Abstract
The caseinolytic protease (Clp) system is essential for survival under stress conditions and for virulence in several pathogenic bacteria. Xanthomonas campestris pv. campestris (Xcc) is a plant pathogen which causes black rot disease in crucifers. In this study, the Xcc clpP gene which is annotated to encode the proteolytic core of Clp was characterized. Mutation of clpP resulted in susceptibility to high temperature and puromycin stresses. Site-directed mutagenesis revealed that S105, H130, and D179 are critical amino acid residues for ClpP function in puromycin tolerance. Inactivation of clpP also revealed an attenuation of virulence on the host plant and a reduction in the production of extracellular cellulase, mannanase, pectinase, and protease. The affected phenotypes of the clpP mutant could be complemented to wild-type levels by the intact clpP gene. Transcriptional analysis revealed that expression of clpP is induced under heat shock condition.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Pathogenic bacteria often encounter different stresses exerted by the external environment. These stresses include antimicrobial chemicals, changes in osmolarity, pH, and temperature [1]. To conquer harmful situations, bacterial cells are equipped with various mechanisms to enable them to adapt to heterogeneous environments. Caseinolytic protease (Clp) system plays an essential role in protein quality control and stress management [2]. In addition to its involvement in protein homeostasis and stress tolerance, the Clp protein has a wide range of functions, such as expression of pathogenicity factor and regulation of developmental process in bacteria [2, 3].
The members of the Clp family composed of two functional units: proteolytic core (ClpP and ClpQ) and ATPase-active chaperone rings (ClpA, ClpC, ClpE, ClpX, and ClpY) [4]. The ClpP protease can interact with different chaperons, namely ClpA, ClpC, ClpE, and ClpX; while ClpQ associates with ClpY to form active proteolytic machinery [4]. The ClpP is a conserved protein that is present in nearly all sequence eubacterial genomes [2]. ClpP is a serine protease characterized by a unique arrangement of the active site triad, Ser-His-Asp and is found to be involved in the proteolysis of damaged and misfolded proteins, regulatory proteins, and ribosome-stalled proteins in several species [5]. In addition, ClpP is important for the degradation of proteins involved in biofilm formation, cell motility, heat stress response, metabolism, nutrient starvation, and stationary phase adaptation [5]. In a number of pathogenic bacteria, ClpP function plays a vital role in infectivity and virulence [5].
Xanthomonas campestris pv. campestris (Xcc), a plant pathogen, is the causal agent of black rot on leaves of economically important crops such as cabbage, cauliflower, and radish [6]. Xcc is known to produce a variety of virulence factors such as exopolysaccharide and extracellular enzymes (cellulase, mannanase, pectinase, and protease) [7, 8]. In the sequenced Xcc genome, several putative clp genes (such as clpA, clpP, clpQ, clpX, and clpY) were annotated [9,10,11]. Despite Clp proteins being studied extensively in other bacteria, the specific functions of Clp proteins in Xcc are nearly unknown. Till now, only one report regarding Xcc Clp proteins was found in the available literature. It was indicated that the Xcc clpX gene is important for bacterial attachment, stress tolerance, and virulence [12]. Therefore, in the present study, we explored the function and transcription of clpP to gain more insight into the role and regulation of Clp proteins in Xcc.
Materials and Methods
Bacterial Strains, Plasmids, and Growth Conditions
Bacterial strains and plasmids used in the present study are listed in Table 1. E. coli strains were cultured in Luria–Bertani (LB) medium [13] at 37 °C. Xcc strains were grown in LB medium or XOLN medium [14] at 28 °C. For liquid cultures, bacteria were shaken at 180 rpm. Agar (1.5%) was added for solid media. The concentrations of the antibiotics used were as follows: ampicillin (50 μg/mL), gentamycin (15 μg/mL), kanamycin (50 μg/mL), and tetracycline (15 μg/mL).
DNA Techniques
Molecular biology protocols were as described by Sambrook et al. [13]. Polymerase chain reaction (PCR) was done as previously described [15] using the primers listed in Table 2. DNA sequencing was performed by Mission Biotech Co., Ltd. (Taipei, Taiwan). Transformation of E. coli and Xcc was performed by the standard method [13] and electroporation [16], respectively.
Mutant Construction and Complementation
The clpP mutant was generated by insertional inactivation of Xc17 and designated as CE17. First, the 717-bp PstI-XbaI fragment containing the Xc17 clpP gene was amplified by PCR using primer pair 1622PstI/2338XbaI and ligated into the yT&A cloning vector (Yeastern), giving pTclpP. After sequencing verification, the 717-bp PstI/XbaI fragment from pTclpP was cloned into pOK12 [17], generating pOKclpP. Then, the gentamycin resistance gene (GmR cartridge) from pUCGM [18] was inserted into the EcoRV site within the pOKclpP insert. The resultant plasmid, pOKclpPG, was electroporated into Xc17 allowing for double crossover. Insertion of GmR cartridge into clpP gene was confirmed by PCR.
For complementation, the 717-bp PstI-XbaI fragment from pTclpP was cloned into pRK415 [19]. The generating plasmid, pRKclpP, was then introduced into the clpP mutant CE17 by electroporation, resulting in the complemented strain CE17(pRKclpP). In parallel, the empty vector pRK415 was transferred into Xc17 and CE17, giving Xc17(pRK415) and CE17(pRK415) for comparison.
Site-Directed Mutagenesis
The site-directed mutagenesis of ClpP was conducted using the QuikChange Lightning Site-Directed Mutagenesis Kit (Agilent Technologies), according to the manufacturer's instructions. The mutation was constructed in the highly conserved active site residues of ClpP (S105, H130, or D179), using pTclpP as a template and the primers listed in Table 2, generating pTclpPS105A, pTclpPH130A, and pTclpPD179A. After DNA sequence confirmation, the mutated clpP was cloned into pRK415 to give pRKclpPS105A, pRKclpPH130A, and pRKclpPD179A. These constructs were separately electroporated into CE17, generating CE17(pRKclpPS105A), CE17(pRKclpPH130A), and CE17(pRKclpPD179A).
Stress Tolerance Assay
The methods for temperature tolerance assay and evaluating the sensitivity of bacteria to hydrogen peroxide (H2O2), sodium dodecyl sulfate (SDS), and puromycin are according to previously study [12]. The effect of temperature on bacterial growth was also determined quantitatively in liquid culture. Strains to be assayed were grown in LB medium and incubated at 28 °C or 37 °C with shaking (180 rpm). Samples were taken at designated intervals, and the growth of each tested strain was evaluated by measuring the OD550 values. These experiments were performed at least three times.
Virulence Assay and Extracellular Enzyme Activity Analysis
The virulence of Xcc to host plant cabbage was assayed by leaf-clipping method [20] and the lesion lengths were measured 14 days post-inoculation. Six replicates were performed for each strain and three independent experiments were carried out. The activities of extracellular enzymes (cellulase, mannanase, pectinase, and protease) were assessed by substrate-supplemented plate assay method as described previously [20]. The substrates supplemented were carboxymethyl cellulose (CMC, for cellulase), locust bean gum (LBG, for mannanase), sodium polypectate (for pectinase), and skim milk (for protease). The activity of cellulase and mannanase was examined by staining CMC- and LBG- supplemented plates with Congo red, and pectinase activity was examined by staining sodium polypectate containing plates with ruthenium red. Substrate degradation by extracellular enzyme depleted the plate of stain-binding material, forming clearing zones. Protease activity was examined directly by the appearance of clear zones surrounding the colonies on plates containing skim milk. Each experiment was done at least three times.
Promoter Activity Determination
The upstream region of clpP gene was PCR-amplified and cloned into pFY13–9, which is a promoter-probing vector using lacZ as the reporter [21]. The resulting constructs pFYclpP1, pFYclpP2, and pFYclpP3 carried nt − 384/− 10, − 254/− 10, and − 196/− 10 regions relative to the clpP translational start site, respectively. These constructs were separately introduced into Xc17 by electroporation. Strains carrying these reporter constructs were cultured overnight and then inoculated into fresh media to adjust an initial OD550 value of 0.35. Sampling interval was designated and promoter activity was monitored by measuring the β-galactosidase activity of the reporter constructs according to previously described method [22]
Statistical Analysis
Data are presented as the mean of triplicate per measurement, and each experiment was repeated at least three times. Student’s t-test was used to determine the statistical significance of differences between means.
Results and Discussion
clpP is Required for Heat and Puromycin Tolerance
In the genome of the Xcc strain Xc17, the locus_tag AAW18_RS04850 was annotated to encode ClpP (GenBank accession no. NZ_CP011946) [9]. The Xcc clpP gene encodes a 22-kDa product and displays similarity to the family of ClpP proteins found in several bacteria, such as E. coli (75.6% identity) and Bacillus subtilis (71.4%). Sequence analysis also revealed that the Xcc ClpP protein contains the conserved amino acid residues Ser-105, His-130, and Asp-179 that have been shown to constitute the catalytic triad in E. coli [5].
To determine the functions of ClpP in Xcc, the clpP mutant designated as CE17 was first constructed by insertional inactivation using wild-type Xc17 as the parental strain. Complementation of the mutant CE17 was performed by transforming plasmid pRKclpP and designated as CE17(pRKclpP). In parallel, parental strain Xc17 and the clpP mutant CE17 were transformed with pRK415 [19] to control for any effects of the vector itself and named as Xc17(pRK415) and CE17(pRK415). Further, the clpP mutant CE17 was transformed with mutated versions pRKclpPS105A, pRKclpPH130A, and pRKclpPD179A, in which the three above-mentioned amino acids (S105, H130, and D179) were substituted by alanine, to evaluate the role of these potential catalytic residues.
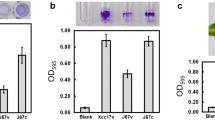
It has indicated that bacteria Clp protease has a multitude of functions, such as stress tolerance and virulence factor expression [3]. To validate the involvement of clpP in stress adaptation of Xcc, the wild-type Xc17(pRK415), clpP mutant CE17(pRK415), and complemented strain CE17(pRKclpP) were subjected to heat (37 °C), oxidative stress (1% H2O2), detergent (5% SDS), and antibiotic (5 mg/mL puromycin) treatments. Under standard culture conditions, i.e., 28 °C on LB plate, the clpP mutant grew as well as the wild-type and the complemented strains (Fig. 1a, upper). However, at 37 °C on LB plate, the growth of clpP mutant was completely arrested (Fig. 1a, lower). This growth deficiency was restored by genetic complementation (Fig. 1a, lower). Comparison of the growth curves of wild-type and mutant strains revealed no significant difference in LB medium at 28 °C (Fig. 1b, upper), whereas the growth of clpP mutant at 37 °C was much slower than that of both wild-type and complemented strains (Fig. 1b, lower). These results demonstrated that ClpP is not required for optimal growth at physiological temperature, whereas it has an important role in growth of Xcc at high temperature. The clpP mutant did not differ from the parent strain in sensitivity to H2O2 and SDS (Fig. 2a, b). However, the clpP mutant CE17(pRK415) was more sensitive to puromycin than its parent Xc17(pRK415) (Fig. 2c). The sensitivity of the clpP mutant to puromycin was complemented in CE17(pRKclpP), whereas the ability to restore bacteria growth was eliminated when CE17 was complemented with mutated versions (pRKclpPS105A, pRKclpPH130A, and pRKclpPD179A) (Fig. 2c). Taken together, these results imply that ClpP is required for heat and puromycin tolerance and the potential catalytic amino acid residues (S105, H130, and D179) are associated with the full function of ClpP in puromycin tolerance.
Effects of mutation of clpP on cell growth under heat stress. a Bacterial cells were spotted on LB plates and incubated at 28 °C or 37 °C for 3 days. b Cells were grown in LB medium at 28 °C or 37 °C and the cell density was measured at OD550 following cell growth. Results are the means of at least three independent experiments. Error bars indicate standard deviations
Sensitivity of clpP mutant to H2O2, SDS, and puromycin stresses. Bacterial cells were spread on LB plates and disks containing 10 μl of 1% H2O2 (a), 5% SDS (b), and 5 mg/mL puromycin (c) were placed on top. After 3 days of incubation, the diameters of inhibition zones were measured. Values under each photographs represent the mean diameter of inhibition zone (in cm) (mean ± standard deviation) from three independent experiments. Different letters following the values indicate significant difference (t-test, P < 0.01)
The ClpP is involved in the degradation of abnormal or damaged proteins that arise in response to stress treatment [23, 24]. Growth at elevated temperature or in the presence of puromycin, an aminoacyl-tRNA analogue which prematurely aborts protein translation, leads to the accumulation of misfolded proteins [25]. Here, we observed that the clpP mutant of Xcc exhibited a reduced growth rate at high temperature and an impaired resistance to puromycin treatment compared to the wild type. The failure of clpP mutant to grow at elevated temperature or in the presence of puromycin suggesting ClpP is required for Xcc to deal with conditions that are known to cause proteins to misfold. Several previous studies have demonstrated growth deficits resulting from ClpP deficiency under heat treatment, including Actinobacillus pleuropneumoniae [26], Lactococcus lactis [24], Staphylococcus aureus [25], Streptococcus pneumonia [23], and Streptococcus suis [27]. The clpP mutants of L. lactis, S. aureus and S. pneumonia were more sensitive to puromycin than wild-type cells [23,24,25].
clpP is Essential for Full Virulence of Xcc and is Involved in Extracellular Enzyme Production
To investigate whether clpP is involved in the pathogenicity of Xcc, the constructed strains were inoculated onto host plant cabbage by leaf-clipping methods. Fourteen days after inoculation, the clpP mutant CE17(pRK415) produced disease symptom with a mean lesion length of 0.93 cm on the infected leaves, whereas typical V-shaped disease symptoms with mean lesion lengths of 1.70 and 1.75 cm were observed on the leaves inoculated with the wild-type strain Xc17(pRK415) and the complemented strain CE17(pRK415), respectively (Fig. 3a). Xcc produces exopolysaccharide and a range of extracellular enzymes which collectively essential for pathogenesis. To evaluate whether clpP influences the synthesis of these virulence factors, the exopolysaccharide yields and the activities of four extracellular enzymes produced by the clpP mutant were determined. The results revealed that the amounts of exopolysaccharide produced by the mutant were similar to those produced by the wild type (data not shown). However, the clpP mutant produced a lower level of extracellular enzymes, including cellulase, mannanase, pectinase, and protease (Fig. 3b). The diameters of hydrolytic zone produced by the clpP mutant were significantly lower than those observed for the wild-type and complemented strains (Fig. 3b). Taken together, these results demonstrated that clpP is essential for full virulence of Xcc and is required for the ability to synthesize extracellular enzymes.
Effects of mutation of clpP on Xcc virulence and extracellular enzyme production. a Black rot symptoms caused by Xcc strains on cabbage leaves inoculated by leaf-clipping method. Images were taken at day 14 post-inoculation. Scale bars = 1 cm. Values under each leaf are the mean lesion length (in cm) (mean ± standard deviation) from three repeats, each with six leaves. Different letters following the values indicate significant difference (t-test, P < 0.01). b The activity of extracellular cellulase, mannanase, pectinase, and protease (from top to down) was evaluated using the substrate-supplemented plate assay. Values under each photographs are the mean diameter of hydrolysis zone (in cm) (mean ± standard deviation) from three independent experiments. Different letters following the values indicate significant difference (t-test, P < 0.01)
The ClpP plays an important role in virulence of several pathogens, such as Dickeya dadantii [28], Listeria monocytogene [29], S. aureus [25], S. pneumonia [23], and S. suis [27]. In D. dadantii, inactivation of clpP reduced the production of pectinolytic enzyme, which has a major role in the pathogenicity of this organism [28]. In L. monocytogene, mutation of clpP reduced the haemolytic activity of the major listeria virulence factor, listeriolysin O [29]. The amount of extracellular enzymes and toxins in S. aureus is reduced in the absence of ClpP [25]. The expression of apuA gene which has been linked to virulence in S. suis is reduced in clpP mutant [27]. Our observations showing decreased virulence of clpP mutant and inactivation of clpP affected the ability of the mutant to synthesize a range of extracellular enzymes suggested that reduced virulence factor synthesis might be the main reason for the attenuated virulence. The ClpP might be involved in the expression of these virulence genes and/or the turnover of them, which remain to be characterized. Xcc genome annotation revealed that several virulence factors are related to bacterial pathogenesis. Whether decreased virulence of Xcc clpP mutant is linked to the expression of other unidentified pathogenicity determinant(s) merits further investigation.
In E. coli, over 60 proteins involved in numerous biological functions were identified as potential substrates of a proteolytically inactive variant of ClpP [30]. A similar repertoire of substrate candidates has been found for B. subtilis ClpP [31]. Identification of potential targets of ClpP in these model bacteria indicates that ClpP has a wide range of impact on the bacterial proteome. In addition to E. coli and B. subtilis, the ClpP is found to be important for the degradation of various proteins, such as damaged and misfolded proteins, regulatory proteins, as well as proteins involved in heat stress response, metabolism, and virulence in several bacteria [5]. Our observations showing ClpP has multiple roles, including heat and puromycin tolerance, extracellular enzyme production, as well as virulence, suggested the involvement of ClpP in several cellular processes in Xcc. Though the potential targets of ClpP are unknown, it is reasonable to suggest that ClpP might be involved in the regulation of the Xcc proteome. Whether ClpP has an impact on Xcc proteome requires further evaluation.
clpP Expression is Induced by Heat Shock
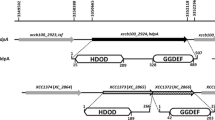
To investigate the expression of clpP in Xcc, the upstream regions of clpP gene were cloned ahead of a promoterless lacZ gene in pFY13–9 [21]. Three reporter constructs (pFYclpP1, pFYclpP2, and pFYclpP3) (Fig. 4a) were introduced into Xc17, generating reporter strains named Xc17(pFYclpP1), Xc17(pFYclpP2) and Xc17(pFYclpP3). The β-galactosidase activity produced by the obtained reporter strains grown in XOLN medium containing glycerol at 28 °C or 37 °C was measured. As shown in Fig. 4b, the β-galactosidase production from the same constructs did not change over the course of different time interval at 28 °C, indicating that the clpP promoter is independent on the growth phase of the cell under physiological temperature. At 28 °C, both Xc17(pFYclpP1) and Xc17(pFYclpP2) produced similar β-galactosidase activities (2124 U and 2273 U at 24 h, respectively), whereas Xc17(pFYclpP3) showed the same levels of activity as those of Xc17 carrying vector pFY13–9 (42 U) (Fig. 4b). At 37 °C, Xc17(pFYclpP1) and Xc17(pFYclpP2) expressed 4762 U and 4716 U of β-galactosidase, representing increases of 2.24- and 2.08-fold over the same strains grown under heat stress treatment, respectively. Taken together, these results demonstrated that the clpP expression is growth phase-independent and is under heat shock control. The expression of clpP is induced after heat treatment suggesting that ClpP is implicated in the degradation of aberrant proteins occurring after heat shock. Sequence analysis revealed that the upstream regions of the clpP gene presence of a putative σ32 promoter elements: a − 35 box (TTGAGA) and a − 10 box (CCCCAACT) located at − 229 and − 207 (with a spacer of 14 nucleotides) relatives to the translation start site, respectively (Fig. 4a). A predicted ribosome-binding site (GGAG) was located 6 nt upstream of the translation stat codon. The predicted − 35/− 10 region (TTGAGA/CCCCAACT) was a close match (underlined bases) to the consensus sequence (TTGAAA/CCCCATNT) of E. coli promoters known to be recognized by the σ32 factor [32]. In addition, the 14 bp spacing between the − 35 and − 10 regions was as well conserved as those of σ32 promoters [32]. The observations that (i) expression from pFYclpP1 and pFYclpP2 (carrying nt − 384/− 10 and − 254/− 10 regions relative to the clpP translation start site, which included potential − 35/− 10 elements) gave similar activity and induction fold; and (ii) the level of Xc17(pFYclpP3) (carrying nt − 196/− 10 regions, which did not include − 35/− 10 boxes) was the same as Xc17(pFY13–9), suggesting the − 254/− 10 regions contain the complete promoter sequence and is required for heat stress response.
Characterization of the clpP gene upstream region. a The upstream regions of the clpP gene cloned into the promoter-probing vector pFY13−9 to form reporter constructs pFYclpP1, pFYclpP2, and pFYclpP3. The regions containing the putative σ32 promoter elements are thickened. b Promoter activities expressed from different reporter constructs in Xc17. Promoter activity was measured as β-galactosidase activity (Miller units). Results are the means of at least three independent experiments. Error bars indicate standard deviations
The expression of clpP has been shown to be induced by heat shock in several Gram-positive bacteria, including Bacillus subtilis [33, 34], L. lactis [24], L. monocytogene [29], S. aureus [25], Streptococcus mutans [35], and S. pneumoniae [23]. A putative CtsR (class three stress gene repressor) DNA binding motif was present in the upstream region of clpP in B. subtilis [36], L. lactis [24], L. monocytogene [29], and S. pneumoniae [23]. Limited reports are available regarding the clpP expression in Gram-negative bacteria. The clpP of E. coli is a σ32-dependent heat shock gene and a twofold induction of transcription is observed after heat treatment [37, 38]. The transcription of Caulobacter crescentus clpP was found to be induced by heat shock and the strength of the clpP promoter is dependent on the growth phase of the cells [39]. In addition, the upstream region of clpP of C. crescentus shown to resemble the previously identified σ32-dependent heat shock promoter [39]. σ32 controls the expression of heat shock genes in E. coli and is broadly distributed in proteobacteria [32]. Here, a typical σ32-dependent promoter was identified upstream of the Xcc clpP and transcription of clpP was under heat control indicating the clpP gene is regulated by σ32 in this organism. Little information is available regarding the σ32-dependent promoter in Xcc. Prior to the present study, it was only known that the transcription of grpE, dnaK, and hspA with σ32-dependent promoter is induced in the presence of heat stress [40, 41]. Our results extend the insights of σ32 regulon in Xcc, although the regulatory mechanism remains to be elucidated.
Conclusion
In this work, we investigated the function and transcription of clpP gene of Xcc, the causative agent of black rot disease brassica corps worldwide. We constructed the clpP mutant by homologous recombination to examine the function of ClpP. The clpP mutant was shown to display a pleiotropic phenotype, including an impaired growth at high temperature, a reduced tolerance in response to puromycin treatment, an attenuation of virulence, and a defect in extracellular enzyme production. Genetic complementation of wild-type clpP could restore these altered phenotypes. Promoter activity assays indicated that clpP expression was induced by heat shock. The ClpP proteolytic activity is activated upon association with members of the ATPase-active chaperons [4], and such members have been annotated in Xcc genome [9,10,11]. The ClpX is involved in bacterial attachment, stress tolerance, and virulence in Xcc [12]. In our future work, we will determine whether ClpX (or ClpA) interacts with ClpP and what role they play in modulating the proteolytic activity in Xcc under physiological and stress conditions.
References
Deng CY, Deng AH, Sun ST, Wang L, Wu J, Wu Y, Chen XY, Fang RX, Wen TY, Qian W (2014) The periplasmic PDZ domain-containing protein Prc modulates full virulence, envelops stress responses, and directly interacts with dipeptidyl peptidase of Xanthomonas oryzae pv. oryzae. Mol Plant Microbe Interact 27(2):101–112. https://doi.org/10.1094/MPMI-08-13-0234-R
Brotz-Oesterhelt H, Sass P (2014) Bacterial caseinolytic proteases as novel targets for antibacterial treatment. Int J Med Microbiol 304(1):23–30. https://doi.org/10.1016/j.ijmm.2013.09.001
Malik IT, Brotz-Oesterhelt H (2017) Conformational control of the bacterial Clp protease by natural product antibiotics. Nat Prod Rep 34(7):815–831. https://doi.org/10.1039/c6np00125d
Kress W, Maglica Z, Weber-Ban E (2009) Clp chaperone-proteases: structure and function. Res Microbiol 160(9):618–628. https://doi.org/10.1016/j.resmic.2009.08.006
Bhandari V, Wong KS, Zhou JL, Mabanglo MF, Batey RA, Houry WA (2018) The role of ClpP protease in bacterial pathogenesis and human diseases. ACS Chem Biol 13(6):1413–1425. https://doi.org/10.1021/acschembio.8b00124
Ryan RP, Vorholter FJ, Potnis N, Jones JB, Van Sluys MA, Bogdanove AJ, Dow JM (2011) Pathogenomics of Xanthomonas: understanding bacterium–plant interactions. Nat Rev Microbiol 9(5):344–355
Dow JM, Crossman L, Findlay K, He YQ, Feng JX, Tang JL (2003) Biofilm dispersal in Xanthomonas campestris is controlled by cell-cell signaling and is required for full virulence to plants. Proc Natl Acad Sci USA 100(19):10995–11000
Chan JW, Goodwin PH (1999) The molecular genetics of virulence of Xanthomonas campestris. Biotechnol Adv 17(6):489–508
Liu YC, Wang SC, Yu YJ, Fung KM, Yang MT, Tseng YH, Tsai SF, Sun HS, Lyu PC, Chou SH (2015) Complete genome sequence of Xanthomonas campestris pv. campestris strain 17 from Taiwan. Genome Announc. https://doi.org/10.1128/genomeA.01466-15
Qian W, Jia Y, Ren SX, He YQ, Feng JX, Lu LF, Sun Q, Ying G, Tang DJ, Tang H, Wu W, Hao P, Wang L, Jiang BL, Zeng S, Gu WY, Lu G, Rong L, Tian Y, Yao Z, Fu G, Chen B, Fang R, Qiang B, Chen Z, Zhao GP, Tang JL, He C (2005) Comparative and functional genomic analyses of the pathogenicity of phytopathogen Xanthomonas campestris pv. campestris. Genome Res 15(6):757–767
da Silva AC, Ferro JA, Reinach FC, Farah CS, Furlan LR, Quaggio RB, Monteiro-Vitorello CB, Van Sluys MA, Almeida NF, Alves LM, do Amaral AM, Bertolini MC, Camargo LE, Camarotte G, Cannavan F, Cardozo J, Chambergo F, Ciapina LP, Cicarelli RM, Coutinho LL, Cursino-Santos JR, El-Dorry H, Faria JB, Ferreira AJ, Ferreira RC, Ferro MI, Formighieri EF, Franco MC, Greggio CC, Gruber A, Katsuyama AM, Kishi LT, Leite RP, Lemos EG, Lemos MV, Locali EC, Machado MA, Madeira AM, Martinez-Rossi NM, Martins EC, Meidanis J, Menck CF, Miyaki CY, Moon DH, Moreira LM, Novo MT, Okura VK, Oliveira MC, Oliveira VR, Pereira HA, Rossi A, Sena JA, Silva C, de Souza RF, Spinola LA, Takita MA, Tamura RE, Teixeira EC, Tezza RI, Trindade dos Santos M, Truffi D, Tsai SM, White FF, Setubal JC, Kitajima JP (2002) Comparison of the genomes of two Xanthomonas pathogens with differing host specificities. Nature 417(6887):459–463
Lo HH, Liao CT, Li CE, Chiang YC, Hsiao YM (2020) The clpX gene plays an important role in bacterial attachment, stress tolerance, and virulence in Xanthomonas campestris pv. campestris. Arch Microbiol 202(3):597–607. https://doi.org/10.1007/s00203-019-01772-3
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Habor Press, Cold Spring Harbor
Fu JF, Tseng YH (1990) Construction of lactose-utilizing Xanthomonas campestris and production of xanthan gum from whey. Appl Environ Microbiol 56(4):919–923
Hsiao YM, Liao HY, Lee MC, Yang TC, Tseng YH (2005) Clp upregulates transcription of engA gene encoding a virulence factor in Xanthomonas campestris by direct binding to the upstream tandem Clp sites. FEBS Lett 579(17):3525–3533
Wang TW, Tseng YH (1992) Electrotransformation of Xanthomonas campestris by RF DNA of filamentous phage fLf. Lett Appl Microbiol 14(2):65–68
Vieira J, Messing J (1991) New pUC-derived cloning vectors with different selectable markers and DNA replication origins. Gene 100:189–194
Schweizer HD (1993) Small broad-host-range gentamycin resistance gene cassettes for site-specific insertion and deletion mutagenesis. Biotechniques 15(5):831–834
Keen NT, Tamaki S, Kobayashi D, Trollinger D (1988) Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene 70(1):191–197
Liao CT, Chiang YC, Hsiao YM (2019) Functional characterization and proteomic analysis of lolA in Xanthomonas campestris pv. campestris. BMC Microbiol 19(1):20. https://doi.org/10.1186/s12866-019-1387-9
Lee TC, Chen ST, Lee MC, Chang CM, Chen CH, Weng SF, Tseng YH (2001) The early stages of filamentous phage fLf infection require the host transcription factor. Clp J Mol Microbiol Biotechnol 3(3):471–481
Chiang YC, Liao CT, Du SC, Hsiao YM (2017) Functional characterization and transcriptional analysis of icd2 gene encoding an isocitrate dehydrogenase of Xanthomonas campestris pv. campestris. Arch Microbiol 199(6):917–929. https://doi.org/10.1007/s00203-017-1370-5
Robertson GT, Ng WL, Foley J, Gilmour R, Winkler ME (2002) Global transcriptional analysis of clpP mutations of type 2 Streptococcus pneumoniae and their effects on physiology and virulence. J Bacteriol 184(13):3508–3520. https://doi.org/10.1128/jb.184.13.3508-3520.2002
Frees D, Ingmer H (1999) ClpP participates in the degradation of misfolded protein in Lactococcus lactis. Mol Microbiol 31(1):79–87. https://doi.org/10.1046/j.1365-2958.1999.01149.x
Frees D, Qazi SN, Hill PJ, Ingmer H (2003) Alternative roles of ClpX and ClpP in Staphylococcus aureus stress tolerance and virulence. Mol Microbiol 48(6):1565–1578. https://doi.org/10.1046/j.1365-2958.2003.03524.x
Xie F, Zhang Y, Li G, Zhou L, Liu S, Wang C (2013) The ClpP protease is required for the stress tolerance and biofilm formation in Actinobacillus pleuropneumoniae. PLoS ONE 8(1):e53600. https://doi.org/10.1371/journal.pone.0053600
Roy S, Zhu Y, Ma J, Roy AC, Zhang Y, Zhong X, Pan Z, Yao H (2019) Role of ClpX and ClpP in Streptococcus suis serotype 2 stress tolerance and virulence. Microbiol Res 223–225:99–109. https://doi.org/10.1016/j.micres.2019.04.003
Li Y, Yamazaki A, Zou L, Biddle E, Zeng Q, Wang Y, Lin H, Wang Q, Yang CH (2010) ClpXP protease regulates the type III secretion system of Dickeya dadantii 3937 and is essential for the bacterial virulence. Mol Plant Microbe Interact 23(7):871–878. https://doi.org/10.1094/MPMI-23-7-0871
Gaillot O, Pellegrini E, Bregenholt S, Nair S, Berche P (2000) The ClpP serine protease is essential for the intracellular parasitism and virulence of Listeria monocytogenes. Mol Microbiol 35(6):1286–1294. https://doi.org/10.1046/j.1365-2958.2000.01773.x
Flynn JM, Neher SB, Kim YI, Sauer RT, Baker TA (2003) Proteomic discovery of cellular substrates of the ClpXP protease reveals five classes of ClpX-recognition signals. Mol Cell 11(3):671–683. https://doi.org/10.1016/s1097-2765(03)00060-1
Kock H, Gerth U, Hecker M (2004) The ClpP peptidase is the major determinant of bulk protein turnover in Bacillus subtilis. J Bacteriol 186(17):5856–5864. https://doi.org/10.1128/JB.186.17.5856-5864.2004
Koo BM, Rhodius VA, Campbell EA, Gross CA (2009) Dissection of recognition determinants of Escherichia coli s32 suggests a composite −10 region with an 'extended −10' motif and a core −10 element. Mol Microbiol 72(4):815–829. https://doi.org/10.1111/j.1365-2958.2009.06690.x
Msadek T, Dartois V, Kunst F, Herbaud ML, Denizot F, Rapoport G (1998) ClpP of Bacillus subtilis is required for competence development, motility, degradative enzyme synthesis, growth at high temperature and sporulation. Mol Microbiol 27(5):899–914. https://doi.org/10.1046/j.1365-2958.1998.00735.x
Gerth U, Kruger E, Derre I, Msadek T, Hecker M (1998) Stress induction of the Bacillus subtilis clpP gene encoding a homologue of the proteolytic component of the Clp protease and the involvement of ClpP and ClpX in stress tolerance. Mol Microbiol 28(4):787–802. https://doi.org/10.1046/j.1365-2958.1998.00840.x
Hou XH, Zhang JQ, Song XY, Ma XB, Zhang SY (2014) Contribution of ClpP to stress tolerance and virulence properties of Streptococcus mutans. J Basic Microbiol 54(11):1222–1232. https://doi.org/10.1002/jobm.201300747
Derre I, Rapoport G, Msadek T (1999) CtsR, a novel regulator of stress and heat shock response, controls clp and molecular chaperone gene expression in gram-positive bacteria. Mol Microbiol 31(1):117–131. https://doi.org/10.1046/j.1365-2958.1999.01152.x
Gottesman S, Clark WP, de Crecy-Lagard V, Maurizi MR (1993) ClpX, an alternative subunit for the ATP-dependent Clp protease of Escherichia coli. Sequence and in vivo activities. J Biol Chem 268(30):22618–22626
Kroh HE, Simon LD (1990) The ClpP component of Clp protease is the sigma 32-dependent heat shock protein F215. J Bacteriol 172(10):6026–6034. https://doi.org/10.1128/jb.172.10.6026-6034.1990
Osteras M, Stotz A, Schmid Nuoffer S, Jenal U (1999) Identification and transcriptional control of the genes encoding the Caulobacter crescentus ClpXP protease. J Bacteriol 181(10):3039–3050
Lin CH, Lee CN, Lin JW, Tsai WJ, Wang SW, Weng SF, Tseng YH (2010) Characterization of Xanthomonas campestris pv. campestris heat shock protein A (HspA), which possesses an intrinsic ability to reactivate inactivated proteins. Appl Microbiol Biotechnol 88(3):699–709. https://doi.org/10.1007/s00253-010-2776-z
Weng SF, Tai PM, Yang CH, Wu CD, Tsai WJ, Lin JW, Tseng YH (2001) Characterization of stress-responsive genes, hrcA-grpE-dnaK-dnaJ, from phytopathogenic Xanthomonas campestris. Arch Microbiol 176(1–2):121–128. https://doi.org/10.1007/s002030100302
Yang BY, Tseng YH (1988) Production of exopolysaccharide and levels of protease and pectinase activity in pathogenic and non-pathogenic strains of Xanthomonas campestris pv. campestris. Bot Bull Acad Sin 29:93–99
Acknowledgements
This study was supported by the Ministry of Science and Technology of Taiwan (Grant No. MOST107-2313-B-166-001-MY3).
Author information
Authors and Affiliations
Contributions
CE carried out the experiments and prepared figures and tables. CE, CT, HH, and YM analyzed the experimental results and interpreted the data. YM managed the grants, supervised the laboratory work, led the design, and coordination of this study and wrote the manuscript draft. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, CE., Liao, CT., Lo, HH. et al. Functional Characterization and Transcriptional Analysis of clpP of Xanthomonas campestris pv. campestris. Curr Microbiol 77, 2876–2885 (2020). https://doi.org/10.1007/s00284-020-02093-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-020-02093-1