Abstract
Familial Mediterranean fever (FMF) is an inherited autoinflammatory disorder that can result in attacks with accompanying recurrent episodes of fever, serositis, and skin rash. MiRNAs are demonstrated to be associated with a number of other diseases; however, no comprehensive study has revealed its association with FMF disease. The aim is to investigate the role of microRNAs in FMF. We included 51 patients with genetically diagnosed FMF who had clinical symptoms and 49 healthy volunteers. Fifteen miRNAs that were found to be associated with autoinflammatory diseases and have a part in immune response were evaluated. The expression levels of 11 miRNAs (miR-125a, miR-132, miR-146a, miR-155, miR-15a, miR-16, miR-181a, miR-21, miR-223, miR-26a, and miR-34a) in the patient group were significantly low, compared with the control group (p < 0.05). The patient group was analyzed and compared within itself, and the expression levels of 5 miRNAs (miR-132, miR-15a, miR-181a, miR-23b, miR-26a) in the patients who took colchicine seemed to have increased and levels of 5 miRNAs (miR-146a, miR-15a, miR-16, miR-26a, miR-34a) in the patients who took colchicine were significantly lower (p < 0.05). Furthermore, the attack patients were compared with the control group, and their expression levels of 4 miRNAs (miR-132, miR-15a, miR-21, miR-34a) were significantly lower (p < 0.05). Levels of 9 miRNAs (miR-132, miR-146a, miR-15a, miR-16, miR-181a, miR-21, miR-223, miR-26a, miR-34a) in non-attack patients decreased significantly (p < 0.05). Our study demonstrates that miRNAs could be effective in the pathogenesis of FMF.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Familial Mediterranean fever (FMF) is an autosomal recessive autoinflammatory disease [1, 2]. The MEFV is encoding the Pyrin protein which controls inflammation. The mutation of MEFV gene results with inflammation and clinical manifestations are caused by pyrin dysfunction. It has been suggested that a physiological consequence of FMF-associated mutations causes an increase in IL-1b production through caspase-1, in which the mechanism is based on the direct interaction of the PRP/SPRY region of the pyrin protein with caspase-1 [3]. Interestingly, pyrin was shown to act both as an inhibitor and as an activator of IL-1b [3]. In addition to this, pyrin mutations have also been shown to be responsible for increased NLRP3-dependent IL-1b secretion [4]. Moreover, PRY/SPRY mutations have been shown to inhibit autophagic degradation of NLRP through the pyrin, contributing to the FMF-associated inflammatory phenotype with IL-1b responses [5]. In the light of this information, modeling data indicated that it is still unclear how several MEFV mutations lead to the proinflammatory phenotype of FMF, suggesting that alternative pathogenic pathways leading to FMF needed to be shown [5]. The vast majority of the clinically diagnosed episodes are accompanied by recurrent fevers and serosal involvement. Increases in leukocyte count, serum amyloid-A (SAA), and acute phase reactants have also been detected during attacks [6]. However, specific factors involved in the triggering of attacks remain elusive [7]. Race, genetic predisposition, and environmental factors have all been suggested in the literature [8, 9]. The genotype-phenotype correlation in FMF is not well understood. However, amyloidosis which is the most important has often been reported to be associated with a specific mutation in exon 10, M694V, especially at the homozygous state [10, 11], while no specific genotype correlation with the other symptoms has been reported [12]. Because the underlying molecular mechanism of FMF etiopathogenesis is not yet fully understood, more recent studies have focused on immunological events in an attempt to identify potential markers. FMF attacks are accompanied by neutrophil-rich serosal inflammation [13]. Interleukin (IL)-1, IL-6, IL-8, and tumor necrosis factor (TNF)-α levels of individuals were found to be higher during attacks [14, 15]. Colchicine therapy has been shown to suppress these cytokines [16]. Since the clinic of FMF is suggested to be variable depending on epigenetic conditions; microRNAs may be the part of these epigenetic mechanisms. [17].
MicroRNAs (miRNA) are small, single-stranded, non-coding endogenous RNA molecules ranging in length from 19 to 25 nucleotides. They affect messenger RNA (mRNA) through degradation and translational inhibition [18], as well as playing an important role in the regulation of transcriptional and posttranscriptional levels, particularly on cellular differentiation, proliferation, morphogenesis, metabolism, and apoptosis [19, 20]. miRNAs have also been shown to regulate innate immune responses to inflammatory stimuli [21]. Through a number of studies, their effect within the pathogenesis of inflammatory and autoimmune diseases such as rheumatoid arthritis (RA), multiple sclerosis (MS), and systemic lupus erythematosus (SLE) has been well documented. It has also revealed that miRNAs have an important function in the development of many cell types, including T and B cell development and differentiation, monocyte-neutrophil proliferation, and regulation of the release of inflammatory mediators [21, 22].
The aims of this study were to identify miRNAs which have the following:
-
1.
Associations with FMF
-
2.
A modulation effect on MEFV gene
-
3.
An impact on phenotypic changes in disease progression
-
4.
A relationship between the mutations detected in the patients
Patients and methods
Study design
In this cross-sectional study, patients were consecutively enrolled from three pediatric rheumatology referral centers of İzmir: Ege University Hospital, Dokuz Eylül University Hospital, and Behçet Uz Children’s Hospital. Following collection of venous blood samples from both patients and control group, RT-PCR was applied to evaluate miRNA expressions. Two patients in the control group were excluded from the study due to poor levels of miRNA expression in their venous blood samples. miRNA expression differences were evaluated in terms of the following criteria:
-
1.
Identification of significantly dysregulated miRNAs in FMF patients
-
2.
miRNA expression level changes in relation to colchicine treatment in FMF
-
3.
The impact of age on identification of miRNAs
-
4.
The impact of attacks on identification of miRNAs
-
5.
Association of miRNAs with the laboratory parameters playing a role in FMF
-
6.
The effect of demographic characteristics on miRNAs
Patients
Fifty-one patients were included in the study. All patients were diagnosed according to the Tell-Hashomer criteria, modified by Livneh et al. [23]. A genetic analysis was performed on all patients. Forty-nine healthy children were enrolled as a control group. Inclusion criteria for the control group included having no clinical findings of FMF disease, no family history of FMF, no MEFV mutation, and being free of any active infection or chronic health problem. Patients who could not be confirmed genetically were excluded from the study. A background sheet was also obtained on each patient; this included information concerning age, gender, use of colchicine, and the state of attack (Table 1). Twenty patients (39.2%) of the study group were included before the colchicine treatment started at the time of diagnosis, and the rest of the patients were already receiving colchicine (Table 1). The study group was divided into the groups based on the attack period and colchicine treatment. The study protocol was approved by the local ethics committee. Parents and children were informed about the procedure and written informed consent was obtained from the participants prior to the study.
Mutation analysis
DNA was extracted from blood samples using a standard method. Direct sequencing of the MEFV gene was performed as described previously [24]. Mutation distributions of patients group are given in Table 2.
RNA preparation and real-time PCR
Real-time RT-PCR was used to detect and quantify the miRNA expression in the samples. Frozen material was thawed and RNA was extracted from the 10 mg of homogenized sample using PureLink® RNA Mini Kit according to the protocol for TRIzol-homogenized samples (Invitrogen). The miRNeasy RNA isolation Kit (Qiagen, Hilden, Germany) was used for the isolation and enrichment of miRNAs in accordance with the manufacturer’s instructions. TaqMan® microRNA assay quantification was performed using two-step RT-PCR. In reverse transcription process, cDNA was obtained from total RNA samples in TaqMan® MicroRNA Reverse Transcription Kit using specific miRNA primers. In the second step, PCR products were amplified using TaqMan® MicroRNA Assay and TaqMan® Universal PCR Master Mix by LightCycler 480 (Roche, Mannheim, Germany). The expressions of 15 miRNAs (miR-15a, miR-146a, miR-155, miR-26, miR-21, miR-223, miR-16, miR-181, miR-125a, miR-34a, miR-124a, miR-203, miR-346, miR-132, miR-23b) were evaluated. The u6-snRNA was used as a control to normalize differences in total RNA levels in each sample. The relative amount of each miRNA to u6-snRNA was expressed using equation 2−ΔΔCt, where ΔΔCt = (Ct miRNA − Ct U6). The value of each control sample was set at 1 and was used to calculate the fold change in targets.
Statistical analysis
For the statistical analysis, miRNA expressions were calculated by using RT2 Profiler PCR Array Software (SABiosciences http://pcrdataanalysis.sabiosciences.com/pcr/arrayanalysis. php) and a 푝 value < 0.05 was considered statistically significant. The data was normalized by geometric mean to the U6 snRNA expression and a threshold cycle (Ct) cut-off was set at 35 cycles. miRNA expression interpreted from Ct of each miRNA was normalized to Ct of U6 snRNA (ΔCt = Ct miRNA − Ct U6 snRNA). Sample size (per group) of 20 is required to ensure that at least 95% of genes have power greater than 80% [25].
Results
There were 29 females and 22 males in the patient group and 28 females and 19 males in the control group. Mean age of the patients was 105.9 ± 23.96 months (min 15, max 211) and controls were 116.23 ± 8.18 months (min 22, max 213). No significant difference considering age and gender was found between the patients and the controls.
The mean age at time of diagnosis was 80.7 ± 21.23 months. At time of inclusion, 39.2% of the patients were newly diagnosed, while 60.8% were the previously diagnosed patients under follow-up.
The most common clinical finding was fever (86.3%); the second was abdominal pain (80.4%), with 74.5% of patients having both abdominal pain and fever symptoms. Other fairly prominent symptoms included joint pain (49%), muscle pain (11.8%), chest pain (9.8%), and rash (7.8%).
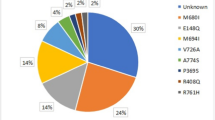
The most frequent MEFV mutation was M694V mutation in the patient group. Homozygote M694V mutation and compound heterozygote M694V mutation were detected in 31.2 and 41.1% of patients, respectively. At time of study, 27.5% were in active stages of FMF attack, while 72.5% were in attack-free period. The attack-free period was defined as “at least 2 weeks after an attack”. Among the patients, 51% were using colchicine while 49% had no history of colchicine usage.
Regarding laboratory parameters in the attack group, the mean value of leukocyte count was 8823.3 ± 856.3 cells/mm3, erythrocyte sedimentation rate 34.6 ± 10.1 mm/h, C-reactive protein 6.8 ± 1.8 mg/dl, and serum amyloid-A 492.3 ± 152.8 mg/L. For the attack-free group, the mean values were leukocyte count 7936 ± 408.27 cells/mm3, erythrocyte sedimentation rate 25.6 ± 5.6 mm/h, C-reactive protein 2.97 ± 0.86 mg/dL, and serum amyloid-A 94.5 ± 68.3 mg/L.
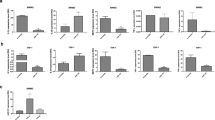
Expression levels from 11 of 15 miRNAs were found to be significantly decreased in the study group (Table 3), (Fig. 1).
In comparison to the control group, colchicine-treated patients revealed statistically significant differences in miR-132, miR-15a, miR-21, miR-26a, and miR-34a (p < 0.05).
When the group using colchicine treatment was compared to groups who were not, miR-132, miR-15a, miR-181a, miR-23b, and miR-26a delta delta Ct values were statistically significant (p < 0.05) (Table 4), (Fig. 2).
To determine the effect of different miRNAs in relation to age, patients were sub-divided into 3 groups: group 1: patients younger than 60 months old (17.6%), group 2: patients between 61 and 120 months old (19.6%), and group 3: patients older than 121 months old (62.7%). Patient subgroups were compared with the age-matched control subgroups and there was no statistically significant difference between group 1 and control group in terms of delta delta Ct values (p > 0.05). However, when groups 2 and 3 were compared to the control group, statistically significant differences were detected regarding delta delta Ct values of miR-146a, miR-155, miR-16, miR-26a, and miR-34a in group 2 and miR-125a, miR-132, miR-146a, miR-155, miR-15a, miR-16, miR-181a, miR-21, miR-223, miR-26a, and miR-34a in group 3 (p < 0.05). When comparisons were made according to mutation type, attack status, and biochemical parameters including acute phase reactants, no significant difference in expressed miRNA was detected.
Discussion
The aim of the study was to evaluate the associations of certain miRNAs with some characteristics of FMF in this study. The results of this study revealed that expression levels of 11 miRNAs were significantly lower in FMF patients compared to healthy controls.
It has been reported that miRNAs are involved in a wide range of biological processes and diseases. The groups of collagen diseases particularly rheumatoid arthritis, systemic lupus erythematosus, Sjögren’s disease, and osteoarthritis are some which come to mind [26,27,28,29,30,31,32,33,34]. Increased levels of miR-155, a very well-known miRNA in many metabolic pathways, have been found in RA and JIA, while decreased levels are seen in SLE [27]. The effects of MiR-155 have been reported as suppression and/or regulation of molecules such as matrix metalloproteinase, protein phosphatase 2A, TLR ligands, and IL-2 [27]. It can be speculated that FMF and SLE may have common etiopathogenesis mechanism associated with miR-155.
miR-146a, another miRNA, has been widely studied in relation to connective tissue disorders. Pauley et al. suggested a possible mechanism contributing to RA pathogenesis, its upregulation followed by TNF-α production [28]. In studies performed on mice, investigating altered expressions of miRNAs in the central nervous system and its association with chronic knee joint pain in osteoarthritis, Li et al. showed that a reduction of miR-146a expression results in an increase in inflammatory cytokines. It has been suggested that miR-146a might offer a protective anti-inflammatory effect in osteoarthritis [29]. However, in studies related to miR-146a in RA [30], conflicting results were revealed. In our study, miR-146a expression decreased in the FMF patient group when compared to control group. A decrease in expression of miR-146a and a negative regulatory effect on IL-1β-induced inflammation associated with endogenous inflammation in FMF is supported by our findings.
miR-223 is thought to have important roles in the regulation of NLRP3 in neutrophils [31]. Downregulation of miR-223 by IL-6 has been reported to support IL-1β and IL-6 production [32]. It has been shown that miR-223, miR-15a, and miR-16 play roles in the downregulation of the NF-κB signaling pathway and show effects as NF-κB kinase inhibitors (IKKα) [33]. Reduced expression of miR-223 has been previously shown to increase the proinflammatory cytokine response and our study confirmed these findings with decreased expression of miR-223 in patients with FMF. Several studies in the literature do contradict our results, reporting increased expression of miR-223 in peripheral T cells and synovium of patients with RA [34]. However, this difference may be attributed to the fact that miR-223 regulates RA and FMF through different pathways.
Downregulated miR-15a and miR-16 expressions may have an effect on NF-κB signal with resulting inflammation in FMF patients, confirming what our results showed. Pauley et al. have demonstrated that elevated miR16 expression in mononuclear cells of RA patients (compared to healthy volunteers) is indicative of miR-146a and miR-16 being disease activity indicators [28].
In our study, when comparing patients undergoing colchicine treatment to those who were not, expression of miR-15a was shown to have increased. It is known that colchicine, the primary drug used in the treatment of FMF disease, has anti-inflammatory, apoptotic, antimitotic, and anti-fibrotic effects. The suppressed inflammation in patients using colchicine could partly be attributed to the increased expression of miR-15a.
miR-125a is a type shown to be expressed in T cells. It has been shown that Kruppel-like factor 13 (KLF13) and tumor necrosis α-induced protein 3 (TNFAIP3) are negative regulators, inhibiting the secretion of inflammatory chemokine RANTES and supporting the NF-κB pathway [35]. Reduced expression of miR-125a in lupus T cells has been reported in conjunction with decreased inhibitor activity on KLF13 and an increase in RANTES, an inflammatory chemokine [35]. In our study, we found that miR-125a expression had decreased in FMF patients when compared to the healthy control group, confirming previously published SLE patient studies.
PDCD4 (Programmed cell death protein 4) is a proinflammatory protein that suppresses IL-10 and is supported by the NF-κB pathway. It is known that miR-21 inhibits NF-κB activation thereby suppressing PDCD4 and leading to an increase in IL-10 production [36]. When there is increased miR-21 expression via TLR (Toll-like receptor), excessive inflammatory cytokine production is prevented by negative feedback. A decreased expression of miR-21 in our patients supports those findings. Previous studies, however, have also shown conflicting results for miR-21. In RA and SLE patients [37], for example, an increase was shown. It can be speculated, therefore, that varying expression patterns across different disease groups may be directly affected by pathway usage.
One of the first miRNAs proven to play a role in the development of immune cells was miR-181a. Studies by Lashine et al. showed decreased miR-181a expression in pediatric SLE patients. However, no difference was noted between the control group and FMF patients in that study [38]. In another study concerning miR-181a, Okuhara et al. showed that expression of miR-181a in mononuclear cells of osteoarthritic (OA) patients had increased [39]. In our study, miR-181a expression decreased in patients with FMF when compared to healthy control group and was found to have increased when compared with patients not receiving colchicine therapy. While the expression of miR-181a is thought to be negatively correlated with severity of inflammation, it remains unclear through which pathway this expression is actually carried out.
Studies by Gandhi et al. and Tufekci et al. have shown that miR-34a is upregulated in multiple sclerosis (MS) active lesions [40, 41]. Niederer et al. also reported that miR-34a is downregulated in patients with RA and OA’s synovium [42]. In our study, we found decreased miR-34a expression in FMF patients when compared to healthy controls.
miR-132 is an miRNA which has received very little attention in the literature in terms of connective tissue disorders. Of its association with inflammation, it is known to offer anti-inflammatory effects through the TLR4-NFkB-TNF-a/IL1B signaling pathway. Murata et al. showed that miR-132 expression had decreased in the plasma samples of OA patients. [43]. Decreased miR-132 expression levels of FMF patients in our study, therefore, may be attributed to the anti-inflammatory reducing effect of this miRNA.
Previous studies have shown that miR-26a is an IL-6 related miRNA. IL-6 is responsible for the regulation of Th17/Treg balance and is believed to be responsible for relapse in MS patients. Zhang et al. emphasized that the reduction of miR-26a expression in MS patients, when compared to remittent and healthy subjects, is inversely linked to disease activation [43]. Increased miR-26a expression in the colchicine users of our study supports these findings. Because miR-26a expression had decreased in our patient group, it can be speculated that this miRNAs causes inflammation to be triggered through different pathways.
It has been reported by Zu et al. that miR-23b inhibits TNFα, IL-17, and NF-KB, while at the same time being downregulated in the inflammatory lesions of SLE and RA patients [44]. In our study, there was no significant expression difference between the patient and control group. The expression of miR-23b, however, was found to have increased in patients who were using colchicine when compared to patients who were not.
Our study is one of the pioneer studies that comprehensively investigate the miRNA association with FMF. There are only two studies related with FMF and circulating miRNAs published so far. Recently, Latsoudis et al. showed the expression change of miR-4520a in FMF patients and they suggested that RHEB which has an important role in FMF by being activator of mTOR signaling is a target of this miRNA. This result supported the view of miRNAs’ interaction in FMF etiopathogenesis [45]. In the other study related to the subject, the scientist showed that expression patterns of circulating miRNAs differ among FMF subgroups based on MEFV mutations between FMF episodes [46].
The biggest limitation in our study was the limited number of available patients. It is known that inflammation is more severe during the attack period. There is a need to increase the number of patients in order to better understand the association of miRNA within the attack period.
Conclusion
The lack of comparable studies investigating miRNA associations with FMF disease has also limited our ability to fully interpret our data. However, the number of studies related to the impact of miRNA’s on collagen tissue disorders has increased rapidly in recent years and there are positive developments in the use of these molecules in the diagnosis, monitoring, and treatment of diseases. The importance of miRNA’s in the pathogenesis of disease is slowly being realized but the journey is only just beginning.
To better understand the relationship of these miRNAs with FMF in terms of pathogenesis and phenotype/genotype correlation, further in depth studies must be added to the literature.
Change history
07 January 2019
The name of the last author of this article was incorrectly presented as “Cogulu Ozgur” this should have been “Ozgur Cogulu”.
References
Peru H, Altun B, Doğan M, Kara F, Elmaci AM, Oran B (2008) The evaluation of carotid intima-media thickness in children with familial Mediterranean fever. Clin Rheumatol 27(6):689–694. https://doi.org/10.1007/s10067-007-0764-1
Zadeh N, Getzug T, Grody WW (2011) Diagnosis and management of familial Mediterranean fever: integrating medical genetics in a dedicated interdisciplinary clinic. Genet Med 13(3):263–269. https://doi.org/10.1097/GIM.0b013e31820e27b1
Papin S, Cuenin S, Agostini L, Martinon F, Werner S, Beer HD, Grütter C, Grütter M, Tschopp J (2007) The SPRY domain of Pyrin, mutated in familial Mediterranean fever patients, interacts with inflammasome components and inhibits proIL-1beta processing. Cell Death Differ 14(8):1457–1466. https://doi.org/10.1038/sj.cdd.4402142
Chae JJ, Cho YH, Lee GS, Cheng J, Liu PP, Feigenbaum L et al (2011) Gain-of-function pyrin mutations induce NLRP3 protein-independent interleukin-1β activation and severe autoinflammation in mice. Immunity 34(5):755–768
de Torre-Minguela C, Mesa Del Castillo P, Pelegrín P (2017) The NLRP3 and pyrin inflammasomes: implications in the pathophysiology of autoinflammatory diseases. Front Immunol 8:43
Ben-Chetrit E, Levy M (1998) Familial Mediterranean fever. Lancet 351(9103):659–364. https://doi.org/10.1016/S0140-6736(97)09408-7
Samuels J, Aksentijevich I, Torosyan Y, Centola M, Deng Z, Sood R, Kastner DL (1998) Familial Mediterranean fever at the millennium. Clinical spectrum, ancient mutations and a survey of 100 American referrals to the National Institutes of Health. Medicine 77(4):268–297. https://doi.org/10.1097/00005792-199807000-00005
Heller H, Shoar E, Gafni J, Heller J (1961) Amyloidosis in FMF: an independent genetically determined character. Arch Intern Med 107(4):539–550. https://doi.org/10.1001/archinte.1961.03620040065007
Rogers DB, Shohat M, Petersen CM, Bickal J, Congleton J, Schwabe AD et al (1989) Familial Mediterranean fever in Armenians: autosomal recessive inheritance with high gene frequency. Am J Med Genet 34(2):168–172. https://doi.org/10.1002/ajmg.1320340206
Ben-Chetrit E, Backenroth R (2001) Amyloidosis induced, end stage renal disease in patients with familial Mediterranean fever is highly associated with point mutations in the MEFV gene. Ann Rheum Dis 60(2):146–149. https://doi.org/10.1136/ard.60.2.146
Mansour I, Delague V, Cazeneuve C, Dodé C, Chouery E, Pêcheux C, Medlej-Hashim M, Salem N, ElZein L, Levan-Petit I, Lefranc G, Goossens M, Delpech M, Amselem S, Loiselet J, Grateau G, Mégarbane A, Naman R (2001) Familial Mediterranean fever in Lebanon: mutation spectrum, evidence for cases in Maronites, Greek orthodoxes, Greek catholics, Syriacs and Chiites and for an association between amyloidosis and M694V and M694I mutations. Eur J Hum Genet 9(1):51–55. https://doi.org/10.1038/sj.ejhg.5200574
Tekin M, Yalcinkaya F, Cakar N, Akar N, Misirlioglu M, Tastan H et al (2000) MEFV mutations in multiplex families with familial Mediterranean fever: is a particular genotype necessary for amyloidosis? Clin Genet 57(6):430–434
Daniel LK (1997) İntermittent and periodic arthritic syndromes. In: Kopman JK (ed) Arthritis and allied conditions: a textbook of rheumatology, 13 edn. Williams and Wilkins Comp., Philadelphia, pp 1279–1306
Baykal Y, Saglam K, Yilmaz MI, Taslipinar A, Akinci SB, Inal A et al (2003) Serum sIL-2r, IL-6, IL-10 and TNF-alpha level in familial Mediterranean fever patients. Clin Rheumatol 22(2):99–101. https://doi.org/10.1007/s10067-002-0682-1
Korkmaz C, Ozdogan H, Kasapcopur O, Yazıcı H (2002) Acute phase response in familial Mediterranean fever. Ann Rheum Dis 61(1):79–81. https://doi.org/10.1136/ard.61.1.79
Ben-Cherit E, Bergmann S, Sood R (2006) Mechanism of the antiinflammatory effect of colchicine in rheumatic diseases :a possible new outlook through microarray analysis. Rheumatology 45(3):274–282. https://doi.org/10.1093/rheumatology/kei140
Álvarez-Errico D, Vento-Tormo R, Ballestar E (2017) Genetic and epigenetic determinants in autoinflammatory diseases. Front Immunol 22(8):318
Bartel DP (2004) MiRNAs: genomics, biogenesis, mechanism, and function. Cell 116(2):281–297. https://doi.org/10.1016/S0092-8674(04)00045-5
Lee RC, Feinbaum RL, Ambros V (1993) The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 75(5):843–854. https://doi.org/10.1016/0092-8674(93)90529-Y
Jackson RJ, Standart N (2007) How do miRNAs regulate gene expression? Sci STKE 1:367
Cobb BS, Hertweck A, Smith J, O'Connor E, Graf D, Cook T, Smale ST, Sakaguchi S, Livesey FJ, Fisher AG, Merkenschlager M (2006) A role for Dicer in immune regulation. J Exp Med 203(11):2519–2527. https://doi.org/10.1084/jem.20061692
Muljo SA, Ansel KM, Kanellopoulou C, Livingston DM, Rao A, Rajewsky K (2005) Aberrant T cell differentiation in the absence of Dicer. J Exp Med 202(2):261–269. https://doi.org/10.1084/jem.20050678
Livneh A, Langevitz P, Zemer D, Zaks N, Kees S, Lidar T, Migdal A, Padeh S, Pras M (1997) Criteria for the diagnosis of familial Mediterranean fever. Arthritis Rheum 40(10):1879–1885. https://doi.org/10.1002/art.1780401023
Tone Y, Toma T, Toga A, Sakakibara Y, Wada T, Yabe M, Kusafuka H, Yachie A (2012) Enhanced exon 2 skipping caused by c.910G>A variant and alternative splicing of MEFV genes in two independent cases of familial Mediterranean fever. Mod Rheumatol 22(1):45–51. https://doi.org/10.3109/s10165-011-0461-4
Jung SH, Young SS (2012) Power and sample size calculation for microarray studies. J Biopharm Stat 22(1):30–42. https://doi.org/10.1080/10543406.2010.500066
Stanczyk J, Pedrioli DM, Brentano F, Sanchez-Pernaute O, Kolling C, Gay RE et al (2008) Altered expression of MiRNA in synovial fibroblasts and synovial tissue in rheumatoid arthritis. Arthritis Rheum 58(4):1001–1009. https://doi.org/10.1002/art.23386
Faraoni I, Antonetti FR, Cardone J, Bonmassar E (2009) miR-155 gene: a typical multifunctional microRNA. Biochim Biophys Acta 1792(6):497–505. https://doi.org/10.1016/j.bbadis.2009.02.013
Pauley KM, Satoh M, Chan AL, Bubb MR, Reeves WH, Chan EK (2008) Upregulated miR-146a expression in peripheral blood mononuclear cells from rheumatoid arthritis patients. Arthritis Res Ther 10(4):R101. https://doi.org/10.1186/ar2493
Li X, Kroin JS, Kc R, Gibson G, Chen D, Corbett GT, Pahan K, Fayyaz S, Kim JS, van Wijnen AJ, Suh J, Kim SG, Im HJ (2013) Altered spinal miRNA-146a and the miRNA-183 cluster contribute to osteoarthritic pain in knee joints. J Bone Miner Res 28(12):2512–2522. https://doi.org/10.1002/jbmr.2002
Tang Y, Luo X, Cui H, Ni X, Yuan M, Guo Y, Huang X, Zhou H, de Vries N, Tak PP, Chen S, Shen N (2009) MiRNA-146A contributes to abnormal activation of the type I interferon pathway in human lupus by targeting the key signaling proteins. Arthritis Rheum 60(4):1065–1075. https://doi.org/10.1002/art.24436
Wang G, Tam LS, Li EK, Kwan BC, Chow KM, Luk CC et al (2010) Serum and urinary cell-free MiR-146a and MiR-155 in patients with systemic lupus erythematosus. J Rheumatol 37(12):2516–2522. https://doi.org/10.3899/jrheum.100308
Bauernfeind F, Rieger A, Schildberg FA, Knolle PA, Schmid-Burgk JL, Hornung V (2012) NLRP3 inflammasome activity is negatively controlled by miR-223. J Immunol 189(8):4175–4181. https://doi.org/10.4049/jimmunol.1201516
Li T, Morgan MJ, Choksi S, Zhang Y, Kim YS, Liu ZG (2010) MiRNAs modulate the noncanonical transcription factor NF-kappaB pathway by regulating expression of the kinase IKKalpha during macrophage differentiation. Nat Immunol 11(9):799–805. https://doi.org/10.1038/ni.1918
Fulci V, Scappucci G, Sebastiani GD, Giannitti C, Franceschini D, Meloni F, Colombo T, Citarella F, Barnaba V, Minisola G, Galeazzi M, Macino G (2010) miR-223 is overexpressed in T-lymphocytes of patients affected by rheumatoid arthritis. Hum Immunol 71(2):206–211. https://doi.org/10.1016/j.humimm.2009.11.008
Murata K, Yoshitomi H, Tanida S, Ishikawa M, Nishitani K, Ito H et al (2010) Plasma and synovial fluid miRNAs as potential biomarkers of rheumatoid arthritis and osteoarthritis. Arthritis Res Ther 12:R86
Kim SW, Ramasamy K, Bouamar H, Lin AP, Jiang D, Aguiar RC (2012) MiRNAs miR-125a and miR-125b constitutively activate the NF-kappaB pathway by targeting the tumor necrosis factor alphainduced protein 3 (TNFAIP3, A20). Proc Natl Acad Sci U S A 109(20):7865–7870. https://doi.org/10.1073/pnas.1200081109
Zhao X, Tang Y, Qu B, Cui H, Wang S, Wang L, Luo X, Huang X, Li J, Chen S, Shen N (2010) MiRNA-125a contributes to elevated inflammatory chemokine RANTES levels via targeting KLF13 in systemic lupus erythematosus. Arthritis Rheum 62(11):3425–3435. https://doi.org/10.1002/art.27632
Lashine YA, Seoudi AM, Salah S, Abdelaziz AI (2011) Expression signature of miRNA 181-a reveals its crucial role in the pathogenesis of paediatric systemic lupus erythematosus. Clin Exp Rheumatol 29(2):351–357
Okuhara A, Nakasa T, Shibuya H, Niimoto T, Adachi N, Deie M, Ochi M (2012) Changes in miRNA expression in peripheral mononuclear cells according to the progression of osteoarthritis. Mod Rheumatol 22(3):446–457. https://doi.org/10.3109/s10165-011-0536-2
Tufekci KU, Oner MG, Genc S, Genc K (2011) MiRNAs and multiple sclerosis. Autoimmun Dis 2011:1–27. https://doi.org/10.4061/2011/807426
Gandhi R, Healy B, Gholipour T, Egorova S, Musallam A, Shuja M et al (2013) Circulating miRNAs as biomarkers for disease staging in multiple sclerosis. Ann Neurol 73(6):729–740. https://doi.org/10.1002/ana.23880
Niederer F, Trenkmann M, Ospelt C, Karouzakis E, Neidhart M, Stanczyk J, Kolling C, Gay RE, Detmar M, Gay S, Jüngel A, Kyburz D (2012) Down-regulation of miRNA-34a* in rheumatoid arthritis synovial fibroblasts promotes apoptosis resistance. Arthritis Rheum 64(6):1771–1779. https://doi.org/10.1002/art.34334
Zhang R, Tian A, Wang J, Shen X, Qi G, Tang Y (2015) miR26a modulates Th17/Treg balance in the EAE model of multiple sclerosis by targeting IL6. NeuroMolecular Med 17(1):24–34. https://doi.org/10.1007/s12017-014-8335-5
Zhu S, Pan W, Song X, Liu Y, Shao XR, Tang YJ et al (2012) The miRNA miR-23b suppresses IL-17-associated autoimmune inflammation by targeting TAB 2, TAB 3 and IKK-alpha. Nat Med 18(7):1077–1086. https://doi.org/10.1038/nm.2815
Wada T, Toma T, Matsuda Y, Yachie A, Itami S, Taguchi Y et al (2017) Microarray analysis of circulating microRNAs in familial Mediterranean fever. Mod Rheumatol 6:1–18
Latsoudis H, Mashreghi MF, Grün JR, Chang HD, Stuhlmüller B, Repa A, Gergiannaki I, Kabouraki E, Vlachos GS, Häupl T, Radbruch A, Sidiropoulos P, Doukoumetzidis K, Kardassis D, Niewold TB, Boumpas DT, Goulielmos GN (2017) Differential expression of miR-4520a associated with pyrin mutations in familial Mediterranean fever (FMF). J Cell Physiol 232(6):1326–1336. https://doi.org/10.1002/jcp.25602
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The current study received ethical approval from the university’s departmental ethics committee.
Disclosures
None.
Additional information
The original version of this article was revised: The name of the second author of this article was incorrectly presented as “Cogulu Ozgur” this should have been “Ozgur Cogulu".
Rights and permissions
About this article
Cite this article
Hortu, H.O., Karaca, E., Sozeri, B. et al. Evaluation of the effects of miRNAs in familial Mediterranean fever. Clin Rheumatol 38, 635–643 (2019). https://doi.org/10.1007/s10067-017-3914-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10067-017-3914-0