmune disorders, were also found as significantly decreased in FMF patients. While patients were grouped according to the mutation type, attack status, and presence of acute-phase reactant elevations, there were no significant differences in these miRNA expressions. The patient group was analyzed and compared within itself, and the expression levels of five miRNAs (miR-132, miR-15a, miR-181a, miR-23b, and miR-26a) in the patients who took colchicine seemed to have increased and levels of 5 miRNAs (miR-146a, miR-15a, miR-16, miR-26a, and miR-34a) in the patients who took colchicine were significantly lower [19]. With respect to this study, we similarly found significantly down-regulated expressions in miR-16-5p, miR-26a-5p, miR-26b-5p, and miR-146a-5p, which suggest their negative regulatory roles on FMF. On the counterpart, we found up-regulated expressions of miR-181a-5p, miR-181b-5p, and miR-181c-5p in FMF patients, which was similar to that study revealing decreased overall expression in FMF patients, but elevated expression in patients under colchicine treatment. In fact, all participants had been received colchicine in our study, and thus, our results may also support colchicine may increase miR-181 expression and thus control excessive inflammation. Besides, it was previously suggested that colchicine induces cell apoptosis in colon cancer and normal liver cells in a dose-dependent manner [31, 32]. We think that these miRNAs somehow could take part in these apoptotic pathways. Further functional studies should be performed to clarify the effects of colchicine on the expression of these miRNAs. With improving knowledge about the effects of miRNAs, miRNAs-based therapies may be even thinkable in FMF in the near future [33].
Amarilyo et al. observed that four miRNAs (miR-144-3p, miR-21-5p, miR-4454, and miR-451a) were up-regulated and three (miR-107, let-7d-5p, and miR-148b-3p) were down-regulated significantly in FMF patients [20]. Moreover, Demir et al. studied four miRNAs, which were previously found deregulated in FMF patients, and found decreased miRNA-155 and miRNA-204 and increased miRNA-16 and miRNA-451 expressions [21]. Studies investigating miRNAs in FMF were identified through a literature search on MEDLINE/PubMed and Scopus databases, and the main results of the studies are summarized in Table 4.
Apoptosis can be defined as a physiological process and programmed cell death resultant from endogenous or exogenous signals. In the last 2 decades, it has been an area of interest whether apoptosis plays a role in inflammatory and rheumatic diseases, particularly by terminating excessive inflammation [34]. FMF is characterized by recurrent autoinflammation due to dysfunctional pyrin protein, which interacts ASC by its PYD domain and thus activation of caspase-1 [26, 35]. Suggesting the self-limiting nature of the disease, patients were found to have increased neutrophil apoptosis and FasL levels during FMF attacks, compared to healthy controls in a preliminary study [36]. On the other hand, another study reported no difference in neutrophil apoptosis between FMF patients and controls [37]. Similarly, another study found similar serum FasL levels between FMF patients on attack, during remission and control group, which thus yielded no diagnostic aid [38]. In a more recent study, both spontaneous and induced neutrophil apoptosis were found significantly higher in FMF patients during an attack or remission than healthy controls and this finding was supported by the elevation of caspase-3 mRNA. The authors induced apoptosis with lipopolysaccharides, TNF-alpha, MDP, CSK4, ATP, and even colchicine; however, they came up with the question of which endogenous stimuli may trigger the spontaneous neutrophil apoptosis in FMF patients [39].
On the other hand, some miRNAs were prominently altered miRNAs in the present study, which were at least 1.5 times increased (miR-181b-5p) or decreased (miR-16-5p, miR-17-5p, miR-25-3p, and miR-195-5p). Moreover, it was attempted to clarify the role of these miRNAs on apoptosis by functional studies previously. First, miR-181b-5p, which was the most prominently up-regulated miRNA in our study, was suggested to inhibit apoptosis through MEK/ERK/p21 pathway [40]. Besides, decreased expression of miR-17-5p and miR-25 were shown to induce apoptosis by upregulation of target genes including phosphatase and tensin homolog and high mobility group box-1, respectively [41, 42]. Our results also showed decreased serum expressions of miR-17-5p and miR-25-3p, suggesting these miRNAs may be involved in FMF pathogenesis via apoptotic pathways.
In contrary, overexpression of miR-16-5p and miR-195-5p were proposed to activate caspases 3 and 9 and lead to apoptosis in other studies [43, 44]. However, we found significant downregulations of miR-16-5p and miR-195-5p, and upregulation of miR-181b-5p which conflicts with the hypothesis of excessive apoptosis in FMF pathogenesis. Nonetheless, we still speculate that these apoptosis-related miRNAs could have a complex interaction in regulating apoptosis by multiple target genes, be affected by colchicine treatment, and individually activate or control inflammation in FMF patients.
The major limitations of our study were the lack of functional annotation of the expressed miRNAs and the evidence showing the neutrophil apoptosis. Additionally, all patients included in the study were under colchicine treatment, and we cannot be sure that miRNA expression was affected by colchicine treatment or not. Moreover, the etiology could be other autoinflammatory diseases in patients who had insignificant MEFV results. Although we performed further genetic analysis, there is still a possibility that the phenotype was caused by mutations in other genes or by novel genes. Thus, further studies are needed to clarify these limitations on this topic in the future.
To our knowledge, this is the first study investigating apoptosis-related miRNAs, widely investigated in the pathogenesis of several cancers before. More intriguingly, we found significant miRNA downregulations and upregulations in FMF patients, regardless of genotype, colchicine response, and having an inflammatory attack during miRNA analysis. We think that the inclusion of genetically heterogeneous FMF patients also makes our study more interesting, since most of the previous studies investigated either M694V homozygote or heterozygote patients. Because, the emerging concerns about FMF pathogenesis, whether epigenetic or environmental factors have an additional effect or not, especially have raised from the presence of a substantial proportion of FMF patients with heterozygote MEFV mutations, exon 2 mutations, or no mutations.
In conclusion, our results revealed significant deregulations of 26 apoptosis-related miRNAs regardless of genotype; therefore, we speculate that apoptosis-related miRNAs might be involved in FMF pathogenesis, by affecting apoptotic pathways. After all, we also highlight that there is a thriving need for more work to clarify epigenetics in FMF.
References
Ben-Chetrit E, Touitou I (2009) Familial Mediterranean fever in the world. Arthritis Rheum 61:1447–1453. https://doi.org/10.1002/art.24458
Tunca M, Akar S, Onen F, Ozdogan H, Kasapcopur O, Yalcinkaya F, Tutar E, Ozen S, Topaloglu R, Yilmaz E, Arici M, Bakkaloglu A, Besbas N, Akpolat T, Dinc A, Erken E, Turkish FMF Study Group (2005) Familial Mediterranean fever (FMF) in Turkey: results of a nationwide multicenter study. Medicine (Baltimore) 84(1):11. https://doi.org/10.1097/01.md.0000152370.84628.0c
Kisla Ekinci RM, Balci S, Dogruel D, Altintas DU, Yilmaz M (2019) Twenty-year experience of a single referral center on pediatric familial Mediterranean fever: what has changed over the last decade? J Clin Rheumatol. https://doi.org/10.1097/RHU.0000000000001146
Hentgen V, Grateau G, Kone-Paut I, Livneh A, Padeh S, Rozenbaum M, Amselem S, Gershoni-Baruch R, Touitou I, Ben-Chetrit E (2013) Evidence-based recommendations for the practical management of familial Mediterranean fever. Semin Arthritis Rheum 43:387–391. https://doi.org/10.1016/j.semarthrit.2013.04.011
Kasifoglu T, Bilge SY, Sari I, Solmaz D, Senel S, Emmungil H, Kilic L, Oner SY, Yildiz F, Yilmaz S, Bakirli DE, Tufan MA, Yilmaz S, Yazisiz V, Pehlivan Y, Bes C, Cetin GY, Erten S, Gonullu E, Temel T, Sahin F, Akar S, Aksu K, Kalyoncu U, Direskeneli H, Erken E, Kisacik B, Sayarlioglu M, Korkmaz C (2014) Amyloidosis and its related factors in Turkish patients with familial Mediterranean fever: a multicentre study. Rheumatology (Oxford) 53:741–745. https://doi.org/10.1093/rheumatology/ket400
Ait-Idir D, Djerdjouri B, Bouldjennet F, Taha RZ, El-Shanti H, Sari-Hamidou R, Khellaf G, Benmansour M, Benabadji M, Haddoum F (2017) The M694I/M694I genotype: a genetic risk factor of AA-amyloidosis in a group of Algerian patients with familial Mediterranean fever. Eur J Med Genet 60:149–153. https://doi.org/10.1016/j.ejmg.2016.12.003
The International FMF Consortium (1997) Ancient missense mutations in a new member of the RoRet gene family are likely to cause familial Mediterranean fever. Cell 90:797–807. https://doi.org/10.1016/s0092-8674(00)80539-5
French FMF Consortium (1997) A candidate gene for familial Mediterranean fever. Nat Genet 17:25–31. https://doi.org/10.1038/ng0997-25
Manukyan G, Aminov R (2016) Update on pyrin functions and mechanisms of familial Mediterranean fever. Front Microbiol 7:456. https://doi.org/10.3389/fmicb.2016.00456
Álvarez-Errico D, Vento-Tormo R, Ballestar E (2017) Genetic and epigenetic determinants in autoinflammatory diseases. Front Immunol 8:318. https://doi.org/10.3389/fimmu.2017.00318
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75:843–854. https://doi.org/10.1016/0092-8674(93)90529-y
Reinhart BJ, Slack FJ, Basson M, Pasquinelli AE, Bettinger JC, Rougvie AE, Horvitz HR, Ruvkun G (2000) The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403:901–906. https://doi.org/10.1038/35002607
Saliminejad K, Khorram Khorshid HR, Soleymani Fard S, Ghaffari SH (2019) An overview of microRNAs: biology, functions, therapeutics, and analysis methods. J Cell Physiol 234:5451–5465. https://doi.org/10.1002/jcp.27486
Ambros V (2004) The functions of animal microRNAs. Nature 431:350–355. https://doi.org/10.1038/nature02871
Latsoudis H, Mashreghi MF, Grün JR, Chang HD, Stuhlmüller B, Repa A, Gergiannaki I, Kabouraki E, Vlachos GS, Häupl T, Radbruch A, Sidiropoulos P, Doukoumetzidis K, Kardassis D, Niewold TB, Boumpas DT, Goulielmos GN (2017) Differential expression of miR-4520a associated with pyrin mutations in familial Mediterranean fever (FMF). J Cell Physiol 232:1326–1336. https://doi.org/10.1002/jcp.25602
Koga T, Migita K, Sato T, Sato S, Umeda M, Nonaka F, Fukui S, Kawashiri SY, Iwamoto N, Ichinose K, Tamai M, Nakamura H, Origuchi T, Ueki Y, Masumoto J, Agematsu K, Yachie A, Yoshiura KI, Eguchi K, Kawakami A (2018) MicroRNA-204-3p inhibits lipopolysaccharide-induced cytokines in familial Mediterranean fever via the phosphoinositide 3-kinase γ pathway. Rheumatology (Oxford) 57:718–726. https://doi.org/10.1093/rheumatology/kex451
Wada T, Toma T, Matsuda Y, Yachie A, Itami S, Taguchi YH, Murakami Y (2017) Microarray analysis of circulating microRNAs in familial Mediterranean fever. Mod Rheumatol 27:1040–1046. https://doi.org/10.1080/14397595.2017.1285845
Akkaya-Ulum YZ, Balci-Peynircioglu B, Karadag O, Eroglu FK, Kalyoncu U, Kiraz S, Ertenli AI, Özen S, Yilmaz E (2017) Alteration of the microRNA expression profile in familial Mediterranean fever patients. Clin Exp Rheumatol 35(Suppl 108):90–94
Hortu HO, Karaca E, Sozeri B, Gulez N, Makay B, Gunduz C, Atik T, Tekin IM, Unsal SE, Cogulu O (2019) Evaluation of the effects of miRNAs in familial Mediterranean fever. Clin Rheumatol 38(3):635–643. https://doi.org/10.1007/s10067-017-3914-0
Amarilyo G, Pillar N, Ben-Zvi I, Weissglas-Volkov D, Zalcman J, Harel L, Livneh A, Shomron N (2018) Analysis of microRNAs in familial Mediterranean fever. PLoS ONE 13:e0197829. https://doi.org/10.1371/journal.pone.0197829
Demir F, Çebi AH, Kalyoncu M (2020) Assessment of circulating microribonucleic acids in patients with familial Mediterranean fever. Arch Rheumatol. https://doi.org/10.5606/ArchRheumatol.2020.7414
de Jesus AA, Canna SW, Liu Y, Goldbach-Mansky R (2015) Molecular mechanisms in genetically defined autoinflammatory diseases: disorders of amplified danger signaling. Annu Rev Immunol 33:823–874. https://doi.org/10.1146/annurev-immunol-032414-112227
Park YH, Wood G, Kastner DL, Chae JJ (2016) Pyrin inflammasome activation and RhoA signaling in the autoinflammatory diseases FMF and HIDS. Nat Immunol 17:914–921. https://doi.org/10.1038/ni.3457
Xu H, Yang J, Gao W, Li L, Li P, Zhang L, Gong YN, Peng X, Xi JJ, Chen S, Wang F, Shao F (2014) Innate immune sensing of bacterial modifications of Rho GTPases by the pyrin inflammasome. Nature 513:237–241. https://doi.org/10.1038/nature13449
Martinon F, Hofmann K, Tschopp J (2001) The pyrin domain: a possible member of the death domain-fold family implicated in apoptosis and inflammation. Curr Biol 11:118–120. https://doi.org/10.1016/s0960-9822(01)00056-2
Richards N, Schaner P, Diaz A, Stuckey J, Shelden E, Wadhwa A, Gumucio DL (2001) Interaction between pyrin and the apoptotic speck protein (ASC) modulates ASC-induced apoptosis. J Biol Chem 276:39320–39329. https://doi.org/10.1074/jbc.M104730200
Gumucio DL, Diaz A, Schaner P, Richards N, Babcock C, Schaller M, Cesena T (2002) Fire and ICE: the role of pyrin domain-containing proteins in inflammation and apoptosis. Clin Exp Rheumatol 20:45–53
Kashyap D, Tuli HS, Garg VK, Goel N, Bishayee A (2018) Oncogenic and tumor-suppressive roles of MicroRNAs with special reference to apoptosis: molecular mechanisms and therapeutic potential. Mol Diagn Ther 22:179–201. https://doi.org/10.1007/s40291-018-0316-1
Pathak S, McDermott M, Savic S (2016) Autoinflammatory diseases: update on classification diagnosis and management. J Clin Pathol 70:1–8. https://doi.org/10.1136/jclinpath-2016-203810
Ozen S, Demirkaya E, Erer B, Livneh A, Ben-Chetrit E, Giancane G, Ozdogan H, Abu I, Gattorno M, Hawkins PN, Yuce S, Kallinich T, Bilginer Y, Kastner D, Carmona L (2016) EULAR recommendations for the management of familial Mediterranean fever. Ann Rheum Dis 75:644–651. https://doi.org/10.1136/annrheumdis-2015-208690
Huang Z, Xu Y, Peng W (2015) Colchicine induces apoptosis in HT-29 human colon cancer cells via the AKT and c-Jun N-terminal kinase signaling pathways. Mol Med Rep 12:5939–5944. https://doi.org/10.3892/mmr.2015.4222
Chen XM, Liu J, Wang T, Shang J (2012) Colchicine-induced apoptosis in human normal liver L-02 cells by mitochondrial mediated pathways. Toxicol In Vitro 26:649–655. https://doi.org/10.1016/j.tiv.2012.01.024
Balci-Peynircioglu B, Akkaya-Ulum YZ, Akbaba TH, Tavukcuoglu Z (2019) Potential of miRNAs to predict and treat inflammation from the perspective of familial Mediterranean fever. Inflamm Res 68:905–913. https://doi.org/10.1007/s00011-019-01272-6
Siegel RM, Fleisher TA (1999) The role of Fas and related death receptors in autoimmune and other disease states. J Allergy Clin Immunol 103:729–738. https://doi.org/10.1016/s0091-6749(99)70412-4
Shiohara M, Taniguchi S, Masumoto J, Yasui K, Koike K, Komiyama A, Sagara J (2002) ASC, which is composed of a PYD and a CARD, is up-regulated by inflammation and apoptosis in human neutrophils. Biochem Biophys Res Commun 293:1314–1318. https://doi.org/10.1016/S0006-291X(02)00384-4
Ozen S, Uckan D, Baskin E, Besbas N, Okur H, Saatci U, Bakkaloglu A (2001) Increased neutrophil apoptosis during attacks of familial Mediterranean fever. Clin Exp Rheumatol 19:68–71
Ceri M, Unverdi S, Senes M, Altay M, Yilmaz R, Yucel D, Duranay M (2013) Serum soluble fas ligand levels in familial Mediterranean fever. Ren Fail 35:835–857. https://doi.org/10.3109/0886022X.2013.794660
Davtyan TK, Harutyunyan VA, Hakobyan GS, Avetisyan SA (2008) Heightened endotoxin susceptibility of monocytes and neutrophils during familial Mediterranean fever. FEMS Immunol Med Microbiol 52:370–378. https://doi.org/10.1111/j.1574-695X.2008.00385.x
Manukyan G, Aminov R, Hakobyan G, Davtyan T (2015) Accelerated apoptosis of neutrophils in familial Mediterranean fever. Front Immunol 6:239. https://doi.org/10.3389/fimmu.2015.00239
Liu B, Guo Z, Gao W (2019) miR-181b-5p promotes proliferation and inhibits apoptosis of hypertrophic scar fibroblasts through regulating the MEK/ERK/p21 pathway. Exp Ther Med 17:1537–1544. https://doi.org/10.3892/etm.2019.7159
Wang X, Li Z, Bai J, Song W, Zhang F (2019) miR-17-5p regulates the proliferation and apoptosis of human trabecular meshwork cells by targeting phosphatase and tensin homolog. Mol Med Rep 19:3132–3138. https://doi.org/10.3892/mmr.2019.9973
Sárközy M, Kahán Z, Csont T (2018) A myriad of roles of miR-25 in health and disease. Oncotarget 9:21580–21612. https://doi.org/10.18632/oncotarget.24662
Zheng C, Zheng Z, Sun J, Zhang Y, Wei C, Ke X, Liu Y, Deng L, Wang H (2017) MiR-16-5p mediates a positive feedback loop in EV71-induced apoptosis and suppresses virus replication. Sci Rep 7:16422. https://doi.org/10.1038/s41598-017-16616-7
Zhao DL, Wu QL (2019) Effect of inhibition to Yes-related proteins-mediated Wnt/β-catenin signaling pathway through miR-195-5p on apoptosis of gastric cancer cells. Eur Rev Med Pharmacol Sci 23:6486–6496. https://doi.org/10.26355/eurrev_201908_18532
Acknowledgements
We specially thank to Aidan Boga (Brisbane, Australia) for his excellent work on language editing of the paper.
Funding
This work was supported by the grants from the Cukurova University Scientific Research Projects Coordination Unit (TTU-2017-7994), Adana, Turkey.
Author information
Authors and Affiliations
Contributions
Dr. Karpuzoglu and Dr. Yilmaz conceptualized and designed the study, drafted the initial manuscript, and reviewed and revised the manuscript. Dr. Kisla Ekinci, Dr. Bisgin, and Dr. Balci collected data carried out the initial analyses, and critically reviewed and revised the manuscript. All authors approved the final manuscript as submitted and agreed to be accountable for all aspects of the work. All co-authors take full responsibility for the integrity of the study.
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that they have no conflicts of interest. All co-authors take full responsibility for the integrity of the study.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from parents of the participants included in the study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Karpuzoglu, E.M., Kisla Ekinci, R.M., Balci, S. et al. Altered expression of apoptosis-related, circulating cell-free miRNAs in children with familial Mediterranean fever: a cross-sectional study. Rheumatol Int 41, 103–111 (2021). https://doi.org/10.1007/s00296-020-04541-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00296-020-04541-4