Abstract
Inorganic polyphosphate (polyP) is a ubiquitous linear polymer of hundreds of orthophosphate (Pi) residues linked by ATP-like, high-energy, phosphoanhydride bonds. The gene Rv1026 in Mycobacterium tuberculosis encodes a putative exopolyphosphatase which progressively hydrolyzes the terminal residues of polyP to liberate Pi. Rv1026 was cloned into the expressive plasmid pMV261. The resulting plasmid pRv1026 and the plasmid pMV261 were transformed into M. smegmatis strain mc2155 by electroporation. The recombinant M. smegmatis (pRv1026) showed relatively decreased polyP concentration and a phenotype different from the M. smegmatis (pMV261) in sliding motility and biofilm formation. The surfactant Tween 80 can enhance this effect on the sliding motility and biofilm formation of M. smegmatis. There are four different peaks between the gas chromatography of cellular wall fatty acid of the M. smegmatis (pRv1026) and the M. smegmatis (pMV261). These results indicate that polyP deficiency can affect the fatty acid composition of cellular wall and these alteration of cell wall might elucidate the reductive ability of strains to slide and form biofilm. This investigation provides novel recognition about the role of Rv1026, which provides novel clues for further study on the physiological role of Rv1026 in M. tuberculosis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A biofilm is multicellular communities in which cells adhere to each other or solid surface. These adhering cells are frequently encapsulated within a self-produced matrix of extracellular polymeric substance. Biofilm cells significantly enhance their resistance to antibiotics and the human immune system than their planktonic counterparts. Biofilm formation is initiated with surface attachment of planktonic bacteria, followed by formation of clusters and microcolonies, and subsequent development of differentiated structures in which individual bacteria and the entire community are enclosed by exopolysaccharides. And when biofilms disperses, the biofilm development cycle comes full circle [17].
While the ability of Mycobacterium tuberculosis to form biofilms remains obscure, many other species of mycobacteria, including M. avium, M. fortuitum, M. marinum, and M. smegmatis, are well-documented biofilm producers. However, little is known about the molecular mechanisms involved in mycobacterial biofilm formation, the nature of the matrix or the biofilm structure. Previous articles reported that M. smegmatis forms biofilms involving in glycopeptidolipids [26, 27] and mycolyl-diacylglycerols [6], and the amphiphilic molecules model elucidated the function of these molecules in biofilm formation [27]. Additionally, some other factors such as undecaprenyl phosphokinase [30], serine/threonine protein kinase [9], iron [19, 41], and GroEL1 are required for biofilm formation. M. smegmatis biofilm formation earlier events involve attachment and spreading, followed by the maturation and matrix formation. Because mycobacterial genomes do not contain the genetic material for exopolysaccharide biosynthesis [41], the matrix is probably primarily composed of fatty acids. For example, GroEL1 is required for fatty acid synthesis, which results in the increased synthesis of C56–C68 fatty acids during the biofilm formation, and a mutant defective in GroEL1 is specifically deficient in the late stages of biofilm formation [18]. Besides, the availability of iron for M. smegmatis biofilm maturation correlates closely with the synthesis of C56–C68 fatty acids [19, 41].
Due to mycobacteria being nonflagellated microorganisms, sliding motility is produced by the expansive forces of the growing bacterial population and cell surface properties that favor reduced friction between the cells and substrate, and the result is the slow movement of a uniform monolayer of cells as a unit [14]. Both non-pathogenic M. smegmatis and the opportunistic pathogen M. avium are able to slide, and this sliding motility correlates with the presence of glycopeptidolipids and mycolyl-diacylglycerols [27]. So the ability of mycobacteria to slide over the surface of motility plates is relative with biofilm formation.
Inorganic polyphosphate (polyP) is a linear polymer of hundreds of orthophosphate (Pi) residues linked by ATP-like, high-energy, phosphoanhydride bonds and found in all organisms [21]. In bacteria, two main kinds of enzymes are involved in the metabolism of polyP: polyphosphate kinases (PPK1 and PPK2) catalyze the reversible conversion of the terminal phosphate of ATP (or GTP) into polyP; exopolyphosphatase (PPX) progressively hydrolyzes the terminal residues of polyP to liberate Pi [2, 12]. Studies with ppk1 bacterial mutants have showed that polyP function might be promiscuous, such as inhibition of RNA degradation, activation of Lon protease during stringent response, engagement in membrane channel structure [28, 29], correlated with the resistance to diverse stresses such as heat, oxidants, osmotic challenge, antibiotics and UV, and influence on motility, biofilm development, quorum sensing, sporulation, and virulence [23–25, 31]. Particularly, polyP is also involved in the M. tuberculosis macrophage survival [34, 35].
However, previous experiments with ppk1 bacterial mutants might be flawed by a neglect of endogenous polyP fluctuation. For example, the stationary phase Escherichia coli ppk mutant was reportedly susceptible to oxidant [7], heat shock, and osmotic stress [22]. However, oxidant resistant visible small-colonies occasionally emerged from the stationary cultures of ppk mutant [22]. One possible cause for these emerging papillate colonies might be the short chain polyP produced by the ppk mutant via an alternative pathway [3]. To preclude the endogenous polyP interference, the introduction of recombinant exopolyphosphatase is advisable [4, 32]. This methodology was pursued in our investigation into the role of PolyP in mycobacterial biofilm and colony morphology. We speculate that two M. tuberculosis genome annotation of conserved hypothetical protein homologs of PPX1 (Rv1026) and PPX2 (Rv1026) might be functional exopolyphosphatases. A structure-based sequence comparison of PPX [13] revealed that the PPX of M. tuberculosis have biological activity. As the first attempt, Rv1026 was cloned into the pMV261, which harbor an expression cassette containing 404 bp of the 5′ regulatory region of the BCG hsp60 gene and can induce the expression of its downstream gene even at 37°C [1, 33]. The resulting plasmid pMV261-Rv1026 (pRv1026) and the plasmid pMV261 were transformed into M. smegmatis strain mc2155 by electroporation. We found that the M. smegmatis (pRV1026) showed relatively decreased polyP concentration, and displayed a phenotype that was different from the M. smegmatis (pMV261) in sliding motility and biofilm formation, the surfactant Tween 80 enhancing this effect on the sliding motility and biofilm formation of the M. smegmatis (pRV1026). Comparison of the fatty acid composition of the M. smegmatis (pRV1026) with the M. smegmatis (pMV261) revealed that there are four different peaks between both strains. These results indicate that polyP can affect the fatty acid composition of cell wall and these alteration can alter the strains ability to slide and form biofilm.
Materials and Methods
Bacterium Strains and Growth Conditions
Mycobacterium smegmatis strain mc2155 was routinely grown in Middlebrook 7H9 broth with 0.05% Tween 80 unless otherwise indicated and plated in the Middlebrook 7H10 supplemented with 1% glucose and 20 μg/ml of kanamycin unless otherwise indicated.
Molecular Cloning and Electroporation
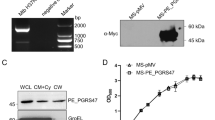
The open reading frame of Rv1026 of M. tuberculosis was PCR amplified using the forward primer 5′-CTGGTGGATCCGCAGTGG-3′ and the reverse primer 5′-CGGGCAGAAGCTTGTCCC-3′. The resulting PCR product was digested with bamHI and HindШ and cloned into pMV261 pretreated with the same enzymes to yield pRv261.
The plasmids pRv1026 and pMV261 were introduced into M. smegmatis mc2 155 by electroporation.
In Vitro Growth Kinetics and the Concentration of polyP of M. smegmatis Strains
Fresh mid log-phase cultures of the M. smegmatis (pRV1026) and the M. smegmatis (pMV261) were inoculated in M7H9 supplemented with 0.05% Tween80 to an initial OD600 of 0.02. Aliquots of 2 ml were removed at suitable intervals up to 29 h and growth was monitored by measuring OD600 values. After the measure of OD600 values, the aliquots of 2 ml were centrifuged with 12,000 r/min for 1 min and the cell pellet was used to assay total intracellular polyP by a modification of the methods of Werner, TP et al. [39], and McGrath, JW and Quinn, JP [15]. Briefly, the cell pellet was extracted for maximal 5 min at room temperature after resuspending the cell in 50 μl 1 M sulfuric acid. The suspension was neutralized with 50 μl of 2 M NaOH. Cell fragments were removed by centrifugation. To determine total intracellular polyP, 100 μl of concentrated HCl was added to 0.5 ml cell extract and heated at 100°C for 45 min; the phosphate liberated was assayed by the method of Werner, TP et al. [39]. 86 μl of 28 mM ammonium heptamolybdate in 2.1 M H2SO4 and 64 μl of 0.76 mM malachite green in 0.35% polyvinyl alcohol were added. The OD595 was measured in a SpectraMax190 produced by molecular device. The polyP concentrations were expressed in OD595 and are given as means of triplicates.
Sliding Motility of Mycobacteria
The M. smegmatis (pRv1026) and M. smegmatis (pMV261) grown on M7H9 broth (with 1% glucose, 0.05% Tween 80, and 20 μg/ml of kanamycin) were inoculated via sterile toothpicks onto M7H9 plates containing 0.3% agarose without any carbon source with/without 0.05% Tween 80. The plates were incubated at 37°C for 4–5 days.
Determination of Biofilm Formation
M. smegmatis strains were inoculated into 96-well flat bottom plate which every cell filled with 1 ml of M7H9 medium supplement with/without 0.05% Tween 80. Then, cells were incubated at 37°C without disturbance for 4–5 days. The M. smegmatis (pRv1026) and M. smegmatis (pMV261) were grown in 96-well plates, stained with 1% crystal violet and assayed for biofilm formation by spectrophotometric reading of the ethanol extract at 570 [5, 19]. Biofilm formation was analyzed by growing the strains in M7H9 media without Tween 80 and with 0.05% Tween 80 in triplicate, respectively, in 96-well plate. Cells were stained with 1% crystal violet, washed with water and crystal violet dissolved using 95% ethanol.
Extraction of Fatty Acid of Mycobacteria and Gas Chromatography Analysis
Lipid extracts from cells and gas chromatography analysis are accomplished following instruction of the Microbial Identification System (version 6.0).
Results and Discussion
The Intracellular polyP Concentration and In Vitro Growth Kinetics of the M. smegmatis
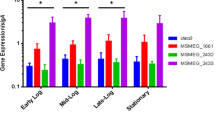
The M. smegmatis (pRv1026) showed reductive intracellular polyP concentration compared with M. smegmatis (pMV261) at early stationary phases, indicating that Rv1026 affects the polyP concentration of stationary phases cell (Fig. 1). However, the M. smegmatis (pRv1026) showed higher polyP concentration in exponential phase compared with the M. smegmatis (pMV261). This phenomenon maybe caused by the polyphoshate kinases (PPK1 and PPK2), which catalyze the reversible conversion of the terminal phosphate of ATP (or GTP) into polyP. Recently, Jagannathan et al. reported that the reversibility of the enzymatic activity of PPK1 may tilt the balance of the reaction one way or the other [11]. They speculated that the forward reaction of PPK1 is predominantly operative during the log phase, and result in polyphosphate production; during the stationary phase, the reversal reaction is triggered to synthesize ATP [11]. The existence of PPK1 and PPK2 in M. smegmatis genome suggests that the M. smegmatis PPK might be functional. So the phenomenon that the concentration of polyP in M. smegmatis (pRv1026) during exponential phase is not reductive, but increasing, may indicate that there are sufficient nutrition to supply PPK with ATP to synthetize polyP. However, during stationary phase, the polyP concentration in M. smegmatis (pRv1026) is more reductive than that in M. smegmatis (pMV261), because not only the reverse reaction of PPK is operative to synthesize ATP, but also the Rv1026 hydrolyzes the terminal residues of polyP to liberate Pi.
Intracellular polyP concentration in M. smetmatis mc2155 and the growth characteristics in vitro. The M. smegmatis (pRv1026) and the M. smegmatis (pMV261) were inoculated at a final OD600 of 0.05 into fresh medium. Aliquots of 2 ml were taken out from the M. smegmatis (pMV261) (filled diamond) and the M. smegmatis (pRv1026) (filled triangle) at time points indicated and measured for OD600. After the measure of OD600, the aliquots of 2 ml were used to extract polyP as described in method. The polyP concentration in the M. smegmatis (pRv1026) (black column) and the M. smegmatis (pMV261) (white column) was denoted as OD595 of Pi. Values plotted are the averages and SD from triplicate experiments. An unhydrolyzed sample was used as a control to determine the background level of Pi
The reduction of polyP concentration in M. smegmatis (pRv1026) have no apparent effect to the planktonic growth (Fig. 1), which is consistent with the growth curves of ppk mutant of E. coli and Pseudomonas aeruginosa [24]. So we analyze the effect of reduction of polyP on sliding motility and morphology of the M. smegmatis (pRv1026) and M. smegmatis (pMV261).
Analysis of Sliding Motility and Morphology of M. smegmatis
Previous studies have shown that polyP-defective mutants like Paeruginosa [8, 23–25], Bacillus cereus [31], Dictyostelium discoideum [42], and Burkholderia pseudomallei [37] have defective motility including swimming motility, swarming and twitching motilities. In spite of being nonflagellated microorganisms, mycobacteria can spread on the surface of solid growth medium by a sliding mechanism [14]. This form of surface motility is produced by the expansive forces of the growing bacterial population and cell surface properties that favor reduced friction between the cells and substrate, and the result is the slow movement of a uniform monolayer of cells as a unit [10]. The lipid components of cell wall including glycopeptidolipids (GPLs) [26, 27] and mycolyl-diacylglycerols (MDAGs) [6] contribute to sliding motility and biofilm formation. We found that the M. smegmatis (pRv1026) showed morphological variations compared with M. smegmatis (pMV261) grown on 0.3% agarose plates without 0.05% Tween80 (Fig. 2a (1,2)) and with 0.05% Tween 80 (Fig. 2b (1,2)), respectively.
Morphology and sliding motility of M. smegmatis. a Morphology of M. smegmatis (pMV261) (1) and the M. smegmatis (pRv1026) (2) grown on 0.3% agarose plate without Tween 80. b Morphology of the M. smegmatis (pMV261) (1) and the M. smegmatis (pRv1026) (2) grown on 0.3% agarose plate with 0.05% Tween 80. c Sliding motility of the M. smegmatis (pMV261) (1) and the M. smegmatis (pRv1026) (2) grown on 0.3% agarose plate without Tween 80. d Sliding motility of the M. smegmatis (pMV261) (1) and the M. smegmatis (pRv1026) (2) grown on 0.3% agarose plate with 0.05% Tween 80
Differences in colony morphology were also shown to be associated with the ability of sliding motility. The motility of the M. smegmatis (pRv1026) was slightly increased when compared with the M. smegmatis (pMV261) on 0.3% agarose plates without Tween80 (Fig. 2c (1,2)). In contrast, colony of the M. smegmatis (pRv1026) grown on 0.3% agarose plate with 0.05% Tween80 showed distinguished reduction in diameter as compared with M. smegmatis (pMV261) (Fig. 2d (1,2)), indicating that the sliding motility of the M. smegmatis (pRv1026) is more sensitive to denaturant such as Tween 80 than the M. smegmatis (pMV261). Previous article had reported that Tween 80 alters the cell surface properties and expose gradually more deeply buried lipids [20]. And recently, many evidences indicate that polyP is an active factor engaged in transcriptional regulation and metabolism [36, 38, 40]. So we hypothesize that the polyP can cause the variation of constituents of bacterial cell envelope by involvement in membrane channel structure or involvement in the transcriptional regulation of genes encoding lipids on the cell envelopes of mycobacteria.
The Altered Biofilm Formation of Recombinant
Since the M. smegmatis (pRv1026) forms drastically different colonies on agarose plates with 0.05% Tween80 from M. smegmatis (pMV261), we also examined that whether the ability of biofilm formation is different between the two strains. Using an assay to determine biofilm formation [5, 19], we found that the M. smegmatis (pRv1026) formed more biofilms in comparison with the M. smegmatis (pMV261) grown on media without Tween 80 (Fig. 3a). However, the M. smegmatis (pRv1026) formed less biofilms compared with the M. smegmatis (pMV261) on media with 0.05% Tween 80 (Fig. 3b). This result was also conformed by Quantitative estimation of crystal violet staining (Fig. 3c).
Biofilm formation associated with polyP depletion. a, b Analysis of biofilm formation. The dissolved crystal violet of the M. smegmatis (pMV261) (row A), the M. smegmatis (pRv1026) (row B) and M7H9 (no strains) (row C) grown without Tween 80 (a) and with 0.05% Tween 80 (b) was transferred to novel 96-well plate in triplicate columns 1, 2, and 3. c Estimation of crystal violet uptake. OD570 was measured to calculate the amount of crystal violet uptake by the M. smegmatis (pMV261) (A), the M. smegmatis (pRv1026) (B) and M7H9 media (C) grown without Tween 80 (black bar) and with 0.05% Tween 80 (blank bar). The experiment war carried out in triplicate and the graph indicates the mean value with standard deviation bars
A previous study proposed a model for the role of GPLs in sliding motility and biofilm formation based on interaction between the mycobacterial cell surface and either hydrophilic (agarose) or hydrophobic (polystyrene) surfaces [27]. In addition, the loss of hydrophobic mycolyl-diacylalycerols (MDAGs) reduces the hydrophobicity of cell surface and biofilm formation [6]. Preliminary studies showed there are four different peaks between the gas chromatography of cellular wall fatty acid of the M. smegmatis (pRv1026) and the M. smegmatis (pMV261) (Fig. 4). These results indicate that polyP deficiency can affect the fatty acid composition of cellular wall. These alteration of cell wall might elucidate the reductive ability of strains to slide and form biofilm. Colony morphology is a complex phenotype influenced by the ability of cells to interact with one another. Cell wall architecture is not only responsible for distinctive colony morphology, but also contributes to virulence, persistence in macrophages, and modulation of the host immune response. Presently, the known lipoids including glycopeptidolipids and mycolyl-diacylglycerols affect mycobacterial phenotypes such as colony morphology, sliding motility, and biofilm formation. Additionally, the nutrition factor such as iron [19] and signal transduction such as two-component system [16] and serine/threonine protein kinase [9] also have effect on mycobacterial phenotypes. As to know, this is the first report on the functional analysis of a putative Rv1026 from M. tuberculosis and might provide new information about the role of polyP in affecting cell wall architecture. Further studies are needed to understand the mechanisms underlying the effect of polyP on cell wall architecture.
References
Andreu N, Soto CY, Roca I, Martin C, Gibert I (2004) Mycobacterium smegmatis displays the Mycobacterium tuberculosis virulence-related neutral red character when expressing the Rv0577 gene. FEMS Microbiol Lett 231:283–289
Brown MR, Kornberg A (2008) The long and short of it—polyphosphate, PPK and bacterial survival. Trends Biochem Sci 33:284–290
Castuma CE, Huang R, Kornberg A, Reusch RN (1995) Inorganic polyphosphates in the acquisition of competence in Escherichia coli. J Biol Chem 270:12980–12983
Chavez FP, Mauriaca C, Jerez CA (2009) Constitutive and regulated expression vectors to construct polyphosphate deficient bacteria. BMC Res Notes 2:50
Chen JM, German GJ, Alexander DC, Ren H, Tan T, Liu J (2005) Roles of Lsr2 in colony morphology and biofilm formation of Mycobacterium smegmatis. J Bacteriol 188:633–641
Chen JM, German GJ, Alexander DC, Ren H, Tan T, Liu J (2006) Roles of Lsr2 in colony morphology and biofilm formation of Mycobacterium smegmatis. J Bacteriol 188:633–641
Crooke E, Akiyama M, Rao NN, Kornberg A (1994) Genetically altered levels of inorganic polyphosphate in Escherichia coli. J Biol Chem 269:6290–6295
Fraley CD, Rashid MH, Lee SS, Gottschalk R, Harrison J, Wood PJ et al (2007) A polyphosphate kinase 1 (ppk1) mutant of Pseudomonas aeruginosa exhibits multiple ultrastructural and functional defects. Proc Natl Acad Sci USA 104:3526–3531
Gopalaswamy R, Narayanan S, Jacobs WR Jr, Av-Gay Y (2008) Mycobacterium smegmatis biofilm formation and sliding motility are affected by the serine/threonine protein kinase PknF. FEMS Microbiol Lett 278:121–127
Henrichsen J (1972) Bacterial surface translocation: a survey and a classification. Bacteriol Rev 36:478–503
Jagannathan V, Kaur P, Datta S (2010) Polyphosphate kinase from M. tuberculosis: an interconnect between the genetic and biochemical role. PLoS One 5:e14336
Kornberg A, Rao NN, Ault-Riche D (1999) Inorganic polyphosphate: a molecule of many functions. Annu Rev Biochem 68:89–125
Lindner SN, Knebel S, Wesseling H, Schoberth SM, Wendisch VF (2009) Exopolyphosphatases PPX1 and PPX2 from Corynebacterium glutamicum. Appl Environ Microbiol 75:3161–3170
Martinez A, Torello S, Kolter R (1999) Sliding motility in mycobacteria. J Bacteriol 181:7331–7338
McGrath JW, Quinn JP (2000) Intracellular accumulation of polyphosphate by the yeast Candida humicola G-1 in response to acid pH. Appl Environ Microbiol 66:4068–4073
Nguyen HT, Wolff KA, Cartabuke RH, Ogwang S, Nguyen L (2010) A lipoprotein modulates activity of the MtrAB two-component system to provide intrinsic multidrug resistance, cytokinetic control and cell wall homeostasis in Mycobacterium. Mol Microbiol 76:348–364
O’Toole G, Kaplan HB, Kolter R (2000) Biofilm formation as microbial development. Annu Rev Microbiol 54:49–79
Ojha A, Anand M, Bhatt A, Kremer L, Jacobs WR Jr, Hatfull GF (2005) GroEL1: a dedicated chaperone involved in mycolic acid biosynthesis during biofilm formation in mycobacteria. Cell 123:861–873
Ojha A, Hatfull GF (2007) The role of iron in Mycobacterium smegmatis biofilm formation: the exochelin siderophore is essential in limiting iron conditions for biofilm formation but not for planktonic growth. Mol Microbiol 66:468–483
Ortalo-Magne A, Lemassu A, Laneelle MA, Bardou F, Silve G, Gounon P et al (1996) Identification of the surface-exposed lipids on the cell envelopes of Mycobacterium tuberculosis and other mycobacterial species. J Bacteriol 178:456–461
Rao NN, Gomez-Garcia MR, Kornberg A (2009) Inorganic polyphosphate: essential for growth and survival. Annu Rev Biochem 78:605–647
Rao NN, Kornberg A (1996) Inorganic polyphosphate supports resistance and survival of stationary-phase Escherichia coli. J Bacteriol 178:1394–1400
Rashid MH, Kornberg A (2000) Inorganic polyphosphate is needed for swimming, swarming, and twitching motilities of Pseudomonas aeruginosa. Proc Natl Acad Sci USA 97:4885–4890
Rashid MH, Rao NN, Kornberg A (2000) Inorganic polyphosphate is required for motility of bacterial pathogens. J Bacteriol 182:225–227
Rashid MH, Rumbaugh K, Passador L, Davies DG, Hamood AN, Iglewski BH et al (2000) Polyphosphate kinase is essential for biofilm development, quorum sensing, and virulence of Pseudomonas aeruginosa. Proc Natl Acad Sci USA 97:9636–9641
Recht J, Kolter R (2001) Glycopeptidolipid acetylation affects sliding motility and biofilm formation in Mycobacterium smegmatis. J Bacteriol 183:5718–5724
Recht J, Martinez A, Torello S, Kolter R (2000) Genetic analysis of sliding motility in Mycobacterium smegmatis. J Bacteriol 182:4348–4351
Reusch RN (1999) Polyphosphate/poly-(R)-3-hydroxybutyrate) ion channels in cell membranes. Prog Mol Subcell Biol 23:151–182
Reusch RN (2000) Transmembrane ion transport by polyphosphate/poly-(R)-3-hydroxybutyrate complexes. Biochemistry (Mosc) 65:280–295
Rose L, Kaufmann SH, Daugelat S (2004) Involvement of Mycobacterium smegmatis undecaprenyl phosphokinase in biofilm and smegma formation. Microbes Infect 6:965–971
Shi X, Rao NN, Kornberg A (2004) Inorganic polyphosphate in Bacillus cereus: motility, biofilm formation, and sporulation. Proc Natl Acad Sci USA 101:17061–17065
Shiba T, Tsutsumi K, Yano H, Ihara Y, Kameda A, Tanaka K et al (1997) Inorganic polyphosphate and the induction of rpoS expression. Proc Natl Acad Sci USA 94:11210–11215
Stover CK, de la Cruz VF, Fuerst TR, Burlein JE, Benson LA, Bennett LT et al (1991) New use of BCG for recombinant vaccines. Nature 351:456–460
Sureka K, Dey S, Datta P, Singh AK, Dasgupta A, Rodrigue S et al (2007) Polyphosphate kinase is involved in stress-induced mprAB-sigE-rel signalling in mycobacteria. Mol Microbiol 65:261–276
Sureka K, Sanyal S, Basu J, Kundu M (2009) Polyphosphate kinase 2: a modulator of nucleoside diphosphate kinase activity in mycobacteria. Mol Microbiol 74:1187–1197
Tsutsumi K, Munekata M, Shiba T (2000) Involvement of inorganic polyphosphate in expression of SOS genes. Biochim Biophys Acta 1493:73–81
Tunpiboonsak S, Mongkolrob R, Kitudomsub K, Thanwatanaying P, Kiettipirodom W, Tungboontina Y et al (2010) Role of a Burkholderia pseudomallei polyphosphate kinase in an oxidative stress response, motilities, and biofilm formation. J Microbiol 48:63–70
Varela C, Mauriaca C, Paradela A, Albar JP, Jerez CA, Chavez FP (2011) New structural and functional defects in polyphosphate deficient bacteria: a cellular and proteomic study. BMC Microbiol 10:7
Werner TP, Amrhein N, Freimoser FM (2005) Novel method for the quantification of inorganic polyphosphate (iPoP) in Saccharomyces cerevisiae shows dependence of iPoP content on the growth phase. Arch Microbiol 184:129–136
Yang ZX, Zhou YN, Yang Y, Jin DJ (2011) Polyphosphate binds to the principal sigma factor of RNA polymerase during starvation response in Helicobacter pylori. Mol Microbiol 77:618–627
Zambrano MM, Kolter R (2005) Mycobacterial biofilms: a greasy way to hold it together. Cell 123:762–764
Zhang H, Gomez-Garcia MR, Brown MR, Kornberg A (2005) Inorganic polyphosphate in Dictyostelium discoideum: influence on development, sporulation, and predation. Proc Natl Acad Sci USA 102:2731–2735
Acknowledgments
The study is Supported by the National key infectious disease project (No. 2008ZX10003-006, No. 2008ZX10003-001), national natural science foundation (No. 81071316,90813019), Excellent PhD thesis fellowship of southwest university (No. kb2009010, No. ky2009009), The Fundamental Research Funds for the Central Universities (XDJK2009A003) and Natural Science Foundation Project of CQ CSTC (CSTC, 2010BB5002).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Shi, T., Fu, T. & Xie, J. Polyphosphate Deficiency Affects the Sliding Motility and Biofilm Formation of Mycobacterium smegmatis . Curr Microbiol 63, 470 (2011). https://doi.org/10.1007/s00284-011-0004-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-011-0004-4