Abstract
Vibrio parahaemolyticus, the leading cause of bacterial seafood-associated gastroenteritis, can form biofilms. In this work, the gene expression profiles of V. parahaemolyticus during biofilm formation were investigated by transcriptome sequencing. A total of 183, 503, and 729 genes were significantly differentially expressed in the bacterial cells at 12, 24 and 48 h, respectively, compared with that at 6 h. Of these, 92 genes were consistently activated or repressed from 6 to 48 h. The genes involved in polar flagellum, chemotaxis, mannose-sensitive haemagglutinin type IV pili, capsular polysaccharide, type III secretion system 1 (T3SS1), T3SS2, thermostable direct hemolysin (TDH), type VI secretion system 1 (T6SS1) and T6SS2 were downregulated, whereas those involved in V. parahaemolyticus pathogenicity island (Vp-PAI) (except for T3SS2 and TDH) and membrane fusion proteins were upregulated. Three extracellular protease genes (vppC, prtA and VPA1071) and a dozen of outer membrane protein encoding genes were also significantly differentially expressed during biofilm formation. In addition, five putative c-di-GMP metabolism-associated genes were significantly differentially expressed, which may account for the drop in c-di-GMP levels after the beginning of biofilm formation. Moreover, many putative regulatory genes were significantly differentially expressed, and more than 1000 putative small non-coding RNAs were detected, suggesting that biofilm formation was tightly regulated by complex regulatory networks. The data provided a global view of gene expression profiles during biofilm formation, showing that the significantly differentially expressed genes were involved in multiple cellular pathways, including virulence, biofilm formation, metabolism, and regulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biofilms are surface-attached aggregates of bacterial cells that are embedded in a self-produced matrix known as extracellular polymeric substances, which are mainly composed of exopolysaccharides (EPS), extracellular proteins, lipids and extracellular DNA [1, 2]. Biofilm matrix protects the embedded microbial cells from stress conditions such as the usage of disinfectants and antibiotics [2]. Biofilm is generally formed through four basic stages: initial attachment, biofilm development, maturation stage and dispersal stage [1]. Each stage is strictly regulated by complex regulatory processes such as quorum sensing (QS), cyclic di-GMP (c-di-GMP) and small non-coding RNA (sRNA) [1].
Vibrio parahaemolyticus, the leading cause of bacterial seafood-borne gastroenteritis, is widely found in marine ecosystems. Human infections are usually caused by consumption of raw or undercooked seafood [3]. V. parahaemolyticus not only produces various virulence factors, including thermostable direct hemolysin (TDH), TDH-related hemolysin (TRH), type III secretion system 1 (T3SS1), T3SS2, type VI secretion system 1 (T6SS1), and T6SS2, but also has a strong ability to form biofilms on the surface [4, 5]. Biofilm formation requires specific structures, including flagella, type IV pili, and EPS [5,6,7,8]. Flagella are required for the initial stages of biofilm formation, and the polar flagellum deficient strains failed to form mature biofilms [5, 8]. V. parahaemolyticus expresses two kinds of type IV pili, mannose-sensitive haemagglutinin type IV pili (MSHA) and chitin-regulated pili (ChiRP), that contribute to surface attachment and agglutination [7]. EPS is the most prevalent component of biofilms that holds bacterial cells together on the surface [5]. The cpsA-K and scvA-O gene clusters are responsible for EPS synthesis in V. parahaemolyticus that are required for the mature biofilm formation [5, 9, 10]. In addition, other substances, including extracellular proteins, extracellular DNA, and membrane fusion proteins (Mfp), also play roles in biofilm formation [6, 8].
Biofilm formation by V. parahaemolyticus is regulated by a number of factors, including positive regulators such as OxyR [11], ToxR [12], RpoN [13], CpsQ [14], and AphA [15], negative regulators such as OpaR [16], QsvR [8], RobA [17], and VP0610 [18], environmental cues such as temperature [19] and salinity [20], cyclic nucleotide messengers such as c-di-GMP [5, 21], and sRNAs such as Qrr1-5 [22]. Moreover, comparative transcriptome analysis also revealed that there were significant differences in the expression levels of 956 genes in biofilm cells compared with planktonic cells in V. parahaemolyticus, including many putative regulatory genes and biofilm-associated genes [23]. In addition, biofilm cells and planktonic cells are interchangeable in the processes of biofilm formation [2]. Therefore, clarifying the gene expression profiles in both biofilm cells and planktonic cells during biofilm formation is helpful to understand the mechanisms of biofilm formation.
In this work, total RNA of V. parahaemolyticus RIMD2210633 were extracted simultaneously from biofilms and planktonic fractions for the RNA sequencing (RNA-seq) analysis to compare the gene expression profiles during biofilm formation. The data showed that the significantly differentially expressed genes (DEGs) were involved in multiple cellular pathways, including motility, virulence, biofilm formation, metabolism, and regulation.
Materials and Methods
Bacterial Growth Conditions
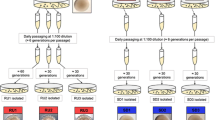
V. parahaemolyticus RIMD2210633, a pandemic O3: K6 strain isolated from a diarrhoea patient in 1996 [24, 25], was used in this study. A total of 20 μl glycerol stock of bacterial cells was inoculated into 5 ml of 2.5% (w/v) Bacto heart infusion (HI; BD Bioscience, USA) broth and incubated at 37 °C with shaking at 200 rpm for 12 h. The cell culture was diluted 50-fold into 2 ml of fresh Difco marine (M) broth 2216 (BD Biosciences, USA) in a 24-well cell culture plate and then grown at 30 °C with shaking at 150 rpm.
Crystal Violet (CV) Staining
CV staining assay was performed similarly as previously described [26]. Briefly, V. parahaemolyticus RIMD2210633 was culture in M broth at 30 °C with shaking at 150 rpm for 6, 12, 24, 48, 60 and 72 h, respectively. The media containing the planktonic cells were collected for absorbance measurements at OD600. The adherent biofilms were gently washed three times with 2 ml of deionized water, followed by incubated at 80 °C for 15 min to fix the attached biofilm cells. The attached cells were stained with 2 ml of 0.1% CV for 15 min, and then washed three times with 2 ml of deionized water. The bound CV was dissolved with 20% acetic acid, and the OD570 values were measured as an index of CV staining. Relative biofilm formation was calculated as OD570/OD600.
RNA Sequencing (RNA-seq)
V. parahaemolyticus RIMD2210633 was incubated at 30 °C with shaking at 150 rpm for 6, 12, 24 and 48 h (here termed as B6, B12, B24 and B48), respectively, with three biological replicates at each time point. Bacterial cells were harvested simultaneously from biofilms and planktonic fractions for the preparation of total RNA, which were extracted using TRIzol Reagent (Invitrogen, USA). One RNA sample was prepared from each biological replicate at each time point, with a total of 12 samples. RNA concentration was measured using a Nanodrop 2000, and RNA integrity was evaluated using agarose gel electrophoresis. RNA amount in each sample was required to be greater than 2 μg with OD260/OD280 value between 1.8 and 2.2. The rRNA removal and mRNA enrichment were performed using an Illumina/Ribo-Zero™ rRNA Removal Kit (bacteria) (Illumina, USA) according to the manufacturer’s instructions. All RNA-related manipulations were performed in GENEWIZ Biotechnology Co. Ltd. (Suzhou, China). cDNA sequencing was performed on an Illumina Hiseq platform [23].
Raw RNA-seq data were filtered using Cutadapt (v1.9.1) [27] to remove adapters, contamination and low quality reads, and alignment of filtered reads was performed using Bowtie2 (v2.2.6) [28] with V. parahaemolyticus RIMD2210633 as the reference genome. Gene expression in bacterial cells harvested at 12, 24 and 48 h (test groups) were compared with that in bacterial cells harvested at 6 h (reference group). HTSeq (v0.6.1) [29] and FPKM (Fragments Per Kilo bases per Million reads) [30] methods were applied to calculate gene expression. DESeq (v1.6.3) [31] of the Bioconductor package was used to analyze the difference in gene expression between the test and reference groups with selection criteria of qvalue (fdr, padj) ≤ 0.05 and absolute log2 fold change (|log2 FC|) ≥ 1. Gene Ontology (GO) functional annotation was performed to analyze DEGs involved in molecular functions, cellular components, and biological processes [32]. The Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis was applied to analyze DEGs involved in the main metabolic pathways [33]. The functions of proteins encoded by DEGs were predicted by the Cluster of Orthologous Groups of proteins (COG) database [34].
Quantification of c-di-GMP
Quantification of c-di-GMP was performed as previously described [35]. Briefly, the planktonic and biofilm cells of V. parahaemolyticus RIMD2210633 were harvested at different time points (6, 12, 24 and 48 h), resuspended in 2 ml ice-cold PBS, incubated at 100 °C for 5 min, and then sonicated for 15 min (power 100%, frequency 37 kHz) in an ice-water bath. The supernatant containing c-di-GMP was collected, and the pellet was resuspended in 2 ml ice-cold PBS and re-extracted another two times. Intracellular c-di-GMP levels were determined using a c-di-GMP Enzyme-linked Immunosorbent Assay (ELISA) Kit (Mskbio, China). Total proteins in the supernatant were determined using a Pierce BCA Protein Assay kit (ThermoFisher Scientific, USA) according to the manufacturer’s instructions. Concentration of c-di-GMP was expressed as pmol/g of protein. The experiment was performed three independent times with three biological replicates in each. The results were expressed as the means ± standard deviation (SD). A two-way ANOVA with Tukey’s post hoc corrections for multiple comparisons was used to calculate statistical significance, with a P value less than 0.05 considered significant.
Validation of RNA-seq Data Using Reverse Transcriptase Quantitative PCR (RT-qPCR)
RT-qPCR was performed as previously described [36]. Briefly, total RNA was extracted from the planktonic and biofilm cells of V. parahaemolyticus RIMD2210633, which were harvested from different time points (6, 12, 24 and 48 h), using TRIzol Reagent. cDNA was generated from 1 μg of total RNA using a FastKing First Strand cDNA Synthesis Kit (Tiangen Biotech, China) according to the manufacturer’s instructions. The relative mRNA levels of each target gene were determined using the classic 2−ΔΔCt method. The 16S rRNA gene was used as the internal control. Primers used in this work are listed in Supplementary Table 1. The experiment was performed three independent times with three biological replicates in each. The results were expressed as the means ± SD. A two-way ANOVA with Tukey’s post hoc corrections for multiple comparisons was used to calculate statistical significance, with a P value less than 0.05 considered significant.
Results and Discussion
Biofilm Formation by V. parahaemolyticus
Bacterial biofilm is generally formed through several steps [2]. Previously, the processes of biofilm formation by V. parahaemolyticus were demonstrated to be divided into three main stages: bacterial attachment and growth stage, biofilm maturation stage, and biofilm diffusion stage [37]. Here, the data showed that biofilm formation by V. parahaemolyticus, measured by CV staining, increased gradually from 6 to 48 h, but decreased from 48 to 72 h (Fig. 1). However, the biofilm formation speed from 6 to 24 h was much faster than that from 24 to 48 h, and the maximum amounts of biofilm occurred at 48 h (Fig. 1). These results are consistent with previous observations [37]. Therefore, although we did not use other methods such as confocal laser scanning microscopy to further observe the development of biofilm formation, we still concluded that the biofilm growth stage might occur between 6 and 36 h, the maturation stage was around 48 h, and the diffusion stage probably appeared after 48 h.
Crystal violet staining assays of time course of biofilm formation by V. parahaemolyticus. Experiment was performed three independent times with three biological replicates in each. The results are expressed as the mean ± standard deviation (SD). The asterisks indicate the statistical significance between biofilms at 12, 24, 48, 60 or 72 h and that at 6 h
Overview of Gene Expression During Biofilm Formation
A total of 12 Illumina libraries were sequenced with more than 14.1 million raw reads for each library, which were deposited in the Sequence Read Archive (accession number: PRJNA874225). The correlations between the transcriptomes of all samples were notably high, with the Pearson correlation coefficient values ranging from 0.919 to 1.000 (Supplementary Figure S1). A total of 183, 503 and 729 DEGs were identified in the samples from 12, 24 and 48 h, respectively, compared with that from 6 h (defined here as regulon 12, 24, and 48 h, respectively), including 82, 208 and 321 downregulated genes and 101, 295 and 408 upregulated genes (Fig. 2a–c). Comparisons between the three regulons showed that 133, 99, 334 and 92 genes with various predicted functions coexist in the regulons 12 and 24 h, 12 and 48 h, 24 and 48 h, and 12, 24 and 48 h, respectively (Fig. 3 and Table 1). The KEGG pathway enrichment analysis demonstrated that the regulon 12 h contained 10 organismal system genes, 118 metabolism genes, 12 human disease genes, 12 environmental information processing genes and 17 cellular process genes (Fig. 2d); the regulon 24 h contained 12 organismal system genes, 273 metabolism genes and 11 human disease genes (Fig. 2e); whereas the regulon 48 h contained 4 organismal system genes, 330 metabolism genes, 17 human disease genes and 54 environmental information processing genes (Fig. 2f). The results of COG enrichment showed that DEGs were divided into approximately 20 functional categories, including several metabolism-associated pathways such as amino acid transport and metabolism, inorganic ion transport and metabolism, lipid transport and metabolism, and coenzyme transport and metabolism (Supplementary Figure S2). The detailed descriptions of DEGs are listed in Supplementary Tables S2, S3 and S4.
Gene expression in different stages of biofilm formation. a–c Volcano plot shows genes that are upregulated (red), downregulated (blue) and non-changed (grey), respectively. d–f Pathways of differentially expressed genes analyzed by Kyoto Encyclopedia of Genes and Genomes (KEGG). The number on the right of each bar indicates the number of differentially expressed genes (Color figure online)
Validation of RNA-seq Data by RT-qPCR
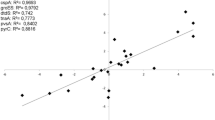
The RT-qPCR assays were employed to validate the results of RNA-seq. Seventeen genes were randomly selected as the target genes. As shown in Table 2, RT-qPCR revealed a consistent trend with the RNA-seq data, confirming the reliability of the transcriptome data.
DEGs Involved in Motility and Chemotaxis
V. parahaemolyticus expresses dual flagellar systems, a sheathed polar flagellum for swimming in liquids and unsheathed lateral flagella for swarming over surfaces [38]. Polar flagellar genes are organized on chromosome 1, whereas lateral flagellar genes are found on chromosome 2 [39, 40]. Polar flagellar genes are expressed continuously, whereas lateral flagellar genes are induced by certain conditions [40, 41]. In addition, flagella-propelled motility is also controlled by various regulators. AphA and OpaR are the master QS regulators operating at low cell density (LCD) and high cell density (HCD), respectively [42, 43]. AphA activates the transcription of polar and lateral flagellar genes at LCD, whereas OpaR represses their transcription at HCD [44,45,46]. ToxR activates both swimming and swarming motility of V. parahaemolyticus [12]. Swimming motility is activated by VapH [47] and OxyR [11], whereas swarming motility is induced by LafK [48], SwrT [49] and ScrABC [50]. Moreover, swarming motility is also repressed by SwrZ [49], H-NS [51] and CalR [52].
In this study, the RNA-seq data showed that expression of 16, 10 and 5 polar flagellar genes including flagellin genes were downregulated in the bacterial cells incubated for 12, 24 and 48 h, respectively, compared to those incubated for 6 h (Supplementary Tables S5, S6 and S7). Flagellar filaments, which consist of self-assembling flagellin, act as propellers to drive motility in liquids [38]. Six polar flagellin genes, flaA, flaB, flaC, flaD and flaE, were found in the polar flagellar gene loci [39]. flaA, flaB and flaD contribute more to the polar flagella filament [53]. However, deletion of any of the six flagellin genes has little or no effect on swimming motility of V. parahaemolyticus [53]. Three polar flagellin genes, flaB, flaD and flaE, were continuously downregulated from the initial attachment stage to biofilm maturation stage (Table 1). These results were consistent with the fact that flagella-mediated motility is required for the initial stages of biofilm formation [5].
In contrast, the RNA-seq data demonstrated similar expression levels of most lateral flagellar genes between the different stages of biofilm formation (data not shown). Only one gene (flgF) encoding the lateral flagellar basal body rod protein was downregulated in the regulon 24 h (Supplementary Table S6). It is generally understood that transcription of flagellar gene is inhibited during the motility-to-biofilm transition [54]. The method of collecting bacterial cells may affect the results of RNA-seq, but the mechanisms behind the observation that the lateral flagellar genes were stably transcribed during biofilm formation need to be further investigated.
A total of 3 (VP0421, VP0183 and VP2827), 5 (VP2159, VP2225, VP2229, VP2827 and VPA1449), and 8 (VP0963, VP1486, VP2225, VP2629, VP2827, VPA0024, VPA0941 and VPA0596) genes involved in chemotaxis were detected in the regulons 12, 24 and 48 h, respectively (Supplementary Tables S5, S6 and S7). All of the putative chemotaxis genes were significantly downregulated during biofilm formation except for VPA0491 and VPA0596. In addition, VP2827 encoding a potential methyl-accepting chemotaxis protein was continuously downregulated during biofilm formation (Table 1). However, the roles of these chemotaxis genes are still unknown in V. parahaemolyticus.
DEGs Involved in Biofilm Formation
The extracellular matrix is a basic ingredient of bacterial biofilms that plays multiple roles in bacterial biofilms [55]. The biofilm matrix predominantly consists of EPS, but also contains extracellular proteins, extracellular DNA and lipids [55]. However, a previous study showed that biofilm formation by V. parahaemolyticus was strongly positively correlated with the contents of extracellular DNA and proteins, but not with that of EPS [6]. The presented data also demonstrated similar expression levels of EPS biosynthesis-associated genes during biofilm formation (data not shown).
Four MSHA-associated genes, VP700, VP2701, VP2702 and VP2703, were downregulated in the maturation stage (48 h) compared with the initial attachment and biofilm development stages (Supplementary Table S7). However, the ChiRP genes were similarly expressed between the different stages of biofilm formation (data not shown). MSHA and ChiRP are positively correlated with V. parahaemolyticus biofilm formation [7]. Defective biofilms formed by MSHA mutants can be compensated for over time [8], which may aid to explain why the expression levels of MSHA genes were decreased at the maturation stage. MSHA also functions as an adherence factor of V. parahaemolyticus to Caco-2 cells, and thus may act as a potential virulence factor [56]. As one of the important factors affecting biofilm formation by V. parahaemolyticus, expression of type IV pili was regulated by several factors, including chitin and QS [57, 58], but its regulatory mechanisms are still far from being elucidated.
The VP0215-0237 locus is related to capsule polysaccharide (CPS) production [9]. V. parahaemolyticus undergoes opaque (OP) and translucent (TR) variation on agar plate based on whether CPS is produced [9, 59]. CPS acts as a negative effector of biofilm formation, but both OP and TR strains can form stable biofilms with different architectures [8, 60]. In this work, two CPS genes, rfbC and wcvB, were downregulated in the maturation stage (48 h) compared with the initial attachment and biofilm development stages (Supplementary Table S7), suggesting that CPS synthesis was repressed in the biofilm maturation stage. OpaR represses biofilm formation by V. parahaemolyticus but activates CPS gene expression [16, 61]. Thus, regulation of CPS gene expression may be one of the most important mechanisms adopted by V. parahaemolyticus to regulate biofilm formation.
The changes of c-di-GMP levels in bacteria influence multiple cellular pathways, including biofilm formation and virulence factor production [62]. c-di-GMP is catalyzed from guanosine triphosphate by diguanylate cyclase (DGC) containing a GGDEF domain, and is degraded by phosphodiesterase (PDE) carrying HD-GYP or EAL domain [62]. V. parahaemolyticus expresses more than 50 proteins that may be involved in c-di-GMP metabolism [24], and several of them, including the GGDEF-EAL-type proteins, ScrG [63] and ScrC [64], GGDEF-type proteins, ScrO [65], GefA [66], ScrJ [67] and ScrL [67], and EAL-type protein, LafV [67], have been confirmed to affect c-di-GMP metabolism, motility and biofilm formation. In this study, a total of 5 genes encoding the GGDEF and/or EAL-type proteins were significantly differentially expressed during biofilm formation (Supplementary Tables S6 and S7). Of these, VP2366 (GGDEF-type) and VPA1735 (GGDEF-EAL-type) were downregulated in regulon 24 h; VP1979 (EAL-type) and VPA0609 (GGDEF-EAL-type) were upregulated in regulon 48 h; VPA0360 (GGDEF-type) was simultaneously upregulated in regulons 24 and 48 h. Furthermore, there was a significant drop in intracellular c-di-GMP levels at the beginning of biofilm formation (Fig. 4), and further work is needed to understand the complexities of how c-di-GMP levels are governed during biofilm formation.
Intracellular c-di-GMP levels of V. parahaemolyticus. Bacterial cells were harvested after 6, 12, 24 and 48 h of incubation at 30 °C with shaking at 150 rpm in a 24-well cell culture plate. The results are expressed as the mean ± SD of three independent experiments. Asterisks represent statistically significant differences (P < 0.05)
The RNA-seq data also showed that mfpB and mfpC were significantly upregulated in regulon 24 h (Supplementary Table S6). mfpB and mfpC, transcribed from the mfpABC operon, are required for the mature biofilm formation of V. parahaemolyticus [8, 68]. Transcription of mfpABC was activated by OpaR [68], CalR [69], QsvR [70] and H-NS [71], but repressed by AphA [68]. Most importantly, OpaR, QsvR, H-NS and AphA are all involved in the regulation of biofilm formation [8, 46, 71, 72]. Therefore, regulation of mfpABC expression should be one of the mechanisms adopted by transcriptional regulators to regulate biofilm formation by V. parahaemolyticus.
DEGs Involved in Virulence
V. parahaemolyticus RIMD2210633 expressed a number of virulence determinants, including TDH, T3SS1, T3SS2, T6SS1, and T6SS2 [24]. TDH contributes to the hemolytic activity, mice lethal toxicity, enterotoxicity and cytotoxicity of V. parahaemolyticus, and can cause β-hemolysis on Wagatsuma agar [73]. T3SS1 is involved in the cytotoxicity, whereas T3SS2 is a major contributor to enterotoxicity and has cytotoxic effects on a limited number of cell lines [74, 75]. The T3SS2 gene cluster and the two copies of tdh genes are located on an 80 kb pathogenicity island on chromosome II, known as Vp-PAI (VPA1312-1398) [24]. T3SS2 delivers TDH from V. parahaemolyticus as an effector into host cells, resulting in the intestinal fluid accumulation in a rabbit diarrhoeal model [76]. T6SS1 is a contributor to the antibacterial activity of V. parahaemolyticus, whereas T6SS2 is an adherence factor of the bacterium [77, 78]. In addition, other factors such as lipopolysaccharide, proteases and outer membrane proteins (OMPs) are also involved in the pathogenesis of V. parahaemolyticus [4, 79].
The RNA-seq data showed that T3SS1 was consistently downregulated during biofilm formation (Supplementary Tables S5, S6 and S7). One, thirteen, and twenty-five T3SS1 genes were included in regulons 12, 24, and 48 h, respectively. Of these, VP1680 encoding the T3SS1 effector VopQ [80] was downregulated continuously (Table 1). In particular, the regulon 48 h contains exsC, exsD and exsE (Supplementary Table S7) encoding proteins that belong to the ExsACDE regulatory cascade. ExsA, an AraC-type transcriptional regulator, activates T3SS1 gene expression [81, 82]. ExsD binds ExsA to prevent ExsA-dependent activation of T3SS1, whereas ExsC binds ExsD to block the binding of ExsD to ExsA [83]. ExsE is a secreted substrate of T3SS1 and interacts with ExsC to prevent the binding of ExsC to ExsD, and thereby negatively regulating T3SS1 [84, 85]. Some regulators including H-NS, OpaR, CalR, QsvR, AphA, and HlyU control T3SS1 expression by directly regulating exsACDE [86,87,88,89]. Moreover, ExsA represses lateral flagellar gene expression and swarming motility [52], suggesting that the ExsACDE cascade may be also involved in the regulation of biofilm formation.
The tdh gene encoding TDH and the vscS2 gene encoding a putative type III secretion apparatus protein were significantly downregulated and upregulated, respectively, in the bacterial cells incubated for 24 h compared to those incubated for 6 h (Supplementary Table S6). In addition, seven genes located in Vp-PAI were significantly differentially expressed in the bacterial cells incubated for 48 h compared to those incubated for 6 h. Of these, VPA1342 encoding the EscR/YscR/HrcR family type III secretion system export apparatus protein was downregulated, and the other six genes (VPA1319, VPA1376, VPA1387, VPA1388, VPA1396 and VPA1379) were upregulated (Supplementary Table S7). These results suggested that the Vp-PAI genes were differentially expressed during biofilm formation, with downregulation of the virulence-related T3SS2 and TDH genes, and upregulation of the other genes.
Each regulon contains several of T6SS1 and T6SS2 genes, which were all downregulated (Supplementary Tables S5, S6 and S7). In particular, a total of 17 T6SS2 genes were downregulated in the regulon 48 h (Supplementary Table S7). T6SS plays multiply physiological roles including bacterial competition, ion transport, host infection, and stress response [90]. T6SS also participates in bacterial biofilm formation. For example, expression of T6SS were remarkably enhanced in biofilm cells of Pseudomonas fluorescens, and the lack of T6SS impeded biofilm formation [91, 92]. The T6SS gene mutants of Acidovorax citrulli produced less biofilms than the wild-type strain [93]. The phosphatase PppA, which is encoded in the T6SS gene cluster of V. alginolyticus, negatively regulates c-di-GMP production and biofilm formation [94]. V. cholerae T6SS is still active within biofilms that kill bacteria lacking T6SS within the interior of biofilm populations [95]. However, whether V. parahaemolyticus T6SSs play roles in biofilm formation remain unclear.
Extracellular proteases can digest host proteins into amino acids, aiding the spread of V. parahaemolyticus into host tissues when wound infection occurs [79]. As shown in Supplementary Table S5, vppC encoding a zinc metalloprotease and prtA encoding a serine protease were up- and down-regulated in the regulon 12 h, respectively. In addition, VPA1071 encoding another serine protease was downregulated in the regulon 24 h (Supplementary Table S6). VppC, which is secreted during the early stationary phase at 26 °C, digests native collagen [96, 97]. PrtA, which is secreted during the late-log phase, contributes to hemolytic activity and cytotoxicity of V. parahaemolyticus [98]. However, roles of VPA1071 are completely unknown.
OMPs can strongly elicit the host immune response and thus can be good targets for vaccine against bacterial infections [99]. However, some OMPs such as OmpF and YfiB also affect biofilm formation and virulence of bacteria [100, 101]. In this work, the data showed that one, six and twelve OMP genes were significantly differentially expressed in regulons 12, 24 and 48 h, respectively (Supplementary Tables S2, S3 and S4), suggesting that OMPs were remodeled during biofilm formation. Of these, VP1008, VP1634, VP1901, VP2212, VP2467, ompW, VPA0860 and VPA1644 were downregulated, while VP0802, VP1218, VP1631, aqpZ, VP2362 and VP2385 were upregulated. However, whether these genes play roles in biofilm formation needs to be further investigated.
DEGs Involved in Regulatory Functions
Five, nineteen and twenty-nine putative regulatory genes were significantly differentially expressed in regulons 12, 24 and 48 h, respectively (Supplementary Tables S5, S6 and S7). Many of them encode global regulators, including the LysR family transcriptional regulators (VPA0602, VPA0387, VPA0388, VPA0717 and VPA0961), MarR family transcriptional regulators (VPA1219 and OhrR), Fis family transcriptional regulator (VP1704), TetR/AcrR family transcriptional regulator (VP1229), PadR family transcriptional regulator (AphA) and two component regulator systems (PhoR/PhoB and BtsS/BtsR). AphA regulates virulence, biofilm formation and motility of V. parahaemolyticus [15, 44, 46, 87, 102]. OhrR, a sensor of organic peroxides, is involved in the resistance of exogenous cumene hydroperoxide stress [103]. PhoR/PhoB represses biofilm formation and c-di-GMP production but activates motility of V. cholerae [104]. BtsS/BtsR represses curli production and biofilm formation of E. coli [105]. VPA0961 is induced in the low-salt growth condition [106], but its roles are still unknown. In addition, scvE encoding a sigma-54 dependent transcriptional regulator was upregulated in regulon 48 h (Supplementary Table S7). ScvE promotes V. parahaemolyticus biofilm formation via activation of scvA-O transcription [10]. It is noteworthy that VPA1682 was continuously downregulated during biofilm formation (Table 1), suggesting that it may play important roles in biofilm formation. Moreover, many other putative regulators with unknown functions, such as VP0252, VPA0717 and VPA0804, were also significantly differentially expressed during biofilm formation.
A total of 1065 putative small non-coding RNAs (sRNAs) were detected during biofilm formation. Of these, 1049 were cis-encoded and 15 were trans-encoded (Supplementary Table S8). However, whether they truly exist remains to be experimentally confirmed. sRNAs regulate gene expression at the posttranscriptional levels [107]. In V. parahaemolyticus, Qrr1-5 sRNAs control OpaR expression, motility, CPS production and metabolism, but Qrr2 alone is also sufficient for OpaR regulation [22]. The sRNA Spot42 interacts with a sequence encompassing the ribosomal binding site and start codon of VP1682 mRNA to inhibit the expression of VP1682 [108]. V. parahaemolyticus RyhB binds to the 5′-UTR of pvsABCDE mRNA to enhance the expression of vibrioferrin [109]. The two sRNAs, svpa1401.1 and svpa1453.1, which are exclusively present in V. parahaemolyticus pandemic strains, are remarkably decreased at 3 h post-infection of Caco-2 cells, and involved in regulating the expression of VPA1379 [110]. A novel sRNA srvg23535 conserved among V. harveyi, V. parahaemolyticus and V. splendidus can affect the utilization of carbon/nitrogen sources and the response to osmotic and pH stress [111]. More studies should be performed in the future to gain more knowledge on sRNAs in V. parahaemolyticus.
Conclusions and Expectations
In this work, we showed that the growth stage of biofilm formation by V. parahaemolyticus occurred from 6 to 36 h, the maturation stage was around 48 h, and the diffusion stage appeared after 48 h of growth. A total of 183, 503 and 729 genes were significantly differentially expressed in the bacterial cells at 12, 24 and 48 h compared with those at 6 h, respectively. Of these, 92 genes were consistently activated or repressed from 6 to 48 h. The genes involved in polar flagellum, chemotaxis, MSHA, CPS, T3SS1, T3SS2, TDH, T6SS1, and T6SS2 were significantly downregulated during biofilm formation from 6 to 48 h, whereas those involved in Vp-PAI (except for the T3SS2 and TDH genes) and mfp genes were upregulated. Three extracellular protease genes (vppC, prtA and VPA1071) and some OMP genes were also significantly differentially expressed during biofilm formation. Five c-di-GMP metabolism-associated genes were also significantly differentially expressed that may account for the drop in c-di-GMP levels after the beginning of biofilm formation. In addition, many putative regulatory genes were significantly differentially expressed, and more than 1000 putative sRNAs were detected during biofilm formation, suggesting that biofilm formation is tightly regulated by complex regulatory networks. In brief, the RNA-seq data provided a global view of gene expression during biofilm formation, showing that the significantly differentially expressed genes were involved in multiple cellular pathways, including virulence, biofilm formation, metabolism and regulation.
Data Availability
The original data presented in the study are included in the article/supplementary materials, further inquiries can be directed to the corresponding authors. The raw data of RNA-seq are deposited in the NCBI repository, accession number PRJNA874225.
References
Toyofuku M, Inaba T, Kiyokawa T, Obana N, Yawata Y, Nomura N (2016) Environmental factors that shape biofilm formation. Biosci Biotechnol Biochem 80(1):7–12. https://doi.org/10.1080/09168451.2015.1058701
Ashrafudoulla M, Mizan MFR, Park SH, Ha SD (2021) Current and future perspectives for controlling Vibrio biofilms in the seafood industry: a comprehensive review. Crit Rev Food Sci Nutr 61(11):1827–1851. https://doi.org/10.1080/10408398.2020.1767031
Letchumanan V, Chan KG, Lee LH (2014) Vibrio parahaemolyticus: a review on the pathogenesis, prevalence, and advance molecular identification techniques. Front Microbiol 5:705. https://doi.org/10.3389/fmicb.2014.00705
Li L, Meng H, Gu D, Li Y, Jia M (2019) Molecular mechanisms of Vibrio parahaemolyticus pathogenesis. Microbiol Res 222:43–51. https://doi.org/10.1016/j.micres.2019.03.003
Yildiz FH, Visick KL (2009) Vibrio biofilms: so much the same yet so different. Trends Microbiol 17(3):109–118. https://doi.org/10.1016/j.tim.2008.12.004
Li W, Wang JJ, Qian H, Tan L, Zhang Z, Liu H, Pan Y, Zhao Y (2020) Insights into the role of extracellular DNA and extracellular proteins in biofilm formation of Vibrio parahaemolyticus. Front Microbiol 11:813. https://doi.org/10.3389/fmicb.2020.00813
Shime-Hattori A, Iida T, Arita M, Park KS, Kodama T, Honda T (2006) Two type IV pili of Vibrio parahaemolyticus play different roles in biofilm formation. FEMS Microbiol Lett 264(1):89–97. https://doi.org/10.1111/j.1574-6968.2006.00438.x
Enos-Berlage JL, Guvener ZT, Keenan CE, McCarter LL (2005) Genetic determinants of biofilm development of opaque and translucent Vibrio parahaemolyticus. Mol Microbiol 55(4):1160–1182. https://doi.org/10.1111/j.1365-2958.2004.04453.x
Chen Y, Dai J, Morris JG Jr, Johnson JA (2010) Genetic analysis of the capsule polysaccharide (K antigen) and exopolysaccharide genes in pandemic Vibrio parahaemolyticus O3:K6. BMC Microbiol 10:274. https://doi.org/10.1186/1471-2180-10-274
Liu M, Nie H, Luo X, Yang S, Chen H, Cai P (2022) A Polysaccharide biosynthesis locus in vibrio parahaemolyticus important for biofilm formation has homologs widely distributed in aquatic bacteria mainly from Gammaproteobacteria. eSystems 7(2):e0122621. https://doi.org/10.1128/msystems.01226-21
Chung CH, Fen SY, Yu SC, Wong HC (2016) Influence of oxyR on growth, biofilm formation, and mobility of Vibrio parahaemolyticus. Appl Environ Microbiol 82(3):788–796. https://doi.org/10.1128/AEM.02818-15
Chen L, Qiu Y, Tang H, Hu LF, Yang WH, Zhu XJ, Huang XX, Wang T, Zhang YQ (2018) ToxR is required for biofilm formation and motility of Vibrio Parahaemolyticus. Biomed Environ Sci 31(11):848–850. https://doi.org/10.3967/bes2018.112
Whitaker WB, Richards GP, Boyd EF (2014) Loss of sigma factor RpoN increases intestinal colonization of Vibrio parahaemolyticus in an adult mouse model. Infect Immun 82(2):544–556. https://doi.org/10.1128/IAI.01210-13
Ferreira RB, Chodur DM, Antunes LC, Trimble MJ, McCarter LL (2012) Output targets and transcriptional regulation by a cyclic dimeric GMP-responsive circuit in the Vibrio parahaemolyticus Scr network. J Bacteriol 194(5):914–924. https://doi.org/10.1128/JB.05807-11
Lu R, Osei-Adjei G, Huang X, Zhang Y (2018) Role and regulation of the orphan AphA protein of quorum sensing in pathogenic Vibrios. Future Microbiol 13:383–391. https://doi.org/10.2217/fmb-2017-0165
Zhang Y, Qiu Y, Gao H, Sun J, Li X, Zhang M, Xue X, Yang W, Ni B, Hu L, Yin Z, Lu R, Zhou D (2021) OpaR controls the metabolism of c-di-GMP in Vibrio parahaemolyticus. Front Microbiol 12:676436. https://doi.org/10.3389/fmicb.2021.676436
Zhong X, Lu R, Liu F, Ye J, Zhao J, Wang F, Yang M (2021) Identification of LuxR family regulators that integrate into quorum sensing circuit in Vibrio parahaemolyticus. Front Microbiol 12:691842. https://doi.org/10.3389/fmicb.2021.691842
Jiang F, Lei T, Wang Z, He M, Zhang J, Wang J, Zeng H, Chen M, Xue L, Ye Q, Pang R, Wu S, Gu Q, Ding Y, Wu Q (2021) A novel gene vp0610 negatively regulates biofilm formation in Vibrio parahaemolyticus. Front Microbiol 12:656380. https://doi.org/10.3389/fmicb.2021.656380
Billaud M, Seneca F, Tambutte E, Czerucka D (2022) An increase of seawater temperature upregulates the expression of Vibrio parahaemolyticus virulence factors implicated in adhesion and biofilm formation. Front Microbiol 13:840628. https://doi.org/10.3389/fmicb.2022.840628
Li X, Sun J, Zhang M, Xue X, Wu Q, Yang W, Yin Z, Zhou D, Lu R, Zhang Y (2021) The effect of salinity on biofilm formation and c-di-GMP production in Vibrio parahaemolyticus. Curr Microbiol 79(1):25. https://doi.org/10.1007/s00284-021-02723-2
Trimble MJ, McCarter LL (2011) Bis-(3’-5’)-cyclic dimeric GMP-linked quorum sensing controls swarming in Vibrio parahaemolyticus. Proc Natl Acad Sci USA 108(44):18079–18084. https://doi.org/10.1073/pnas.1113790108
Tague JG, Hong J, Kalburge SS, Boyd EF (2022) Regulatory small RNA Qrr2 is expressed independently of sigma factor-54 and can function as the sole Qrr small RNA to control quorum sensing in Vibrio parahaemolyticus. J Bacteriol 204(1):0035021. https://doi.org/10.1128/JB.00350-21
Wang Q, Wang P, Liu P, Ou J (2022) Comparative transcriptome analysis reveals regulatory factors involved in Vibrio parahaemolyticus biofilm formation. Front Cell Infect Microbiol 12:917131. https://doi.org/10.3389/fcimb.2022.917131
Makino K, Oshima K, Kurokawa K, Yokoyama K, Uda T, Tagomori K, Iijima Y, Najima M, Nakano M, Yamashita A, Kubota Y, Kimura S, Yasunaga T, Honda T, Shinagawa H, Hattori M, Iida T (2003) Genome sequence of Vibrio parahaemolyticus: a pathogenic mechanism distinct from that of V cholerae. Lancet 361(9359):743–749. https://doi.org/10.1016/S0140-6736(03)12659-1
Nasu H, Iida T, Sugahara T, Yamaichi Y, Park KS, Yokoyama K, Makino K, Shinagawa H, Honda T (2000) A filamentous phage associated with recent pandemic Vibrio parahaemolyticus O3:K6 strains. J Clin Microbiol 38(6):2156–2161. https://doi.org/10.1128/JCM.38.6.2156-2161.2000
Fang N, Gao H, Wang L, Qu S, Zhang YQ, Yang RF, Zhou DS (2013) Optimized methods for biofilm analysis in Yersinia pestis. Biomed Environ Sci 26(5):408–411. https://doi.org/10.3967/0895-3988.2013.05.012
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17(1):10–12. https://doi.org/10.14806/ej.17.1.200
Bray NL, Pimentel H, Melsted P, Pachter L (2016) Near-optimal probabilistic RNA-seq quantification. Nat Biotechnol 34(5):525–527. https://doi.org/10.1038/nbt.3519
Anders S, Pyl PT, Huber W (2015) HTSeq–a python framework to work with high-throughput sequencing data. Bioinformatics 31(2):166–169. https://doi.org/10.1093/bioinformatics/btu638
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5(7):621–628. https://doi.org/10.1038/nmeth.1226
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15(12):550. https://doi.org/10.1186/s13059-014-0550-8
Harris MA, Clark J, Ireland A, Lomax J, Ashburner M, Foulger R, Eilbeck K, Lewis S, Marshall B, Mungall C, Richter J, Rubin GM, Blake JA, Bult C, Dolan M, Drabkin H, Eppig JT, Hill DP, Ni L, Ringwald M, Balakrishnan R, Cherry JM, Christie KR, Costanzo MC, Dwight SS, Engel S, Fisk DG, Hirschman JE, Hong EL, Nash RS, Sethuraman A, Theesfeld CL, Botstein D, Dolinski K, Feierbach B, Berardini T, Mundodi S, Rhee SY, Apweiler R, Barrell D, Camon E, Dimmer E, Lee V, Chisholm R, Gaudet P, Kibbe W, Kishore R, Schwarz EM, Sternberg P, Gwinn M, Hannick L, Wortman J, Berriman M, Wood V, de la Cruz N, Tonellato P, Jaiswal P, Seigfried T, White R, Gene Ontology C (2004) The gene ontology (GO) database and informatics resource. Nucleic Acids Res. https://doi.org/10.1093/nar/gkh036
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28(1):27–30. https://doi.org/10.1093/nar/28.1.27
Tatusov RL, Galperin MY, Natale DA, Koonin EV (2000) The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 28(1):33–36. https://doi.org/10.1093/nar/28.1.33
Gao H, Ma L, Qin Q, Qiu Y, Zhang J, Li J, Lou J, Diao B, Zhao H, Shi Q, Zhang Y, Kan B (2020) Fur represses vibrio cholerae biofilm formation via direct regulation of vieSAB, cdgD, vpsU, and vpsA-K transcription. Front Microbiol 11:587159. https://doi.org/10.3389/fmicb.2020.587159
Gao H, Zhang Y, Yang L, Liu X, Guo Z, Tan Y, Han Y, Huang X, Zhou D, Yang R (2011) Regulatory effects of cAMP receptor protein (CRP) on porin genes and its own gene in Yersinia pestis. BMC Microbiol 11:40. https://doi.org/10.1186/1471-2180-11-40
Tan L, Zhao F, Han Q, Zhao A, Malakar PK, Liu H, Pan Y, Zhao Y (2018) High correlation between structure development and chemical variation during biofilm formation by Vibrio parahaemolyticus. Front Microbiol 9:1881. https://doi.org/10.3389/fmicb.2018.01881
McCarter LL (2004) Dual flagellar systems enable motility under different circumstances. J Mol Microbiol Biotechnol 7(1–2):18–29. https://doi.org/10.1159/000077866
Kim YK, McCarter LL (2000) Analysis of the polar flagellar gene system of Vibrio parahaemolyticus. J Bacteriol 182(13):3693–3704. https://doi.org/10.1128/JB.182.13.3693-3704.2000
Stewart BJ, McCarter LL (2003) Lateral flagellar gene system of Vibrio parahaemolyticus. J Bacteriol 185(15):4508–4518. https://doi.org/10.1128/JB.185.15.4508-4518.2003
Kawagishi I, Imagawa M, Imae Y, McCarter L, Homma M (1996) The sodium-driven polar flagellar motor of marine Vibrio as the mechanosensor that regulates lateral flagellar expression. Mol Microbiol 20(4):693–699. https://doi.org/10.1111/j.1365-2958.1996.tb02509.x
Sun F, Zhang Y, Wang L, Yan X, Tan Y, Guo Z, Qiu J, Yang R, Xia P, Zhou D (2012) Molecular characterization of direct target genes and cis-acting consensus recognized by quorum-sensing regulator AphA in Vibrio parahaemolyticus. PLoS ONE 7(9):e44210. https://doi.org/10.1371/journal.pone.0044210
Zhang Y, Qiu Y, Tan Y, Guo Z, Yang R, Zhou D (2012) Transcriptional regulation of opaR, qrr2–4 and aphA by the master quorum-sensing regulator OpaR in Vibrio parahaemolyticus. PLoS ONE 7(4):e34622. https://doi.org/10.1371/journal.pone.0034622
Lu R, Tang H, Qiu Y, Yang W, Yang H, Zhou D, Huang X, Hu L, Zhang Y (2019) Quorum sensing regulates the transcription of lateral flagellar genes in Vibrio parahaemolyticus. Future Microbiol 14:1043–1053. https://doi.org/10.2217/fmb-2019-0048
Lu R, Sun J, Qiu Y, Zhang M, Xue X, Li X, Yang W, Zhou D, Hu L, Zhang Y (2021) The quorum sensing regulator OpaR is a repressor of polar flagellum genes in Vibrio parahaemolyticus. J Microbiol 59(7):651–657. https://doi.org/10.1007/s12275-021-0629-3
Wang L, Ling Y, Jiang H, Qiu Y, Qiu J, Chen H, Yang R, Zhou D (2013) AphA is required for biofilm formation, motility, and virulence in pandemic Vibrio parahaemolyticus. Int J Food Microbiol 160(3):245–251. https://doi.org/10.1016/j.ijfoodmicro.2012.11.004
Park KS, Arita M, Iida T, Honda T (2005) vpaH, a gene encoding a novel histone-like nucleoid structure-like protein that was possibly horizontally acquired, regulates the biogenesis of lateral flagella in trh-positive Vibrio parahaemolyticus TH3996. Infect Immun 73(9):5754–5761. https://doi.org/10.1128/IAI.73.9.5754-5761.2005
Gode-Potratz CJ, Kustusch RJ, Breheny PJ, Weiss DS, McCarter LL (2011) Surface sensing in Vibrio parahaemolyticus triggers a programme of gene expression that promotes colonization and virulence. Mol Microbiol 79(1):240–263. https://doi.org/10.1111/j.1365-2958.2010.07445.x
Jaques S, McCarter LL (2006) Three new regulators of swarming in Vibrio parahaemolyticus. J Bacteriol 188(7):2625–2635. https://doi.org/10.1128/JB.188.7.2625-2635.2006
Boles BR, McCarter LL (2002) Vibrio parahaemolyticus scrABC, a novel operon affecting swarming and capsular polysaccharide regulation. J Bacteriol 184(21):5946–5954. https://doi.org/10.1128/JB.184.21.5946-5954.2002
Wang Y, Zhang Y, Yin Z, Wang J, Zhu Y, Peng H, Zhou D, Qi Z, Yang W (2018) H-NS represses transcription of the flagellin gene lafA of lateral flagella in Vibrio parahaemolyticus. Can J Microbiol 64(1):69–74. https://doi.org/10.1139/cjm-2017-0315
Gode-Potratz CJ, Chodur DM, McCarter LL (2010) Calcium and iron regulate swarming and type III secretion in Vibrio parahaemolyticus. J Bacteriol 192(22):6025–6038. https://doi.org/10.1128/JB.00654-10
McCarter LL (2001) Polar flagellar motility of the Vibrionaceae. Microbiol Mol Biol Rev 65(3):445–462. https://doi.org/10.1128/MMBR.65.3.445-462.2001
Guttenplan SB, Kearns DB (2013) Regulation of flagellar motility during biofilm formation. FEMS Microbiol Rev 37(6):849–871. https://doi.org/10.1111/1574-6976.12018
Flemming HC, Wingender J (2010) The biofilm matrix. Nat Rev Microbiol 8(9):623–633. https://doi.org/10.1038/nrmicro2415
O’Boyle N, Houeix B, Kilcoyne M, Joshi L, Boyd A (2013) The MSHA pilus of Vibrio parahaemolyticus has lectin functionality and enables TTSS-mediated pathogenicity. Int J Med Microbiol 303(8):563–573. https://doi.org/10.1016/j.ijmm.2013.07.010
Frischkorn KR, Stojanovski A, Paranjpye R (2013) Vibrio parahaemolyticus type IV pili mediate interactions with diatom-derived chitin and point to an unexplored mechanism of environmental persistence. Environ Microbiol 15(5):1416–1427. https://doi.org/10.1111/1462-2920.12093
Sun J, Li X, Qiu Y, Xue X, Zhang M, Yang W, Zhou D, Hu L, Lu R, Zhang Y (2022) Quorum sensing regulates transcription of the pilin gene mshA1 of MSHA pilus in Vibrio parahaemolyticus. Gene 807:145961. https://doi.org/10.1016/j.gene.2021.145961
McCarter LL (1998) OpaR, a homolog of Vibrio harveyi LuxR, controls opacity of Vibrio parahaemolyticus. J Bacteriol 180(12):3166–3173. https://doi.org/10.1128/JB.180.12.3166-3173.1998
Joseph LA, Wright AC (2004) Expression of Vibrio vulnificus capsular polysaccharide inhibits biofilm formation. J Bacteriol 186(3):889–893. https://doi.org/10.1128/JB.186.3.889-893.2004
Zhang M, Xue X, Li X, Wu Q, Zhang T, Yang W, Hu L, Zhou D, Lu R, Zhang Y (2023) QsvR and OpaR coordinately repress biofilm formation by Vibrio parahaemolyticus. Front Microbiol 14:1079653. https://doi.org/10.3389/fmicb.2023.1079653
Biswas S, Chouhan OP, Bandekar D (2020) Diguanylate cyclases in Vibrio cholerae: essential regulators of lifestyle switching. Front Cell Infect Microbiol 10:582947. https://doi.org/10.3389/fcimb.2020.582947
Kim YK, McCarter LL (2007) ScrG, a GGDEF-EAL protein, participates in regulating swarming and sticking in Vibrio parahaemolyticus. J Bacteriol 189(11):4094–4107. https://doi.org/10.1128/JB.01510-06
Ferreira RB, Antunes LC, Greenberg EP, McCarter LL (2008) Vibrio parahaemolyticus ScrC modulates cyclic dimeric GMP regulation of gene expression relevant to growth on surfaces. J Bacteriol 190(3):851–860. https://doi.org/10.1128/JB.01462-07
Kimbrough JH, Cribbs JT, McCarter LL (2020) Homologous c-di-GMP-binding scr transcription factors orchestrate biofilm development in Vibrio parahaemolyticus. J Bacteriol. https://doi.org/10.1128/JB.00723-19
Zhong X, Lu Z, Wang F, Yao N, Shi M, Yang M (2022) Characterization of GefA, a GGEEF domain-containing protein that modulates Vibrio parahaemolyticus motility, biofilm formation, and virulence. Appl Environ Microbiol 88(6):e0223921. https://doi.org/10.1128/aem.02239-21
Kimbrough JH, McCarter LL (2021) Identification of three new GGDEF and EAL domain-containing proteins participating in the scr surface colonization regulatory network in Vibrio parahaemolyticus. J Bacteriol. https://doi.org/10.1128/JB.00409-20
Zhou D, Yan X, Qu F, Wang L, Zhang Y, Hou J, Hu Y, Li J, Xin S, Qiu J, Yang R, Mao P (2013) Quorum sensing modulates transcription of cpsQ-mfpABC and mfpABC in Vibrio parahaemolyticus. Int J Food Microbiol 166(3):458–463. https://doi.org/10.1016/j.ijfoodmicro.2013.07.008
Gao H, Zhang L, Osei-Adjei G, Yang W, Zhou D, Huang X, Yang H, Yin Z, Zhang Y (2017) Transcriptional regulation of cpsQ-mfpABC and mfpABC by CalR in Vibrio parahaemolyticus. Microbiologyopen. https://doi.org/10.1002/mbo3.470
Zhang Y, Qiu Y, Xue X, Zhang M, Sun J, Li X, Hu L, Yin Z, Yang W, Lu R, Zhou D (2021) Transcriptional regulation of the virulence genes and the biofilm formation associated operons in Vibrio parahaemolyticus. Gut Pathog 13(1):15. https://doi.org/10.1186/s13099-021-00410-y
Zhang L, Weng Y, Wu Y, Wang X, Yin Z, Yang H, Yang W, Zhang Y (2018) H-NS is an activator of exopolysaccharide biosynthesis genes transcription in Vibrio parahaemolyticus. Microb Pathog 116:164–167. https://doi.org/10.1016/j.micpath.2018.01.025
Kernell Burke A, Guthrie LT, Modise T, Cormier G, Jensen RV, McCarter LL, Stevens AM (2015) OpaR controls a network of downstream transcription factors in Vibrio parahaemolyticus BB22OP. PLoS ONE 10(4):e0121863. https://doi.org/10.1371/journal.pone.0121863
Cai Q, Zhang Y (2018) Structure, function and regulation of the thermostable direct hemolysin (TDH) in pandemic Vibrio parahaemolyticus. Microb Pathog 123:242–245. https://doi.org/10.1016/j.micpath.2018.07.021
Park KS, Ono T, Rokuda M, Jang MH, Okada K, Iida T, Honda T (2004) Functional characterization of two type III secretion systems of Vibrio parahaemolyticus. Infect Immun 72(11):6659–6665. https://doi.org/10.1128/IAI.72.11.6659-6665.2004
Hiyoshi H, Kodama T, Iida T, Honda T (2010) Contribution of Vibrio parahaemolyticus virulence factors to cytotoxicity, enterotoxicity, and lethality in mice. Infect Immun 78(4):1772–1780. https://doi.org/10.1128/IAI.01051-09
Matsuda S, Okada R, Tandhavanant S, Hiyoshi H, Gotoh K, Iida T, Kodama T (2019) Export of a Vibrio parahaemolyticus toxin by the sec and type III secretion machineries in tandem. Nat Microbiol 4(5):781–788. https://doi.org/10.1038/s41564-019-0368-y
Yu Y, Yang H, Li J, Zhang P, Wu B, Zhu B, Zhang Y, Fang W (2012) Putative type VI secretion systems of Vibrio parahaemolyticus contribute to adhesion to cultured cell monolayers. Arch Microbiol 194(10):827–835. https://doi.org/10.1007/s00203-012-0816-z
Salomon D, Gonzalez H, Updegraff BL, Orth K (2013) Vibrio parahaemolyticus type VI secretion system 1 is activated in marine conditions to target bacteria, and is differentially regulated from system 2. PLoS ONE 8(4):e61086. https://doi.org/10.1371/journal.pone.0061086
Osei-Adjei G, Huang X, Zhang Y (2018) The extracellular proteases produced by Vibrio parahaemolyticus. World J Microbiol Biotechnol 34(5):68. https://doi.org/10.1007/s11274-018-2453-4
Shimohata T, Nakano M, Lian X, Shigeyama T, Iba H, Hamamoto A, Yoshida M, Harada N, Yamamoto H, Yamato M, Mawatari K, Tamaki T, Nakaya Y, Takahashi A (2011) Vibrio parahaemolyticus infection induces modulation of IL-8 secretion through dual pathway via VP1680 in Caco-2 cells. J Infect Dis 203(4):537–544. https://doi.org/10.1093/infdis/jiq070
Liu AC, Thomas NA (2015) Transcriptional profiling of Vibrio parahaemolyticus exsA reveals a complex activation network for type III secretion. Front Microbiol 6:1089. https://doi.org/10.3389/fmicb.2015.01089
Zhou X, Shah DH, Konkel ME, Call DR (2008) Type III secretion system 1 genes in Vibrio parahaemolyticus are positively regulated by ExsA and negatively regulated by ExsD. Mol Microbiol 69(3):747–764. https://doi.org/10.1111/j.1365-2958.2008.06326.x
Zhou X, Konkel ME, Call DR (2010) Regulation of type III secretion system 1 gene expression in Vibrio parahaemolyticus is dependent on interactions between ExsA, ExsC, and ExsD. Virulence 1(4):260–272. https://doi.org/10.4161/viru.1.4.12318
Kodama T, Yamazaki C, Park KS, Akeda Y, Iida T, Honda T (2010) Transcription of Vibrio parahaemolyticus T3SS1 genes is regulated by a dual regulation system consisting of the ExsACDE regulatory cascade and H-NS. FEMS Microbiol Lett 311(1):10–17. https://doi.org/10.1111/j.1574-6968.2010.02066.x
Erwin DP, Nydam SD, Call DR (2012) Vibrio parahaemolyticus ExsE is requisite for initial adhesion and subsequent type III secretion system 1-dependent autophagy in HeLa cells. Microbiology 158(Pt 9):2303–2314. https://doi.org/10.1099/mic.0.059931-0
Sun F, Zhang Y, Qiu Y, Yang H, Yang W, Yin Z, Wang J, Yang R, Xia P, Zhou D (2014) H-NS is a repressor of major virulence gene loci in Vibrio parahaemolyticus. Front Microbiol 5:675. https://doi.org/10.3389/fmicb.2014.00675
Zhang Y, Hu L, Qiu Y, Osei-Adjei G, Tang H, Zhang Y, Zhang R, Sheng X, Xu S, Yang W, Yang H, Yin Z, Yang R, Huang X, Zhou D (2019) QsvR integrates into quorum sensing circuit to control Vibrio parahaemolyticus virulence. Environ Microbiol 21(3):1054–1067. https://doi.org/10.1111/j.1365-2958.2010.07445.x
Osei-Adjei G, Gao H, Zhang Y, Zhang L, Yang W, Yang H, Yin Z, Huang X, Zhou D (2017) Regulatory actions of Tox and Cal on their own genes and type III secretion system 1 in Vibrio parahaemolyticus. Oncotarget 8(39):65809–65822. https://doi.org/10.18632/oncotarget.19498
Getz LJ, Thomas NA (2018) The transcriptional regulator HlyU positively regulates expression of exsA, leading to type III secretion system 1 activation in Vibrio parahaemolyticus. J Bacteriol. https://doi.org/10.1128/JB.00653-17
Yang X, Pan J, Wang Y, Shen X (2018) Type VI secretion systems present new insights on pathogenic yersinia. Front Cell Infect Microbiol 8:260. https://doi.org/10.3389/fcimb.2018.00260
Gallique M, Decoin V, Barbey C, Rosay T, Feuilloley MG, Orange N, Merieau A (2017) Contribution of the pseudomonas fluorescens MFE01 type VI secretion system to biofilm formation. PLoS ONE 12(1):0170770. https://doi.org/10.1371/journal.pone.0170770
Chen L, Zou Y, Kronfl AA, Wu Y (2020) Type VI secretion system of Pseudomonas aeruginosa is associated with biofilm formation but not environmental adaptation. Microbiologyopen 9(3):e991. https://doi.org/10.1002/mbo3.991
Tian Y, Zhao Y, Wu X, Liu F, Hu B, Walcott RR (2015) The type VI protein secretion system contributes to biofilm formation and seed-to-seedling transmission of Acidovorax citrulli on melon. Mol Plant Pathol 16(1):38–47. https://doi.org/10.1111/mpp.12159
Sheng L, Lv Y, Liu Q, Wang Q, Zhang Y (2013) Connecting type VI secretion, quorum sensing, and c-di-GMP production in fish pathogen Vibrio alginolyticus through phosphatase PppA. Vet Microbiol 162(2–4):652–662. https://doi.org/10.1016/j.vetmic.2012.09.009
Teschler JK, Jimenez-Siebert E, Jeckel H, Singh PK, Park JH, Pukatzki S, Nadell CD, Drescher K, Yildiz FH (2022) VxrB influences antagonism within biofilms by controlling competition through extracellular matrix production and type 6 secretion. MBio. https://doi.org/10.1128/mbio.01885-22
Kim SK, Yang JY, Cha J (2002) Cloning and sequence analysis of a novel metalloprotease gene from Vibrio parahaemolyticus 04. Gene 283(1–2):277–286. https://doi.org/10.1016/s0378-1119(01)00882-4
Miyoshi S, Nitanda Y, Fujii K, Kawahara K, Li T, Maehara Y, Ramamurthy T, Takeda Y, Shinoda S (2008) Differential gene expression and extracellular secretion of the collagenolytic enzymes by the pathogen Vibrio parahaemolyticus. FEMS Microbiol Lett 283(2):176–181. https://doi.org/10.1111/j.1574-6968.2008.01159.x
Chang SC, Lee CY (2018) OpaR and RpoS are positive regulators of a virulence factor PrtA in Vibrio parahaemolyticus. Microbiology 164(2):221–231. https://doi.org/10.1099/mic.0.000591
Wang W, Liu J, Guo S, Liu L, Yuan Q, Guo L, Pan S (2020) Identification of Vibrio parahaemolyticus and vibrio spp specific outer membrane proteins by reverse vaccinology and surface proteome. Front Microbiol 11:625315. https://doi.org/10.3389/fmicb.2020.625315
Gao J, Han Z, Li P, Zhang H, Du X, Wang S (2021) Outer membrane protein F is involved in biofilm formation, virulence and antibiotic resistance in Cronobacter sakazakii. Microorganisms. https://doi.org/10.3390/microorganisms9112338
Sen T, Verma NK (2022) YfiB: an outer membrane protein involved in the virulence of Shigella flexneri. Microorganisms. https://doi.org/10.3390/microorganisms10030653
Zhang Y, Gao H, Osei-Adjei G, Yang W, Yang H, Yin Z, Huang X, Zhou D (2017) Transcriptional regulation of the Type VI secretion system 1 genes by quorum Sensing and ToxR in Vibrio parahaemolyticus. Front Microbiol 8:2005. https://doi.org/10.3389/fmicb.2017.02005
Chen NX, Chu YJ, Ni B, Hsu P, Wong HC (2021) Organic hydroperoxide resistance gene ohr (VPA1681) confers protection against organic peroxides in the presence of alkyl hydroperoxide reductase genes in Vibrio parahaemolyticus. Appl Environ Microbiol 87(21):0086121. https://doi.org/10.1128/AEM.00861-21
Pratt JT, McDonough E, Camilli A (2009) PhoB regulates motility, biofilms, and cyclic di-GMP in Vibrio cholerae. J Bacteriol 191(21):6632–6642. https://doi.org/10.1128/JB.00708-09
Ogasawara H, Ishizuka T, Yamaji K, Kato Y, Shimada T, Ishihama A (2019) Regulatory role of pyruvate-sensing BtsSR in biofilm formation by Escherichia coli K-12. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fnz251
Yang L, Zhan L, Han H, Gao H, Guo Z, Qin C, Yang R, Liu X, Zhou D (2010) The low-salt stimulon in Vibrio parahaemolyticus. Int J Food Microbiol 137(1):49–54. https://doi.org/10.1016/j.ijfoodmicro.2009.11.006
Perez-Reytor D, Plaza N, Espejo RT, Navarrete P, Bastias R, Garcia K (2016) Role of non-coding regulatory RNA in the virulence of human pathogenic vibrios. Front Microbiol 7:2160. https://doi.org/10.3389/fmicb.2016.02160
Tanabe T, Miyamoto K, Tsujibo H, Yamamoto S, Funahashi T (2015) The small RNA spot 42 regulates the expression of the type III secretion system 1 (T3SS1) chaperone protein VP1682 in Vibrio parahaemolyticus. FEMS Microbiol Lett. https://doi.org/10.1093/femsle/fnv173
Tanabe T, Funahashi T, Nakao H, Maki J, Yamamoto S (2013) The Vibrio parahaemolyticus small RNA RyhB promotes production of the siderophore vibrioferrin by stabilizing the polycistronic mRNA. J Bacteriol 195(16):3692–3703. https://doi.org/10.1128/JB.00162-13
Plaza N, Perez-Reytor D, Ramirez-Araya S, Pavon A, Corsini G, Loyola DE, Jana V, Pavez L, Navarrete P, Bastias R, Castillo D, Garcia K (2019) Conservation of small regulatory RNAs in Vibrio parahaemolyticus: possible role of RNA-OUT encoded by the pathogenicity island (VPaI-7) of pandemic strains. Int J Mol Sci. https://doi.org/10.3390/ijms20112827
Deng Y, Su Y, Liu S, Guo Z, Cheng C, Ma H, Wu J, Feng J, Chen C (2018) Identification of a novel small RNA srvg23535 in Vibrio alginolyticus ZJ-T and its characterization with phenotype microarray technology. Front Microbiol 9:2394. https://doi.org/10.3389/fmicb.2018.02394
Funding
This study was supported by the Natural Science Research Project of Nantong Science and Technology Bureau (JC2021027), and the National Natural Science Foundation of China (82072239).
Author information
Authors and Affiliations
Contributions
YZ, BG and RL designed, organized and supervised the experiments, interpreted the results, and edited the manuscript. TZ, YQ, MZ, XL, WY, LH and DZ performed or supported the laboratory experiments. YZ and TZ drafted the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.


Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Zhang, T., Qiu, Y. et al. Transcriptomic Profiles of Vibrio parahaemolyticus During Biofilm Formation. Curr Microbiol 80, 371 (2023). https://doi.org/10.1007/s00284-023-03425-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-023-03425-7