Abstract
To determine the effect of organics (yeast extract) on microbial community during chalcopyrite bioleaching at different temperature, real-time polymerase chain reaction (PCR) was employed to analyze community dynamics of major bacteria applied in bioleaching. The results showed that yeast extract exerted great impact on microbial community, and therefore influencing bioleaching rate. To be specific, yeast extract was adverse to this bioleaching process at 30°C due to decreased proportion of important chemolithotrophs such as Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans. However, yeast extract could promote bioleaching rate at 40°C on account of the increased number and enhanced work of Ferroplasma thermophilum, a kind of facultative bacteria. Similarly, bioleaching rate was enhanced under the effect of yeast extract at 50°C owing to the work of Acidianus brierleyi. At 60°C, bioleaching rate was close to 100% and temperature was the dominant factor determining bioleaching rate. Interestingly, the existence of yeast extract greatly enhanced the relative competitiveness of Ferroplasma thermophilum in this complex bioleaching microbial community.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The recent years witnessed serious depletion of copper resources, especially the high grade ones. As a result, it is inevitable for people to process great number of intractable low grade ones. A lot of researches concerning the copper processing technology have been conducted [23–25]. In comparison with the conventional methods, bioleaching shows a number of advantages including simple process flows, low investment and operation cost, and environmentally friendly character [29].
The bioleaching microorganisms can be classified into three categories according to their adaptation to temperature: mesophiles, moderate thermophiles, and extreme thermophiles [1, 12]. Because various kinds of bacteria show distinct metabolic activity at different temperature, the environmental temperature exercises great impact on the function and structure of microbial community. Apart from the factor of temperature, the effect of organics should never be overlooked. Yeast extract often serves as the energy source for obligative or facultative heterotrophic microorganism [4]. In some experimental researches, to evaluate the effect of organics on bioleaching process, yeast extract is usually adopted as the typical organics [22]. According to some researches, a certain amount of organics is adverse to some autotrophic bioleaching bacteria [3, 5]. It is also reported that relatively high concentration (0.2% w/v) of yeast extract would inhibit the growth of some facultative autotrophic bacteria [20]. On the contrary, heterotrophic and facultative microorganisms can use organics to support their growth [34]. Obviously, the role played by organics in bioleaching is very complicated. It is the fact that bioleaching microbial community is extremely complex in actual situation, which consists of the autotrophic, heterotrophic, and facultative microorganisms. In this artificially assembled microbial community, what the role played by the organics in the complex bioleaching microbial community? How it influence the structure and function of the complex bioleaching microbial community? So far, others only focused on one strain to study the effect of organics on bioleaching [15]. And no comprehensive and systematic researches have been conducted to clarify the effect of organics on the structure and function of complex bioleaching microbial community within so large temperature range.
High levels of bacterial diversity make quantifying and characterizing microbial communities a daunting task. In recent years, real-time PCR has emerged as a strongly and widely used method for biological investigation for its outstanding advantages [27]. This method provides very accurate and reproducible quantization of gene copies and can detect and quantify extremely small amounts of specific nucleic acid sequences. Besides, because the real-time PCR does not require post-PCR sample handling, it can prevent potential PCR product carry-over contamination, resulting in much faster and higher throughput assays. Moreover, real-time PCR method has a very large dynamic range of starting target molecule determination [9]. As a result, the real-time PCR method has been widely applied to assess microbial community in some particular environment [6, 31]. Also, the application of real-time PCR has been demonstrated valid and accurate to monitor population dynamics of mixed bioleaching bacteria [32].
In this study, yeast extract was adopted to study the effect of organics on the population dynamics of bioleaching microbial community including Acidithiobacillus ferrooxidans, Leptospirillum ferriphilum, Acidithiobacillus thiooxidans, Acidithiobacillus caldus, Acidiphilium spp., Ferroplasma thermophilum, Acidianus brierleyi, and Sulfobacillus thermosulfidooxidans using the real-time PCR method. The structure and function of complex bioleaching microbial community were determined under the effect of temperature and organics.
Materials and Methods
Mineral Composition, Bacterial Strains, and Culture Conditions
The low grade mineral contained 0.3% copper and chalcopyrite accounted for 62.2% of the minerals containing copper. The mineral samples were crushed and then sieved through the 75 μm size pore for later use. All strains were provided by China Center for Type Culture Collection (CCTCC). Each strain was cultivated in appropriate medium under different conditions (Table 1). After being harvested by centrifugation at 10,000×g for 20 min and being made into bacterial suspension with sterilized water (pH 2.0), these eight bacteria strains served as the inocula for chalcopyrite bioleaching. Then, each strain was equally (1.0 × 107 per/ml) inoculated into 250 ml flask containing sterilized chalcopyrite medium (pH 1.5). We carried out the enumeration through direct counting method using blood counting chamber. The sterilized chalcopyrite medium consisted of the processed minerals (previously mentioned) and sterilized 9K basic medium without FeSO4. The special 9K culture medium included (NH4)4SO4 (3.0 g/l), Ca(NO3)2 (0.01 g/l), MgSO4·7H2O (0.5 g/l), K2HPO4 (0.5 g/l), and KCl (0.1 g/l). After this step, bacterial mixture was cultivated at 30, 40, 50, and 60°C under the aerobic condition at 170 r/min (in the rotary platform). There were two experiment groups at each temperature, one group was added with 0.01% yeast extract and the other group was not. The abiotic controls were also designed at each temperature. The leaching experiments lasted for 28 days. All tests were conducted in triplicate.
Physicochemical Analysis
Samples were collected at regular intervals to be measured for dissolved copper, total dissolved iron, and pH. Copper and total iron concentration in solution were measured by inductively coupled plasma (ICP) total analysis method at 4th, 8th, 12th, 16th, 20th, 24th, and 28th day. The pH of the bioleaching systems was measured with a pH meter, and the pH of the system was kept stable around 1.5.
Extraction of Genomic DNA
Genomic DNA was extracted from bacteria using a TIANamp Bacteria DNA kit (Tiangen, Beijing, China). The acquired genomic DNA stained by ethidium bromide was checked through 1.0% agarose gel electrophoresis. Purified genomic DNA concentration was measured spectrophotometrically using a DanoDrop ND-1000 spectrophotometer (NanoDrop Technologies, Wilmington, USA) and adjusted to proper concentration for the PCR use.
Design and Specificity of PCR Primers
Primers used in the study are summarized in Table 2. PCR primers were designed by using Primer Primier 5.0 and then synthesized by Shanghai Sangon Company (China). According to the size of the specific amplified products stained by ethidium bromide through 1.5% agarose gel electrophoresis, the specificity of primers can be determined. DNA sequencing was conducted by Shanghai Sangon Company (China) and BLAST analysis in GenBank. Specificity of these primers was also evaluated by real-time PCR (see below).
The PCR reactions were carried out under the following condition: 30 s denaturation at 95°C, 30 s annealing at 59°C, and 30 s extension at 72°C, along with an initial 3 min denaturation at 95°C, and a final 10 min extension reaction at 72°C. Then, the PCR products were analyzed through agarose gel electrophoresis and were purified using QIAquick-spin PCR purification kit (Qiagen, Hilden, Germany). The PCR product was cloned to T-Vector for sequence analysis. The real-time PCR was carried out with iCycler iQ Real-time PCR detection system (Bio-Rad, Hercules, Calif.). Each reaction mixture (final volume, 25 μl) contained 12.5 μl of SYBR Green Real-time PCR Master Mix (Toyobo) which consisted of Taq DNA polymerase, dNTP, MgCl2, SYBR Green I dye, 5 pmol/μl the forward primer, 5 pmol/μl the reverse primer, and 5 μl DNA template. Then nuclease-free water was used to make up the whole system to 25 μl. Under the monitoring of an iCycler iO Real-time PCR detection system, the real-time PCR reactions were carried out under the following condition: initial 3 min denaturation at 95°C, and then 40 cycles of 95°C for 30 s, 59°C for 30 s, and 72°C for 30 s. After the completion of each run, melting curves for the amplicons were measured by raising the temperature 0.5°C from 59 to 99°C while monitoring the change of fluorescence rate. All tests were conducted in triplicate.
Determination of Microbial Community Structure
At 4th, 12th, 20th, and 28th day, leaching solution combined with the elution from chalcopyrite surface were added into centrifugal flask, then the bacteria were harvested through centrifugation at 10,000×g for 20 min at 4°C in a 5804R centrifuge (Eppendorf, Wesbury, NY). Genomic DNA was extracted using a TIANamp Bacteria DNA kit (Tiangen, Beijing, China). To determine the microbial community structure, real-time PCR method was carried out using specific primers based on gyrB gene, 16S rRNA gene, and arsB gene. The real-time PCR mixture and reaction condition was the same just as previously mentioned. The microbial community structure was determined according to the proportion of the targeted gene copies of each strain in the same sample.
Results and Discussion
Evaluation of the Specificity of Primers and Real-time PCR Assay
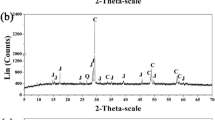
The quality of the amplified products stained by ethidium bromide was checked through 1.5% agarose gel electrophoresis. Amplified DNA fragments were correct because each amplified products was a single segment matching the expected size (Fig. 1). DNA sequencing was carried out by BLAST analysis in GenBank, and they matched the sequence of the gene that has been deposited in GenBank. So, we do not re-put the existed sequence into GenBank. GenBank numbers of these sequences have been shown in the Table 2. These results suggested that the conserved 16S rRNA gene, less conserved gyrB gene, and functional gene such as arsB gene are suitable and accurate to analyze this microbial community [11, 30]. In order to examine the bacteria growth kinetics and microbial community dynamics, copy number of the specific gene can be regarded as a proxy for cell counts using quantitative PCR [8]. No matter the genes are single-copy or multi-copy in their genomes, copy number of the specific gene can serve as the relatively quantitative indicator that can reflect the alternation tendency of this microbial community.
PCR amplification products of selected genes deriving from eight strains on 2% agarose gel: lane 2 A. ferrooxidans F1, lane 3 L. ferriphilum YSK, lane 4 At. thiooxidans A01, lane 5 At. caldus S1, lane 6 Acidiphilium spp. DX1-1, lane 7 F. thermophilum L1, lane 8 A. brierleyi, lane 9 Sulfobacillus thermosulfidooxidans, and lanes 1, 10 marker
The Effect of Yeast Extract on Microbial Community Structure at Different Temperature
The Microbial Community Shift Under the Effect of 0.01% Yeast Extract at 30°C
In the group without being added yeast extract, L. ferriphilum YSK accounting for 51% of the whole community and At. thiooxidans A01 accounting for 28% of the whole community were the dominant strains at 4th day. Then, the population proportion of L. ferriphilum YSK and At. thiooxidans, respectively, decreased to 34 and 18% at 12th day. The microbial community structure became increasingly simple in the following period. L. ferriphilum YSK was the only dominant strain in later period together with little number of other strains such as At. thiooxidans A01, At. caldus S1, and F. thermophilum L1 (Fig. 2a). The predominance of L. ferriphilum at 30°C was because it optimally utilizes energy from oxidizing Fe2+ at warm temperatures (25–42°C) [18]. Under the effect of 0.01% yeast extract, microbial community structure was different (Fig. 2a). F. thermophilum L1 accounting for 23% of the whole community, At. thiooxidans A01 accounting for 29% of the whole community, and Acidiphilium spp. DX1-1 accounting for 35% of the whole were the dominant strains at 4th day. Total seven strains except for A. brierleyi were found at 4th day. From 4th day, the number of F. thermophilum L1 began to increase. As a result, this strain became the dominant strain in the following period. This was because yeast extract could promote the growth of F. thermophilum L1 that belongs to a kind of facultative bacteria [33]. It was found that the microbial community under the effect of 0.01% yeast extract was more complex than the community without being added yeast extract at 28th day (Fig. 2a). According to the Simpson’ diversity index equation, the diversity index value of the community added with yeast extract (0.6320) was much greater than the value of the community without being added yeast extract (0.09674). Obviously, the existence of yeast extract increased the number of facultative heterotrophically bacteria such as Acidiphilium spp. DX1-1 and F. thermophilum L1, and thereby generating high microbial community diversity index. Interestingly, it seemed that At. thiooxidans A01 could perform better under the effect of 0.01% yeast extract compared with other autotrophic bacteria at 30°C.
Microbial community shift under the effect of yeast extract at different temperature. a Microbial community shift of the group added with 0.01% yeast extract and the group without being added yeast extract at 30°C; b microbial community shift of the group added with 0.01% yeast extract and the group without being added yeast extract at 40°C; c microbial community shift of the group added with 0.01% yeast extract and the group without being added yeast extract at 50°C; d microbial community shift of the group added with 0.01% yeast extract and the group without being added yeast extract at 60°C. The left bar stands for the microbial community composition of the group added with yeast extract at a specific day; the right bar stands for the microbial community composition of the group without being added yeast extract at a specific day
The Microbial Community Shift Under the Effect of 0.01% Yeast Extract at 40°C
In the group without being added 0.01% yeast extract, seven strains except for A. brierleyi were found at 4th and 12th day. The thermophile character of A. brierleyi [14] accounts for its absence in the medium at 40°C. At. caldus S1 and L. ferriphilum YSK were the dominant bacteria at 4th and 12th day and their population proportion, respectively, increased from 34 to 37% and from 21 to 27%. From 12th day, the number of F. thermophilum L1 began to increase. Its population proportion increased from 27% at 20th day to 29% at 28th day. In the final period (from 20th to 28th day), F. thermophilum L1, At. caldus S1, and L. ferriphilum YSK were the dominant strains (Fig. 2b). This was because F. thermophilum L1 performs optimally at 45°C, At. caldus is a moderate thermophile with a growth temperature optimum of 45°C [13], and L. ferriphilum YSK has the ability to grow at 45°C [7]. Under the effect of 0.01% yeast extract, microbial community structure shift was distinct (Fig. 2b). At the 4th day, the microbial community, containing F. thermophilum L1 (33%), Acidiphilium spp. DX1-1 (15%), At. caldus S1 (28%), and other little number of strains, was relatively complex. At the 12th day, the dominant bacteria were F. thermophilum L1 (45%) and Acidiphilium spp. DX1-1 (27%). In the following period, the number of F. thermophilum L1 made a further increase, and this strain became the only dominant bacteria. According to Fig. 2b, it is clear that the proportion of F. thermophilum L1 was higher in the group added with yeast extract than the other group without being added yeast extract at 40°C. The strong competitive power of F. thermophilum stemmed from the fact that F. thermophilum L1 has a temperature optimum of 45°C and grows well utilizing both ferrous iron and yeast extract [33].
The Microbial Community Shift Under the Effect of 0.01% Yeast Extract at 50°C
Acidithiobacillus caldus S1 (56%) and S. thermosulfidooxidans (34%) were the dominant strains at 4th day in the group without being added yeast extract. From the 4th day, the number of At. caldus S1 was dramatically decreased, however, the number of S. thermosulfidooxidans and F. thermophilum L1 was considerably increased. As a result, these two strains became the dominant bacteria (Fig. 2c). This was because S. thermosulfidooxidans belongs to moderate thermophile and actively functions in chalcopyrite dissolution at this temperature [28]. Under the effect of 0.01% yeast extract, microbial community structure shift was different at 50°C (Fig. 2c). During the whole experiment period, only four kinds of strains such as F. thermophilum L1, S. thermosulfidooxidans, A. brierleyi, and At. caldus S1 were found. F. thermophilum L1 (55%) and S. thermosulfidooxidans (24%) were the dominant bacteria at 4th day. From the 4th day, the number of At. caldus S1 began to fall down, and this strain was almost vanished in the medium finally. From the 20th day, the number of F. thermophilum L1 began to fall, however, the number of A. brierleyi started to increase significantly. As a result, A. brierleyi (54%) became the dominant group together with previous dominant strains, F. thermophilum L1 (30%) and S. thermosulfidooxidans (16%), at 28th day. This was due to the fact that F. thermophilum L1 and S. thermosulfidooxidans are moderate thermophiles [10] and A. brierleyi belongs to thermophiles, however, other strains might be inhibited by high temperature. Besides, A. brierleyi demonstrated strong competitive power at 50°C even its optimal temperature (around 65°C) is far beyond 50°C. This was probably because yeast extract abounding with protein, amino acid, vitamin, and trace element is beneficial for its growth.
The Microbial Community Shift under the Effect of 0.01% Yeast Extract at 60°C
In the group without being added 0.01% yeast extract, F. thermophilum L1 (28%), S. thermosulfidooxidans (29%), and A. brierleyi(43%) were the dominant strains at 4th day. From 4th day, the number of F. thermophilum L1 was dramatically decreased, and A. brierleyi became the only dominant bacteria together with a little proportion of S. thermosulfidooxidans (Fig. 2d). This was because A. brierleyi showed high activity at 60°C. Under the effect of 0.01% yeast extract, microbial community comprised only three strains, namely, F. thermophilum L1, S. thermosulfidooxidans, and A. brierleyi (Fig. 2d). In the beginning, these three strains were dominant groups. From 4th day, the number of A. brierleyi began to increase, however, other two strains started to decrease. During the mid-term and later period (from 12th to 28th day), A. brierleyi grew up to the only dominant strain, which was attributed to the fact that A. brierleyi belongs to thermophile that could adapt to extremely high temperature [14]. At 60°C, the existence of yeast extract also decreased competitive power of S. thermosulfidooxidans, and temperature was the major factor determining the microbial community structure.
Surprisingly, the F. thermophilum L1 was the dominant group at 30, 40, and 50°C and was existed at 60°C in the group added with 0.01% yeast extract. As a result, the existence of yeast extract greatly enhanced the relative competitiveness of F. thermophilum L1 in this environment containing available organics and inorganics. This was because F. thermophilum L1 has the special ability of utilizing both ferrous iron and yeast extract [33].
Exploration Toward the Effect of Yeast Extract on Fe3+ Concentration and Bioleaching Rate Based on Microbial Community Shift at Different Temperatures
The concentration of Fe3+ was dramatically increased during the first 4 days (Figs. 3, 4). Fe3+ derived from the oxidation of Fe2+ [19], during this process bacteria could produce energy for their growth and metabolism. At 30°C, Fe3+ concentration of the groups without being added yeast extract was 498, 768, 930, 982, 1014, 1132, and 1290 mg/l, ,respectively at 4th, 8th, 12th, 16th, 20th, 24th, and 28th day. Obviously, it kept increasing all the time, and it was higher than the concentration of the group added with yeast extract (Figs. 3, 4). This could be explained in two parts. First, organics exhibited toxicity to chemolithotrophs that could oxidize Fe2+ to Fe3+ [16]. To be specific, organics might inhibit oxidation of ferrous iron and sulfur by A. ferrooxidans and At. thiooxidans, the major functional strains at 30°C [5]. Besides, in the group added with 0.01% yeast extract, the increasing number of Acidiphilium spp. (Fig. 2a) reduced Fe3+ to Fe2+ utilizing energy from organics [12] at 30°C. For the reason that Fe3+ can function as oxidizer to accelerate mineral dissolution and exclusive Fe3+ is beneficial to mineral oxidation [25, 26], high Fe3+ concentration gave rise to the high bioleaching rate (from 17.2% at 4th day to 27.6% at 28th day) of the group without being added yeast extract compared with the rate (from 19.2 to 32.7%) of the group added with yeast extract at 30°C (Fig. 5).
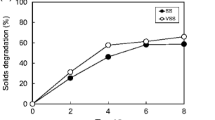
Comparison of bioleaching rate between the groups added with 0.01% yeast extract and the group without being added 0.01% yeast extract. a Change of bioleaching rate of the group without being added 0.01% yeast extract at different temperature; b change of bioleaching rate of the group added with 0.01% yeast extract at different temperature
At 40°C, Fe3+ concentration of the group added with yeast extract kept increasing during the whole bioleaching process. To be specific, it was 697 mg/l at 4th day, 875 mg/l at 8th day, 1004 mg/l at 12th day, 1124 mg/l at 16th day, 1244 mg/l at 20th day, 1301 mg/l at 24th day, and 1351 mg/l at 28th day. However, Fe3+ concentration of the group without being added yeast extract turned to decrease at 16th day, and it was lower than the concentration of the group added with yeast extract all the time (Figs. 3, 4). F. thermophilum L1, a strain has a temperature optimum of 45°C and is capable of chemomixotrophic growth on ferrous ion and yeast extract [33], took the dominant place of At. caldus S1 that is a sulfur oxidizer [17] under the effect of yeast extract at 40°C (Fig. 2b). For the reason that F. thermophilum L1, could oxidize Fe2+ into Fe3+, the increasing number of F. thermophilum L1 in the group added with yeast extract led to the high Fe3+ concentration in solution. Because Fe3+ is functional in mineral oxidation and dissolution [25, 26], the relatively high Fe3+ concentration promoted the bioleaching rate (from 24.9 to 49.8%) of the group added with yeast extract (Fig. 5).
At 50°C, Fe3+ concentration of the group added with some yeast extract reached the peak (1317 mg/l) at 20th day, then it began to fall down (Fig. 4). Fe3+ concentration of the group without being added 0.01% yeast extract turned to fall down at 12th day (Fig. 3). Fe3+ concentration of the group added with yeast extract was higher than that of the other group in later period probably because organics increased jarosite dissolution [2]. The microbial community of the group added with yeast extract mainly consisted of A. brierleyi, F. thermophilum L1, S. thermosulfidooxidans, and the community of the group without being added yeast extract primarily consisted of F. thermophilum L1 and S. thermosulfidooxidans (Fig. 2c). Obviously, the existence of A. brierleyi functioned as the differential factor. It is possible that the combined work of three facultative bacteria strains, namely, A. brierleyi, F. thermophilum L1, and S. thermosulfidooxidans was superior to the work of F. thermophilum L1 and S. thermosulfidooxidans. This was probably because A. brierleyi, a bacteria that is capable of oxidizing sulfur, plays an important role in mineral dissolution. During the oxidization process, chalcopyrite abounded with metal sulfide was broken up under the work of A. brierleyi. The solubilization of metal sulfides was to provide sulfuric acid for a proton attack and to keep the iron in the oxidized ferric state for an oxidative attack on the mineral, and therefore improving bioleaching rate [23]. Consequently, the high Fe3+ concentration and strong combined work of A. brierleyi, F. thermophilum L1, and S. thermosulfidooxidans might account for the high bioleaching rate (from 33.0 to 78.1%) of the group added with 0.01% yeast extract in comparison with the rate (from 31.5 to 71.2%) of the other group at 50°C (Fig. 5).
At 60°C, Fe3+ concentration of the group without being added yeast extract began to decrease from 8th day and Fe3+ concentration of the other group started to fall down from 4th day (Figs. 3, 4). The dramatic decrease of concentration might be caused by jarosite formation [21]. Even though microbial community structure was different between the two groups (Fig. 2d), Figure 5 indicated that the bioleaching rate of the two groups were both close to 100% at 28th day, respectively, 97.1 and 98.0%. This was probably because chalcopyrite lattice was broken under the effect of high temperature. Consequently, temperature was the dominant factor determining bioleaching rate at 60°C.
References
Bond PL, Druschel GK, Banfield JF (2000) Comparison of acid mine drainage microbial community in physically and geochemically distinct ecosystem. Appl Environ Microbiol 66:4962–4971
Chu C, Lin C, Wu Y, Lu W, Long J (2006) Organic matter increases jarosite dissolution in acid sulfate soils under inundation conditions. Aust J Soil Res 44:11–16
Coram NJ, Rawlings DE (2002) Molecular relationship between two groups of the genus Leptospirillum and the finding that Leptospirillum ferriphilum sp. nov. dominates South African commercial biooxidation tanks that operate at 40°C. Appl Environ Microbiol 68:838–845
Cruz FLS, Oliveira VA, Guimarães D, Souza AD, Leão VA (2010) High-temperature bioleaching of nickel sulfides: thermodynamic and kinetic implications. Hydrometallurgy 105:103–109
Fang D, Zhou L (2006) Effect of sludge dissolved organic matter on oxidation of ferrous iron and sulfur by Acidithiobacillus ferrooxidans and Acidithiobacillus thiooxidans. Water Air Soil Pollut 171:81–94
Fierer N, Jackson JA, Vilgalys R, Jackson RB (2005) Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71:4117–4120
Gao J, Zhang C, Wu X, Wang H, Qiu G (2007) Isolation and identification of a strain of Leptospirillum ferriphilum from an extreme acid mine drainage site. Ann Microbiol 57:171–176
Girguis PR, Cozen AE, DeLong EF (2005) Growth and population dynamics of anaerobic methane-oxidizing archaea and sulfate-reducing bacteria in a continuous-flow bioreactor. Appl Environ Microbiol 71:3725–3733
Heid CA, Stevens JK, Livak J, Williams PM (1996) Real time quantitative PCR. Genome Res 6:986–994
Ilyasa S, Anwarb MA, Niazia SB, Ghauri MA (2007) Bioleaching of metals from electronic scrap by moderately thermophilic acidophilic bacteria. Hydrometallurgy 88:180–188
Inskeep WP, Rusch DB, Jay ZJ, Herrgard MJ, Kozubal MA (2010) Metagenomes from high-temperature chemotrophic systems reveal geochemical controls on microbial community structure and function. PLoS One 5:e9773
Johnson DB, Hallberg KB (2003) The microbiology of acidic mine waters. Res Microbiol 154:466–473
Kirby BM, Vengadajellum CJ, Burton SG, Cowan DA (2010) Anthropogenically-created habitats—coal, coal mines and spoil heaps. In: Timmis KN (ed) Handbook of hydrocarbon and lipid Microbiology, vol 3. Springer, Heidelberg, pp 2277–2292
Konishi Y, Yoshida S, Asai S (1995) Bioleaching of pyrite by acidophilic thermophile Acidianus brierleyi. Biotechnol Bioeng 48:592–600
Konishi Y, Yoshida S, Asai S (1998) Effect of yeast extract supplementation in leach solution on bioleaching rate of pyrite by acidophilic thermophile Acidianus brierleyi. Biotechnol Bioeng 58:663–667
Ñancucheo I, Johnson DB (2010) Production of glycolic acid by chemolithotrophic iron- and sulfur-oxidizing bacteria and its role in delineating and sustaining acidophilic sulfide mineral-oxidizing consortia. Appl Environ Microbiol 76:461–467
Norris PR (2007) Acidophile diversity in mineral sulfide oxidation. Biomining 10:199–216
Ojumu TV, Hansford GS, Petersen J (2009) The kinetics of ferrous-iron oxidation by Leptospirillum ferriphilum in continuous culture: the effect of temperature. Biochem Eng J 46:161–168
Ozkaya B, Sahinkaya E, Nurmi P, Kaksonen AH, Puhakka JA (2008) Biologically Fe2+ oxidizing fluidized bed reactor performance and controlling of Fe3+ recycle during heap bioleaching: an artificial neural network-based model. Bioprocess Biosyst Eng 31:111–117
Peng J, Zhang R, Zhang Q, Zhang L, Zhou H (2008) Screening and characterization of Acidiphilium sp. PJH and its role in bioleaching. Trans Nonferr Met Soc China 18:1443–1449
Pradhan N, Nathsarma KC, Srinivasa Rao KL, Sukla B, Mishra BK (2008) Heap bioleaching of chalcopyrite: a review. Miner Eng 21:355–365
Puhakka J, Tuovinen OH (1987) Effect of organic compounds on the microbiological leaching of a complex sulphide ore material. World J Microbiol Biotechnol 3:429–436
Rawlings DE (2005) Characteristics and adaptability of iron- and sulfur-oxidizing microorganisms used for the recovery of metals from minerals and their concentrates. Microb Cell Fact 4:13
Rodríguez Y, Ballester A, Blázquez ML, González F, Muñoz JA (2003) New information on the pyrite bioleaching mechanism at low and high temperature. Hydrometallurgy 71:37–46
Sand W, Gehrke TP, Jozsa G (2001) (Bio)chemistry of bacterial leaching—direct vs. indirect bioleaching. Hydrometallurgy 59:159–175
Tributsch H (2001) Direct versus indirect bioleaching. Hydrometallurgy 59:177–185
Valasek MA, Repa JJ (2005) The power of real-time PCR. Adv Physiol Educ 29:151–159
Xia J, Yang Y, He H, Liang C, Zhao X, Zheng L, Ma C, Zhao Y, Nie Z, Qiu G (2010) Investigation of the sulfur speciation during chalcopyrite leaching by moderate thermophile Sulfobacillus thermosulfidooxidans. Int J Miner Process 94:52–57
Yang S, Xie J, Qiu G (2002) Research and application of bioleaching and biooxidation technologies in China. Miner Eng 15:361–363
Yin H, Cao L, Qiu G, Wang D, Kellogg L, Zhou J, Liu X, Dai Z, Ding J, Liu X (2008) Molecular diversity of 16S rRNA and gyrB genes in copper mines. Arch Microbiol 189:101–110
Zeng W, Qiu G, Zhou H, Peng J, Chen M, Tan S, Chao W, Liu X, Zhang Y (2010) Community structure and dynamics of the free and attached microorganisms during moderately thermophilic bioleaching of chalcopyrite concentrate. Bioresour Technol 101:7068–7075
Zhang L, Qiu G, Hu Y, Sun X, Li J, Gu G (2008) Bioleaching of pyrite by A. ferrooxidans and L. ferriphilum. Trans Nonferr Met Soc China 18:1415–1420
Zhou H, Zhang R, Hu P, Zeng W, Xie Y, Wu C, Qiu G (2008) Isolation and characterization of Ferroplasma thermophilum sp. nov., a novel extremely acidophilic, moderately thermophilic archaeon and its role in bioleaching of chalcopyrite. J Appl Microbiol 105:591–601
Zou L, Qian L, Zhang Y, Wan M, Qiu G, Yang Y (2008) Isolation and identification of Acidiphilium strain DY from complex sulfide mines and its bioleaching characterization. Chin J Nonferr Met 18:336–341
Acknowledgments
This research was supported by the National Basic Research Program (No. 2010CB630901), and the National Natural Science Foundation of China (Nos. 50621063, 30428014, and 30900203).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Qihou Li, Ye Tian, and Xian Fu contributed equally to this study.
Rights and permissions
About this article
Cite this article
Li, Q., Tian, Y., Fu, X. et al. The Community Dynamics of Major Bioleaching Microorganisms During Chalcopyrite Leaching Under the Effect of Organics. Curr Microbiol 63, 164–172 (2011). https://doi.org/10.1007/s00284-011-9960-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-011-9960-y