Abstract
Bacteria have evolved to adapt to various conditions and respond to certain stress conditions. The ability to sense and efficiently reply to these environmental effects involve versatile array of sensors and global or specific regulators. Interestingly, modulation of the levels of active global regulators enables bacteria to respond to diverse signals via a single central transcriptional regulator and to activate or repress certain differentiation pathways at a spatio-temporal manner. The Gram-positive Bacillus subtilis is an ideal bacterium to study how membrane bound and cytoplasmic sensor kinases affect the level of phosphorylated global regulator, Spo0A which in response activates genes related to sliding, biofilm formation, and sporulation. In addition, other global regulators, including the two-component system DegS-DegU, modulate overlapping and complementary genes in B. subtilis related to surface colonization and biofilm formation. The intertwinement of global regulatory systems also allows the accurate modulation of differentiation pathways. Studies in the last decade enable us to get a deeper insight into the role of global regulators on the smooth transition of developmental processes in B. subtilis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacteria appeared on earth billions of years ago and have well customized to diverse environmental conditions. Studying the microbial responses and the adjustments of bacterial life styles helps us to understand how certain signals are perceived by these tiny organisms and how certain regulatory proteins contribute to the adaptation processes. Deeper insights into the control of gene regulatory networks reveal how the complexity of signal perception and fine-tuning the transcription of hundreds of genes affect the bacterial differentiation processes.
Soil dwelling bacteria are well adapted to respond to the environmental changes and commit to a certain differentiation pathway depending on the ecological conditions. The Gram-positive bacterium, Bacillus subtilis shows various differentiation traits under laboratory conditions that are related to its survival in the environment. For example, extracellular matrix dependent biofilm formation of B. subtilis enables the bacterium to attach and colonize the root surface (Bais et al. 2004). Depending on the circumstances, B. subtilis can also colonize surfaces using flagellum-driven single cell based movement (swimming), motility-dependent multicellular spreading (swarming), or growth-powered passive surface translocation (sliding) (Kearns 2010). Swarming and sliding are generally observed in wild isolates of B. subtilis and lost in domesticated laboratory strains (Kinsinger et al. 2003; Patrick and Kearns 2009; Pollak et al. 2015). While the cultivating conditions determine which process is activated, it seems that the activation of cellular machineries required for translocation is modulated by precise level of certain sets of global regulators (Grau et al. 2015).
Global regulators influence the differentiation of B. subtilis
Intriguingly, the transition among the differentiation pathways to slide, to develop highly wrinkled colony biofilms, or to enter sporulation depends on the level of the phosphorylated Spo0A (Spo0A~P) protein in B. subtilis (Grau et al. 2015). The global regulator, Spo0A has been originally identified due to its requirement for sporulation (Trowsdale et al. 1978). Analysis of the Spo0A~P modulated genes revealed that the list of genes up- or downregulated depends on the level of Spo0A~P in the cells. While the transcription of abrB gene that codes for a transition state regulator in B. subtilis is repressed already at low levels of Spo0A~P, activation of sporulation related gene expression requires high levels of Spo0A~P (Fujita et al. 2005). Such a differential control of genes is achieved by variance in the Spo0A~P binding sites located in the promoter and regulatory regions of directly affected genes that defines the binding properties of Spo0A~P to these sites and therefore transcriptional activation or repression. The gradual increase in the phosphorylation level of Spo0A is mediated by various histidine kinases in B. subtilis (Fujita and Losick 2005). B. subtilis harbours five histidine kinases, the cytoplasmic KinA and the membrane-bound kinases, KinB to KinE (Jiang et al. 2000). Increased levels of KinA, KinB, or KinC trigger the entry into the sporulation pathway (Fujita and Losick 2005). Upon activation of these kinases, phosphorylation of Spo0A is achieved via a phosphorelay including Spo0F and Spo0B proteins (Hoch 1993). Numerous signals have been described that are perceived by these kinases (Mhatre et al. 2014), including anaerobic conditions [KinA and KinB (Kolodkin-Gal et al. 2013)], plant-derived signals [KinC and KinD (Beauregard et al. 2013; Chen et al. 2013)], polyketides [KinC (López et al. 2009)], combination of glycerol and manganese [KinD (Shemesh and Chai 2013)], osmotic conditions [KinD (Rubinstein et al. 2012)], and potassium level [KinB (Grau et al. 2015)]. In response to these or other so far unknown signals affecting the kinase activity of Kin proteins, Spo0A~P level increases in the cells and modulates the transcription of certain sets of genes. Importantly, the increase in the Spo0A~P level is heterogeneous within a clonal population (Veening et al. 2005). Positive feedback loops contribute to the hetero-chronic timing of Spo0A~P dependent activation (Chastanet et al. 2010; de Jong et al. 2010) that greatly determine the level of phenotypic heterogeneity between individual cells. Heterogeneity is proposed to originate from the variable gene activities of phosphorelay components that result in high cell–cell variability (de Jong et al. 2010). The versatility of single master regulator modulated pathways is not restricted to Spo0A. The level of phosphorylated DegU (DegU~P) determines whether B. subtilis commits to competence for DNA uptake (DegU in the non-phosphorylated form), swims or swarms (low DegU~P level), activate genes related to biofilm formation (medium DegU~P level) or increase protease production (high DegU~P level) (Kobayashi 2007; Verhamme et al. 2007). Importantly, several genes and phenotypes are affected by both Spo0A~P and DegU~P, including biofilm formation (Marlow et al. 2014) and protease secretion (Veening et al. 2008a) presenting a naturally occurring logic-AND gates for the expression of certain genes (Veening et al. 2008b).
The level of phosphorylated Spo0A defines the activation of sliding, biofilm formation or sporulation
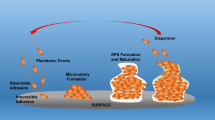
The seemingly continuous transition between the different differentiation pathways was recently reported in B. subtilis using identical environmental conditions, but synthetically altering the level of Spo0A~P (Grau et al. 2015). Depending on the induction level of the sad67 gene that codes for a phosphorylation-independent active Spo0A protein (Spo0A*), distinct phenotypic traits could be stimulated (Fig. 1). Using semi-solid nutrient rich medium (lysogeny broth medium with 0.7 % agar), low, medium, or high levels of Spo0A* activate sliding, architecturally complex colony biofilm development or high level of spore production, respectively. Both sliding and biofilm formation require the production of exopolysaccharide (EPS) and an amphiphilic protein (BslA). Expansion of colony biofilm is proposed to be driven through the secretion of EPS (Seminara et al. 2012; van Gestel et al. 2014) by generating osmotic pressure gradient in the extracellular space. Similar mechanism may act during sliding, where the EPS driven colony expansion additionally requires hydrophobins, BslA and surfactin (Grau et al. 2015). It is appealing to hypothesize that precise levels of these components determine whether cells colonize a surface via sliding or through slight expansion of the colony biofilm. Alternatively, sliding perceives spatial fixing of cells in the biofilm, i.e., surface attachment and fruiting body formation. However, it is clear that gradual activation of a global activator controls the transition between surface spreading and biofilm formation, previously believed to be antagonistic and independent prokaryotic social behaviours (Grau et al. 2015). The importance of single cell based motility has recently been reported during B. subtilis biofilm development on the air-medium interface verifying the ecological connection of distinctive pathways (Hölscher et al. 2015).
Gradual increase in the level of Spo0A~P defines which developmental pathways are activated in B. subtilis. Sliding, biofilm development or production of resistant spores are induced at certain levels of Spo0A~P protein in the bacterial cells (Grau et al. 2015)
Sensor kinases influence which differentiation pathways are activated by Spo0A~P
The gradual activation of Spo0A~P is believed to be mediated by the differential activation of histidine kinases of B. subtilis. While effective sliding was shown to be dependent on both KinB and KinC (Grau et al. 2015), the KinC and KinD proteins seems to mainly trigger biofilm development under aerobic conditions (Mhatre et al. 2014), and KinA and KinB present the major input during entry into sporulation. KinB-initiated sliding of B. subtilis was proposed be triggered by the level of potassium (Grau et al. 2015). Sensing of potassium in the medium requires a protein domain of KinB that shows an intriguing homology to the conserved K+-filter sequence of eukaryotic and prokaryotic potassium channels. It is hypothesized that while both KinB and KinC are required for the activation of sliding, these kinases might respond to differential levels of potassium (Grau et al. 2015). Finally, it is reasonable to assume that these kinases control the spatial activation of these pathways to guarantee that these traits are sequentially activated during surface colonization. Indeed, colony biofilms were reported to enclose spatially and temporally separated populations with activation of genes related to flagellum-dependent motility, matrix production or spore development (Vlamakis et al. 2008). Thus, it will be fascinating to examine how the activation of these differentiation pathways is mediated during sliding at the single cell levels and determine how heterogeneity contribute to the smooth transition between these phenotypes.
Protein phosphorylation is not only restricted to regulatory proteins in bacteria, but also found in diverse processes, including proteins with enzymatic function or those involved in cell division (Derouiche et al. 2015). By adjusting protein phosphorylation, bacteria have implemented a general way to modulate the function of global regulators depending on the environmental cues. Sensory kinases take an important role in differentiating the signals and adequately modifying activities of regulators that initiate certain bacterial differentiation processes.
References
Bais HP, Fall R, Vivanco JM (2004) Biocontrol of Bacillus subtilis against infection of Arabidopsis roots by Pseudomonas syringae is facilitated by biofilm formation and surfactin production. Plant Physiol 134:307–319
Beauregard PB, Chai Y, Vlamakis H, Losick R, Kolter R (2013) Bacillus subtilis biofilm induction by plant polysaccharides. Proc Natl Acad Sci USA 110:E1621–E1630
Chastanet A, Vitkup D, Yuan GC, Norman TM, Liu JS, Losick RM (2010) Broadly heterogeneous activation of the master regulator for sporulation in Bacillus subtilis. Proc Natl Acad Sci USA 107:8486–8491. doi:10.1073/pnas.1002499107
Chen Y, Yan F, Chai Y, Liu H, Kolter R, Losick R, Guo Jh (2013) Biocontrol of tomato wilt disease by Bacillus subtilis isolates from natural environments depends on conserved genes mediating biofilm formation. Environ Microbiol 15:848–864
de Jong IG, Veening JW, Kuipers OP (2010) Heterochronic phosphorelay gene expression as a source of heterogeneity in Bacillus subtilis spore formation. J Bacteriol 192:2053–2067. doi:10.1128/JB.01484-09
Derouiche A, Shi L, Kalantari A, Mijakovic I (2015) Evolution and tinkering: what do a protein kinase, a transcriptional regulator and chromosome segregation/cell division proteins have in common? Curr Genet. doi:10.1007/s00294-015-0513-y
Fujita M, Losick R (2005) Evidence that entry into sporulation in Bacillus subtilis is governed by a gradual increase in the level and activity of the master regulator Spo0A. Genes Dev 19:2236–2244
Fujita M, Gonzalez-Pastor JE, Losick R (2005) High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis. J Bacteriol 187:1357–1368. doi:10.1128/JB.187.4.1357-1368.2005
Grau RR et al (2015) A duo of potassium-responsive histidine kinases govern the multicellular destiny of Bacillus subtilis. mBio 6:e00581. doi:10.1128/mBio.00581-15
Hoch JA (1993) The phosphorelay signal transduction pathway in the initiation of Bacillus subtilis sporulation. J Cell Biochem 51:55–61. doi:10.1002/jcb.240510111
Hölscher T et al (2015) Motility, chemotaxis and aerotaxis contribute to competitiveness during bacterial pellicle biofilm development. J Mol Biol. doi:10.1016/j.jmb.2015.06.014
Jiang M, Shao W, Perego M, Hoch JA (2000) Multiple histidine kinases regulate entry into stationary phase and sporulation in Bacillus subtilis. Mol Microbiol 38:535–542
Kearns DB (2010) A field guide to bacterial swarming motility. Nat Rev Microbiol 8:634–644
Kinsinger RF, Shirk MC, Fall R (2003) Rapid surface motility in Bacillus subtilis is dependent on extracellular surfactin and potassium ion. J Bacteriol 185:5627–5631
Kobayashi K (2007) Gradual activation of the response regulator DegU controls serial expression of genes for flagellum formation and biofilm formation in Bacillus subtilis. Mol Microbiol 66:395–409. doi:10.1111/j.1365-2958.2007.05923.x
Kolodkin-Gal I, Elsholz AK, Muth C, Girguis PR, Kolter R, Losick R (2013) Respiration control of multicellularity in Bacillus subtilis by a complex of the cytochrome chain with a membrane-embedded histidine kinase. Genes Dev 27:887–899. doi:10.1101/gad.215244.113
López D, Fischbach MA, Chu F, Losick R, Kolter R (2009) Structurally diverse natural products that cause potassium leakage trigger multicellularity in Bacillus subtilis. Proc Natl Acad Sci USA 106:280–285. doi:10.1073/pnas.0810940106
Marlow VL, Porter M, Hobley L, Kiley TB, Swedlow JR, Davidson FA, Stanley-Wall NR (2014) Phosphorylated DegU manipulates cell fate differentiation in the Bacillus subtilis biofilm. J Bacteriol 196:16–27. doi:10.1128/JB.00930-13
Mhatre E, Monterrosa RG, Kovács ÁT (2014) From environmental signals to regulators: modulation of biofilm development in Gram-positive bacteria. J Basic Microbiol 54:616–632. doi:10.1002/jobm.201400175
Patrick JE, Kearns DB (2009) Laboratory strains of Bacillus subtilis do not exhibit swarming motility. J Bacteriol 191:7129–7133. doi:10.1128/JB.00905-09
Pollak S, Omer Bendori S, Eldar A (2015) A complex path for domestication of B. subtilis sociality. Curr Genet. doi:10.1007/s00294-015-0479-9
Rubinstein SM, Kolodkin-Gal I, McLoon A, Chai L, Kolter R, Losick R, Weitz DA (2012) Osmotic pressure can regulate matrix gene expression in Bacillus subtilis. Mol Microbiol 86:426–436. doi:10.1111/j.1365-2958.2012.08201.x
Seminara A et al (2012) Osmotic spreading of Bacillus subtilis biofilms driven by an extracellular matrix. Proc Natl Acad Sci USA 109:1116–1121
Shemesh M, Chai Y (2013) A combination of glycerol and manganese promotes biofilm formation in Bacillus subtilis via histidine kinase KinD signaling. J Bacteriol 195:2747–2754. doi:10.1128/JB.00028-13
Trowsdale J, Chen SM, Hoch JA (1978) Evidence that spo0A mutations are recessive in spo0A-/spo0A + merodiploid strains of Bacillus subtilis. J Bacteriol 135:99–113
van Gestel J, Weissing FJ, Kuipers OP, Kovács ÁT (2014) Density of founder cells affects spatial pattern formation and cooperation in Bacillus subtilis biofilms. ISME J 8:2069–2079. doi:10.1038/ismej.2014.52
Veening JW, Hamoen LW, Kuipers OP (2005) Phosphatases modulate the bistable sporulation gene expression pattern in Bacillus subtilis. Mol Microbiol 56:1481–1494. doi:10.1111/j.1365-2958.2005.04659.x
Veening JW, Igoshin OA, Eijlander RT, Nijland R, Hamoen LW, Kuipers OP (2008a) Transient heterogeneity in extracellular protease production by Bacillus subtilis. Mol Syst Biol 4:184. doi:10.1038/msb.2008.18
Veening JW, Smits WK, Kuipers OP (2008b) Bistability, epigenetics, and bet-hedging in bacteria. Annu Rev Microbiol 62:193–210. doi:10.1146/annurev.micro.62.081307.163002
Verhamme DT, Kiley TB, Stanley-Wall NR (2007) DegU co-ordinates multicellular behaviour exhibited by Bacillus subtilis. Mol Microbiol 65:554–568. doi:10.1111/j.1365-2958.2007.05810.x
Vlamakis H, Aguilar C, Losick R, Kolter R (2008) Control of cell fate by the formation of an architecturally complex bacterial community. Genes Dev 22:945–953. doi:10.1101/gad.1645008
Acknowledgments
Work in the laboratory of A. T. K. is supported by a Marie Skłodowska Curie career integration grant (PheHetBacBiofilm), Grants KO4741/2-1 and KO4741/3-1 from the Deutsche Forschungsgemeinschaft (DFG), and a Jena School for Microbial Communication (JSMC) start-up fund.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Rights and permissions
About this article
Cite this article
Kovács, Á.T. Bacterial differentiation via gradual activation of global regulators. Curr Genet 62, 125–128 (2016). https://doi.org/10.1007/s00294-015-0524-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-015-0524-8