Abstract
Botrytis cinerea is a plant pathogenic fungus with a broad host range. Due to its rapid growth and reproduction by asexual spores (conidia), which increases the inoculum pressure, the fungus is a serious problem in different fields of agriculture. The formation of the conidia is promoted by light, whereas the formation of sclerotia as survival structures occurs in its absence. Based on this observation, putative transcription factors (TFs) whose expression is induced upon light exposure have been considered as candidates for activating conidiation and/or repressing sclerotial development. Previous studies reported on the identification of six light-responsive TFs (LTFs), and two of them have been confirmed as crucial developmental regulators: BcLTF2 is the positive regulator of conidiation, whose expression is negatively regulated by BcLTF1. Here, the functional characterization of the four remaining LTFs is reported. BcLTF3 has a dual function, as it represses conidiophore development by repressing bcltf2 in light and darkness, and is moreover essential for conidiogenesis. In bcltf3 deletion mutants conidium initials grow out to hyphae, which develop secondary conidiophores. In contrast, no obvious functions could be assigned to BcLTF4, BcLTF5 and BcLTF6 in these experiments. BcREG1, previously reported to be required for virulence and conidiogenesis, has been re-identified as light-responsive transcriptional regulator. Studies with bcreg1 overexpression strains indicated that BcREG1 differentially affects conidiation by acting as a repressor of BcLTF2-induced conidiation in the light and as an activator of a BcLTF2-independent conidiation program in the dark.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Filamentous fungi are sessile organisms that produce high numbers of asexual and/or sexual spores for ensuring survival (Wyatt et al. 2013). The production of spores in a sexual cycle, which allows for recombination of genetic material and thereby adaptation to environmental changes, appears less relevant in several fungi that have lost the cycle or undergo it only during very specific, yet unknown conditions (Taylor et al. 1999). When fungi can reproduce in both ways, the balance between the two programs has to be tightly regulated because both are energy-requiring processes.
The asexual spores, called conidia, are developed at specialized stalks, the conidiophores (Cole 1986). In many fungi, conidiophore development and subsequent conidiogenesis (conidiation) is induced by specific environmental cues, such as nutrient starvation, injury or light. The latter factor is a meaningful signal, as it conveys that the fungus is exposed to air allowing for distribution of the conidia by abiotic and biotic vectors (Cole 1986; Kumagai 1988). Besides, light is a general stress signal as it may be accompanied by damaging UV light, production of reactive oxygen species (ROS), heat and osmotic stress (Braga et al. 2015; Fuller et al. 2015). Fungi may sense different light qualities, e.g. UV, blue, green, red and far-red light by different photoreceptors; however, not all fungi do so. Apart from the other receptors, the blue light-sensing transcriptional activator (white collar) complex (WCC) is highly conserved in the fungal kingdom (Fischer et al. 2016; Idnurm et al. 2010).
Regulation of conidiation by light is best known in the Eurotiomycete Aspergillus nidulans and the Sordariomycete Neurospora crassa. Both systems comprise a central transcription factor (TF) whose expression is essential and sufficient to induce conidiation. Thus, expression of brlA (bristle), encoding a C2H2 TF, is induced in a red light-dependent manner in A. nidulans (Adams et al. 1988; Mooney and Yager 1990), while fl (fluffy), encoding a Zn2Cys6 TF, is induced by blue light through the WCC in N. crassa (Bailey-Shrode and Ebbole 2004; Bailey and Ebbole 1998; Olmedo et al. 2010). BrlA is embedded in a complex regulatory network and is transcriptionally regulated by the fluffy low bristle TFs (FlbB, FlbC, FlbD) and regulates the expression of further TF-encoding genes (abaA, wetA) acting during advanced stages of conidiation (Krijgsheld et al. 2013; Park and Yu 2012). The hierarchy of TFs in N. crassa is not that well understood. Nevertheless, the involvement of a number of TFs in certain stages of conidiophore development and conidiogenesis is known. For instance, mutants of the C2H2 TFs SAH-1 (short aerial hyphae) and CSP-1 (conidial separation) are affected in aerial hyphae formation and the onset of conidiation and in formation of free macroconidia from proconidial chains, respectively (Colot et al. 2006; Lambreghts et al. 2009; Sun et al. 2012). Orthologs of the latter two TFs exist in A. nidulans, but they have not been functionally characterized. In contrast, the GATA-type TFs termed NsdD (never in sexual development) and SUB-1 (submerged protoperithecia) in A. nidulans and N. crassa, respectively, have been identified as key regulators of development in both systems. NsdD is crucial for balancing asexual and sexual development as its deletion results in hyper-conidiation and loss of fruiting body formation (Han et al. 2001; Lee et al. 2014, 2016). SUB-1 is involved in sexual development (protoperithecia formation) and the modulation of light-responsive gene expression (Chen et al. 2009; Colot et al. 2006).
Likewise, light is a vital environmental signal for regulation of development in the gray mold fungus Botrytis cinerea (Epton and Richmond 1980). The fungus belongs to the class of the Leotiomycetes, and causes diseases on many unrelated plant species (Fillinger and Elad 2016). At the end of the infection cycle, B. cinerea produces different reproduction structures, either multinucleate (macro)conidia or sclerotia and uninucleate microconidia. The conidia that are formed holoblastically at branched conidiophores are designated as inoculum for further infections while the sclerotia allow for the survival of unfavorable conditions such as the absence of the host during the winter. In next spring, the sclerotia may germinate asexually by bearing conidiophores and conidia or sexually by bearing apothecia containing the ascospores (Williamson et al. 2007). Light is a decision-making tool, as it triggers conidiation (asexual reproduction) and at the same time represses sclerotial development (asexual/sexual reproduction). Upon fertilization of sclerotia by microconidia, light induces the formation of the apothecia. The species B. cinerea is characterized by a high degree of variation. Though the majority of wild strains are light responsive in terms of reproduction, i.e. they form conidia in the light and sclerotia in the dark, many blind strains are found that exhibit the same morphologies under different light conditions (always conidia, always sclerotia, always mycelia) (Canessa et al. 2013; Schumacher et al. 2012).
To identify the genes involved in (photoinduced) conidiation in B. cinerea, transcriptional responses of vegetative mycelia of the light-responsive strain B05.10 upon exposure to white light was studied, resulting in the identification of six TF-encoding genes that were significantly induced and thereafter termed light-responsive TFs (LTFs) (Table 1). BcLTF1, initially identified as virulence-associated gene by a random mutagenesis approach, is the ortholog of NsdD and SUB-1 and has overlapping but also specific functions. Thus, like NsdD, it represses conidiation and its deletion results in hyper-conidiation (always conidia) (Schumacher et al. 2014). BcLTF2, the putative ortholog of SAH-1, has subsequently been identified as positive regulator of conidiation in B. cinerea and thereby as the functional counterpart of A. nidulans BrlA and N. crassa FL (Cohrs et al. 2016). The WCC, formed by the GATA-type TFs called BcWCL1 and BcWCL2, mediates responses to light in different ways, e.g. it activates and represses the expression of light-responsive genes. Bcltf2 belongs to the latter group of genes; consequently, the deletion of bcwcl1 results in elevated bcltf2 transcript levels and de-repressed conidiation in light and darkness (always conidia) (Canessa et al. 2013; Schumacher 2012).
In this study, the functional characterization of the other four LTFs is described. It revealed that the C2H2 TF BcLTF3, the ortholog of CSP-1, is required for conidiation while the Zn2Cys6 TFs BcLTF4-6 have no obvious functions. The previously identified transcriptional regulator BcREG1, being involved amongst others in conidiogenesis (Michielse et al. 2011), was found to be light responsive as well and therefore included in this study. Both BcLTF3 and BcREG1, apart from their need for conidiogenesis, have impact on regulation of conidiophore development by influencing the expression levels of bcltf2.
Materials and methods
Bioinformatics
Sequences of B. cinerea B05.10 were obtained from the B. cinerea Database at EnsemblFungi (http://fungi.ensembl.org/Botrytis_cinerea). DNA and protein sequences from other fungi were obtained from the public databases at the National Center for Biotechnology (NCBI) (http://www.ncbi.nlm.nih.gov). Protein sequence alignments were generated using the ‘one click’ method at Phylogeny.fr (http://phylogeny.lirmm.fr/). Conserved protein domains and putative nuclear localization signals (NLS) were identified by PfamSearch (http://pfam.xfam.org/search) and WoLF Psort (http://www.genscript.com/psort/wolf_psort.html). CCCCT motifs in the promoters of annotated B. cinerea genes (a maximum of 1000 bp of the 5′ non-coding regions) were identified by running fuzznuc (http://emboss.bioinformatics.nl/cgi-bin/emboss/fuzznuc). For sequence analyses and protein alignments (Clustal V), programs of the DNASTAR® Lasergene software package were used.
NimbleGen 12-plex arrays comprising 62,478 60-mer specific probes covering all the 20,885 predicted gene models and non-mapping ESTs of B. cinerea, as well as 15,707 random probes as negative controls, were used (Amselem et al. 2011). Data processing, quality controls and differential expression analysis were performed using ANAIS methods (Simon and Biot 2010). Genes featuring intensities exceeding the threshold (1.5 × 95th percentile of random probes) in at least half of the hybridizations were considered as expressed and kept for further analyses. BcLTF3-dependent genes (in darkness or after a light pulse) were identified using a one-way ANOVA with the False Discovery Rate (FDR) correction of p values. Transcripts with a corrected p value <0.05 and more than a twofold change in transcript levels were considered as significantly differentially expressed (DE). Details on the experiments, raw and normalized values are available in the Gene Expression Omnibus Database (GEO) (https://www.ncbi.nlm.nih.gov/geo/) (accession number: GSE95738).
Cultivation of B. cinerea
Botrytis cinerea B05.10 is an isolate from Vitis vinifera and is used as the recipient strain for genetic modifications (Table 2). Strains were cultivated in Petri dishes containing solid synthetic complete medium (CM) (Pontecorvo et al. 1953). Cultures were incubated at 20 °C under white light [12 h light/12 h darkness (LD) and 24 h light (LL)] for induction of conidiation or in continuous darkness (DD) for induction of sclerotia formation. White light, 9000 lx at culture level, was generated by using Sylvania Standard F18W/29-530 ‘warm white’ and F36W/33-640 ‘cool white’ fluorescent bulbs. For DNA and RNA isolation, mycelia were grown on solid CM with cellophane overlays. Resistant strains were cultivated on CM supplemented with hygromycin B (Invitrogen) or nourseothricin (Werner BioAgents GmbH) in a concentration of 70 µg/ml. Protocols for protoplast formation and transformation of B. cinerea were described previously (Schumacher 2012). Virulence assays on French bean (Phaseolus vulgaris) were performed as described (Schumacher 2016a).
Standard molecular methods
Fungal genomic DNA was prepared as described previously (Cenis 1992). For Southern blot analysis, fungal genomic DNA was digested with restriction enzymes (Thermo Scientific), separated on 1% (w/v) agarose gels and transferred to Whatman® Nytran™ SuPerCharge (SPC) nylon blotting membranes. Blot hybridizations with random-primed α-32P-dCTP-labeled probes were performed as described previously (Schumacher 2012). Total RNA was isolated making use of the TRI Reagent® RNA Isolation Reagent (Sigma-Aldrich). For northern blot analyses, samples (25 µg) of total RNA were transferred to Nytran membranes after electrophoresis on a 1% (w/v) agarose gel containing formaldehyde (Sambrook et al. 1989). For cDNA synthesis, total RNA was treated with RNAse-free DNAse (Macherey–Nagel GmbH & Co. KG). cDNA for real-time quantitative reverse transcription PCR (RT-qPCR) analyses was obtained from 1 µg of total RNA by a retro-transcription step realized with the SuperScript® II Reverse Transcriptase (Life Technologies) using the primer oligo(dT20). RT-qPCR reactions were performed on 1:5- or 1:7.5-diluted cDNA using iQ™SYBR®Green Supermix and an iCycler Thermal Cycler (Bio-Rad Laboratories Inc.). Used primers are listed in Table S1. The data were analyzed with the iQ™5 Optical System Software version 2.1 (Bio-Rad Laboratories Inc.). Gene expression levels were calculated according to the 2−ΔCT method (Livak and Schmittgen 2001). Technical duplicates were run for each biological replicate. Synthesis of cDNA, Cy3-labeling and hybridizations of B. cinerea microarrays (Amselem et al. 2011) were performed by Arrows Biomedical Deutschland GmbH using the procedures and reagents established by Nimblegen (Roche). PCR reactions were performed using the Phusion high-fidelity DNA polymerase (ThermoFisher Scientific) for cloning purposes and the BioTherm™ Taq DNA Polymerase (GeneCraft) for diagnostic applications. Used primers are listed in Table S2. Replacement fragments and expression vectors were assembled in the uracil-auxotrophic Saccharomyces cerevisiae strain FY834 by exploiting its homologous recombination machinery (Colot et al. 2006; Oldenburg et al. 1997; Schumacher 2012). Sequencing of DNA fragments was performed with the BigDye® Terminator v3.1 cycle sequencing kit (ThermoFisher Scientific) in an ABI Prism capillary sequencer (Applied Biosystems).
Generation of B. cinerea mutants
For construction of gene replacement fragments according to the strategy for bcltf3 shown in Fig. S1a, the 5′- and 3′-noncoding regions of bcltf3, bcltf4, bcltf5 and bcltf6 (genes of interest, GOIs) were assembled with the nourseothricin resistance cassette (PtrpC::nat1) and the shuttle vector pRS426 (Christianson et al. 1992) by yeast recombinational cloning. The gene flanks were amplified using primer combinations GOI-5F/-5R and GOI-3F/-3R and DNA of B. cinerea B05.10 as template, and the resistance cassette was amplified with primers hph-F and nat1-R using pZPnat1 (GenBank: AY631958.2) as template. PCR fragments were subsequently co-transformed with the EcoRI- and XhoI-digested pRS426 resulting in pΔGOI_natR. The replacement fragments were amplified using primer combinations GOI-5F/-3R and the plasmids pΔGOI_natR as template and used for transformation. Protoplasts of B. cinerea B05.10 (wild type) were transformed for the generation of Δbcltf3, Δbcltf4, Δbcltf5 and Δbcltf6 mutants, those of the Δbcltf1 mutant with the bcltf3 deletion fragment for the generation of ΔΔbcltf1/3 mutants. Homologous integration events in resistant transformants were detected by diagnostic PCR using primers nat1-hiF and PtrpC-P2 binding in the resistance cassette, and the primers GOI-hi5F and GOI-hi3R binding up- and downstream of the flanking regions. Protoplast (Δbcltf3, ΔΔbcltf1/3) or conidial (Δbcltf4-6) isolates derived from independent primary transformants (heterokaryons) were screened for the absence of wild-type alleles by using primers (GOI-WT-F/-WT-R). Southern blot analyses were performed to detect additional ectopic integration events (Fig. S1b; data not shown).
The complementation of the bcltf3 deletion mutant (Δbcltf3-T9) was accomplished by targeted integration of bcltf3 at the native gene locus resulting in the replacement of the nat1 resistance cassette. The vector pbcltf3 CiL_hygR was assembled in yeast by co-transformation of the following fragments: (1) pRS426 digested with EcoRI and XhoI, (2) 5′ flank + bcltf3 (primers bcltf3-5F/-Tgluc-R), (3) Tgluc [terminator of a glucanase-encoding gene of B. cinerea (Schumacher 2012)] fused to the PtrpC::hph cassette derived from pCSN44 (Staben et al. 1989) [primers Tgluc-F2/hph-F; template: pbclae1 CiL_hygR (Schumacher et al. 2015)] and (4) 3′ flank of bcltf3 (primers bcltf3-3F/-3R). Targeted integration of the construct that results in the restoration of the 5′ region was detected by diagnostic PCR using primer combinations bcltf3-hi5F/-hi5R and hph-hiR/bcltf3-hi3R (Fig. S1a; data not shown).
Overexpression strains for bcltf3, bcltf4, bcltf5, bcltf6 and bcreg1 were generated by integration of second gene copies under control of the constitutive PoliC at the nitrate reductase locus (bcniaD). The open reading frames (ORFs) were amplified with primers GOI-PoliC-F/-Tgluc-R and co-transformed with the NcoI- and NotI-digested pNDH-OGG (Schumacher 2012) in yeast yielding pNDH_PoliC::GOI. Transformants were tested for targeted integration of the overexpressing constructs using the primer combinations bcniaD-hi5F/Tgluc-hiF and bcniaD-hi3R/hph-hiF. Increased expression levels in independent transformants were demonstrated by northern blot analyses (Fig. S1d; data not shown).
For localization of BcLTF3 and BcREG1 inside the cell, the genes were fused to gfp. For this, ORFs were amplified with primers bcreg1-PoliC-F/-GFP-R, bcltf3-PoliC-F/-GFP-R and bcltf3-gfp-F/-Tgluc-R and assembled with the NotI- or NcoI-digested pNAN-OGG or pNAH-OGG (Schumacher 2012), resulting in pNAN-PoliC::bcreg1-gfp, pNAH-PoliC::bcltf3-gfp and pNAH-PoliC::gfp-bcltf3. The constructs were introduced into the corresponding deletion mutants (Δbcreg1-T16, Δbcltf3-T9) to confirm the functionality of the fusion proteins. Targeted integration at bcniiA (nitrite reductase locus) of the constructs was detected using primer combinations bcniiA-hi5F/Tgluc-hiF and bcniiA-hi3R/hph-hiF or nat1-hiF. Expression of BcLTF3 fusion proteins failed, as neither fluorescence nor the restoration of the wild-type phenotype could be observed (data not shown).
Microscopy
Close-up images of B. cinerea colonies were captured with a SteREO Discovery V.20 stereomicroscope equipped with an AxioCamMRc camera and the Axiovision Rel 4.8 software package (Zeiss). Fluorescence and light microscopy of growing hyphae was performed with an AxioImager M1 microscope equipped with the ApoTome.2 technology for optical sectioning with structured illumination (Zeiss). Differential interference microscopy (DIC) was used for bright field images. For visualizing the nuclei, the hyphae were stained using the fluorescent dye Hoechst 33342 (Sigma-Aldrich) as described previously (Schumacher 2012). Hoechst staining was examined using the filter set 49 DAPI shift free (excitation G 365, beam splitter FT 395, emission BP 445/50) and GFP fluorescence with filter set 38 (excitation BP 470/40, beam splitter FT 495, emission BP 525/50). Images were captured with an AxioCam MRm camera and further processed using the Axiovision Rel 4.8 software package (Zeiss).
Results
Bcltf3 encodes a putative C2H2 transcription factor
Expression of bcltf3 (Bcin11g01720) was induced by light in the wild-type B05.10 as well as in the ∆bcltf1 mutant in an earlier study (Table 1) (Schumacher et al. 2014). Bcltf3 has an ORF of 1.059 kb and does not contain introns. The transcript is significantly larger (about 3 kb in size) due to long 5′- and 3′-untranslated regions (UTRs) of 0.8 and 1.1 kb (Fig. S2a). Only few genes are located upstream in close proximity to bcltf3: gene calls Bcin11g01730 and Bcin11g01740 are dubious (no orthologs in other fungi), and Bcin11g01750 is located ~13 kb upstream of bcltf3 and encodes a putative pre-mRNA splicing factor. Strikingly, transcription of the region upstream of bcltf3 occurred, though no gene is predicted in this area (Fig. S2a).
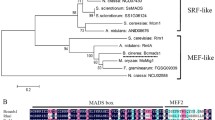
Bcltf3 encodes a protein of 352 amino acids (aa) and comprises two C2H2 zinc finger domains (181–208 and 214–238 aa) (Fig. 1). No further conserved motifs or nuclear localization signals (NLS) were predicted. Overall sequence identities with proteins from other Ascomycota containing similar C2H2 domains vary between 19% (Saccharomyces cerevisiae) and 60% (Sclerotinia sclerotiorum). Nevertheless, the two C2H2 domains are highly conserved, from 50% (S. cerevisiae) to 88% aa identity (S. sclerotiorum), suggesting that these proteins are orthologs of BcLTF3. N. crassa CSP-1 (287 aa), one of the few characterized orthologs, shares 32 and 75% aa identity with the entire BcLTF3 and the C2H2 domains, respectively. Besides, all orthologs of the Leotiomycetes are longer (321–360 aa) than those from the Sordariomycetes (287–306 aa) due to C-terminal extensions.
BcLTF3 is a putative C2H2 transcription factor with orthologs in yeast and filamentous fungi. a Phylogenetic tree of BcLTF3 orthologs. Species names, geneIDs and sizes of the proteins are indicated. Tree construction and calculation of the amino acid (aa) identities (%) between BcLTF3 and the other proteins (shown in brackets) were conducted using Clustal V. b BcLTF3 and its orthologs contain two conserved C2H2-type zinc finger domains. Indicated are the aa identities shared by the N-termini, the regions of the C2H2 domains and the C-termini in the proteins shown above. The alignment of the putative DNA-binding region is shown below. Amino acids that are identical in all sequences are shaded black; amino acids shared in three or more proteins are shaded gray. The C2H2 domains (IPR015880) are underlined; the conserved Cys and His residues are indicated
To demonstrate the putative nuclear localization of BcLTF3 as a potential TF, N- and C-terminal GFP fusion proteins under control of a constitutive promoter were expressed in B. cinerea; however, no fluorescence could be observed in several independent transformants containing the respective constructs (data not shown). For studying the functions of BcLTF3, single (∆bcltf3) and double (∆∆bcltf1/3) deletion mutants as well as overexpression (OE::bcltf3) strains were constructed (Table 2) (see “Materials and methods” for details). For all strategies, at least two independent mutants exhibiting identical phenotypes were obtained. The data shown correspond to one arbitrarily chosen mutant for each construct. Furthermore, it was demonstrated that re-introduction of bcltf3 into ∆bcltf3 (bcltf3 CiL strains) restored the wild-type phenotype (Fig. S1c), confirming the gene-phenotype linkage.
The putative transcriptional regulator BcREG1 localizes to the nuclei
A previous study reported on the requirement of BcREG1 for conidiogenesis (formation of conidia at the denticles of regularly developed conidiophores), virulence (invasive growth) and secondary metabolism [production of the toxins botrydial (BOT) and botcinic acid (BOA)] (Michielse et al. 2011). Genome-wide approaches revealed that expression levels of bcreg1 significantly responded to light treatment in the wild type, though the fold change of 1.4 was below the defined threshold (FC > 2) (Table 1). RNA sequencing data indicate that the transcript of bcreg1, similar to that of bcltf3, contains large 5′ and 3′ UTRs of ~0.9 and ~0.5 kb and that there is a long region (8 kb) without any genes and significant transcription upstream of the start codon of bcreg1 (Fig. S2a).
Orthologs of BcREG1 are found in several but not all Ascomycetes (Fig. S2b). For instance, no ortholog is present in N. crassa, while orthologs are found in Fusarium spp. (SGE1, 48–49% aa identity) and A. nidulans (OsaA, 48% aa identity). In general, the N-terminal regions of the orthologous proteins containing the Gti1_Pac2 domain exhibit higher similarity (44–85%) than the C-terminal regions (7–67%). Two putative nuclear localization signals in BcREG1 are predicted (Fig. S2c).
Based on the previous observations that bcreg1 is essential for conidiogenesis and may be under light control, the transcriptional regulator was included into this study for more detailed analyses. Besides using the already described deletion mutant (∆bcreg1), overexpression strains (OE::bcreg1) were generated by integrating a second copy of bcreg1 under control of PoliC at the bcniaD locus (Table 2). Nuclear localization of BcREG1 was demonstrated by expressing a BcREG1-GFP fusion protein in the ∆bcreg1 background. Its expression under control of the strong PoliC failed to restore the wild-type phenotype but resulted in the phenotype found for OE::bcreg1 strains, indicating that the fusion protein is functional (Fig. 2).
BcREG1 as putative transcriptional regulator is localized in the nuclei. a BcREG1-GFP fusion protein under control of the constitutive PoliC promoter was expressed in the ∆bcreg1 mutant. a The BcREG1-GFP fusion protein is functional as its overexpression phenocopies OE::bcreg1. Indicated strains were cultivated for 2 weeks on solid CM in DD or LD. b The BcREG1-GFP fusion protein is localized in the nuclei. Conidia of the bcreg1-gfp strain were incubated for 16 h on a microscope slide in liquid Gamborg B5 medium supplemented with 2% glucose and 0.02% ammonium phosphate. GFP signals co-localized with the Hoechst staining patterns in parallel experiments indicating that BcREG1 is localized in the nuclei (data not shown)
Bcltf3 and bcreg1 are induced by white light in a WCC-dependent manner
When cultures of B. cinerea B05.10 are incubated in continuous light (LL), the first conidiophores and conidia are visible after 2 days. In contrast, cultures that are kept in constant darkness (DD) stay vegetative (competent stage) for a longer time and finally differentiate sclerotia when no light signals are detected within one week of growth (Fig. 3a).
Expression of bcltf3 and bcreg1 is induced by white light in a WCC-dependent manner. Gene expression was analyzed by RT-qPCR and is given as ratio between genes of interest (GOIs) and reference genes (REFs). Data shown are derived from three biological replicates with two technical replicates each. Statistical tests revealed significant differences (*p < 0.05, **p < 0.001). a Growth phenotypes of WT:B05.10 in DD and LL. The strain was grown on solid complete medium (CM) with cellophane overlays in DD for induction of sclerotia formation, and in LL for induction of conidiation. b Bcltf3 and bcreg1 expression is induced in vegetative mycelia upon light exposure. WT:B05.10 was incubated on solid CM with cellophane overlays for 2 days in DD or LL. Vegetative mycelia (grown in DD) were exposed to white light for the indicated periods. Asterisks indicate differences between light treatments and DD. c Bcltf3 and bcreg1 are highly expressed during conidiation in the light. Mycelia and reproduction structures of WT:B05.10 were harvested after 2, 4 and 6 days of incubation (see a). Asterisks indicate differences to 2-day-old cultures in each light condition. d Expression in bcltf2 strains. Indicated strains were cultivated for 4 days on solid CM with cellophane overlays in DD or LL. Bcltf2 deletion mutants do not produce conidia, OE::bcltf2 strains produce conidia even in the dark and exhibit increased conidiation in the light. Asterisks indicate differences to WT:B05.10 in each light condition. e Light induction of bcltf3 and bcreg1 expression depends on the WCC, BcLTF1 and the VELVET complex. Indicated strains were incubated on solid CM for 2 days in DD or LL. DD cultures were additionally exposed to white light pulses of 15, 60 and 180′. BcWCL1 is a component of the White collar complex (WCC), BcVEL1 and BcLAE1 are components of the VELVET complex. Asterisks indicate differences between mutants and WT:B05.10 in each light condition
Thus, young vegetative mycelia obtained by growing the fungus in DD exhibit high responsiveness to light. A time course experiment applying light pulses (LP) of 5–300 min to vegetative mycelia (grown for two days in DD) was performed to study the expression levels of bcltf3 and bcreg1 by RT-qPCR analyses over time. The experiment confirmed the light induction found by the genome-wide microarray approaches for LP 60′ and showed furthermore that the expression profiles of both genes exhibit different kinetics. Bcltf3 expression was already significantly induced at LP 5′ (2.7-fold) and reached its maximal levels at LP 15′ (4.9-fold). Then, expression levels decreased to the DD level after LP 300′. Notably, expression levels were lower during incubation in LL than in DD. In contrast, bcreg1 expression levels remained constant until LP 300′ after reaching the optimum at LP 15′ (2.4-fold) and were still elevated in LL compared to DD (1.7-fold) (Fig. 3b).
Next, the expression of the genes was followed during conidiation and sclerotial development in LL and DD, respectively. Expression levels of both bcltf3 and bcreg1 increased during cultivation in LL/conidial development; however, while those of bcreg1 remained low in DD, those of bcltf3 also increased during sclerotial development in DD (Fig. 3c). The data were confirmed in a second experiment, in which the expression levels were determined in bcltf2 mutants after 4 days of cultivation in DD or LL. ∆bcltf2 mutants do not produce conidia in the light which is accompanied by the formation of sclerotial initials, whereas OE::bcltf2 strains exhibit hyper-conidiation accompanied by the loss of sclerotial development in DD (Cohrs et al. 2016). Expression levels of bcltf3 were slightly decreased in the OE::bcltf2 strain grown in LL, those of bcreg1 in the conidiation-deficient ∆bcltf2 mutant in LL (Fig. 3d).
Finally, the light induction of bcltf3 and bcreg1 was studied in different mutant backgrounds, i.e. in mutants of the blue light-sensing WCC (∆bcwcl1), of BcLTF1 (∆bcltf1) and the VELVET complex (∆bcvel1, ∆bclae1) (Table 2). Induction of both genes relies on the WCC, as bcltf3 expression levels remained unchanged and those of bcreg1 were only slightly increased in ∆bcwcl1 upon light exposure (1.3-fold compared to 2.6-fold in wild type at LP 60′). Notably, expression levels of both genes were slightly affected in DD as well. In the other mutants, light induction of bcltf3 generally occurred, but basal expression levels and amplitudes (LP 15′) were slightly altered. Noticeable were the increased expression levels of bcreg1 in ∆bcltf1 and ∆bcvel1 in DD, accounting for the lower induction by subsequent light treatment (1.7- and 1.2-fold at LP 60′), and the lower levels after light induction in the ∆bclae1 mutant (1.3-fold at LP 60′) (Fig. 3e).
Taken together, bcltf3 and bcreg1 are light-responsive genes whose induction depends on the WCC. Furthermore, their expression levels are slightly modulated by BcLTF1, BcVEL1 and BcLAE1. Expression levels of bcltf3 increased during conidial and sclerotial development, suggesting a role of BcLTF3 in light as well as in darkness. In contrast, the substantial expression of bcreg1 exclusively during conidiation is in agreement with the previous finding that deletion of bcreg1 affects conidiation but not sclerotial development (Michielse et al. 2011).
BcLTF3 is required for conidiogenesis and repressing conidiophore development in darkness
Deletion and overexpression of bcltf3 did not result in drastically changed growth rates neither under standard nor stress conditions; however, the OE::bcltf3 strains grew a bit slower than the wild type and the ∆bcltf3 mutants. The ∆∆bcltf1/3 mutants exhibited reduced growth in the light, under neutral/alkaline conditions, and during exposure to oxidative and osmotic stress similarly to the ∆bcltf1 single mutant (Fig. 4).
BcREG1, but not BcLTF3, has influence on responses to pH and oxidative stress. a Overexpression, but not the deletion of bcltf3 or bcreg1, results in reduced radial growth rates. Indicated strains were grown for 3 days on solid CM pH5 in DD, LD or LL. Mean values and standard deviations were calculated from three colonies per strain and condition. Asterisks indicate significant differences compared to wild-type B05.10 in the three light conditions (*p ≤ 0.001). b Mutations of BcREG1 affect the ability to acidify the culture medium. Strains were cultivated for 4 days on solid CM pH 7.5 supplemented with 0.01% of the pH indicator bromothymol blue. The green coloration of the medium indicates pH values around seven, and the yellowish coloration pH values below six. c Bcreg1 mutants differentially respond to pH and oxidative stress. Diameters of three colonies (two measurements per colony) per strain and condition were determined after 3 days of incubation in LD. Mean values and standard deviations were calculated. Growth rates under the indicated stress conditions are given as percentages of the controls (CM pH5). Asterisks indicate significant differences compared to wild-type B05.10 under the tested conditions (*p ≤ 0.005)
Conidiophore development started earlier in the ∆bcltf3 mutants in LD than in the wild type. This effect was not observed in the ∆∆bcltf1/3 mutants due to their reduced growth rates in LD at that time. However, when comparing the colonies of ∆∆bcltf1/3 and the recipient strain ∆bcltf1 grown in DD, the ∆bcltf3 effect, i.e. the de-repression of conidiophore development becomes apparent (Fig. 5a). The same effect was observed for older colonies that have been cultivated for 2 weeks. The ∆bcltf3 mutants developed abundant conidiophores and fewer sclerotia in DD (~50% of wild type) (Fig. 5b; Table 3). Notable was the coloration of the ∆bcltf3 and ∆∆bcltf1/3 colonies that was more grayish compared to that of the wild type and the other mutants. Closer inspection of the conidiophores of the ∆bcltf3 single and double mutants showed that de facto no mature conidia were formed (Fig. 5c; Table 3). Thus, the denticles at the young conidiophores developed conidial initials; however, these initials subsequently grew out to new hyphae (Fig. 5d). In sum, the deletion of bcltf3 abolishes the formation of mature conidia and promotes the formation of conidiophores in both light conditions. In accordance, the overexpression of bcltf3 had an opposing effect by leading to decreased conidiophore development and conidiation in LD (~50% of wild type).
Mutations of BcLTF3 and BcREG1 affect the light-dependent differentiation programs. WT:B05.10 and the indicated deletion mutants and overexpression strains were cultivated on solid CM in DD and LD. a Overexpression of BcREG1 results in altered pigmentation of young colonies. Pictures were taken after 3 days of incubation. Note that ∆bcltf1 and ∆bcltf1/3 mutants exhibit significantly reduced radial growth rates in LL and consequently less pigmentation. See Fig. 4a for details. b Overexpression strains and deletion mutants of bcltf3 and bcreg1 exhibit altered colony morphologies. Pictures were taken after 2 weeks of incubation (top views). All mutants were affected in proper conidiation. Numbers of sclerotia and conidia formed by the mutants were determined; see Table 3. c ∆bcltf3 and ∆bcreg1 mutants form malformed conidiophores. Cross sections of 2-week-old colonies, cultivated in LD, are shown. Scale bar represents 200 µm. d BcLTF3 and BcREG1 are essential for conidiogenesis. Close-up images of the conidiophores of WT:B05.10 and the two single deletion mutants are shown. While ∆bcreg1 mutants do not develop conidia at the denticles, ∆bcltf3 develop conidial initials (young conidiophore on the left), which grow out to sterile hyphae (older conidiophore shown on the right). Scale bar represents 40 µm
As conidiation depends on BcLTF2, its expression levels in the Δbcltf3 and OE::bcltf3 strains were studied. Young cultures grown in DD were harvested directly or exposed to light for 60′ or 180′ before harvest. Moreover, cultures grown in LL were included (Fig. 6). Expression levels of bcltf2 were elevated in the ∆bcltf3 mutant during growth in DD and in LL (2.7- and 2.0-fold compared to wild type), which is in accordance with the increased conidiophore development. On the other hand, induction of bcltf2 expression by light (LP 60′, 180′) was decreased in OE::bcltf3 strains (7.9-fold compared to 33.4-fold in wild type).
BcLTF3 and BcREG1 influence gene expression. Wild-type B05.10 and the indicated strains were cultivated on solid CM for 2 days in DD or LL. Mycelia (DD) were then additionally exposed for 60 or 180 min to white light. Gene expression analyzed by RT-qPCR is given as the ratio GOI/bcact1. Mean values and standard deviations shown are derived from three biological replicates with two technical replicates each. Statistical analyses revealed significant differences between mutants and wild type under the different conditions (*p ≤ 0.05). On the left: genes encoding putative transcriptional regulators, on the right: secondary metabolism-related genes encoding the key enzyme for conidial melanogenesis (bcpks13), the oxaloacetate hydrolase for oxalic acid formation (bcoahA), the key enzymes for biosynthesis of the phytotoxic compounds botrydial (bcbot2) and botcinic acid (bcboa6) and a sesquiterpene cyclase (bcstc5)
In sum, BcLTF3 is essential for conidiogenesis and for repressing conidiophore development in light and dark by affecting the expression levels of bcltf2 encoding the key regulator of conidiation.
Overexpression of BcREG1 affects developmental programs differentially in light and dark
The deletion of bcreg1 was shown to prevent conidiogenesis (Michielse et al. 2011). In this case, regularly shaped conidiophores are formed but no conidial initials develop at the denticles. In contrast to the ∆bcltf3 mutants, the ∆bcreg1 mutants produced fewer conidiophores and instead few sclerotia even in the light, suggesting a negative feedback regulation of conidiophore development. However, bcltf2 expression levels were similar in ∆bcreg1 mutants and the wild type (Fig. 6).
The overexpression of bcreg1 resulted in drastic phenotypic changes, including reduced radial growth rates in LL (Fig. 4a). Thus, young cultures exhibited a yellowish and reddish pigmentation when incubated in DD and LD, respectively (Fig. 5a). Formation of aerial hyphae and conidia in the light was severely reduced (~70% of wild type), while significant numbers of conidia were produced in DD (Fig. 5b; Table 3). Conidiation in DD was accompanied by increased aerial hyphae formation and loss of sclerotial development. The elevated transcript levels of bcpks13 encoding the key enzyme in conidial melanogenesis (Schumacher 2016a) in young cultures grown in DD with/without LP (Fig. 6) suggests that the observed pigmentation is due to the accumulation of DHN melanin intermediates. The yellow pigment (perhaps scytalone) was light sensitive, as the pigmentation vanished after transfer of the DD cultures to light. The reddish pigmentation in LD on the other hand may be the result of the oxidation of T4HN resulting in the orange-colored flaviolin. Expression of bcltf2 was not induced upon light exposure in OE::bcreg1 strains, which is in accordance with the reduced conidiation in LD (Fig. 6).
The deletion and overexpression of bcreg1 differentially affected the abilities to grow under neutral/alkaline conditions and to acidify the culture medium. While ∆bcreg1 mutants exhibited similar acidification in DD (similar to wild type) and LD (reduced compared to wild type), the OE::bcreg1 strains acidified the culture medium faster than the wild type in both light conditions. Remarkably, acidification—at least in the wild type—is a light-dependent process. Both deletion and overexpression of bcreg1 released the process from light control (Fig. 4b). The reduced ability of the ∆bcreg1 and ∆bcltf1 mutants to establish an acidic environment was in accordance with the reduced growth rates under neutral and alkaline conditions (Fig. 4c). However, the expression levels of bcoahA encoding the oxaloacetate hydrolase (Han et al. 2007) were not significantly affected in the mutants, suggesting that the observed phenotypes are not due to a deregulated oxalic acid formation (Fig. 6).
In conclusion, BcREG1 is crucial for conidiogenesis and it represses and promotes conidiation in light and darkness, respectively. Furthermore, it regulates BcPKS13-derived melanogenesis and the processes leading to acidic environments.
BcREG1 but not BcLTF3 is relevant for virulence
Virulence of the bcltf3 and bcreg1 mutants was studied on primary leaves of French bean (Phaseolus vulgaris) using mycelial plugs for inoculation. For quantification, the lesion diameters were determined after 2 days (Fig. 7a). As previously reported, the ∆bcreg1 mutants were severely hampered in colonizing the host tissue. A slightly increased colonization rate was observed for the OE::bcreg1 strains; however, deviations between independent experiments were high, possibly due to the fact that the strains were heterokaryons. Nevertheless, the colonization of the host tissue was accompanied by an increased proliferation of aerial hyphae and reduced conidiation.
BcREG1, but not BcLTF3, contributes to full virulence on French bean plants. a Bcreg1 deletion mutants are impaired in invasive growth. Primary leaves of living Phaseolus vulgaris (French bean) plants were inoculated with plugs of vegetative mycelia of the indicated strains. Mean values and standard deviations were calculated from eight lesions per strain with two measurements per lesion. Asterisks indicate significant differences compared to WT:B05.10 (*p ≤ 0.05, **p ≤ 0.001). b Bcltf3 and bcreg1 mutants exhibit altered differentiation phenotypes on colonized host tissues. Infected leaves were detached after 2 days of inoculation and incubated on water agar for 2 weeks in LD and DD for induction of conidiation and sclerotia formation, respectively. Scale bars represent 1 mm
As the full capacity to colonize the plant tissue requires the two phytotoxins BOT and BOA (Dalmais et al. 2011), the expression of the genes encoding the key enzymes of the biosynthetic pathways was studied. As shown in Fig. 6, moderate expression levels of bcbot2 and bcboa6 were found in the wild type under the in vitro conditions only when cultures were incubated in LL. Expression levels were decreased in the ∆bcreg1 mutants, but significantly increased in the OE::bcreg1 strains, which indicated that the deletion and overexpression of bcreg1 resulted in opposing phenotypes.
Deletion of bcltf3 had no impact on virulence, neither in the wild-type background nor in the ∆bcltf1 background as the ∆bcltf3 mutants colonized the host tissue in a similar manner as the recipient strains did. The slightly reduced lesion diameters observed for the OE::bcltf3 strains were likely due to the generally reduced growth rates rather than due to a distinct effect in the fungus–host interaction (Fig. 7a).
For following the light-dependent differentiation of the reproduction structures on the host tissue, infected bean leaves were detached and incubated for 2 weeks in LD or DD. The strains behaved similar to the in vitro assay on solid CM: all strains produced conidiophores in LD (the ∆bcltf3 and ∆bcreg1 mutants failed to develop conidia), and few of them, i.e. the wild type, OE::bcltf3 and ∆bcreg1 produced sclerotia in DD. The other mutants developed abundant conidiophores in DD (Fig. 7b). In sum, BcREG1, but not BcLTF3, contributes to virulence and all strains exhibit similar developmental defects in planta and in vitro.
BcLTF3 has only a minor role in regulating gene expression upon light exposure
To gain insight into the regulatory function of BcLTF3, a genome-wide transcriptomics approach was performed. Wild-type B05.10 and the ∆bcltf3 mutant were cultivated for 2 days in DD, and subsequently the cultures were exposed to white light (LP 60′) or were kept in the dark for 60′ (DD). RNA from three biological replicates was isolated, labeled and hybridized to NimbleGen microarrays containing oligonucleotides comprising all predicted B. cinerea genes (Amselem et al. 2011).
BcLTF3-dependent genes and the effect of light on gene expression in the ∆bcltf3 mutant were determined by statistical tests comparing ∆bcltf3 effects in DD (∆bcltf3/WT in DD) and upon exposure to light (∆bcltf3/WT in LP). In total, 40 genes including bcltf3 were identified as differentially expressed in the ∆bcltf3 mutant and the wild type [fold change (FC) > 2, p < 0.05] (Table S3; Fig. S3). The majority of these genes are categorized as ‘hypothetical proteins’, which makes it difficult to speculate about their function and their putative contribution to the ∆bcltf3 phenotype. Among the light-repressed genes, bcstc5 encoding a sesquiterpene synthase was found. Though the light effect was evident in both WT (−7.2-fold) and ∆bcltf3 (−3.6-fold), bcstc5 exhibited increased expression levels in ∆bcltf3 in both light conditions (2.0- and 3.4-fold). By testing the expression of bcstc5 by RT-qPCR analyses in a time course experiment, the data were confirmed. Furthermore, bcstc5 expression levels were slightly decreased in DD in OE::bcltf3 and ∆bcreg1, and were almost undetectable in the OE::bcreg1 strains (Fig. 6).
The search for putative TF-encoding genes that were possibly deregulated in ∆bcltf3, but below the defined threshold revealed one candidate (Bcin13g04470) with elevated expression levels in DD and LP 60′ (1.9- and 1.8-fold) (data not shown). Due to its light responsiveness in WT (2.4-fold), the TF was named BcLTF7. RT-qPCR data confirmed the regulation of bcltf7 expression levels by light in the wild type and ∆bcltf3, and demonstrated that they were not affected in the bcreg1 mutants (Fig. 6). However, light induction of bcltf7 expression was absent in ∆bcltf1 (1.1-fold) and ∆bcwcl1 (1.3-fold) mutants (Table 1).
In conclusion, very few genes depend on BcLTF3, indicating that the TF has minor influence on the expression of light-responsive genes in contrast to BcLTF1—at least under the tested conditions.
BcLTF4, BcLTF5 and BcLTF6 do not have obvious functions
A first genome-wide transcriptomics approach revealed three further light-responsive genes encoding putative TFs named BcLTF4-6 (Schumacher et al. 2014). Their gene expression levels were all induced upon LP of 60′ in the wild type (4.5-, 2.5-, and 3.2-fold) and were altered in ∆bcltf1 and/or ∆bcwcl1 backgrounds (Table 1). Bcltf4 and bcltf5 exhibited similar expression patterns in the wild type and the ∆bcltf1 mutant (3.4-fold and 2.1-fold), while light induction was abolished in the ∆bcwcl1 mutant (1.0-fold). Expression levels of bcltf6 were generally increased in the absence of bcltf1 (3.9- and 1.9-fold of wild type in DD and LP 60′, respectively), but light induction of bcltf6 expression was only marginally affected in ∆bcltf1 (1.6-fold) or ∆bcwcl1 (2.6-fold). In a time course experiment of the wild type, all three genes showed maximal expression around LP 180′ and their expression levels decreased after prolonged exposure to light (photoadaptation) (Fig. 8a).
BcLTF4, BcLTF5 and BcLTF6 do not have obvious functions. a Expression levels of the three genes increase upon light exposure and are subjected to photoadaptation. WT:B05.10 was incubated on solid CM with cellophane overlays for 2 days in DD or LL. Vegetative mycelia (DD) were exposed to light as indicated. Gene expression studied by RT-qPCR is given as ratio between GOIs and bcact1. Data shown are derived from three biological replicates with two technical replicates each. Asterisks indicate significant differences between light-treated and DD samples (*p < 0.05, **p < 0.001). b The three proteins are putative Zn2Cys6 TFs. Shown are the protein domains identified by Pfam Search; DNA-binding domains (Fungal Zn(2)-Cys(6) binuclear cluster domains) and further TF-related domains. c Deletion and overexpression of the genes did not affect light-dependent differentiation and virulence. Deletion mutants were generated by replacing the genes by nourseothricin resistance cassettes. For overexpressing the genes, second copies under control of the constitutive PoliC were integrated at bcniaD (see Materials and methods for details). Differentiation of conidia and sclerotia: strains were cultivated on solid CM in DD and LD. Top views of the colonies after 1 week (LD) and 2 weeks (DD) are shown. Virulence: primary leaves of P. vulgaris plants were inoculated with plugs of vegetative mycelia of the wild type and the mutants. Average lesion sizes and standard deviations (in mm) were determined after 2 days of inoculation for 12 lesions with two measurements per lesion. No significant differences were found between wild type, deletion mutants and overexpression strains
The three proteins with sizes of 410, 670 and 714 aa contain N-terminal Zn2Cys6 binuclear domains (pfam00172). BcLTF5 and BcLTF6 additionally comprise fungal specific TF domains (pfam04082, pfam04082), and putative NLSs were predicted in BcLTF4 (20-36, 18-24, 41-47 aa) and BcLTF5 (40-43 aa), suggesting that they are TFs (Fig. 8b). BlastP analyses revealed only few orthologs outside the Leotiomycetes. Putative orthologs of BcLTF4 and BcLTF6, but not of BcLTF5, exist in the close relative S. sclerotiorum. The B. cinerea-specific LTF5 exhibits similarity with Bcin15g05000, suggesting a gene duplication event. However, the latter gene is not under light control (data not shown).
To gain insight into the function of the putative TFs, deletion mutants and overexpression strains were generated and phenotypically characterized (Table 2). However, the strains exhibited phenotypes similar to those of the wild type with regard to light-dependent differentiation (conidiation in LD vs. sclerotial development in DD) and virulence (Fig. 8c). Sensitivities to stresses, i.e. to oxidative stress induced by 7.5 mM H2O2 or 500 µM menadione, to osmotic stress induced by 1.4 M sorbitol or 0.7 M NaCl, and to membrane stress induced by 0.02% SDS, were not significantly altered compared to the wild type (data not shown). Thus, no functions could be assigned to these three light-responsive TFs.
Discussion
Botrytis cinerea is a plant pathogen of high relevance, and is used as model to study the fundamental principles of necrotrophic life styles as well as plant immunity in Arabidopsis thaliana (Dean et al. 2012; Mengiste 2012). A number of genes, especially those involved in signal transduction, have been functionally characterized with the focus on their relevance for the pathogenic programs (Schumacher 2016b; Schumacher and Tudzynski 2012). Though several mutants showed different growth characteristics, including those lacking components of the MAP kinase or cAMP cascades, not much is known about the transcriptional regulation of the developmental programs.
The involved TFs and their regulation by upstream signaling cascades differ significantly in various fungi belonging to classes of the Eurotio-, Sordario-, and Leotiomycetes. For instance, A. nidulans, N. crassa and B. cinerea use unrelated TFs (BrlA, Fl, BcLTF2) as positive regulators of conidiation. Moreover, closely related species may already exhibit different behaviors due to the loss of specific programs such as the formation of asexual/sexual spores. Examples are Sordaria macrospora and S. sclerotiorum as close relatives of N. crassa and B. cinerea, respectively. In both species, the lack of conidiation is accompanied by homothallism, which means that the sexual cycle does not require a mating partner. Therefore, it is necessary to elucidate the regulatory networks in the different fungi de novo.
Light triggers conidiation in light-responsive B. cinerea strains, such as the widely used strain B05.10. Consequently, it is assumed that the involved TFs are transcriptionally up-regulated upon light exposure. A first genome-wide transcriptomics approach revealed six LTFs, which have now been all functionally characterized. BcLTF1 and BcLTF2 are involved in the induction of the conidiophore development, whereby BcLTF2 represents the positive regulator whose transcription is negatively regulated by BcLTF1 (Cohrs et al. 2016; Schumacher et al. 2014). In this study, a dual function of the third TF (BcLTF3) in conidiation, i.e. in regulation of conidiophore development and conidiogenesis, is reported. Expression of bcltf3 in vegetative mycelia is highly induced after 15′ of light exposure and decreases after prolonged exposure (Fig. 3b), which is contradictory to the expression profile of bcltf2 and the onset of conidiation. Bcltf2 expression starts to increase from 30′ on and even higher expression levels are observed during cultivation in LL (Cohrs et al. 2016). The fact that the expression of bcltf3 increases over time in aging cultures incubated in DD and LD (Fig. 3c) furthermore suggested a role of BcLTF3 in the dark during sclerotial development. In accordance with this expression pattern, the deletion of bcltf3 resulted in altered phenotypes in both light conditions. Δbcltf3 mutants form abundant conidiophores accompanied by fewer sclerotia in the dark and exhibit an earlier onset of conidiophore development accompanied by increased formation of conidiophores in the light (Fig. 5a, b), indicating that BcLTF3 represses conidiophore development in both conditions. The altered expression levels of bcltf2 in the bcltf3 mutants (Fig. 6) suggest that BcLTF3 affects conidiophore development at least partially through regulation of bcltf2. This hypothesis is in agreement with the opposing expression profiles of both genes in the time course experiments: bcltf3 encoding a repressor is induced immediately after a light signal thereby counteracting the light induction of bcltf2 expression in the early stages. During prolonged light exposure, the repression by BcLTF3 is lifted (decreased bcltf3 expression) allowing for increased expression of bcltf2 and subsequent conidiation. By this, BcLTF3 whose light-induced transcription requires the WCC (Fig. 3e) contributes to the repressing effect of the WCC/blue light on bcltf2/conidiation. But BcLTF3 is only a weak repressor of bcltf2 in contrast to BcLTF1 whose deletion almost completely prevents vegetative growth as the overexpression of bcltf2 does (Cohrs et al. 2016; Schumacher et al. 2014). Nevertheless, an additive effect in the ΔΔbcltf1/3 mutants was observed (earlier onset of conidiation in DD compared to the single mutants) (Fig. 5a), indicating that these two TFs may function independently in repressing bcltf2 expression.
Besides its function during the early stages of conidiation, BcLTF3 is crucial for the formation of mature conidia. Thus, young conidiophores appear normal at the beginning and conidium initials develop but they grow out to hyphae, which differentiate later on new (secondary) conidiophores (Fig. 5c, d, not shown). No other B. cinerea mutants with similar phenotypes are known; however, this phenotype is phenocopied by blue light treatment. Detailed studies on conidiophore development reported on the competence of blue light to inhibit conidiation and to overrule induced differentiation programs by forcing the formation of sterile hyphae: a process that was called “de-differentiation” (Suzuki et al. 1977; Suzuki and Oda 1979; Tan 1974). Hence, exposure of young conidiophores to blue light caused the outgrowth of denticles and conidium initials (Suzuki et al. 1977) resulting in similarly misshaped conidiophores as those produced by the Δbcltf3 mutant. But how blue light and the deletion of bcltf3 affect conidiogenesis in a similar fashion remains an open question at this time.
In sum, BcLTF3 is a crucial regulator of conidiation and is dispensable for virulence and stress responses. Considering the fact that the conidiation program got lost in the close relative S. sclerotiorum, which is caused/accompanied by the loss of light-responsive expression of ssltf2 (Cohrs et al. 2016), SsLTF3 may have become functionless at least in terms of regulating development. In fact, BcLTF3 and SsLTF3 share only 60% aa identity (Fig. 1a), which is below the overall aa identity of 83% (Amselem et al. 2011). Nevertheless, ssltf3 is expressed in young cultures of S. sclerotiorum in a light-dependent fashion (data not shown).
Only limited information is available on the functions of the orthologs in other fungi. CSP-1 in N. crassa is likewise transcriptionally controlled by the WCC, does not significantly affect the expression of other light-responsive genes (Chen et al. 2009) and is required for conidiogenesis, as no conidia are released from the proconidial chains (Lambreghts et al. 2009). However, the reason for the latter phenotype is still unknown. A role of CSP-1 was furthermore described in adaptive responses to antifungal azoles (Chen et al. 2016) and in the modulation of the circadian clock (Sancar et al. 2012). The orthologs Nrg1p and Nrg2p in S. cerevisiae regulate glucose metabolism and stress responses (Berkey et al. 2004; Vyas et al. 2005); NRG1 of Candida albicans is also involved in regulation of development by being critical for invasive growth and the morphological switch between yeast and hyphal growth (Lu et al. 2014). Both Nrg1p and CSP-1 bind to the consensus motif CCCCT (Berkey et al. 2004; Sancar et al. 2011), which suggests—based on the high conservation of the C2H2 domains—that BcLTF3 may recognize the same motifs in promoters of its target genes. Indeed, at least single CCCCT motifs were identified in the promoter regions of 30 out of the 40 BcLTF3-dependent genes (Table S3).
BcREG1 is a transcriptional regulator whose orthologs in different fungi have distinct roles. For instance, WOR1 and RYP1 in C. albicans and Histoplasma capsulatum regulate morphological switching (Huang et al. 2006; Nguyen and Sil 2008), SGE1 orthologs in plant pathogenic fungi such as Fusarium spp. regulate the expression of virulence-associated genes, virulence and/or secondary metabolism (Brown et al. 2014; Jonkers et al. 2012; Michielse et al. 2009, 2014) and OsaA in A. nidulans balances asexual and sexual development (Alkahyyat et al. 2015). A previous study reported on the importance of BcREG1 for virulence, phytotoxin production (BOT, BOA) and conidiogenesis (Michielse et al. 2011). Here, the role of BcREG1 in regulation of conidiation was investigated in more detail, because the expression of bcreg1 is light responsive and significantly increased during early conidiophore development. In fact, the overexpression of bcreg1 resulted in an opposing phenotype to the corresponding deletion regarding the expression of the BOT and BOA genes, virulence and the ability to establish an acidic environment (Figs. 4, 6, 7). However, the effect of the overexpression on development was more striking. Thus, conidiation in the OE::bcreg1 strains was severely reduced in the light (Fig. 5a, b), which was accompanied/caused by the low expression of bcltf2 (no light induction) (Fig. 6). On the other hand, the overexpression promoted the formation of aerial hyphae and conidia in the dark, which was moreover associated with the loss of sclerotial development (Fig. 5a, b). This indicates that BcREG1 differentially affects, by acting as repressor or activator, the conidiation programs in the two light conditions. While the effect in the light is likely due to the repression of bcltf2, the effect observed in the dark may be independent of BcLTF2 as expression of bcltf2 was not significantly altered. The latter observation suggests that a conidiation program can be induced in a parallel, BcLTF2-independent way. The BcLTF2-independent program appears distinctive as it is accompanied by increased proliferation of aerial hyphae, a phenotype that was previously described for deletion mutants of the bZIP TF BcATF1 and BcLAE1, a component of the VELVET complex (Schumacher et al. 2015; Temme et al. 2012). Considering the increasing expression levels of bcreg1 after exposure to light, BcREG1 may contribute to balance vegetative growth and conidiophore development by counteracting BcLTF2, which forces the latter program.
Conidiation is a highly regulated process and most likely requires more regulators. However, as shown here, the other three identified LTFs are dispensable for regulation of development, virulence and to cope with oxidative and osmotic stresses (Fig. 8). Another candidate TF was identified within the framework of this study. Expression of bcltf7 is highly induced by light, remains elevated after onset of conidiation and is deregulated in the ∆bcltf3 mutant (Fig. 6). BcLTF7 is a C2H2 TF orthologous to Magnaporthe oryzae ConX7, for which a role in conidiation has been recently reported (Cao et al. 2016). The characterization of bcltf7 mutants is currently in progress and will reveal whether BcLTF7 is part of the network of TFs regulating conidiophore development and conidiogenesis in B. cinerea.
References
Adams TH, Boylan MT, Timberlake WE (1988) brlA is necessary and sufficient to direct conidiophore development in Aspergillus nidulans. Cell 54:353–362
Alkahyyat F, Ni M, Kim SC, Yu JH (2015) The WOPR domain protein OsaA orchestrates development in Aspergillus nidulans. PLoS One 10:e0137554. doi:10.1371/journal.pone.0137554
Amselem J, Cuomo CA, van Kan JA, Viaud M, Benito EP, Couloux A, Coutinho PM, de Vries RP, Dyer PS, Fillinger S, Fournier E, Gout L, Hahn M, Kohn L, Lapalu N, Plummer KM, Pradier JM, Quevillon E, Sharon A, Simon A, ten Have A, Tudzynski B, Tudzynski P, Wincker P, Andrew M, Anthouard V, Beever RE, Beffa R, Benoit I, Bouzid O, Brault B, Chen Z, Choquer M, Collemare J, Cotton P, Danchin EG, Da Silva C, Gautier A, Giraud C, Giraud T, Gonzalez C, Grossetete S, Guldener U, Henrissat B, Howlett BJ, Kodira C, Kretschmer M, Lappartient A, Leroch M, Levis C, Mauceli E, Neuveglise C, Oeser B, Pearson M, Poulain J, Poussereau N, Quesneville H, Rascle C, Schumacher J, Segurens B, Sexton A, Silva E, Sirven C, Soanes DM, Talbot NJ, Templeton M, Yandava C, Yarden O, Zeng Q, Rollins JA, Lebrun MH, Dickman M (2011) Genomic analysis of the necrotrophic fungal pathogens Sclerotinia sclerotiorum and Botrytis cinerea. PLoS Genet 7:e1002230. doi:10.1371/journal.pgen.1002230
Bailey LA, Ebbole DJ (1998) The fluffy gene of Neurospora crassa encodes a Gal4p-type C6 zinc cluster protein required for conidial development. Genetics 148:1813–1820
Bailey-Shrode L, Ebbole DJ (2004) The fluffy gene of Neurospora crassa is necessary and sufficient to induce conidiophore development. Genetics 166:1741–1749
Berkey CD, Vyas VK, Carlson M (2004) Nrg1 and Nrg2 transcriptional repressors Are differently regulated in response to carbon source. Eukaryot Cell 3:311–317. doi:10.1128/ec.3.2.311-317.2004
Braga GU, Rangel DE, Fernandes EK, Flint SD, Roberts DW (2015) Molecular and physiological effects of environmental UV radiation on fungal conidia. Curr Genet 61:405–425. doi:10.1007/s00294-015-0483-0
Brown DW, Busman M, Proctor RH (2014) Fusarium verticillioides SGE1 is required for full virulence and regulates expression of protein effector and secondary metabolite biosynthetic genes. Mol Plant Microbe Interact 27:809–823. doi:10.1094/MPMI-09-13-0281-R
Büttner P, Koch F, Voigt K, Quidde T, Risch S, Blaich R, Bruckner B, Tudzynski P (1994) Variations in ploidy among isolates of Botrytis cinerea: implications for genetic and molecular analyses. Curr Genet 25:445–450
Canessa P, Schumacher J, Hevia MA, Tudzynski P, Larrondo LF (2013) Assessing the effects of light on differentiation and virulence of the plant pathogen Botrytis cinerea: characterization of the White Collar Complex. PLoS One 8:e84223. doi:10.1371/journal.pone.0084223
Cao H, Huang P, Zhang L, Shi Y, Sun D, Yan Y, Liu X, Dong B, Chen G, Snyder JH, Lin F, Lu J (2016) Characterization of 47 Cys -His zinc finger proteins required for the development and pathogenicity of the rice blast fungus Magnaporthe oryzae. New Phytol. doi:10.1111/nph.13948
Cenis JL (1992) Rapid extraction of fungal DNA for PCR amplification. Nucleic Acids Res 20:2380
Chen CH, Ringelberg CS, Gross RH, Dunlap JC, Loros JJ (2009) Genome-wide analysis of light-inducible responses reveals hierarchical light signalling in Neurospora. EMBO J 28:1029–1042. doi:10.1038/emboj.2009.54
Chen X, Xue W, Zhou J, Zhang Z, Wei S, Liu X, Sun X, Wang W, Li S (2016) De-repression of CSP-1 activates adaptive responses to antifungal azoles. Sci Rep 6:19447. doi:10.1038/srep19447
Christianson TW, Sikorski RS, Dante M, Shero JH, Hieter P (1992) Multifunctional yeast high-copy-number shuttle vectors. Gene 110:119–122. doi:10.1016/0378-1119(92)90454-W
Cohrs KC, Simon A, Viaud M, Schumacher J (2016) Light governs asexual differentiation in the grey mould fungus Botrytis cinerea via the putative transcription factor BcLTF2. Environ Microbiol 18:4068–4086. doi:10.1111/1462-2920.13431
Cole GT (1986) Models of cell differentiation in conidial fungi. Microbiol Rev 50:95
Colot HV, Park G, Turner GE, Ringelberg C, Crew CM, Litvinkova L, Weiss RL, Borkovich KA, Dunlap JC (2006) A high-throughput gene knockout procedure for Neurospora reveals functions for multiple transcription factors. Proc Natl Acad Sci USA 103:10352–10357. doi:10.1073/pnas.0601456103
Dalmais B, Schumacher J, Moraga J, P LEP, Tudzynski B, Collado IG, Viaud M (2011) The Botrytis cinerea phytotoxin botcinic acid requires two polyketide synthases for production and has a redundant role in virulence with botrydial. Mol Plant Pathol 12:564–579 doi:10.1111/j.1364-3703.2010.00692.x
Dean R, Van Kan JA, Pretorius ZA, Hammond-Kosack KE, Di Pietro A, Spanu PD, Rudd JJ, Dickman M, Kahmann R, Ellis J, Foster GD (2012) The Top 10 fungal pathogens in molecular plant pathology. Mol Plant Pathol 13:414–430. doi:10.1111/j.1364-3703.2011.00783.x
Epton HAS, Richmond DV (1980) Formation, structure and germination of conidia. In: Coley-Smith JR, Verhoeff K, Jarvis WR (eds) The biology of Botrytis. Academic Press, London, pp 41–83
Fillinger S, Elad Y (2016) Botrytis—the fungus, the pathogen and its management in agricultural systems. Springer
Fischer R, Aguirre J, Herrera-Estrella A, Corrochano LM (2016) The complexity of fungal vision. Microbiol Spectr 410.1128/microbiolspec.FUNK-0020-2016
Fuller KK, Loros JJ, Dunlap JC (2015) Fungal photobiology: visible light as a signal for stress, space and time. Current genetics 61:275–288
Han KH, Han KY, Yu JH, Chae KS, Jahng KY, Han DM (2001) The nsdD gene encodes a putative GATA-type transcription factor necessary for sexual development of Aspergillus nidulans. Mol Microbiol 41:299–309
Han Y, Joosten HJ, Niu W, Zhao Z, Mariano PS, McCalman M, van Kan J, Schaap PJ, Dunaway-Mariano D (2007) Oxaloacetate hydrolase, the C–C bond lyase of oxalate secreting fungi. J Biol Chem 282:9581–9590. doi:10.1074/jbc.M608961200
Huang G, Wang H, Chou S, Nie X, Chen J, Liu H (2006) Bistable expression of WOR1, a master regulator of white-opaque switching in Candida albicans. Proc Natl Acad Sci USA 103:12813–12818. doi:10.1073/pnas.0605270103
Idnurm A, Verma S, Corrochano LM (2010) A glimpse into the basis of vision in the kingdom Mycota. Fungal Genet Biol 47:881–892. doi:10.1016/j.fgb.2010.04.009
Jonkers W, Dong Y, Broz K, Kistler HC (2012) The Wor1-like protein Fgp1 regulates pathogenicity, toxin synthesis and reproduction in the phytopathogenic fungus Fusarium graminearum. PLoS Pathog 8:e1002724. doi:10.1371/journal.ppat.1002724
Krijgsheld P, Bleichrodt R, Van Veluw G, Wang F, Müller W, Dijksterhuis J, Wösten H (2013) Development in Aspergillus. Stud Mycol 74:1–29
Kumagai T (1988) Photocontrol of fungal development. Photochem Photobiol 47:889–896
Lambreghts R, Shi M, Belden WJ, Park D, Henn MR, Galagan JE, Baştürkmen M, Birren BW, Sachs MS, Dunlap JC (2009) A high-density single nucleotide polymorphism map for Neurospora crassa. Genetics 181:767–781
Lee MK, Kwon NJ, Choi JM, Lee IS, Jung S, Yu JH (2014) NsdD is a key repressor of asexual development in Aspergillus nidulans. Genetics 197:159–173. doi:10.1534/genetics.114.161430
Lee MK, Kwon NJ, Lee IS, Jung S, Kim SC, Yu JH (2016) Negative regulation and developmental competence in Aspergillus. Sci Rep 6:28874. doi:10.1038/srep28874
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Lu Y, Su C, Liu H (2014) Candida albicans hyphal initiation and elongation. Trends Microbiol 22:707–714. doi:10.1016/j.tim.2014.09.001
Mengiste T (2012) Plant immunity to necrotrophs. Annu Rev Phytopathol 50:267–294
Michielse CB, van Wijk R, Reijnen L, Manders EM, Boas S, Olivain C, Alabouvette C, Rep M (2009) The nuclear protein Sge1 of Fusarium oxysporum is required for parasitic growth. PLoS Pathog 5:e1000637. doi:10.1371/journal.ppat.1000637
Michielse CB, Becker M, Heller J, Moraga J, Collado IG, Tudzynski P (2011) The Botrytis cinerea Reg1 protein, a putative transcriptional regulator, is required for pathogenicity, conidiogenesis, and the production of secondary metabolites. Mol Plant Microbe Interact 24:1074–1085. doi:10.1094/MPMI-01-11-0007
Michielse CB, Studt L, Janevska S, Sieber CM, Arndt B, Espino JJ, Humpf HU, Guldener U, Tudzynski B (2014) The global regulator FfSge1 is required for expression of secondary metabolite gene clusters but not for pathogenicity in Fusarium fujikuroi. Environ Microbiol. doi:10.1111/1462-2920.12592
Mooney JL, Yager LN (1990) Light is required for conidiation in Aspergillus nidulans. Genes Dev 4:1473–1482
Nguyen VQ, Sil A (2008) Temperature-induced switch to the pathogenic yeast form of Histoplasma capsulatum requires Ryp1, a conserved transcriptional regulator. Proc Natl Acad Sci USA 105:4880–4885. doi:10.1073/pnas.0710448105
Oldenburg KR, Vo KT, Michaelis S, Paddon C (1997) Recombination-mediated PCR-directed plasmid construction in vivo in yeast. Nucleic Acids Res 25:451–452. doi:10.1093/nar/25.2.451
Olmedo M, Ruger-Herreros C, Corrochano LM (2010) Regulation by blue light of the fluffy gene encoding a major regulator of conidiation in Neurospora crassa. Genetics 184:651–658. doi:10.1534/genetics.109.109975
Park HS, Yu JH (2012) Genetic control of asexual sporulation in filamentous fungi. Curr Opin Microbiol 15:669–677. doi:10.1016/j.mib.2012.09.006
Pontecorvo G, Roper JA, Hemmons LM, Macdonald KD, Bufton AWJ (1953) The genetics of Aspergillus nidulans. Adv Genet Incorp Mol Genet Med 5:141–238. doi:10.1016/S0065-2660(08)60408-3
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning Cold Spring Harbor Laboratory Press, New York, USA
Sancar G, Sancar C, Brugger B, Ha N, Sachsenheimer T, Gin E, Wdowik S, Lohmann I, Wieland F, Hofer T, Diernfellner A, Brunner M (2011) A global circadian repressor controls antiphasic expression of metabolic genes in Neurospora. Mol Cell 44:687–697. doi:10.1016/j.molcel.2011.10.019
Sancar G, Sancar C, Brunner M (2012) Metabolic compensation of the Neurospora clock by a glucose-dependent feedback of the circadian repressor CSP1 on the core oscillator. Genes Dev 26:2435–2442. doi:10.1101/gad.199547.112
Schumacher J (2012) Tools for Botrytis cinerea: new expression vectors make the gray mold fungus more accessible to cell biology approaches. Fungal Genet Biol 49:483–497. doi:10.1016/j.fgb.2012.03.005
Schumacher J (2016a) DHN melanin biosynthesis in the plant pathogenic fungus Botrytis cinerea is based on two developmentally regulated key enzyme (PKS)-encoding genes. Mol Microbiol 99:729–748. doi:10.1111/mmi.13262
Schumacher J (2016b) Signal transduction cascades regulating differentiation and virulence in Botrytis cinerea Botrytis—the fungus, the pathogen and its management in agricultural systems. Springer, pp 247-267
Schumacher J, Tudzynski P (2012) Morphogenesis and infection in Botrytis cinerea. In: Pérez-Martín J, Di Pietro A (eds) Morphogenesis and Pathogenicity in Fungi. Springer-Verlag, Berlin Heidelberg, pp 225–241
Schumacher J, Pradier JM, Simon A, Traeger S, Moraga J, Collado IG, Viaud M, Tudzynski B (2012) Natural variation in the VELVET gene bcvel1 affects virulence and light-dependent differentiation in Botrytis cinerea. PLoS One 7:e47840. doi:10.1371/journal.pone.0047840
Schumacher J, Simon A, Cohrs KC, Viaud M, Tudzynski P (2014) The transcription factor BcLTF1 regulates virulence and light responses in the necrotrophic plant pathogen Botrytis cinerea. PLoS Genet 10:e1004040. doi:10.1371/journal.pgen.1004040
Schumacher J, Simon A, Cohrs KC, Traeger S, Porquier A, Dalmais B, Viaud M, Tudzynski B (2015) The VELVET complex in the gray mold fungus Botrytis cinerea: impact of BcLAE1 on differentiation, secondary metabolism, and virulence. Mol Plant Microbe Interact 28:659–674. doi:10.1094/MPMI-12-14-0411-R
Simon A, Biot E (2010) ANAIS: analysis of NimbleGen arrays interface. Bioinformatics 26:2468–2469. doi:10.1093/bioinformatics/btq410
Staben C, Jensen B, Singer M, Pollock J, Schechtman M, Kinsey J, Selker E (1989) Use of a bacterial hygromycin B resistance gene as a dominant selectable marker in Neurospora crassa transformation. Fungal Genet Newsl 36:79–81
Sun X, Yu L, Lan N, Wei S, Yu Y, Zhang H, Zhang X, Li S (2012) Analysis of the role of transcription factor VAD-5 in conidiation of Neurospora crassa. Fungal Genet Biol 49:379–387. doi:10.1016/j.fgb.2012.03.003
Suzuki Y, Oda Y (1979) Inhibitory loci of both blue and near ultraviolet lights on lateral-type sclerotial development in Botrytis cinerea. Ann Phytopath Soc Japan 45:54–61
Suzuki Y, Kumagai T, Oda Y (1977) Locus of blue and near ultraviolet reversible photoreaction in the stages of conidial development in Botrytis cinerea. J Gen Microbiol 98:199–204
Tan KK (1974) Blue light inhibition of sporulation in Botrytis cinerea. J Gener Microbiol 82:191–200
Taylor J, Jacobson D, Fisher M (1999) The evolution of asexual fungi: reproduction, speciation and classification. Annu Rev Phytopathol 37:197–246
Temme N, Oeser B, Massaroli M, Heller J, Simon A, Collado IG, Viaud M, Tudzynski P (2012) BcAtf1, a global regulator, controls various differentiation processes and phytotoxin production in Botrytis cinerea. Mol Plant Pathol 13:704–718. doi:10.1111/j.1364-3703.2011.00778.x
Vyas VK, Berkey CD, Miyao T, Carlson M (2005) Repressors Nrg1 and Nrg2 regulate a set of stress-responsive genes in Saccharomyces cerevisiae. Eukaryot Cell 4:1882–1891. doi:10.1128/EC.4.11.1882-1891.2005
Williamson B, Tudzynski B, Tudzynski P, van Kan JA (2007) Botrytis cinerea: the cause of grey mould disease. Mol Plant Pathol 8:561–580. doi:10.1111/j.1364-3703.2007.00417.x
Wyatt TT, Wösten HA, Dijksterhuis J (2013) Fungal spores for dispersion in space and time. Adv Appl Microbiol 85:43–91
Acknowledgements
We thank Paul Tudzynski for support and discussion, Kim Cohrs for providing the expression data on the bctlf2 mutants and for help with microscopy, Bettina Richter and Charlotte Kaiser for their contributions to mutant generation, and Dominik Wagner for help with the microarray experiments. This study was supported by the Deutsche Forschungsgemeinschaft (SCHU 2833/4-1).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Brandhoff, B., Simon, A., Dornieden, A. et al. Regulation of conidiation in Botrytis cinerea involves the light-responsive transcriptional regulators BcLTF3 and BcREG1. Curr Genet 63, 931–949 (2017). https://doi.org/10.1007/s00294-017-0692-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-017-0692-9