Abstract
The antibiotic nisin, produced by Lactococcus lactis subsp. lactis N8, offers an extensive commercial prospect as natural food preservatives. The nisin immunity of the L. lactis strains is regulated by a variety of mechanisms. In this study, we isolated a L. lactis L31 strain with increased nisin resistance from a mini-Mu transposon mutant pool of strain N8. The single Mu insertion in strain L31 was in the irpT gene with unknown function. By comparing the proteomic profiles of L. lactis L31 and its parental strain, we found that changes occurred in the synthesis of a protein involved in cell wall biosynthesis (RmlD). Strain L31 had 13.7% higher content of rhamnose in the cell wall than the N8 strain. Overexpression of RmlD involved in the synthesis of dTDP-l-rhamnose in the nisin-sensitive MG1363 strain increased nisin resistance of the strain. The results indicate that these cellular proteins effected nisin resistance in L. lactis N8.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nisin is a post-translationally modified lanthionine-containing peptide antibiotic, which is produced by certain strains of Lactococcus lactis and Streptococcus uberis [1, 2]. In order to avoid self-destruction, the nisin-producing L. lactis strains protect themselves against the bactericidal activity of nisin, which is called nisin immunity [3]. Full nisin immunity requires four polypeptides, the membrane-bound lipoprotein NisI, an ABC transporter complex containing a cytoplasmic NisF, and two integral membrane proteins, NisE and NisG [4, 5]. Functional analysis of these four components showed that protection mediated by NisI occurs at the outer side of the cytoplasmic membrane, the location of lipoproteins, and seems to function partly by intercepting nisin. In addition, part of the translocated NisI escapes lipid modification and is secreted to the medium contributing to nisin immunity. NisI seems to cooperate with the ABC type transporter NisFEG that exports nisin from the cell surfaces into the medium.
Non-nisin-producing strains can develop nisin resistance. For example, through stepwise exposure of cells to increasing concentrations of nisin, the generation of nisin-resistant (Nisr) strains of Listeria. monocytogenes and L. lactis was achieved [6, 7]. Through genetic engineering of Bacillus subtilis, nisin resistance was increased [8]. Transcriptome analysis of nisin-resistant strains revealed many differentially expressed genes involved in nisin resistance [9]. So far, five major nisin resistance mechanisms could be concluded from these analyses. The first mechanism is the prevention of nisin from reaching the membrane and the lipid II molecule [10]. Second, other L. lactis Nisr strains could change the local pH at the outer surface of the cytoplasmic membrane. Third, acquiring nisin resistance may come from changing the elongation and saturation of phospholipids. Fourth, ABC transporters might be involved in expelling nisin from the cytoplasmic membrane, preventing nisin–lipid II binding and pores forming. Finally, expression of nisin-inactivating enzymes could protect strains [11].
Since nisin immunity/resistance is a complex system, several challenges still exist in understanding this system [3]. In this study, through mini-Mu transposon technique, a gene with unknown function, named irpT in L. lactis N8 genome, was found to increase nisin resistance. Proteome analysis of the irpT mutant identified several proteins potentially responsible for this resistance increase. One of these, RmlD involved in the synthesis of dTDP-l-rhamnose and needed for the biosynthesis of the cell wall, mediated nisin resistance upon overexpression in a nisin-sensitive L. lactis strain.
Materials and Methods
Strains and Plasmids
All bacterial strains and plasmids used in this study are presented in Table 1. The Lactococcus lactis strains were grown at 30°C without agitation in M17 medium (Oxoid, Hampshire, UK) supplemented with 0.5% glucose and 0.5% sucrose (M17GS). For trans-complementation, the lactococcal expression plasmid pLEV16 was constructed by PCR amplification, which included in the pWV01 replicon, the constitutive P45 promoter, the chloramphenicol resistance gene of pSG13 and the kanamycin resistance gene of pHTH2. The complete irpT gene PCR product was cloned into the plasmid pLEV16, downstream from the promoter P45, yielding pLEV17. The plasmid pLEV16 carried a 1,039 bp rmlD gene fragment, yielding pLEV18.
Mini-Mu Transposon Technique and Southern Blotting
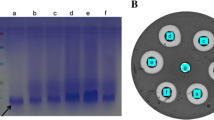
Lactococcus lactis N8 was used as a target bacterium for the Mu insertion mutagenesis. The methods for preparation of electrocompetent cells, gene cloning, and sequencing were described previously [12]. Using the mini-Mu transposition technique in L. lactis N8, we generated a genomic random insertion mutant library of approximately 1,800 member clones. Genomic DNA, digested with PvuII and EcoRI, respectively, was electrophoresed on a 1% agarose gel and blotted onto a nylon membrane. Southern hybridization was carried out with DIG-High Prime labeled (Roche, Mannheim, Germany) Em-Mu was used as a probe. Restriction enzymes and Taq DNA polymerases were purchased from TaKaRa (Dalian, China).
Determination of Minimum Inhibitory Concentrations
The insertion mutant pool was screened for altered minimum inhibitory concentrations of nisin, which were estimated from the growth curves similarly as in Takala and Saris [13]. The L. lactis strains were grown overnight in antibiotic-free M17GS medium, diluted 1:100, and nisin was added to concentrations of 0–7,000 IU ml−1 for screening the ones with changed MIC of nisin. The MIC was determined as the minimum nisin needed to ensure that the culture did not grow to over 10% of the relative cell density at 600 nm in 16 h. L. lactis L31 was identified from the mutant pool that portrayed an increased Nisin resistance. Nisin standard sample was purchased from Sigma (Steinheim, Germany).
Two-Dimensional Gel Electrophoresis
For two-dimensional gel electrophoresis (2-DE) analysis, 50 ml of the lactococcal culture was harvested in the logarithmic period of growth by centrifugation. The cell pellets were suspended in 3 ml of lysis buffer (7.0 mol l−1 urea, 2.0 mol l−1 thiourea, 4% CHAPS). The cells were disrupted with Scientz JY92-II sonifier (Ningbo, China). 2-DE was carried out as described by Görg et al. [14] using the 2-DE system from Bio-Rad (Hercules, CA, USA). The first dimension (IEF) was performed using a 17 cm IPG strips with a linear gradient ranging from 4 to 7. The second dimension was carried out on 12% polyacrylamide gels. PDQuest Analysis Software was used for image analysis.
Real-Time Quantitative PCR
All primer pairs were designed based on the corresponding gene sequence of L. lactis SK11 [15]. Total RNA of L. lactis strains was extracted with TRIzol Reagent from Invitrogen (Carlsbad, CA, USA), according to the manufacturer’s protocol. RT-qPCR was performed on the iQ5 real-time system (Bio-Rad, Hercules, CA, USA), using TranStart Green qPCR SuperMix (TransGen, Beijing, China). All RT-qPCR reactions were repeated independently three times and each repeat contained three biological replicates. Data analysis was carried out using the comparative CT (2−ΔΔCT) method with 16S rRNA as the internal reference gene [16].
Isolation and Characterization of Cell Wall Sugars
Isolation of cell wall sugars was performed as described by Looijesteijn et al. [17]. The monomeric sugar composition after hydrolysis was determined by high-performance liquid chromatography [18]. The values presented below are averages based on at least three independent experiments.
Results
Isolation and Phenotypic Characterization of L. lactis L31
Through the MIC determination of L. lactis N8 strain and its derivatives, a mutant strain with improved nisin resistance was isolated and named L31. The result showed that L31 with a Mu transposon had significantly higher nisin resistance than N8 (Table 2).
Identification and Sequence Analysis of Hypothetical irpT Gene
The flanking region of the Mu insertion site in L. lactis L31 was cloned and sequenced, which revealed that the disrupted gene was irpT in L. lactis N8. The irpT gene (708 bp) with unknown function shares 91% identity with a gene encoding a hypothetical protein in L. lactis SK11 [15]. The complete nucleotide sequence of the irpT gene has been submitted to GenBank databases, and has been assigned the accession number GQ386851.
Southern blotting was performed to investigate whether the L. lactis L31 had only one Mu transposon DNA copy in the genomic DNA. No signal was obtained from the control genome of the parental strain N8, whereas the genome of the L. lactis L31 had only one hybridization band (data not shown). This result showed that only a single copy of Em-Mu transposon DNA had inserted into the genomic DNA of L. lactis L31.
Complementation of the Inactive irpT Gene in Strain L31
The vector pLEV17 as well as the empty vector pLEV16 was electroporated into the irpT mutant strain L31, generating the strain L31-pLEV17, and the control strain L31-pLEV16, respectively. The resulting strains were grown in antibiotic-free medium and tested for MIC of nisin. The assays showed that nisin resistance of the L31 mutant was restored to that of the parent strain N8 when exogenous irpT was expressed in it, which further confirmed the relationship between irpT and nisin resistance.
Changes in the Proteome of L. lactis L31
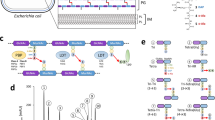
In order to investigate the effects of the irpT gene disruption, two-dimensional polyacrylamide gel electrophoresis (2-DE) was used to compare the proteomic profiles of L. lactis L31 and L. lactis N8. The number of distinct spots detected for L. lactis L31 was highly similar to that found for the parental strain N8 (approx. 390). However, five spots from these two strains showed about 6-fold difference in intensity. The five proteins are dTDP-l-rhamnose synthase (RmlD), phosphocarrier protein Hpr (PtsH), phosphopyruvate hydratase (Eno), glyceraldehyde 3-phosphate dehydrogenase (GapB), and hypothetical protein (YahB), respectively. One of them (PtsH) was considered to be decreased in intensity in L. lactis L31, and the four other spots were increased (Eno, RmlD, YahB, and GapB; Fig. 1a, b).
Comparison of the protein pattern of L. lactis strains by 2-DE (pH gradient 4–7) and the mRNA level by real-time quantitative PCR. Subsection views of differentially expressed proteins on the 2-D gels of L. lactis N8 (a) and L. lactis L31 (b); (c) The mRNA levels of five genes in L. lactis L31 (filled bars) and strain L31-pLEV17 (chromosomal non-functional irpT gene and functional irpT in plasmid; open bars) relative to their corresponding genes in N8 and L31-pLEV16, respectively. The gene 16S rRNA was used as the internal reference gene. All real-time quantitative PCR results were expressed relative to 16S rRNA. The error bars represent the standard deviations based on three experiments
In order to verify that the expression of these five genes was changed in L. lactis L31, the RT-qPCR was done to detect the mRNA levels of these genes. The analysis showed that the ptsH gene was expressed to a lower extent in L. lactis L31, and the eno, rmlD, yahB, and gapB genes showed that higher expression in L31 than in N8 (filled bars in Fig. 1c). Moreover, the mRNA levels were regulated in opposite directions in the complementary strain L31-pLEV17 (open bars in Fig. 1c), in which the ptsH gene was expressed to a higher extent, and other four genes showed that lower expression than in L31. The results of RT-qPCR confirmed up- and down-regulation in the transcription level of five genes, which were consistent with their changes shown in 2-DE.
Influence of the irpT Gene Mutant on Cell Wall Polysaccharides
The proteins with altered expression levels in L31 belong to three functional groups, cell wall biosynthesis, central and energy metabolism, and stress-related proteins. Among them, the rmlD gene is involved in the synthesis of dTDP-l-rhamnose, which is an immediate precursor for cell wall polysaccharides backbone production [19]. Cell wall polysaccharides may act as a barrier that prevents some antibiotics from reaching their target molecules in the cell wall and may partly contribute to the resistance to some antibiotics [20]. Therefore, we focused on the rmlD gene to investigate if this gene can influence cell wall polysaccharides and nisin resistance.
In order to test if the cell wall polysaccharides produced by strain L31 were changed, its sugar composition was analyzed by high-performance liquid chromatography. The result showed that the rhamnose content of cell wall polysaccharides was increased by 13.7% (P < 0.05) in L. lactis L31 compared with its parental strain. The result is consistent with the up-regulation of the rmlD gene expression level.
In order to investigate the relationship between nisin resistance and rmlD gene, RmlD was overexpressed in L. lactis MG1363 [21], followed by analysis of nisin resistance. The result demonstrated that RmlD overexpression improves the nisin resistance (Table 2).
Discussion
In this study, we identified and sequenced the irpT gene that upon inactivation increased nisin resistance of L. lactis N8. In the genome of L. lactis SK11, homologous gene of irpT is present at the penultimate position within the same direction of nine genes, which may generate a gene cluster. The downstream gene of irpT is xerS, encoding a site-specific tyrosine recombinase XerS [22]. Sequence analysis showed that the xerS gene has its own promoter, therefore the disruption of irpT gene may not affect xerS gene expression, which was also confirmed by complementation analysis and RT-PCR (data not shown).
The IrpT protein of unknown function is a membrane protein predicted by the online software SOSUI (http://bp.nuap.nagoya-u.ac.jp/sosui/). In order to determine whether the IrpT protein is directly involved in promoting nisin binding to the cell surface, we performed the nisin adsorption experiment and found that loss of IrpT function in L. lactis L31 does not affect nisin binding to the cell surface (data not shown). Moreover, to find out whether the improvement of nisin resistance caused by irpT disruption is through affecting the level of membrane-bound lipoprotein NisI, we detected the amount of lipoprotein NisI. Western blot analysis showed that the disruption of irpT did not change the level of the lipoprotein NisI (data not shown). As the nisin immunity genes (nisI and NisFEG) are all in the same regulon the effect of irpT knockout seems not be a result of influence on regulation of the nisin operons.
In order to study the mechanisms involved in the increased nisin resistance of L31, we compared the proteomic profiles of L. lactis L31 and its parental strain N8 by 2-DE. The results showed that difference in the expression levels of five proteins, which are not present in the gene cluster of irpT. Four (Hpr, Eno, YahB, and GapB) of the five proteins have been investigated by previous studies to show indirect influence on nisin resistance [9]. For example, YahB is a hypothetical protein, which was defined as a universal stress protein, a UspA-like nucleotide-binding protein. Multiple members of the UspA family are found in a variety of organisms, such as bacteria, archaea, fungi, protozoa, and plants, and their synthesis is induced in response to a large number of stress conditions, including starvation for carbon, nitrogen, phosphate, sulfate, and amino acid and exposure to heat, oxidants, metals, uncouplers of the electron transport chain, polymyxin, cycloserine, ethanol, and antibiotics [23, 24]. Nisin is an antibiotic, a kind of stress for the nisin-producing L. lactis strains. Perhaps, YahB is contributing to the increased nisin resistance in L. lactis L31.
Several recent reports have suggested that the composition of the cell wall can directly influence nisin resistance [10]. For this reason, we focused on the rhamnose content of cell wall polysaccharides to investigate how the rmlD gene, which is overexpressed in L. lactis L31, influences nisin resistance. The rml operon is composed of four genes, rmlABCD, encoding glucose-1-phosphate thymidylyltransferase, dTDP-glucose-4,6-dehydratase, dTDP-4-keto-6-deoxyglucose-3,5-epimerase and dTDP-l-rhamnose synthase, respectively. Through a four-step enzymic reaction, glucose-1-phosphate turns into dTDP-l-rhamnose [25], which is a precursor of cell wall polysaccharides backbone in L. lactis. It has been reported that cell wall polysaccharides was absent in the cell wall of a rmlD mutant strain, which exhibited about 5-fold-higher sensitivity to nisin than wild strain [20], suggesting that the presence of cell wall polysaccharides may confer resistance to nisin in Streptococcus mutans. Although, there are many differences between streptococci and lactococci in the metabolic regulation [26], the involvement of the rmlD gene in S. mutans nisin resistance suggests that the corresponding gene in L. lactis may also influence nisin resistance in a similar way. Over-expression of RmlD, which is the case in L31, may increase nisin resistance potentially via the rhamnose content of the cell wall. Thus, we analyzed the sugar composition of cell wall polysaccharides in L. lactis L31 and L. lactis N8. Indeed, the rhamnose content was increased in L. lactis L31. Moreover, RmlD overexpression in nisin-sensitive L. lactis MG1363 increased the resistance level to nisin. These results implied that the disruption of the irpT gene may increase the nisin resistance level partially by up-regulation of RmlD protein expression.
In conclusion, nisin immunity/resistance in the nisin-producing L. lactis strains is comprehensively regulated by a variety of mechanisms. In this study, we discovered a novel gene irpT, whose disruption increased nisin resistance. IrpT did not seem to be directly involved in nisin resistance. Rather it has a regulatory role as its lack of function resulted in up-regulation of the rmlD gene. These results may be useful for constructing strains with increased production of nisin as the nisin resistance level is one factor restricting the concentration of produced nisin.
References
Immonen T, Ye S, Ra R, Qiao M, Paulin L, Saris PEJ (1995) The codon usage of the nisZ operon in Lactococcus lactis N8 suggests a non-lactococcal origin of the conjugative nisin-sucrose transposon. DNA Seq 5:203–218
Wirawan RE, Klesse NA, Jack RW, Tagg JR (2006) Molecular and genetic characterization of a novel nisin variant produced by Streptococcus uberis. Appl Environ Microbiol 72:1148–1156
Lubelski J, Rink R, Khusainov R, Moll GN, Kuipers OP (2008) Biosynthesis, immunity, regulation, mode of action and engineering of the model lantibiotic nisin. Cell Mol Life Sci 65:455–476
Takala TM, Koponen O, Qiao M, Saris PEJ (2004) Lipid-free NisI: interaction with nisin and contribution to nisin immunity via secretion. FEMS Microbiol Lett 237:171–177
Immonen T, Saris PEJ (1998) Characterization of the nisFEG operon of the nisin Z producing Lactococcus lactis subsp. lactis N8 strain. DNA Seq 9:263–274
Kramer NE, Smid EJ, Kok J, de Kruijff B, Kuipers OP, Breukink E (2004) Resistance of Gram-positive bacteria to nisin is not determined by lipid II levels. FEMS Microbiol Lett 239:157–161
Ming X, Daeschel MA (1993) Nisin resistance of foodborne bacteria and the specific resistance responses of Listeria monocytogenes Scott-A. J Food Protect 56:944–948
Hansen ME, Wangari R, Hansen EB, Mijakovic I, Jensen PR (2009) Engineering of Bacillus subtilis 168 for increased nisin resistance. Appl Environ Microbiol 75:6688–6695
Kramer NE, van Hijum SA, Knol J, Kok J, Kuipers OP (2006) Transcriptome analysis reveals mechanisms by which Lactococcus lactis acquires nisin resistance. Antimicrob Agents Chemother 50:1753–1761
Kramer NE, Hasper HE, van den Bogaard PT, Morath S, de Kruijff B, Hartung T, Smid EJ, Breukink E, Kok J, Kuipers OP (2008) Increased D-alanylation of lipoteichoic acid and a thickened septum are main determinants in the nisin resistance mechanism of Lactococcus lactis. Microbiology 154:1755–1762
Sun Z, Zhong J, Liang X, Liu J, Chen X, Huan L (2009) Novel mechanism for nisin resistance via proteolytic degradation of nisin by the nisin resistance protein NSR. Antimicrob Agents Chemother 53:1964–1973
Wu Z, Xuanyuan Z, Li R, Jiang D, Li C, Xu H, Bai Y, Zhang X, Turakainen H, Saris PEJ (2009) Mu transposition complex mutagenesis in Lactococcus lactis—identification of genes affecting nisin production. J Appl Microbiol 106:41–48
Takala TM, Saris PEJ (2006) C terminus of NisI provides specificity to nisin. Microbiology 152:3543–3549
Gorg A, Obermaier C, Boguth G, Harder A, Scheibe B, Wildgruber R, Weiss W (2000) The current state of two-dimensional electrophoresis with immobilized pH gradients. Electrophoresis 21:1037–1053
Makarova K, Slesarev A, Wolf Y, Sorokin A, Mirkin B, Koonin E, Pavlov A, Pavlova N, Karamychev V, Polouchine N (2006) Comparative genomics of the lactic acid bacteria. Proc Natl Acad Sci USA 103:15611–15616
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods 25:402–408
Looijesteijn PJ, Boels IC, Kleerebezem M, Hugenholtz J (1999) Regulation of exopolysaccharide production by Lactococcus lactis subsp. cremoris by the sugar source. Appl Environ Microbiol 65:5003–5008
Parrish FW, Hicks K, Doner L (1980) Analysis of lactulose preparations by spectrophotometric and high performance liquid chromatographic methods. J Dairy Sci 63:1809–1814
Tsukioka Y, Yamashita Y, Nakano Y, Oho T, Koga T (1997) Identification of a fourth gene involved in dTDP-rhamnose synthesis in Streptococcus mutans. J Bacteriol 179:4411–4414
Tsuda H, Yamashita Y, Shibata Y, Nakano Y, Koga T (2002) Genes involved in bacitracin resistance in Streptococcus mutans. Antimicrob Agents Chemother 46:3756–3764
Gasson MJ (1983) Plasmid complements of Streptococcus lactis NCDO 712 and other lactic streptococci after protoplast-induced curing. J Bacteriol 154:1–9
Le Bourgeois P, Bugarel M, Campo N, Daveran-Mingot ML, Labonte J, Lanfranchi D, Lautier T, Pages C, Ritzenthaler P (2007) The unconventional Xer recombination machinery of Streptococci/Lactococci. PLoS Genet 3:e117
Siegele DA (2005) Universal stress proteins in Escherichia coli. J Bacteriol 187:6253–6254
Nachin L, Nannmark U, Nystrom T (2005) Differential roles of the universal stress proteins of Escherichia coli in oxidative stress resistance, adhesion, and motility. J Bacteriol 187:6265–6272
Boels IC, Beerthuyzen MM, Kosters MH, Van Kaauwen MP, Kleerebezem M, De Vos WM (2004) Identification and functional characterization of the Lactococcus lactis rfb operon, required for dTDP-rhamnose biosynthesis. J Bacteriol 186:1239–1248
Burne RA, Bessen DE, Broadbent JR, Claverys JP (2007) The seventh international conference on the genetics of streptococci, lactococci, and enterococci. J Bacteriol 189:1209–1218
Acknowledgments
The authors thank Dr Harri Savilahti (University of Helsinki) for providing Mu transposon DNA, Prof. Deling Kong (Nankai University) for providing the L. lactis MG1363. This study was supported by program for New Century Excellent Talents in University of China (Grant No. NCET-06-0212).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xuanyuan, Z., Wu, Z., Li, R. et al. Loss of IrpT Function in Lactococcus lactis subsp. lactis N8 Results in Increased Nisin Resistance. Curr Microbiol 61, 329–334 (2010). https://doi.org/10.1007/s00284-010-9615-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-010-9615-4