Abstract
Identification of susceptibility genes in systemic lupus erythematosus (SLE) has recently become a topic of interest. The IL-10 promoter contains three single base-pair substitutions at −627C > A, −854C > T and −1117G > A. These single base-pair substitutions produce three different haplotypes, GCC, ACC and ATA, which affect IL-10 expression. We examined the distribution of −627C > A, −854C > T and −1117G > A IL-10 promoter polymorphisms in patients with SLE (n = 103, women only) and matched controls (n = 300). Despite the higher prevalence of the GCC/GCC, GCC/ATA and ATA/ATA genotypes in SLE patients than in controls, we observed that only GCC/GCC genotype frequency distribution was significant between these groups. We observed that women with the GCC/GCC genotype displayed an approximately twofold increased risk of SLE OR = 2.245 (95% CI = 1.354–3.721, P = 0.0022). We did not find any associations between various genotypes of IL-10 promoter haplotypes and clinical manifestations or autoantibody production in patients with SLE. Our observations indicate that the GCC/GCC promoter genotype may contribute to SLE incidence in Polish patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Systemic lupus erythematosus (SLE) is an autoimmune disease that affects virtually every organ [1, 2]. SLE is accompanied by an abundant production of antibodies directed against self-antigens and deposition of immune complexes in multiple organs [1, 2]. Immune cells from patients with SLE display various aberrations including abnormal cytokine production, reduction of cytotoxic T cell function and enhancement of humoral response [3, 4].
Despite intensive research in recent years, the etiology of SLE is still unknown. The identification of susceptibility genes has recently become a hot topic. Many genes encoding proteins relevant to the immune system or genes encoding proteins affecting SLE manifestations have been linked to the disease as candidate susceptibility genes [5–9].
It has been reported that interleukin 10 (IL-10) is biosynthesized in high amounts in the B cells and monocytes of patients with SLE [10]. Moreover, the abundant biosynthesis of IL-10 affects the abnormal biosynthesis of autoantibodies that occurs in patients with SLE [10]. IL-10 is an anti-inflammatory cytokine that inhibits the production of proinflammatory cytokines in activated macrophages [11].
It has been reported that IL-10 biosynthesis is regulated at a transcriptional level; so, polymorphic forms of the promoter may alter binding transcription factors and affect the activity of the IL-10 promoter [12–14]. The IL-10 promoter contains three single base-pair substitutions at −627C > A, −854C > T and −1117G > A, which produce the three different haplotypes GCC, ACC and ATA [13, 15]. The haplotypes GCC, ACC and ATA were associated with high, intermediate and low IL-10 production, respectively, according to studies of this promoter’s variants [16, 17]. Numerous investigations have been undertaken to determine the association of different IL-10 promoter variants with SLE. Moreover, contribution of the IL-10 promoter variants in the development of SLE and the promoter’s association to several clinical features have been inconsistent [18–29]. We examined the distribution of −627C > A, −854C > T and −1117G > A IL-10 promoter polymorphic variants in Polish patients with SLE.
Patients and methods
Patients and controls
One hundred and three patients (women only) fulfilling the American College of Rheumatology Classification (ACRC) criteria for SLE [30, 31] were chosen for investigation at the Institute of Rheumatology, Warsaw, Poland. The controls included 300 healthy women. No male patients were available for the study. The protocol of the study was approved by the Local Ethical Committee of Poznań University of Medical Sciences. Written consent was obtained from all participating subjects. Both patients and control groups were of Polish Caucasian origin. The mean age of patients with SLE at diagnosis was 37 ± 12 years, and of controls, 34 ± 13 years.
Clinical symptoms in the patient group are found in the central nervous system (20%), as well as having vascular (13%), renal (51%), musculoskeletal (63%), serosal (17%), dermal (54%), immunologic (24%), constitutional (fever) (10%) and hematologic (29%) components.
Genotyping
DNA was isolated from peripheral blood lymphocytes by salt extraction. Polymorphic variants of −627 IL-10 promoter were identified using PCR and the primer pairs 5′ACCTGACTAGCATATAAGAAGC3′ and 5′AGCCACAATCAAGGTTTCCC3′; enzyme digestion followed the identification process [18]. The PCR-amplified fragments of the IL-10 promoter that were 669 base pairs (bp) in length were subjected to digestion with RsaI (GT/AC). The −627A allele was cleaved into 503-bp and 166-bp fragments, whereas the −627C allele remained uncut. DNA fragments were separated by electrophoresis on 2.5% agarose gel and visualized by ethidium bromide staining [18]. Confirmation of −627C > A polymorphism was performed by sequencing analysis. The 854C > T and −1117G > A IL-10 promoter polymorphic variants were identified using PCR with the primer pair 5′AACACTCCTCGCCGCAACC3′ and 5′CCCCTACCGTCTCTATTTTAT3′, followed by direct sequencing analysis of PCR-amplified IL-10 promoter fragments of 430 bp.
Statistical analysis
The distribution of genotypes in all groups was tested for deviation from Hardy–Weinberg equilibrium. Fisher exact test was used to determine differences in the genotypic and allelic distribution between patients and controls. Moreover, the odds ratio (OR) and 95% confidence intervals (CI) were calculated. A P value <0.05 was considered statistically significant. Power analysis was performed using uncorrected chi-square test available from an on-line internet service, http://biostat.mc.vanderbilt.edu/twiki/bin/view/Main/PowerSampleSize.
Results
Distribution of −627C > A, −854C > T and −1117G > A IL-10 promoter polymorphisms in patients with SLE
Genotype analysis of IL-10 promoter polymorphisms did not show a significant deviation from Hardy–Weinberg equilibrium in any group. The frequency of the −627A and −854T alleles was higher in patients with SLE compared to controls and reached 26 and 24%, respectively, in these groups (Table 1). The homozygous −627CC and −854CC genotype frequency was lower in patients than in controls (Table 1). However, the frequency of the homozygous −627AA and −854TT genotypes was slightly higher in patients with SLE (Table 1).
The −1117G IL-10 allele frequency was significantly higher in patients than in controls (P = 0.0035) and amounted 55 and 43%, respectively (Table 1). We also found a higher distribution of homozygous −1117GG in patients compared to controls. The frequency of homozygous −1117GG reached 33 and 18% in these groups, respectively (Table 1). However, the heterozygous −1117GA and homozygous −1117AA prevalence was lower in patients than in controls (Table 1).
Genotype distribution of promoter’s haplotype in patients with SLE and controls
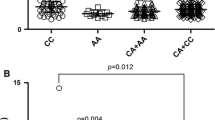
The genotypes GCC/GCC, GCC/ATA and ATA/ATA exhibited higher prevalence in patients with SLE than in healthy individuals (Table 2). However, only the GCC/GCC genotype frequency distribution was significant between these groups (Table 2). OR for patients with SLE with the GCC/GCC genotype was OR = 2.245 (95% CI = 1.354–3.721, P = 0.0022) (Table 2). We did not find any associations between various genotypes of the IL-10 promoter haplotype with clinical manifestations or autoantibody production in patients with SLE. The statistical power of this study amounted to 87% for GCC/GCC genotypes.
Discussion
IL-10 is a significant cytokine that suppresses both immunoproliferative and inflammatory responses [32]. The IL-10 gene is localized to the junction of 1q31–q32 [15]. IL-10 is a 36-kDa homodimeric cytokine and is mainly biosynthesized by macrophages, monocytes and lymphocytes [10, 32]. It has been reported that IL-10 production after lipopolysaccharide whole blood stimulation differs significantly between humans as a result of genetic variations acting at the transcriptional level [12, 33].
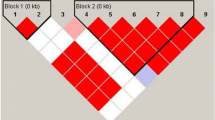
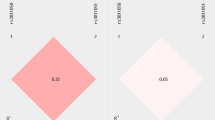
The nucleotides A and G are present at the IL-10 promoter in similar frequencies at position −1,117, while a C → A substitution at position −627 is present in 21–23% of the population and is in complete linkage disequilibrium with C → T at position −854 [13, 15]. Investigation of IL-10 expression in various diseases revealed an increased IL-10 production in individuals bearing the GCC haplotype [12, 16, 34]. Transient transfection studies have also confirmed higher transcriptional promoter activity for the GCC haplotype as compared to the ATA IL-10 promoter haplotype [17].
We observed that the homozygous GCC haplotype increased the risk of SLE development approximately twofold; however, we did not observe an association of this variant or other promoter variants with clinical manifestations of SLE or with autoantibody production.
Our results are different from findings in other populations showing lack of promoter variant association with SLE incidence or ATA haplotype contribution to SLE [18, 20–24, 35, 36]. Lin et al. reported that the −627A IL-10 genotype and allelic frequency was significantly increased in patients with SLE compared to controls [18]. In Chinese patients, the ATA haplotype was associated with kidney disorders but not with autoantibody production [24]. Rood et al. observed a significant association between the ATA haplotype and neuropsychiatric manifestations in patients with SLE [23].
However, our results were consistent with observations in other investigations suggesting GCC haplotype contribution to SLE [19, 26, 28, 29]. Khoa et al. found that the allele frequency of −1117G in patients with SLE was significantly higher than that in healthy controls [19]. Morover, Chung et al. found that the homozygous GCC haplotype was associated with greater SLE severity [28]. Recently, Rosado et al. have suggested that the IL-10 promoter haplotype that biosynthesizes higher levels of IL-10 is linked with the severity of SLE in patients [29]. Lazarus et al. observed that only the IL-10 promoter haplotype GCC was linked with renal disease and Ro autoantibody production in Caucasian patients with SLE [25].
Since the GCC haplotype of the IL-10 promoter is associated with high transcriptional activity [12, 15, 17], we can assume that a high production of IL-10 may make an individual more susceptible to an abundant production of autoantibodies during inflammatory processes, which may favor SLE incidence.
The differences in the effects of IL-10 promoter variants on incidence or clinical manifestations of SLE may be due to distinct racial structure and environmental factors acting on investigated populations. It is possible that different polymorphisms in linkage disequilibrium with the investigated IL-10 promoter variant may contribute to SLE incidence. To determine more precisely the associations of the IL-10 promoter genotype with patients with SLE, further examination of these variants’ distribution in other populations are needed.
References
Herrmann M, Winkler T, Gaipl U, Lorenz H, Geiler T, Kalden JR (2000) Etiopathogenesis of systemic lupus erythematosus. Int Arch Allergy Immunol 123:28–35. doi:10.1159/000024421
Sekigawa I, Naito T, Hira K, Mitsuishi K, Ogasawara H, Hashimoto H et al (2004) Possible mechanisms of gender bias in SLE: a new hypothesis involving a comparison of SLE with atopy. Lupus 13:217–222. doi:10.1191/0961203304lu1012ed
Krishnan S, Farber DL, Tsokos GC (2003) T cell rewiring in differentiation and disease. J Immunol 171:3325–3331
Januchowski R, Wudarski M, Chwalińska-Sadowska H, Jagodzinski PP (2008) Prevalence of ZAP-70, LAT, SLP-76, and DNA methyltransferase 1 expression in CD4(+) T cells of patients with systemic lupus erythematosus. Clin Rheumatol 27:21–27. doi:10.1007/s10067-007-0644-8
Wong M, Tsao BP (2006) Current topics in human SLE genetics. Springer Semin Immunopathol 28:97–107. doi:10.1007/s00281-006-0031-6
Harley JB, Kelly JA, Kaufman KM (2006) Unraveling the genetics of systemic lupus erythematosus. Springer Semin Immunopathol 28:119–130. doi:10.1007/s00281-006-0040-5
Piotrowski PC, Duriagin S, Jagodzinski P (2005) Expression of human endogenous retrovirus clone 4-1 may correlate with blood plasma concentration of anti-U1 RNP and anti-Sm nuclear antibodies. Clin Rheumatol 24:620–624. doi:10.1007/s10067-005-1123-8
Al-Awadhi AM, Haider MZ, Sharma PN, Hasan EA, Botaiban F, Al-Herz A et al (2007) Angiotensin-converting enzyme gene polymorphism in Kuwaiti patients with systemic lupus erythematosus. Clin Exp Rheumatol 25:437–442
Burzynski M, Duriagin S, Mostowska M, Wudarski M, Chwalinska-Sadowska H, Jagodzinski P (2007) MTR 2756 A > G polymorphism is associated with the risk of systemic lupus erythematosus in the Polish population. Lupus 16:450–454. doi:10.1177/0961203307077988
Llorente L, Zou W, Levy Y, Richaud-Patin Y, Wijdenes J, Alcocer-Varela J et al (1995) Role of interleukin 10 in the B lymphocyte hyperactivity and autoantibody production of human systemic lupus erythematosus. J Exp Med 181:839–844. doi:10.1084/jem.181.3.839
Chernoff AE, Granowitz EV, Shapiro L, Vannier E, Lonnemann G, Angel JB et al (1995) A randomized, controlled trial of IL-10 in humans. Inhibition of inflammatory cytokine production and immune responses. J Immunol 154:5492–5499
Lim S, Crawley E, Woo P, Barnes PJ (1998) Haplotype associated with low interleukin-10 production in patients with severe asthma. Lancet 352:113. doi:10.1016/S0140-6736(98)85018-6
Turner DM, Williams DM, Sankaran D, Lazarus M, Sinnott PJ, Hutchinson IV (1997) An investigation of polymorphism in the interleukin-10 gene promoter. Eur J Immunogenet 24:1–8
Kube D, Platzer C, von Knethen A, Straub H, Bohlen H, Hafner M et al (1995) Isolation of the human interleukin 10 promoter. Characterization of the promoter activity in Burkitt’s lymphoma cell lines. Cytokine 7:1–7. doi:10.1006/cyto.1995.1001
Eskdale J, Kube D, Tesch H, Gallagher G (1997) Mapping of the human IL10 gene and further characterization of the 5′ flanking sequence. Immunogenetics 46:120–128. doi:10.1007/s002510050250
Edwards-Smith CJ, Jonsson JR, Purdie DM, Bansal A, Shorthouse C, Powell EE (1999) Interleukin-10 promoter polymorphism predicts initial response of chronic hepatitis C to interferon alfa. Hepatology 30:526–530. doi:10.1002/hep.510300207
Crawley E, Kay R, Sillibourne J, Patel P, Hutchinson I, Woo P (1999) Polymorphic haplotypes of the interleukin-10 5′ flanking region determine variable interleukin-10 transcription and are associated with particular phenotypes of juvenile rheumatoid arthritis. Arthritis Rheum 42:1101–1108 doi:10.1002/1529-0131(199906)42:6<1101::AID-ANR6>3.0.CO;2-Y
Lin PW, Huang CM, Huang CC, Tsai CH, Tsai JJ, Chang CP et al (2007) The association of −627 interleukin-10 promoter polymorphism in Chinese patients with systemic lupus erythematosus. Clin Rheumatol 26:298–301. doi:10.1007/s10067-006-0329-8
Khoa PD, Sugiyama T, Yokochi T (2005) Polymorphism of interleukin-10 promoter and tumor necrosis factor receptor II in Vietnamese patients with systemic lupus erythematosus. Clin Rheumatol 24:11–13. doi:10.1007/s10067-004-0952-1
Van der Linden MW, Westendorp RG, Sturk A, Bergman W, Huizinga TW (2000) High interleukin-10 production in first-degree relatives of patients with generalized but not cutaneous lupus erythematosus. J Investig Med 48:327–334
Dijstelbloem HM, Hepkema BG, Kallenberg CG, van der Linden MW, Keijsers V, Huizinga TW et al (2002) The R-H polymorphism of FCgamma receptor IIa as a risk factor for systemic lupus erythematosus is independent of single-nucleotide polymorphisms in the interleukin-10 gene promoter. Arthritis Rheum 46:1125–1126. doi:10.1002/art.518
Alarcón-Riquelme ME, Lindqvist AK, Jonasson I, Johanneson B, Sandino S, Alcocer-Varela J et al (1999) Genetic analysis of the contribution of IL10 to systemic lupus erythematosus. J Rheumatol 26:2148–2152
Rood MJ, Keijsers V, van der Linden MW, Tong TQ, Borggreve SE, Verweij CL et al (1999) Neuropsychiatric systemic lupus erythematosus is associated with imbalance in interleukin 10 promoter haplotypes. Ann Rheum Dis 58:85–89
Mok CC, Lanchbury JS, Chan DW, Lau CS (1998) Interleukin-10 promoter polymorphisms in Southern Chinese patients with systemic lupus erythematosus. Arthritis Rheum 41:1090–1095. doi:10.1002/1529-0131(199806)41:6<1090::AID-ART16>3.0.CO;2-6
Lazarus M, Hajeer AH, Turner D, Sinnott P, Worthington J, Ollier WE et al (1997) Genetic variation in the interleukin 10 gene promoter and systemic lupus erythematosus. J Rheumatol 24:2314–2317
Chong WP, Ip WK, Wong WH, Lau CS, Chan TM, Lau YL (2004) Association of interleukin-10 promoter polymorphisms with systemic lupus erythematosus. Genes Immun 5:484–492. doi:10.1038/sj.gene.6364119
Hirankarn N, Wongpiyabovorn J, Hanvivatvong O, Netsawang J, Akkasilpa S, Wongchinsri J et al (2006) The synergistic effect of FC gamma receptor IIa and interleukin-10 genes on the risk to develop systemic lupus erythematosus in Thai population. Tissue Antigens 68:399–406. doi:10.1111/j.1399-0039.2006.00681.x
Chung EY, Liu J, Zhang Y, Ma X (2007) Differential expression in lupus-associated IL-10 promoter single-nucleotide polymorphisms is mediated by poly(ADP-ribose) polymerase-1. Genes Immun 8:577–589. doi:10.1038/sj.gene.6364420
Rosado S, Rua-Figueroa I, Vargas JA, Garcia-Laorden MI, Losada-Fernandez I, Martin-Donaire T et al (2008) Interleukin-10 promoter polymorphisms in patients with systemic lupus erythematosus from the Canary Islands. Int J Immunogenet 35:235–242. doi:10.1111/j.1744-313X.2008.00762.x
Tan EM, Cohen AS, Fries JF, Masi AT, McShane DJ, Rothfield NF et al (1982) The 1982 revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum 25:1271. doi:10.1002/art.1780251101
Hochberg MC (1997) Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum 40:1725. doi:10.1002/art.1780400928
Moore KW, O’Garra A, de Waal Malefyt R, Vieira P, Mosmann TR (1993) Interleukin-10. Annu Rev Immunol 11:165–190. doi:10.1146/annurev.iy.11.040193.001121
Westendorp RG, Langermans JA, Huizinga TW, Elouali AH, Verweij CL, Boomsma DI et al (1997) Genetic influence on cytokine production and fatal meningococcal disease. Lancet 349:170–173. doi:10.1016/S0140-6736(96)06413-6
Rosenwasser LJ, Borish L (1997) Genetics of atopy and asthma: the rationale behind promoter-based candidate gene studies (IL-4 and IL-10). Am J Respir Crit Care Med 156:152–155
Crawley E, Woo P, Isenberg DA (1999) Single nucleotide polymorphic haplotypes of the interleukin-10 5’ flanking region are not associated with renal disease or serology in Caucasian patients with systemic lupus erythematosus. Arthritis Rheum 42:2017–2018. doi:10.1002/1529-0131(199909)42:9<2017::AID-ANR34>3.0.CO;2-I
Guseva IA, Omarbekova ZE, Miakotkin VA (2003) Polymorphism of Fc gamma RIIIA-158F/V gene and promoter region of IL-10 gene in systemic lupus erythematosus in Kazakhs. Ter Arkh 75:36–41
Acknowledgments
This study was supported by Poznan University of Medical Sciences (grant number 502-01-01124182-07474).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sobkowiak, A., Lianeri, M., Wudarski, M. et al. Genetic variation in the interleukin-10 gene promoter in Polish patients with systemic lupus erythematosus. Rheumatol Int 29, 921–925 (2009). https://doi.org/10.1007/s00296-008-0776-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00296-008-0776-4