Abstract
Systemic lupus erythematosus (SLE) is the prototype autoimmune rheumatic disease. The etiology of this disease is incompletely understood; however, environmental factors and genetic predisposition are involved. Cytokine-mediated immunity plays a crucial role in the pathogenesis of SLE. We investigate the association of interleukin-10 (IL-10) promoter polymorphisms and their haplotypes in SLE patients from the western Mexico. One hundred and twenty-five SLE patients fulfilling the 1997 ACR criteria and 260 unrelated healthy subjects (HS), both Mexican mestizos, were genotyped for IL-10 −1082A>G, −819C>T, and −592C>A polymorphisms. Haplotypes were inferred using the expectation–maximization algorithm, then allele and haplotype distributions were compared between patients and HS, as well as patients with different clinical variables. We identified at −1082, −819, and −592 four predominant haplotypes ACC (43.70 % in patients vs 46.55 % in HS), ATA (21.45 vs 22.97 %), GCC (16.28 vs 14.21 %), and GTA (14.12 vs 14.12 %). The ATC haplotype was more frequent in SLE respect to HS, suggesting a risk effect (3.23 vs 1.05 %; OR 3.55, CI 1.14–11.11; p = 0.0293). SLE patient carriers of −592 CC genotype as well as the dominant model of inheritance showed higher sIL-10 respect to AA genotype, suggesting that −592 C allele is associated with increased production of the cytokine (p < 0.05). The ACC haplotype had higher IL-10 serum levels and higher values of Mexican version of the Systemic Lupus Erythematosus Disease Activity Index compared with the other haplotype carriers; however, no association was found regarding autoantibodies. Our data suggest that the IL-10 promoter haplotypes play an important role in the risk of developing SLE and influence the production of IL-10 in Mexican population. Nevertheless, further studies are required to analyze the expression of mRNA as well as to investigate the interacting epigenetic factors that could help to define the true contribution of this marker in SLE pathogenesis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Systemic lupus erythematosus (SLE) is a prototype autoimmune disease, in which immune regulation is disrupted, and it is characterized by the production of autoantibodies leading to intense inflammation and multiple organ damage. Although the etiopathology of SLE is still not fully clear, genetic susceptibility and environmental factors have been associated with the initiation and promotion of this complex disorder.
It has been suggested that cytokine-mediated immunity plays a crucial role in the pathogenesis of SLE [1, 2]. Interleukin-10 (IL-10) is a multifunctional cytokine that can be produced by almost all leukocytes. It is a potent inhibitor of both antigen-presenting cell and T lymphocyte functions. Moreover, IL-10 is a key mediator in the interferon-γ signaling pathway, thereby inhibits the production of Th1 cytokines and promotes the development of a type 2 cytokines [1, 2]. In addition, IL-10 is defined as a potent stimulator of B lymphocytes and stimulates the production of anti-DNA autoantibodies in SLE patients [3]. The over-production of IL-10 has been observed in SLE patients and in some of their unaffected relatives [4, 5]. Additionally, SLE activity was correlated with serum IL-10 levels, suggesting that overexpression of IL-10 might play a pathogenic role in severe lupus disease [6, 7].
The IL-10 gene is located on chromosome 1 at position 1q31–1q32 [8]. Three single nucleotide polymorphisms (SNPs) at −1082A>G, −819C>T, and −592C>A in the promoter region have been associated with the pathogenesis of SLE. However, a number of studies have been examined the potential contributions of these polymorphisms and their haplotypes with susceptibility to SLE, but these findings are not consistent [9–13], possibly because of the ethnic and racial differences [14–16].
In the present case–control study, we investigated whether there is an association between IL-10 promoter polymorphisms and their haplotypes with susceptibility to SLE in Mexican population.
Materials and methods
Patients
One hundred and twenty-five systemic lupus erythematosus patients, whom fulfilled the 1997 American College of Rheumatology Criteria [17], were recruited from the Rheumatology Department of the Hospital General de Occidente, Zapopan, Jalisco, Mexico. The Mexican version of the Systemic Lupus Erythematosus Disease Activity Index (Mex-SLEDAI) [18] and the Systemic Lupus International Collaborating Clinics index (SLICC) [19] were applied to the SLE patients at the enrollment of the study. Furthermore, the clinical variables and treatment data were taken from clinical records. As a control group, 260 unrelated healthy subjects (HS) were included. SLE patients and HS were from the same ethnical origin region, and we considered as Mexican mestizos only those individuals, who for three generations, including their own, had been born in Mexico having Spanish-derived last name [20]. The institutional Ethics and Research Committees approved the study, and all subjects provided written informed consent.
Laboratory assessments
A complete blood count, including the lymphocyte count, was performed to SLE patients and HS (CELL-DYN 3500R; Abbott Diagnostics, Abbott Park; IL, USA). Double strain DNA (dsDNA), anti-Ro, anti-La, and anti-Sm antibodies were tested in SLE patients using an ELISA test (Binazyme, The Binding Site Ltd.; Birmingham, UK). For dsDNA antibodies, the range of detection was 12.3–1,000 IU/mL, and sensitivity of the assay was 4.6 IU/mL, while for anti-Ro, anti-La, and anti-Sm, the sensitivity of the assay was 4.0 IU/mL. Rheumatoid factor levels (RF) were performed by nephelometry (IMMAGE 4700 Immunochemistry System, Beckman Coulter Inc.).
IL-10 quantification
Soluble interleukin-10 (sIL-10) levels were quantified from sera of SLE patients and HS using a high sensibility ELISA kit (R&D Systems, Minneapolis, MN, USA). The range of detection was 0.78–50 pg/mL, and the sensitivity of the assay was 0.17 pg/mL.
IL-10 polymorphisms genotypification
Genomic DNA was extracted from 3 mL of peripheral blood leukocyte according to Miller’s method [21]. Amplification of interleukin-10 promoter region was performed by PCR in a Techne Thermal Cycler (TC-312; Cambridge, UK) using the following primers, one of them containing a single base-pair mismatch (underlined) in order to introduce a restriction site: for IL-10 −1082A>G polymorphism: 5′-AACACTACTAAGGCTCCTTTGGGA-3′ (Forward) and 5′-CAAGGAAAAGAAGTCAGGATTCCATGGA-3′ (Reverse) [22], for IL-10 −819C>T polymorphism: 5′-CTACTAAGGCTTCTTTGGG-3′ (Forward) and 5′-AGGTAGTGCTCACCATGACC-3′ (Reverse) [23], and for IL-10 −519C>A polymorphism: 5′-ATCCAAGACAACACTACTAA-3′ (Forward) and 5′-TAAATATCCTCAAAGTTCC-3′ (Reverse) [11]. The PCR was carried out in a final volume of 25 μL containing 500 ng of DNA, 2.5 μM of each primer, 0.5 U/μL Taq DNA polymerase, 1X of supplied 10X buffer enzyme, 2.5 mM MgCl2, and 2.5 mM of each dNTP (Invitrogen™; Carlsbad, CA, USA). The PCR was performed by initial denaturation at 94 °C during 3 min, followed by 35 cycles of amplification at: 94 °C during 30 s for denaturation (95 °C for IL-10 −592C>A), 60 °C during 30 s for annealing (56 °C for IL-10 −592C>A), and 72 °C during 30 s for extension. Finally, 72 °C during 1 min for ending extension was applied. In order to identify each polymorphism, 5 μl of PCR product was digested with three units of EcoNI (New England Biolabs; Beverly, MA, USA) (IL-10 −1082A>G), MsII (New England Biolabs Beverly; MA, USA) (IL-10 −819C>T), and RsaI (New England Biolabs; Beverly, MA, USA) (IL-10 −592C>A) for 3 h at 37 °C. Digested products were observed on a 6 % polyacrylamide gel stained with 2 % AgNO3. For IL-10 −1082A>G polymorphism, AA homozygote was identified by 102 bp fragment, GG homozygote by 82 and 20 bp, and AG heterozygote by 102, 80 and 20 bp fragments. The genotypes for IL-10 −819C>T polymorphism were identified by the next fragments: CC homozygote 288 and 60 bp, TT homozygote 348 bp, and CT heterozygote by 348, 288 and 60 bp. Respect to IL-10 −592C>A polymorphism, fragments of 306, 232, 42, and 8 bp correspond to CC homozygote; 240, 232, 66, 42 and 8 bp identify the AA homozygote, and the CA heterozygote was genotyped by 306, 240, 232, 66, 42 and 8 bp fragments.
Statistical analysis
Statistical analysis was performed using PASW Statistics 18 software (IBM Corporation; Armonk, NY) and GraphPad Prism 5 software (GraphPad Software Inc.; San Diego, CA). Allele and genotype frequencies of IL-10 polymorphisms were obtained by direct counting. Hardy–Weinberg equilibrium was calculated using chi-square test as well as allele and genotype analysis. Mann–Whitney U test and Kruskal–Wallis analysis of variance were used for nonparametric distribution data. P values <0.05 were considered significant. Haplotypes frequencies were inferred using EMHAPFRE software [24]. Pairwise linkage disequilibrium (LD) was expressed as Lewontin’s D’ corrected coefficient [25]. Comparison data was evaluated by χ2 test or the Fisher exact test when applicable. LD at three loci was evaluated according to Gavrilets [26]. The p corrected (pC) values for multiple tests were adjusted by the Bonferroni test [27].
Results
Demographic and clinical characteristics
One hundred and twenty-five SLE patients were included (5 men and 120 women). The average age was 36 years old (range of 16–72) and 7.5 years of disease duration (0–36 years). The SLE patients presented an activity disease score of 2.50 (0.0–11.0) for Mex-SLEDAI and 0.62 (0.0–12.0) score for SLICC damage index. The main clinical manifestations were hematologic, cutaneous, and neurologic (65, 30, and 29 % respectively). The treatment of SLE patients included immunosuppressive drugs (azathioprine, methotrexate, cyclophosphamide, and mycophenolate), chloroquine, and prednisone (Table 1).
IL-10 −1082A>G, −819C>T, and −592C>A polymorphisms
Analysis of IL-10 −1082A>G, −819C>T, and −592C>A polymorphisms were performed in SLE patients and HS. Observed and expected frequencies were in Hardy–Weinberg equilibrium (p>0.05). The AA genotype was more frequent in SLE patients and HS for IL-10 −1082A>G polymorphism (50 and 51 %, respectively). Respect to IL-10 −819C>T polymorphisms, the heterozygote genotype (CT) was more frequent in both SLE and HS (49 and 48 %, respectively). Finally, for IL-10 −592C>A polymorphism, the heterozygote genotype (CA) was also more frequent in SLE patients and HS (48 %). The more frequent allele was the wild in both SLE patients and HS in the three IL-10 polymorphisms analyzed. No statistical significant differences were observed between SLE patients and HS, even when IL-10 polymorphisms were analyzed by dominant or recessive models of inheritance (Table 2).
IL-10 −1082A>G, −819C>T, and −592C>A haplotypes
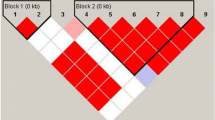
A strong linkage disequilibrium (LD) was observed for IL-10 −1082A>G, −819C>T, −592C>A haplotypes (100 %, pC=3.673 × 10−18). The most frequent haplotype in SLE and HS was ACC (43.70 vs 46.55 %, respectively). ATA, GCC, and GTA frequencies in SLE were 21.45, 16.28, and 14.12 % while in HS 22.97, 14.21, 14.14 %, respectively. ATC haplotype was more frequent in SLE respect to HS (3.23 vs 1.05 %; OR 3.55, CI 1.14–11.11; p = 0.0293).
IL-10 serum levels
The soluble IL-10 levels showed differences in SLE patients, according to genotypes, dominant, and recessive model of the IL-10 –592C>A polymorphism. SLE patient carriers of CC genotype showed higher sIL-10 levels than AA carriers and also than recessive model (2.71 vs 1.86 pg/mL, p = 0.019; 2.71 vs 2.1 pg/mL, p = 0.011, respectively; Fig. 1a) as well as the dominant model showed higher sIL-10 respect to AA genotype (2.39 vs 1.86 pg/mL, p = 0.041; Fig. 1a). Analyzing sIL-10 levels in relation to haplotype present in SLE patients, ACC carriers showed higher levels respect to GCC, and ATA carriers (3.15 vs 2.27 pg/mL, p = 0.012; 3.15 vs 1.86 pg/mL, p = 0.04, respectively; Fig. 1b, Table 3).
Soluble IL-10 levels in systemic lupus erythematosus patients. a Soluble IL-10 levels according to genotypes, dominant, and recessive model of the IL-10 −592C>A polymorphism in SLE patients. b Soluble IL-10 levels in relation to homozygote haplotypes presents in SLE patients. SLE systemic lupus erythematosus, HS healthy subjects. Statistical analysis was performed using Mann–Whitney U test. Median with interquartile range is represented by horizontal lines and error bars, respectively
IL-10 −1082A>G, −819C>T, −592C>A haplotypes and clinical associations
Systemic lupus erythematosus patients were classified according to IL-10 haplotypes (Table 3). ACC carriers showed higher activity disease (Mex-SLEDAI index) respect to ATA carriers (3.0 vs 1.0, p = 0.03). In addition, ACC carriers presented high levels of sIL-10 (p = 0.04 vs ATA haplotype; 3.15 vs 1.86 pg/mL, respectively). However, patients with ATA haplotype showed high levels of anti-Sm antibodies (p = 0.02 vs ACC carriers; 3.05 vs 0.08 IU/mL, respectively). Whereas, ATC haplotype carriers showed an increase of rheumatoid factor levels (p = 0.02 vs ATA haplotype; 74 vs 12 IU/mL, respectively).
Discussion
IL-10 is a potent stimulator of B cells and a strong inhibitor of antigen-presenting cells and T cells. The aberrant expression of IL-10 contributes to development of autoimmune diseases. Previously, several studies have suggested an association of genetic polymorphisms within the IL-10 promoter with the constitutive cytokine secretion capacity [28, 29].
In the present work, we evaluated whether polymorphisms of IL-10 promoter −1082A>G, −819C>T and −592C>A are involved in susceptibility to SLE in Mexicans. Several case–control studies addressed this question showed diverging results [9–13]. Recent meta-analysis indicates that these differences are due largely to the racial and ethnic origin. Regarding IL-10 −1082A>G polymorphism, the meta-analysis reveals an association between SLE and the IL-10 −1082G allele in Europeans, but not in Asians [14–16].
In a large group of unrelated Mexican SLE patients, no association with the IL-10.G microsatellites was detected, and an analysis of multicase families from Mexico, Sweden, and Iceland yielded no evidence for linkage between the IL-10 gene and SLE [30]. However, to our knowledge there are no studies assessing the association for SNPs at positions −1082, −819, and −592 with SLE in Mexican population. A previous study analyzed the distribution of these SNPs in healthy unrelated Mexican mestizo individuals and shown genotypic and allelic frequencies are similar to those reported in our study [31]. We found no differences when comparing genotypic and allelic frequencies of the selected polymorphisms in 125 SLE patients and 260 unrelated HS, even evaluating genetic models, suggesting that these loci individually analyzed, do not confer susceptibility to SLE in our population.
Furthermore, the haplotype analysis showed strong linkage disequilibrium among the three IL-10 gene polymorphisms, similar to other populations studied [11–13, 32]. In our population besides the three most frequent haplotypes GCC, ACC, and ATA, we found that the haplotype GTA is present in 14 % in both studied groups. The distribution of these frequencies is similar to those reported in previous studies of Mexican, even of Italian population, which is explained by our ethnic origin. Ancestry studies in Mexican mestizos from Western, underscored the predominance of the European male contribution (~60–64 %), followed by Amerindian (~25–21 %), and Africans (~15 %) [33]. In general, the stratification by ethnicity indicate that GCC haplotype shows a significant association with SLE in Europeans whereas ATA haplotype with SLE in Asians [16]. However, in this study we found no differences in SLE haplotype frequencies when compared with healthy subjects, except for increased frequency of ATC haplotype in SLE, suggesting that it is a risk haplotype for developing SLE in Mexican population (OR 3.55, CI 1.14–11.11; p=0.0293). This haplotype includes the −592 C allele associated with disease activity and lupus nephritis in Chinese patients [34].
In addition, we evaluated IL-10 serum levels according to the genotypes and found that carriers of CC genotype as well as the dominant model of inheritance showed higher sIL-10 respect to AA genotype, suggesting that the −592 C allele is associated with increased production of the cytokine; however, no differences with clinical variables or autoantibodies were observed. Furthermore, when performing the analysis regarding haplotypes, we found that ACC haplotype had higher IL-10 serum levels and higher values of Mex-SLEDAI index compared with the other haplotype carriers, but no association was found regarding autoantibodies. Carriers of GCC and ATA haplotypes are considered as high and low producers of IL-10, respectively [13, 16]. However, these findings are inconsistent, possibly due to the complexity of the in vivo condition that could regulate IL-10 gene expression.
The role of IL-10 in the pathogenesis of SLE is unknown, but theoretically, IL-10 could stimulate peripheral blood mononuclear cells of SLE patients to produce autoantibodies, and thus be important in the excessive autoantibody production that characterized to SLE [3]. However, other cytokines have been related with SLE activity and the production of autoantibodies, such as cytokine BAFF which also acts in synergy with IL-17, and both are involved in the pathogenesis of SLE [35, 36]. Concerning environmental factors, it has been suggested that infectious agents and even diet play roles in SLE pathogenesis. It has been hypothesized that Epstein–Barr virus and cytomegalovirus may be involved in SLE. In a previous study published by our group, we found an association between high levels of IgG anti-CMV with production of lupus-related autoantibodies to RNA or DNA-protein complex [37].
On the other hand, the immune dysregulation is an important aspect in SLE: T cell dysfunction is pathophysiologically relevant, and evidence suggests that T cells play an important role. T cells are key modulators of B cell activation and class switching necessary for the production of IgG autoantibodies seen in SLE. Several studies suggest that T cells fail to execute their regulatory functions. The CD4+ CD25+ and CD8+ CD28− T regulatory lymphocytes are important in SLE pathogenesis, since both cells are decreased in SLE patients [38, 39]. Specially, CD8+ T suppressor lymphocytes mediate its function through the secretion of IL-10, which affects naïve CD8 T cells activation [40]. It could be interesting to know if the IL-10 promoter polymorphisms are associated with CD8+ T suppressor lymphocytes, since low IL-10 producer haplotypes may affect their generation or functions. Recently, the expression level of IL-10, CD70 (expressed on activated T cells), and OAS2 (and interferon-inducible gene) genes have shown increased in SLE patients with disease activity, and it has been proposed that can differentiate SLE from healthy individuals and patients with other autoimmune diseases [41, 42].
In summary, our data suggest that the IL-10 promoter haplotypes play an important role in the risk of developing SLE and influence the production of IL-10 in Mexican population. However, further studies are required to analyze the expression levels of specific pathogenesis-linked genes, as well as the interacting epigenetic factors that could help to define the true contribution of these markers in SLE pathogenesis.
References
Kyttaris VC, Juang Y-T, Tsokos GC. Immune cells and cytokines in systemic lupus erythematosus: an update. Curr Opin Rheumatol. 2005;17:518–22.
Su D-L, Lu Z-M, Shen M-N, et al. Roles of pro- and anti-inflammatory cytokines in the pathogenesis of SLE. J Biomed Biotechnol. 2012;2012:1–15.
Llorente L, Zou W, Levy Y, et al. Role of interleukin 10 in the B lymphocyte hyperactivity and autoantibody production of human systemic lupus erythematosus. J Exp Med. 1995;181:839–44.
Llorente L, Richaud-Patin Y, Couderc J, et al. Dysregulation of interleukin-10 production in relatives of patients with systemic lupus erythematosus. Arthritis Rheum. 1997;40:1429–35.
Gröndal G, Kristjansdottir H, Gunnlaugsdottir B, et al. Increased number of interleukin-10-producing cells in systemic lupus erythematosus patients and their first-degree relatives and spouses in Icelandic multicase families. Arthritis Rheum. 1999;42:1649–54.
Gröndal G, Gunnarsson I, Rönnelid J, et al. Cytokine production, serum levels and disease activity in systemic lupus erythematosus. Clin Exp Rheumatol. 2000;18:565–70.
Houssiau FA, Lefebvre C, Vanden Berghe M, et al. Serum interleukin 10 titers in systemic lupus erythematosus reflect disease activity. Lupus. 1995;4:393–5.
Iyer SS, Cheng G. Role of interleukin 10 transcriptional regulation in inflammation and autoimmune disease. Crit Rev Immunol. 2012;32:23–63.
Suárez A, López P, Mozo L, et al. Differential effect of IL10 and TNF{alpha} genotypes on determining susceptibility to discoid and systemic lupus erythematosus. Ann Rheum Dis. 2005;64:1605–10.
Chong WP, Ip WK, Wong WH-S, et al. Association of interleukin-10 promoter polymorphisms with systemic lupus erythematosus. Genes Immun. 2004;5:484–92.
Mok CC, Lanchbury JS, Chan DW, et al. Interleukin-10 promoter polymorphisms in Southern Chinese patients with systemic lupus erythematosus. Arthritis Rheum. 1998;41:1090–5.
Lazarus M, Hajeer AH, Turner D, et al. Genetic variation in the interleukin 10 gene promoter and systemic lupus erythematosus. J Rheumatol. 1997;24:2314–7.
Eskdale J, Wordsworth P, Bowman S, et al. Association between polymorphisms at the human IL-10 locus and systemic lupus erythematosus. Tissue Antigens. 1997;49:635–9.
Zhou M, Ding L, Peng H, et al. Association of the interleukin-10 gene polymorphism (−1082A/G) with systemic lupus erythematosus: a meta-analysis. Lupus. 2013;22:128–35.
Wang B, Zhu J-M, Fan Y-G, et al. Association of the −1082G/A polymorphism in the interleukin-10 gene with systemic lupus erythematosus: a meta-analysis. Gene. 2013;519:209–16.
Song GG, Choi SJ, Ji JD, et al. Associations between interleukin-10 polymorphisms and susceptibility to systemic lupus erythematosus: a meta-analysis. Hum Immunol. 2013;74:364–70.
Hochberg MC. Updating the American College of Rheumatology revised criteria for the classification of systemic lupus erythematosus. Arthritis Rheum. 1997;40:1725.
Guzmán J, Cardiel MH, Arce-Salinas A, et al. Measurement of disease activity in systemic lupus erythematosus. Prospective validation of 3 clinical indices. J Rheumatol. 1992;19:1551–8.
Gladman DD, Urowitz MB, Goldsmith CH, et al. The reliability of the Systemic Lupus International Collaborating Clinics/American College of Rheumatology Damage Index in patients with systemic lupus erythematosus. Arthritis Rheum. 1997;40:809–13.
Gorodezky C, Alaez C, Vázquez-García MN, et al. The genetic structure of Mexican Mestizos of different locations: tracking back their origins through MHC genes, blood group systems, and microsatellites. Hum Immunol. 2001;62:979–91.
Miller SA, Dykes DD, Polesky HF. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 1988;16:1215.
Boiardi L, Casali B, Farnetti E, et al. Interleukin-10 promoter polymorphisms in giant cell arteritis. Arthritis Rheum. 2006;54:4011–7.
Santos AR, Suffys PN, Vanderborght PR, et al. Role of tumor necrosis factor-alpha and interleukin-10 promoter gene polymorphisms in leprosy. J Infect Dis. 2002;186:1687–91.
Excoffier L, Slatkin M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol Biol Evol. 1995;12:921–7.
Lewontin RC. The interaction of selection and linkage. I. General considerations; heterotic models. Genetics. 1964;49:49–67.
Gavrilets S. Population genetics: multilocus. In: Encyclopedia of life sciences, 1st ed. Wiley; 2001.
Lander E, Kruglyak L. Genetic dissection of complex traits: guidelines for interpreting and reporting linkage results. Nat Genet. 1995;11:241–7.
Turner DM, Williams DM, Sankaran D, et al. An investigation of polymorphism in the interleukin-10 gene promoter. Eur J Immunogenet Off J Br Soc Histocompat Immunogenet. 1997;24:1–8.
Edwards-Smith CJ, Jonsson JR, Purdie DM, et al. Interleukin-10 promoter polymorphism predicts initial response of chronic hepatitis C to interferon alfa. Hepatology. 1999;30:526–30.
Alarcón-Riquelme ME, Lindqvist AK, Jonasson I, et al. Genetic analysis of the contribution of IL10 to systemic lupus erythematosus. J Rheumatol. 1999;26:2148–52.
Vargas-Alarcon G, Ramírez-Bello J, Juárez-Cedillo T, et al. Distribution of the IL-1RN, IL-6, IL-10, INF-γ, and TNF-α gene polymorphisms in the Mexican population. Genet Test Mol Biomarkers. 2012;16:1246–53.
Font J, García-Carrasco M, Ramos-Casals M, et al. The role of interleukin-10 promoter polymorphisms in the clinical expression of primary Sjögren’s syndrome. Rheumatology. 2002;41:1025–30.
Rangel-Villalobos H, Muñoz-Valle JF, González-Martín A, et al. Genetic admixture, relatedness, and structure patterns among Mexican populations revealed by the Y-chromosome. Am J Phys Anthropol. 2008;135:448–61.
Zhu LJ, Liu ZH, Zeng CH, et al. Association of interleukin-10 gene −592 A/C polymorphism with the clinical and pathological diversity of lupus nephritis. Clin Exp Rheumatol. 2005;23:854–60.
Yu S-L, Kuan W-P, Wong C-K, et al. Immunopathological roles of cytokines, chemokines, signaling molecules, and pattern-recognition receptors in systemic lupus erythematosus. Clin Dev Immunol. 2012;2012:715190.
Doreau A, Belot A, Bastid J, et al. Interleukin 17 acts in synergy with B cell-activating factor to influence B cell biology and the pathophysiology of systemic lupus erythematosus. Nat Immunol. 2009;10:778–85.
Palafox Sánchez CA, Satoh M, Chan EK, et al. Reduced IgG anti-small nuclear ribonucleoprotein autoantibody production in systemic lupus erythematosus patients with positive IgM anti-cytomegalovirus antibodies. Arthritis Res Ther. 2009;11:1–12.
Filaci G, Bacilieri S, Fravega M, et al. Impairment of CD8+ T suppressor cell function in patients with active systemic lupus erythematosus. J Immunol. 2001;166:6452–7.
Konya C, Paz Z, Tsokos GC. The role of T cells in systemic lupus erythematosus: an update. Curr Opin Rheumatol. 2014;26:493–501.
Filaci G, Fravega M, Negrini S, et al. Nonantigen specific CD8+ T suppressor lymphocytes originate from CD8+CD28− T cells and inhibit both T-Cell proliferation and CTL function. Hum Immunol. 2004;65:142–56.
Grammatikos AP, Kyttaris VC, Kis-Toth K, et al. A T cell gene expression panel for the diagnosis and monitoring of disease activity in patients with systemic lupus erythematosus. Clin Immunol. 2014;150:192–200.
Csiszar A, Nagy GY, Gergely P, et al. Increased interferon-gamma (IFN-γ), IL-10 and decreased IL-4 mRNA expression in peripheral blood mononuclear cells (PBMC) from patients with systemic lupus erythematosus (SLE). Clin Exp Immunol. 2000;122:464–70.
Acknowledgments
This work was supported by Grant No. 115567 to CAPS of the National Council of Science and Technology (CONACyT, México-Universidad de Guadalajara).
Conflict of interest
The authors declare no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Palafox-Sánchez, C.A., Oregon-Romero, E., Salazar-Camarena, D.C. et al. Association of interleukin-10 promoter haplotypes with disease susceptibility and IL-10 levels in Mexican patients with systemic lupus erythematosus. Clin Exp Med 15, 439–446 (2015). https://doi.org/10.1007/s10238-014-0315-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10238-014-0315-4